Abstract
Long non-coding RNAs (lncRNAs) influence pathetiology of breast cancer. Besides, VDR and ESR1 signaling pathways are two important pathways in this malignancy. In the present mixed bioinformatics and expression assay study, we have identified lncRNAs that are co-expressed with VDR and ESR1 in breast cancer tissues and analyzed their expression in 42 paired breast cancer and non-cancerous specimens. Expression of SLC16A-AS1 was significantly lower in breast cancer tissues compared with paired non-cancerous samples (expression ratio = 0.27, P value < 0.001). Similarly, LINC00900 was down-regulated in cancer tissues compared with non-cancerous ones (expression ratio = 0.26, P value = 0.01). There were no significant differences in the expressions of VDR and AATBC between these two sets of samples. Expression levels of VDR and AATBC were associated with histological grade (P values = 0.02 and 0.03, respectively). Moreover, expression of VDR was associated with tumor size (P value = 0.02). Finally, expression levels of SLC16A-AS1 were associated with first pregnancy age (P value = 0.006). In brief, the results of current study further support involvement of VDR and ESR1-associated lncRNAs in breast cancer.
Introduction
As the most common cancer, female breast cancer affects 2.3 million cases, based on Global Cancer Statistics. Besides, it is the fifth major source of cancer death [1]. This malignancy is associated with high mortality and morbidity rates necessitating recognition of molecular pathways participating in its pathogenesis to design proper therapeutic modalities as well as diagnostic markers. Vitamin D receptor (VDR) signaling and estrogen receptor (ER) are two important participants in the pathoetiology of breast cancer. Over-expression of VDR and ER-α gene increases risk of this malignancy [2]. The role of estrogen-related receptor α (ERRα)/VDR axis in enhancement of estrogen signaling has been confirmed in this type of cancer. Moreover, overexpression of VDR/CYP24A1/ERRα has been correlated with poor clinical outcome in a certain type of breast cancer [3]. Another study has reported association between VDR over-expression in the nucleus of breast cancer cells and favorable prognostic parameters and lower possibility of breast cancer mortality [4]. VDR has also been suggested as a target for treatment of breast cancer [5]. Calcitriol and inecalcitol have been found to suppress growth of breast cancer cells, particularly those expressing ER and VDR [5]. Another study in breast cancer cells has shown regulation of VDR expression through estrogen-associated activation of the ERK1/2 signaling [6]. We have also validated participation of VDR signaling and a number of VDR-associated long non-coding RNAs (lncRNAs) in the pathoetiology of breast cancer in Iranian patients [7–10]. We have also introduced a combined in silico and literature based strategy for recognition of lncRNAs that influence VDR signaling in breast cancer [11]. We aimed at identification of the role of some VDR- and ESR-related lncRNAs in breast cancer. Based on the importance of VDR and ESR signaling in the pathogenesis of breast cancer, in the present mixed bioinformatics and expression assay study, we have identified lncRNAs that are co-expressed with VDR and ESR1 in breast cancer tissues and analyzed their expression in 42 paired breast cancer and non-tumoral tissues. Among the identified lncRNAs is LINC00900 which is a Toll-like receptor (TLR)-related lncRNA whose expression has been correlated with survival of patients with esophageal cancer [12]. SLC16A1-AS1 has a role in induction of metabolic reprogramming. Moreover, this lncRNA has been shown to be a target and co-activator of E2F1 [13]. However, functions of other lncRNAs have not been elaborated.
Materials and methods
Selection of genes
First, lncRNAs that are co-expressed with VDR and ESR1 in breast cancer tissues were identified using Co-LncRNA tool (http://www.bio-bigdata.com/CoLncRNA/). Then, lncRNAs that show co-expression with both VDR and ESR1 was selected manually. Afterwards, expression pattern of selected lncRNA in breast cancer tissues was assessed using TANRIC (http://ibl.mdanderson.org/tanric/_design/basic/index.html) and those with differential expression were selected. In the next step, genomic changes of lncRNAs in breast cancer samples were assessed by means of the cBioPortal for Cancer Genomics (http://www.cbioportal.org) and the Catalogue Of Somatic Mutations In Cancer (COSMIC) and those with genomic changes were selected. In the final step, interactions between selected lncRNAs and oncogenes, tumor suppressors and miRNAs were assessed using AnnoLnc and miRcode tools. Based on last step results, VDR, AATBC, SLC16A-AS1 and LINC00900 were selected for further study.
Patients
Expression levels of VDR, AATBC, SLC16A-AS1 and LINC00900 were measured in paired tumoral and non-tumoral samples obtained from 42 Iranian patients with breast cancer. Non-tumoral samples from the same patients were considered as control samples. Tissues were obtained from Farmanieh and Sina centers, Tehran, Iran in the period of 2017–2020. The study protocol was confirmed by the ethical committee of Shahid Beheshti University of Medical Science. Informed consent forms were signed by patients. Breast samples were obtained prior to conduction of any chemo/radiotherapy.
Experiments
Frozen tumoral and non-tumoral tissues were chopped into small pieces. RNA was retrieved from tissues using the RiboEx kit (GeneAll, Seoul, South Korea). ExcelRT TM Reverse Transcription Kit II (SMOBIO, Taiwan) was used for making cDNA. Expression of VDR, AATBC, SLC16A-AS1 and LINC00900 was measured in all paired samples in the ABI step one plus PCR system. Expressions of VDR, AATBC, SLC16A-AS1 and LINC00900 were normalized to B2M. RealQ Plus 2x PCR Master Mix (Ampliqon, Odense, Denmark) was used for preparation of reactions. Primers sequences are demonstrated in Table 1.
Primer sequences
Primer sequences
Fold changes of expression levels of VDR, AATBC, SLC16A-AS1 and LINC00900 in tumoral tissues versus non-tumoral tissues were calculated using the efficiency corrected model. Differences in the expression of VDR, AATBC, SLC16A-AS1 and LINC00900 transcripts between paired samples were assessed using Student’s paired t test. Associations between clinical/demographic data and transcript levels of VDR, AATBC, SLC16A-AS1 and LINC00900 genes were appraised using chi square test. The difference between mean values of expression of VDR, AATBC, SLC16A-AS1 and LINC00900 gene between distinct subgroups of patients was appraised using Tukey’s honest significance test. Pairwise correlations between expression levels of VDR, AATBC, SLC16A-AS1 and LINC00900 were calculated using the regression model. P values less than 0.05 were considered as significant. Receiver operating characteristic (ROC) curves were plotted to assess the properness of gene expression levels for differentiating tumoral tissues from non-tumoral ones. Sensitivity, specificity and AUC values were calculated.
Results
General data of patients are shown in Table 2.
General data of patients
General data of patients
Figure 1 Depicts expression of VDR, AATBC, SLC16A-AS1 and LINC00900 transcripts in tumoral and non-tumoral samples of breast cancer patients.
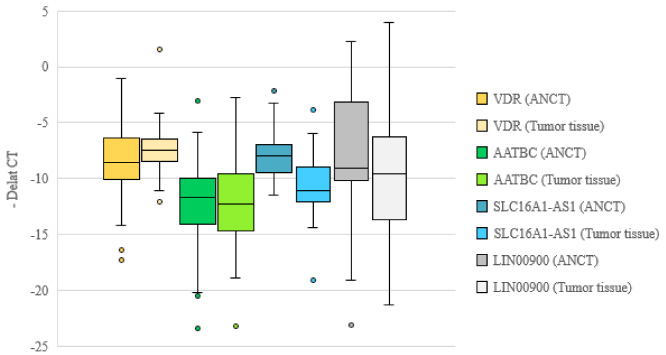
Relative expression of genes in tumoral and non-tumoral tissues as revealed by -delta CT values for each gene (CT reference gene – CT target gene) in each set of specimens.
Expression of SLC16A-AS1 was significantly lower in breast cancer tissues compared with paired non-cancerous samples (expression ratio = 0.27, P value < 0.001). Similarly, LINC00900 was down-regulated in cancer tissues compared with non-cancerous ones (expression ratio = 0.26, P value = 0.01). There were no significant differences in the expressions of VDR and AATBC between these two sets of samples (Table 3).
Relative expression of selected genes in breast cancer samples versus non-cancerous samples
Expression levels of all assumed pairs of genes were correlated with each other in both sets of samples except for VDR/AATBC and VDR/SLC16A-AS1 in non-tumoral samples (Table 4).
Correlations between expression levels of genes in tumoral and non-tumoral tissues (∗: Correlation is significant at P < 0.05 level)
Expression levels of VDR and AATBC were associated with histological grade (P values = 0.02 and 0.03, respectively). Moreover, expression of VDR was associated with tumor size (P value = 0.02). Finally, expression levels of SLC16A-AS1 were associated with first pregnancy age (P value = 0.006) (Table 5).
Associations between relative expression of VDR, AATBC, SLC16A-AS1 and LINC00900 genes and medical data of patients
Expression of AATBC was higher in ER positive samples compared with ER negative ones (P value = 0.01). Moreover, expression of this lncRNA was higher in HER2 positive samples compared with HER2 negative ones (P value = 0.01). Finally, its expression was higher in patients that breastfed their children less than 5 months compared with those with more than 60 months history of breastfeeding (Table 6).
Association between expression of genes in tumors and clinical data
Expression levels of SLC16A-AS1 and LIC00900 could differentiate tumoral tissues from non-tumoral one with AUC values of 0.77 and 0.66, respectively. Both lncRNAs had high sensitivity values (96.7 and 94.1, respectively). Table 7 and Fig. 2 show details of ROC curve analysis.
Detailed parameters of ROC curves
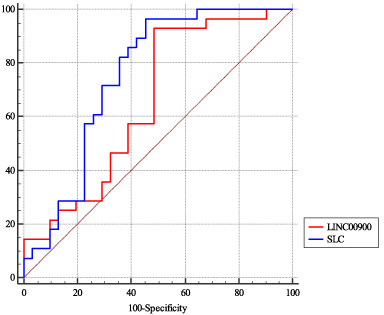
ROC curves presenting diagnostic value of SLC16A-AS1 and LIC00900 in breast cancer.
We measured expression of lncRNAs that are co-expressed with ESR1 and VDR in breast cancer. Notably, expression levels of SLC16A-AS1 and LINC00900 were lower in cancer tissues compared with non-cancerous ones. A single study in bladder cancer has indicated the role of SLC16A1-AS1 in induction of metabolic reprogramming. Moreover, this lncRNA has been shown to be a target and co-activator of E2F1 [13]. This lncRNA is transcribed from the antisense strand of SLC16A1, a gene whose over-expression predicts poor prognosis in urological malignancies [14]. However, the function of SLC16A-AS1 in regulation of expression of the sense transcript is not known.
LINC00900 has been shown to be a TLR-related lncRNA whose expression has been correlated with survival of patients with esophageal cancer [12]. Other studies have shown association between these pattern recognition receptors and evolution of breast cancer. Therefore, TLR targeting has been suggested as a possible method for treatment of breast cancer [15]. Thus, in addition to influencing VDR and ESR signaling, LINC00900 can affect breast cancer pathogenesis through influencing TLRs.
There were no significant differences in the expressions of VDR and AATBC between these two sets of samples. Since previous studies have reported alterations in VDR levels in breast cancer tissues [2], further assessment of VDR protein levels is needed to validate our results.
Expression levels of all assumed pairs of genes were correlated with each other in both sets of samples except for VDR/AATBC and VDR/SLC16A-AS1 in non-tumoral samples. This finding shows that the correlation between these genes is influenced by the presence of cancer.
Expression levels of VDR and AATBC were associated with histological grade. Moreover, expression of VDR was associated with tumor size. Finally, expression levels of SLC16A-AS1 were associated with first pregnancy age. These findings imply that lncRNAs which are co-expressed with VDR and ESR participate in the pathogenesis of breast cancer.
Expression levels of SLC16A-AS1 and LIC00900 could differentiate tumoral tissues from non-tumoral one with AUC values of 0.77 and 0.66, respectively. Both lncRNAs had high sensitivity values, but low specificity. Thus, they might be incorporated in screening panels.
Expression of AATBC was significantly higher in ER positive samples compared with ER negative ones as well as in HER2 positive samples compared with HER2 negative ones. Thus, it might have interactions with breast cancer markers.
In brief, the results of current study further support involvement of VDR and ESR1-associated lncRNAs in breast cancer. Since reproductive and lifestyle factors can affect expression of genes, we have assessed expression of genes in paired samples from patients to control these factors. Taken together, dysregulation of VDR and ESR1-associated lncRNAs might be an important event in the course of breast cancer. The study has a limitation regarding the sample size.
