Abstract
Attenuated total reflection Fourier-transform infrared (ATR-FTIR) spectroscopy is a surface-sensitive and label-free technique, which is applied to obtain dynamic structural information of biomolecules. The study of proteins by ATR-FTIR spectroscopy can be impeded by their tendency to adsorb to solid surfaces. Furthermore, the adsorption process of proteins is often accompanied with conformational changes, which can interfere with the intended structural analysis. We efficiently modified a silicon ATR crystal surface with polyethylene glycol and thereby create a protein-repellent surface. To achieve a high sensitivity, which enables the study of small conformational changes of biomolecules, we combine surface passivation with specific immobilization. This is accomplished via the biotin-streptavidin interaction, which is one of the strongest known non-covalent protein-ligand interactions. As a proof of concept we present the specific immobilization of DNA. The modified surface is stable against elevated temperatures and 8 M urea and can therefore be used to study a wide range of biochemical systems and reactions. The surface chemistry is simple and performed under mild conditions, which leads to a high applicability of the presented approach.
Keywords
Introduction
Thousands of protein structures have been determined to date, which fundamentally improved our understanding of biochemical and cellular processes. Yet, most often those structures give only static snapshots and lack information on structural dynamics. Infrared (IR) spectroscopy is a label-free method, which reveals dynamic structural information of biomolecules and proteins including their protonation state, hydrogen bonding and orientation. However, the strong absorption of water in the amide I region (1600–1700 cm−1), which is sensitive to protein conformations, makes IR-studies of proteins in aqueous solutions challenging. To achieve reliable signals by transmission IR spectroscopy, high protein concentrations (>3 mg/ml) in combination with very short optical pathlengths (<10
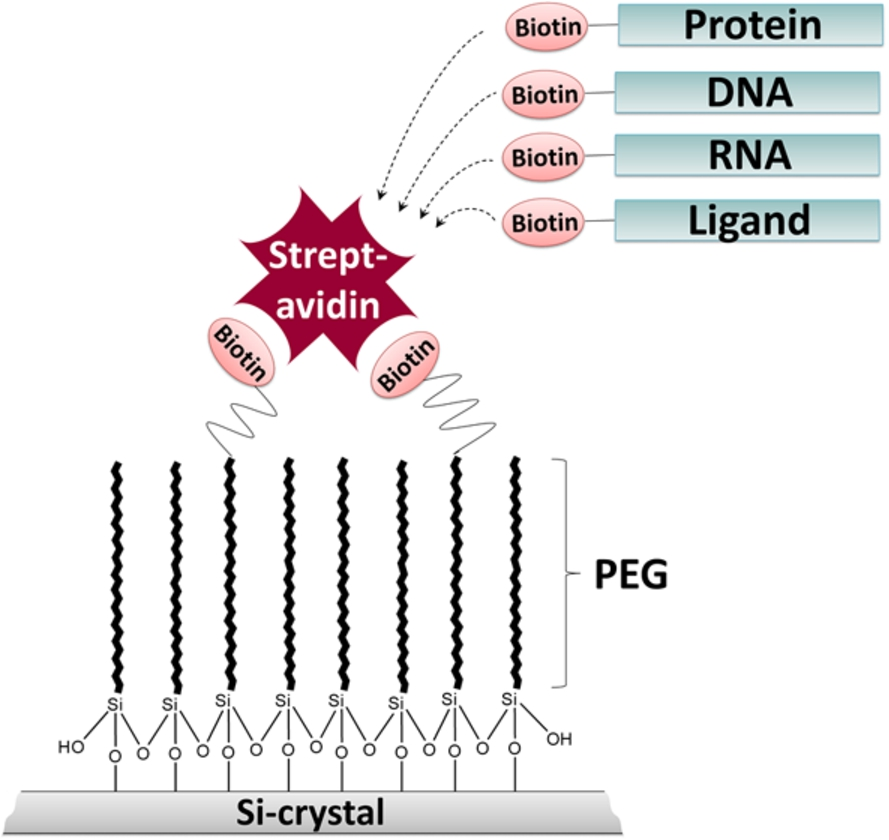
Schematic representation of the ATR-FTIR approach. The silicon (Si) crystal surface is modified with biotin-PEG-silane linkers to prevent unspecific protein adsorption and to enable specific immobilization of biotinylated biomolecules, such as proteins, nucleic acids and ligands via streptavidin.
We developed an ATR-FTIR spectroscopic approach which combines a protein-repellent surface and site-specific immobilization of biomolecules (Fig. 1). We demonstrate that the surface modification with PEG-silane linkers efficiently prevents the adsorption of BSA. By using PEG-linkers modified with biotin at their headgroups, streptavidin can be specifically immobilized at the passivated surface. The tetrameric streptavidin itself can serve as a bridge for the specific immobilization of other biotinylated biomolecules. Here, we demonstrate the efficient immobilization of biotinylated DNA via streptavidin at the ATR crystal. Immobilization strongly increased signal intensities due to higher local DNA concentrations at the surface. The streptavidin-biotin interaction is one of the strongest known non-covalent protein-ligand interactions (Kd value of
ATR-FTIR spectroscopy
ATR-FTIR measurements were performed with a Vertex 70V spectrometer equipped with a BioATR cell II (Bruker). Sample temperature was controlled by an external water bath and set to 20°C. The spectral resolution was 4 cm−1 and each spectrum was an average of 100 scans. The cell was equipped with a multi-reflection silicon crystal. Before each measurement the sample was equilibrated for 20 min.
Surface passivation of the ATR crystal
To modify the crystal with
Adsorption of BSA
The adsorption behavior of BSA was tested by incubating 25
Immobilization of streptavidin
To immobilize streptavidin on the biotinylated surface, 20
Stability of the modified ATR crystal surface
The temperature stability of the modified surface including immobilized streptavidin was analyzed by increasing gradually the temperature of the ATR cell to 45°C, keeping it at this temperature for 10 min and cooling it down to 20°C again. The stability against urea treatment was tested by incubating immobilized streptavidin in 8 M urea for 20 min and washing it afterwards thoroughly with Tris buffer.
Immobilization of biotinylated DNA
Biotinylated double stranded DNA (Sigma: (Btn)5′-CGAGGAACATGTCCCAACATGTTGCTCGAG-3′; 5′-CTCGAGCAACATGTTGGGACATGTTCCTCG-3′) was immobilized via a streptavidin bridge at the biotinylated surface. Therefore, 50 pmol of streptavidin and 100 pmol of DNA were pre-incubated in a total volume of 15
Results and discussion
Surface passivation of the ATR crystal
Protein adsorption is a common phenomenon, which is often accompanied with structural rearrangements of proteins. Small conformational changes of proteins, occurring for example upon binding reactions, can only be studied by ATR-FTIR spectroscopy if the adsorption process to the ATR crystal surface is prevented. To passivate the silicon surface of the ATR crystal, the surface was modified using PEG-silane linkers. Many protocols exist for silanization of surfaces. However, many of them use harsh conditions, which might damage the ATR crystal used as optical internal reflection element or which are not applicable as many crystals are permanently installed in the ATR cell. The method we developed was modified from Kumar et al. [21]. They show the efficient immobilization of silanized nucleic acids on glass slides for the generation of DNA microarrays under mild conditions. We demonstrate that similar mild conditions can be applied to modify the silicon ATR crystal with silanized PEG linkers. While Kumar et al. [21] could only indirectly determine the efficiency of immobilization via fluorescence, we could directly follow all surface modification steps by characteristic vibrational modes in the IR spectrum. First, the surface was washed with a mixture of hydrogen peroxide and sulfuric acid to clean the surface and maximize the number of silanol groups at the surface. We tested silanization at room temperature as well as at 50°C and observed that silanization at higher temperature strongly increased the stability of the silane layer. While the IR bands of the PEG-silane layer produced at room temperature gradually decreased over time and were no longer detectable after 12 h, the spectrum of a silane layer produced at 50°C remained stable over this time period (
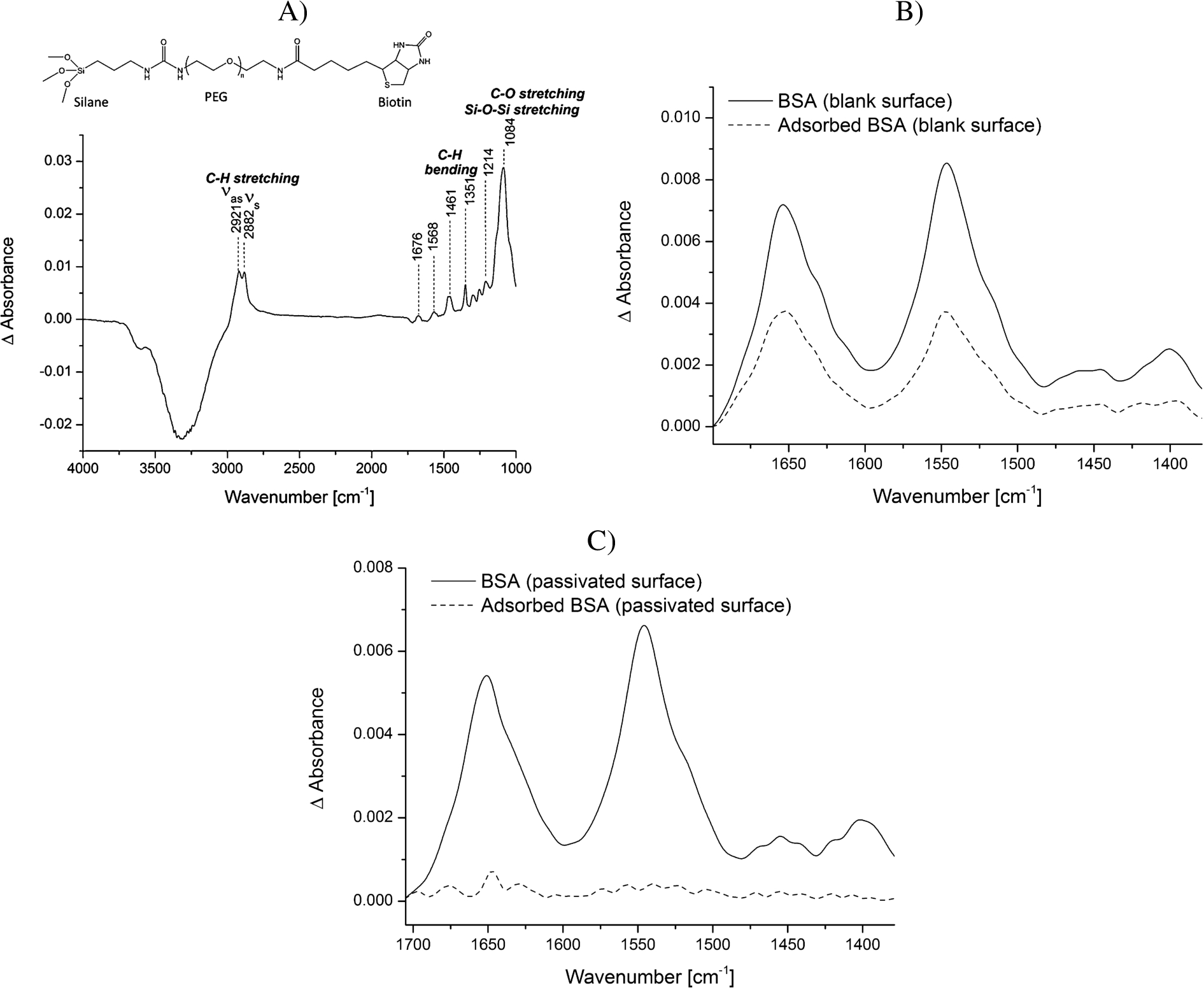
Analysis of the adsorption of BSA to the blank vs. the PEG-silane passivated ATR crystal surface.
By modifying the surface with biotin-PEG-silane groups the biotin headgroups can be used to specifically immobilize streptavidin at the surface. The tetrameric streptavidin itself can then be employed to immobilize specifically other biotinylated biomolecules such as proteins, DNA or RNA at the surface (Fig. 1). This increases the local surface concentration of the studied biomolecule and thereby improves the signal-to-noise ratio while making it possible to use low concentrations (in the low
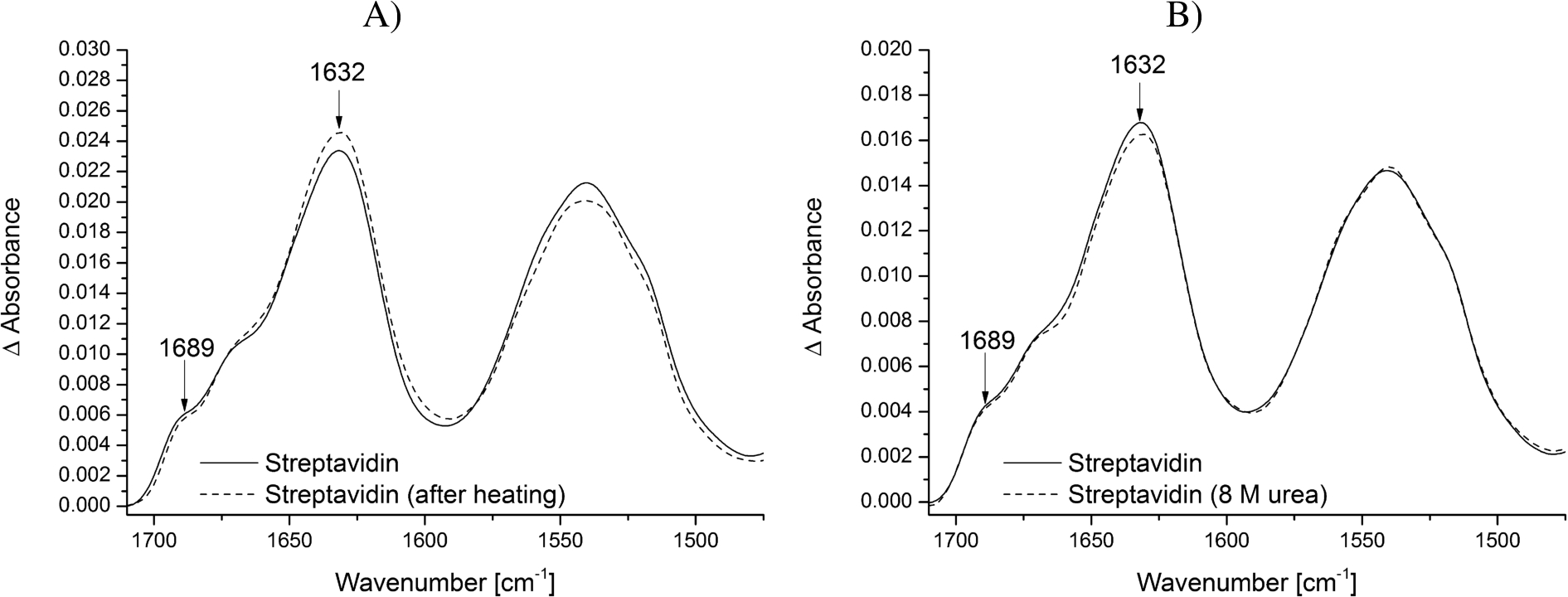
Stability of the modified ATR crystal surface at different temperatures and biochemical conditions. Streptavidin was immobilized on the biotin-PEG-silane modified surface and tested for its stability. The reference spectrum is the PEG-silane modified surface. The typical vibrational modes of
The combination of surface passivation and specific immobilization aims to study small conformational changes of proteins and other biomolecules under native conditions. Only if the modified ATR crystal surface including immobilized streptavidin does not change during the measurements, conclusions about structural changes of the studied biomolecule can be drawn. Therefore, we tested the modified ATR crystal surface including immobilized streptavidin for its stability regarding temperature and denaturing buffer additives. Neither the increase of temperature to 45°C nor the addition of 8 M urea affected the structure and intensity of the immobilized streptavidin (Fig. 3A and B). This is in line with other work showing that the streptavidin-biotin interaction is unaffected by extremes of pH, temperature and denaturants [12,22]. Therefore, it can be assumed that the spectrum of the surface modification including streptavidin does not interfere with the spectrum of the biomolecule of interest, even under harsh conditions.
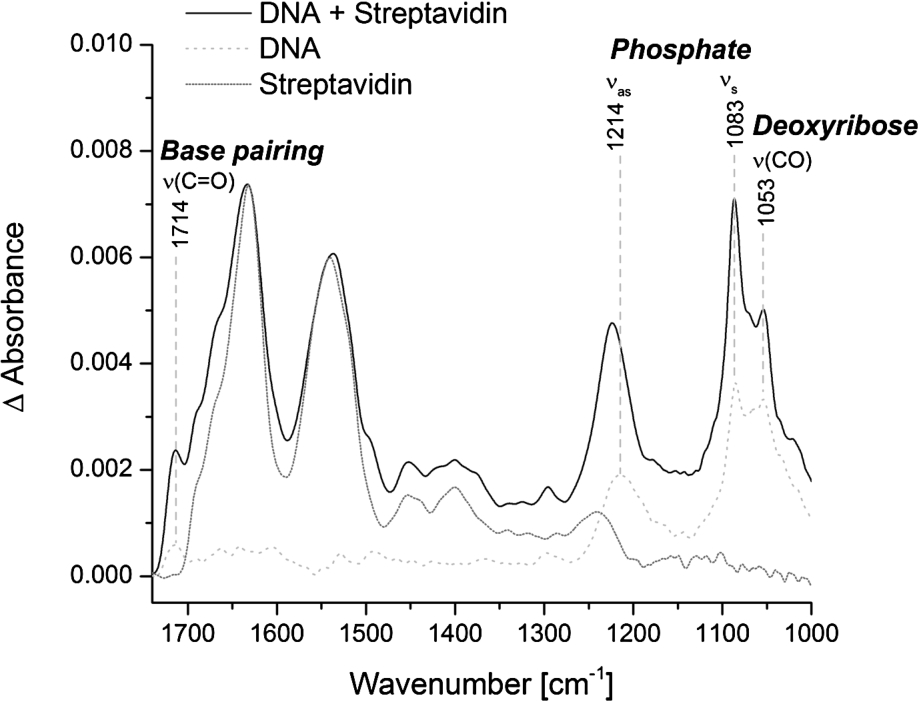
Specific immobilization of biotinylated double-stranded DNA at the ATR crystal. Representative spectra of immobilized streptavidin alone (dotted line) and in combination with biotinylated double-stranded DNA (solid line) are shown. The reference spectrum is the PEG-silane modified surface. The spectrum of streptavidin alone is normalized to the intensity of the amide I band of the combined spectrum of DNA and streptavidin. The spectrum of free DNA was measured on the blank crystal (dashed line). Typical vibrational modes of DNA are highlighted.
The specific immobilization of biomolecules at the ATR crystal increases the local surface concentration and thereby the signal-to-noise ratio. We tested the specific immobilization of biotinylated double-stranded DNA via streptavidin at the PEG modified surface. Surprisingly, the addition of biotinylated DNA to the immobilized streptavidin did not result in a significant DNA signal. Only the pre-incubation of biotinylated DNA with a distinct amount of streptavidin and the subsequent addition to the PEG-biotin modified surface resulted in a distinct and stable DNA signal (Fig. 4). This is probably due to the very strong binding affinity of biotin and streptavidin. As soon as streptavidin alone is added to the PEG-biotin modified surface, all four biotin binding sites of streptavidin are bound to biotin and thereby prevent the additional binding of biotinylated DNA. In contrast, when the biotinylated DNA is pre-incubated with a distinct amount of streptavidin, some biotin-binding sites within the streptavidin tetramer are bound to the biotinylated DNA and some remain free. Those free biotin-binding sites can subsequently bind to the PEG-biotin modified surface and result in the specific immobilization of DNA. The best DNA signal could be achieved by pre-incubating biotinylated DNA and streptavidin tetramers in a 2:1 ratio (Fig. 4). The spectrum of DNA is dominated by the anti-symmetric (1214 cm−1) and symmetric (1083 cm−1) phosphate vibrations. Other DNA characteristic bands are the carbonyl stretching vibration at 1714 cm−1, which is attributed to nucleic bases with strong base pairing and stacking interactions, and the C–O stretching vibration of the backbone at 1053 cm−1 [4,31]. The IR band intensities of immobilized DNA were significantly stronger in comparison to DNA, which was not immobilized, even though the applied concentration of DNA was approximately ten times lower (6.7
Conclusions
The here presented ATR-FTIR approach has many potential biological and biochemical applications and can be used for a variety of biomolecules studied in a range of different conditions. The efficient passivation of the ATR crystal surface with PEG prevents unspecific adsorption of proteins, which is a common problem of surface-sensitive techniques. The direct and specific immobilization of biomolecules via the streptavidin-biotin interaction minimizes the required sample amount and at the same time increases the signal-to-noise ratio. Besides the direct immobilization of biotinylated biomolecules, the method can be adapted to immobilize proteins indirectly via biotin-NTA or biotinylated antibodies. The versatile availability of biotinylated molecules as well as the easy process of biotinylating biomolecules enable the here presented approach to become an universal tool for monitoring a wide spectrum of biochemical processes via ATR-FTIR spectroscopy.
Footnotes
Acknowledgements
This work was supported by the German Research Foundation through the Konstanz Research School Chemical Biology (KoRS-CB), SFB 1214 (A3) and SFB 969 (A2).
