Abstract
BACKGROUND AND AIM:
DCIS is the most common type of non-invasive breast cancer, accounting for about 15 to 30%. Proteome profile is used to detect biomarkers in the tissues of breast cancer patients by mass spectrometry. This study aimed to obtain the expression profile of DCIS proteome, and the expression profile of invasive biomarkers, and finally to introduce a dedicated biomarker panel to facilitate the prognosis and early detection for in situ breast cancer patients.
METHODS AND MATERIALS:
In this study, 10 patients with breast cancer (DCIS) were studied. Benign (marginal) and cancerous tissue samples were obtained from patients for proteomics experiments. Initially, all tissue proteins were extracted using standard methods, and the proteins were separated using two-dimensional electrophoresis. Then, the expression amount of the extracted proteins was determined by ITRAQ. The data were analysed by R software, and gene ontology was utilised for describing the protein in detail.
RESULTS:
30 spots on gel electrophoresis were found in the tumor tissue group (sample), and 15 spots in the margin group (control) with P < 0.05. Healthy and cancerous tissue gels showed that 5 spots had different expression. VWF, MMP9, ITGAM, MPO and PLG protein spots were identified using the site www.ebi.ac.uk/IPI. Finally, protein biomarkers for breast tumor tissue with margin were introduced with the names of P04406, P49915, P05323, P06733, and P02768.
DISCUSSION:
There are 5 critical proteins in inducing cancer pathways especially complement and coagulation cascades. The hall markers of a healthy cell to be cancerous are proliferation, invasion, angiogenesis, and changes in the immune system. Hence, regulation of protein plays a key role in developing recurrence to breast cancer in margins.
Introduction
Breast cancer (BC) is the most common cancer diagnosed among women worldwide, and affects 2.1 million women annually. BC, the leading cause of cancer deaths in women in more than 100 countries, accounted for the death of 684,996 women in 2020, [1]. In Iran, studies were identified as the most common cancer among women [2]. BC is a heterogeneous clinical and genetic disease [3]. According to epidemiological and morphological observations, BC classified as carcinoma in situ and invasive carcinoma [4]. Ductal carcinoma present the most common type in breast cancer, which is invasive with stages [5]. Carcinoma in situ is a clonal proliferation of seemingly malignant cells for which there is no evidence of invasion below the basement membrane in adjacent stromal tissue. It also represents the first detectable cancerous lesion in which tumor cells are confined to the duct surrounded by my epithelial cells and basement membrane [6]. Advances in molecular biology have provided important insights into the relationship between carcinoma in situ and invasive carcinoma: examination of the molecular profile for receptors of estrogen, progesterone and human epidermal growth factor receptor 2 (HER2) shows an increased risk of advancement to invasive carcinoma. Next generation sequencing studies identified nine specific microRNAs that differentiate between DCIS and IDC [7]. Common therapies include surgery, radiotherapy, chemotherapy, and hormone therapy. Surgery is performed as mastectomy or breast preservation. In cases where the breast is preserved, we must use radiotherapy as an adjunct treatment [9]. Therefore, this disease cannot be recognized and treated based on pathology and surgery, radiotherapy, or chemotherapy separately. Proteomic [11], genomic, epigenetic, and metabolomic studies have been performed on a variety of cancers, including BC. Many genes and proteins involved in the disease and how they are altered in the disease process have been identified by these studies [12]. Genomic and proteomic techniques can measure expression profiles and functional interactions in the cells and tissues of various organisms [13], and help us identify disease-causing biomarkers [14]. Therefore, by having a biomarker panel model, molecular transitions and cellular changes in cancer cells can be studied years later [15]. Treatment of cancer is not sufficient if the type of tumor, the nature of its cells and stem cells, the environment around the tumor, cellular and molecular connections between the tumor and its surroundings, the type of treatment, and how to use it are neglected [8]. This study attempts to obtain the proteome expression profile of DCIS tissue, and to study the expression profile of invasive biomarkers, and finally to introduce a specific biomarker panel to help the prognosis and early measures in breast cancer in situ patients in situ.
Material and methods
Sampling
After obtaining the informed consent form, 10 volunteers entered the study for surgery. Then, the relevant surgeon took 150 mg of the breast tumor in situ. Concurrently, 10 tissue samples were taken from their margin of about 2 cm. After rapid transfer inside the liquid nitrogen, a part of the samples was sent to the Proteomics Laboratory of Shahid Beheshti University, and another part was sent to the pathology laboratory for pathological examination.
Experimental group
At first, all participants included in the study from 2020 to 2021 were referred to Shohada hospital of Shahid Beheshti University of Medical Sciences in Tehran. They were examined for the level and type of cancer by an independent pathologist.
Preparation
Protein extraction of Benign and cancerous tissues of patients was frozen under liquid nitrogen. Then, the frozen extracted was powdered completely. The resulting powder with lubricating buffer containing 100 mM Tris-HCl, 3 mM magnesium chloride, 5 mM EDTA and 1 mM.phenyl methyl sulfonyl fluoride (PMSF) and 5 mm beta-mercaptoethanol, 0.5% CHAPS, and 10% glycerol was kept in ice for 30 min. Then, the solution was centrifuged in 16000 rpm at 4 °C for 30 min, and protein assay was performed by Bradford technique [16]. The sample was also taken during the dewatering. After quantification of proteins, the supernatants were kept at −20 °C until used for electrophoresis.
Two-dimensional gel electrophoresis
In each group, 400 μg of the extracted protein was separately mixed with rehydration buffer. The PH = 3 to 10 was applied for IPG strip, and was passively rehydrated with above sample solution overnight at room temperature. Isoelectrofocusing (IEF) was performed by increasing the voltage from 500 to 8000 V during the first 3 h, and then a gradient pattern was used to achieve 8000 V for 3 h. Following the IEF, IPG strips were incubated in equilibration buffer containing 6 M urea, 30% glycerol, 2% SDS, 2% DTT and then alkylated for 20 min in the same buffer with 2.5% iodoacetamide instead of DTT, to separate the second dimension; the treated strips were transferred onto 12% SDS Polyacrylamide slab gel and sealed with 1% agarose. The gels run in 15 W for 30 min as far as the blue front of the bromophenol reaches the end of the gel. The analytical gels were stained with Coomassie blue. The gels were scanned using Bio-Rad Image Scanner and Spot detection, matching, and, quantitative gel analyses were carried out with Nonlinear Progenesis software.
Identification of protein spots
Using protein electrophoresis, the proteins were scanned and the resulted images were compared with gel images in different articles, and matched in molecular weight and isoelectric pH. Then, referring to www.ebi.ac.uk/IPI site, several protein spots were identified.
Extraction of proteins related to breast tumor tissue with a margin
Using ITARQ with powerful techniques and data analysis software R, protein biomarkers with margins were extracted in breast tumor tissues.
Statistical analysis
Scanned 2DE gels were analysed by using Non-linear Prognosis Same Spot software to compare gels together and compare the spots in one statement in gels and get the density of the same spot in each gel. To detect significant differences between the experimental groups, analysis of variance (ANOVAs) was used by SPSS (version 16). A p-value < 0.05 was considered to be statistically significant. Statistics were presented as means ± SE.
Bioinformatics
Identified proteins were used to determine predicted interactions with other proteins. This functional protein association network for each entry was obtained by searching “the string” online database (http://string-db.org). The sub-networks of QQPPI were constructed and visualized by Cytoscape software (Version 3.7.1) [17]. The following parameters were calculated to determine biologically significant nodes. Hub and bottleneck nodes were extracted from the networks in two steps; (first) In the networks, nodes with a degree greater than or equal to the sum of mean and twice the standard Page 3 of 14 deviation (S.D.), i.e., mean C 2*S.D. of the degree distribution, were considered as hubs [18]. (Second) We defined bottlenecks as the proteins that were in the top 5% in terms of betweenness centrality. After all the identified proteins were matched to specific processes or functions by searching the GO in CluGO/Clupedia.
Results
After breast tissue extraction, proteome profile of DCIS was determined and analysed by using Prognosis Same Spots software. The results showed that spots had a statistically significant variation with relative abundance (p < 0.05). As is shown in the Fig. 1, 30 proteins had differentially changed expression (FC > 2). 17 proteins had upregulation and 8 had downregulation. 5 Protein and their biomarkers altered expression of tumor tissue, and margins were investigated by ITRAQ (Table 1). Of these, 1 protein upregulated (ITGAM) and 4 proteins downregulated (VWF, MPO, MMp9, PLG) (Table 2). Changes were calculated by mean and SD standard deviation. A network of protein-protein using common software version 3.4.0 Cytoscape was derived (Fig. 2). Gene ontology analysis counting biological processes, molecular function, and cellular component via Clupedia/CluGO was applied for the four common proteins in the central nodes and the significant differentially expressed proteins (Fig. 3).
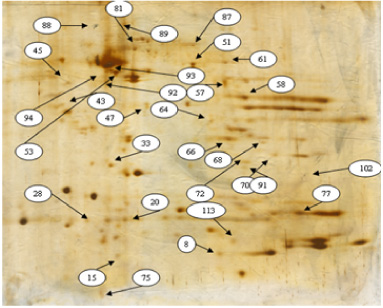
Electrophoresis gel stains have altered expression of breast tumor tissue group.
Five protein and their biomarkers altered expression of tumor tissue and its margin were investigated by ITRAQ
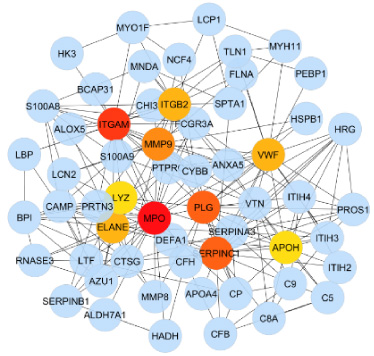
Network interface between tissue proteins in tumor tissue with margin free.
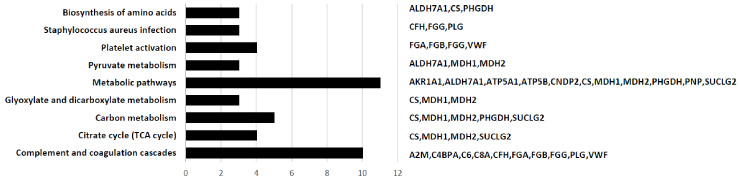
Enriched biological pathways in breast tumor tissue with margin, which were induced by DEPs.
Proteins with altered expression between tumor tissue and its margin
Today, cancer is considered a major health problem in all parts of the world [19]. Among all cancers, breast malignancies have been extensively studied [20]. Every year in the world, more than 1.1 million new cases of breast cancer are detected among women, which is equivalent to 10% of all new cases of cancer and 23% of all cancers in women [70]. Over the past few decades, the prevalence of breast cancer has increased in Iran, and about 24.6% of all cancers in the Iranian population are related to this type of cancer [71]. If the diagnosis of the disease in the first stage can be increased from 25% to 75%, the mortality rate will reduce to 50%. The overall 5-year survival rate of breast cancer is 90–95% in the early stages and limited to the organ, and 30% in the advanced stages [21].
It is very important to study the expression of proteome in patients with breast cancer, since proteome mediates all biological processes and molecular targets of most drugs. In addition, proteome integrates cell genetic information and affects environment [22]. Using proteomics technology in the study of cancer and the identification of proteomes in healthy and cancerous tissues, many researchers have found appropriate biochemical markers to diagnose the degree, type and extent of disease damage [23].
According to WHO, 80% of local recurrence of breast cancer occurs in the same area as the previous lesion; therefore, marginal screening is of importance. Bed tissue proteome in breast cancer is a potential biomarker source for early detection of cancer recurrence [24]. At present, despite effective methods such as ITRAQ, steps can be taken to identify the diagnostic markers of breast cancer recurrence in order to adopt the most efficient treatment methods before the disease progress and relapse [25,26]. Since it is the first study to examine DCIS and compare it with margin by proteomics in humans, we were not able to compare the results with previous studies [27].
In our study, 5 differentially expressed proteins of tumour’s tissue and margin were identified using ITRAQ technique. Of the 5 proteins in samples, 1 protein underwent upregulation and 4 proteins were downregulated.
In 2015, a study was performed on the rat cancer proteome, in which 41 different proteomes from cancerous and normal rat tissue showed a commonality in changes in tumour molecular transitions and healthy tissue around the tumour, which is consistent with the results of our study [15]. Another study by Jones who performed a metabolomics study on serum profile and diagnosis in the early stages of breast cancer in a murine model, and introduced 18 metabolites as a metabolic panel that was associated with breast cancer [28]. In other words, the difference in tumorigenic proteomes indicates that the regulation of a large number of proteins has changed. Similarity in proteomic expression of tumour and margin group can be useful for obtaining proteomic markers for early detection of recurrence of cancer. Some approaches were adopted for describing proteome expression differentiating breast cancer tissues [29]. Simultaneous modification of many proteomes in this disease may indicate biochemical mechanisms involved, and may be beneficial for effective drug purposes in the treatment process [30]. Since novel diagnoses are mostly based on proteomic identification, changes in protein expression can be very helpful in determining the biomarkers of the disease [31].
On the other hand, this study reviewed the pathogenesis of breast cancer using proteomic method to determine and identify changes in the expression of breast protein and normal tissue. To date, several approaches have been developed to describe the differential features of proteomic expression between primary and recurrent breast cancer tissues [32]. Pan et al. compared expression in a chemotherapy-sensitive and resistant tissue using quantitative proteomics through ICAT [33]. In our study, 5 key proteins from VWF, MPO, MMp9, ITGAM, PLG families were identified according to the results of Gene Ontology analysis.
A ligand and a key protein in cellular biological pathways, VWF (von Willebrand Factor Protein), is one of the differentially expressed proteins in this study. The patients with ILC have a higher VWF expression than patients with invasive ductal carcinoma of IDC [34]. VWF has been shown that it plays an important role in the development and metastasis of tumours [35]. Plasma VWF levels have also been reported higher in a variety of cancers, including breast, bladder, prostate, and ovary cancers, compared with benign and controlled disease [36]. In particular, a relationship has been seen between increased VWF concentration and higher tumour grade in breast cancer. Thus, VWF could potentially be a biological indicator of recurrence [37]. Elevated plasma VWF in cancer patients is thought to originate from platelet-activated endothelial cells [38–41]. VWF may regulate cancer spread through a number of pathways, including angiogenesis and hyper coagulopathy [42–44]. A study (2021) reported a relationship between VWF upregulation and breast cancer [45].
In this study, plg protein was identified by downregulating a biomarker for breast cancer studies. This protein plays a key role in regulating the physiology and pathology of tissue, coagulation system, immune responses, angiogenesis, and cell invasion [46].
Myeloperoxidase (MPO) is an enzyme secreted by neutrophil granulocytes as a result of phagocytosis during inflammation [47]. In colorectal cancer, it has been shown that tumour penetration by MPO-expressing cells is independently associated with a favourable prognosis [48]. MPO may play a role in the treatment of BC, in part, because of its prominent role in breast physiology, primarily as an antibacterial enzyme to protect breast milk during lactation [49,50].
MPO is produce due to steroids hormones. Testosterone and tamoxifen bind to the estrogen receptor (ER), thereby reducing dimerization and binding to DNA sites, where it absorbs them as a central activator depending on the tissue or promoter complex. Estrogen inhibits the MPO gene instead of activating, and tamoxifen may absorb activating complexes in the ER, activating MPO gene transcription. Functional polymorphism in MPO leads to higher transcription in G allele carriers and binding site in G/A 463 allele. The G allele is much more expressed in human macrophages and monocytes than the A allele, leading to higher levels of MPO expression [50–54].
A study on 928 breast cancer samples in 2019 reported a relationship between MPO with breast cancer as a biomarker to improve overall survival in human breast cancer [55]. Other studies on mice have shown that mpo stimulates breast tumour growth and lung metastases [56].
The matrix metalloproteinase are a family of zinc-dependent proteases. MMP-2 and MMP-9 are the main members of the MMP family [57,58]. Some studies have shown that MMP-2 and MMP-9 play a major role in the destruction of extracellular matrices and the enhancement of invasion and tumor metastasis [59,60]. Another study found that TBBPA can induce cancer cell metastasis with releasing MMP-9 through ROS-dependent MAPK and AKT pathways in MCF-7 cells [61]. A further study reported overexpression of MMP-9 in breast cancer [62]. Another study on 80 patients with breast cancer showed an increase in upregulation of MMP-9 in breast cancer cell lines [63].
All of these proteins enrich the complement and coagulation cascades pathway, indicating its importance in pharmacotherapy and targeted treatment of breast cancer patients subject to recurrence.
The relationship between cancer and coagulation dates back to 1865 year [64]. The underlying mechanism by which coagulation factors induce tumor cells growth, invasion, metastasis, and angiogenesis is a hot topic in cancer research [65]. Extrinsic and intrinsic coagulation cascade factors play a complex and important role in the progression of cancer by developing blood clots. The intrinsic cascade occurs when the blood contacts with the surface of endothelial cells, and the external cascade works when the factor X is activated in response to the release of TF at the site of injury. TF and factor V11a contribute to develop external cascade and possibly cancer [66]. Blood coagulation cascade can be activated in cancer patients by different mechanisms and at different levels [67]. The importance of this pathway in relation to cancer was investigated in a study on ovarian cancer [68]. Another study in 2020 reported an association between this pathway and the factors involved in DCIS [69].
Conclusion
In inducing cancer pathways, especially the complement and coagulation cascades, 5 critical proteins plat a major role. By studying this pathway, hall markers of cancer such as proliferation, invasion, angiogenesis, and changes in the immune system are well found [70]. It can be concluded that regulating the expression of proteins plays a key role in developing recurrence to breast cancer in margins.
Footnotes
Limitations
It is clear that more research is needed. Further studies should also be performed on a larger community of patients with DCIS in this geographical area.
Compliance with ethical standards
All procedures performed in the study involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. Also, ethics committee of Azad University of Medical Sciences, Iran approved this study by (Code no: IR.IAU.CHALUS.REC.1400.008).
Informed consent
An informed consent was obtained from each individual participating in the study.
Conflict of interest
The authors declare no conflict of interest.
Funding/Support
This study was funded by Chalous Islamic Azad University.
Authors’ contributions
All authors had equal role in design, work, statistical analysis, and manuscript writing.
