Abstract
BACKGROUND:
Accumulating evidence highlights that long noncoding RNA (lncRNA) DSCAM-AS1 play a key regulatory role in different stages of cancer development and progression. This study aimed to investigate whether the expression of DSCAM-AS1 is deregulated in breast cancer.
MATERIALS AND METHODS:
The relative expression of DSCAM-AS1 was measured in fifty breast cancerous and matched adjacent non-neoplastic tissue samples using quantitative real-time polymerase chain reaction (qPCR) technique. The association between DSCAM-AS1 expression and patients’ clinicopathological features was evaluated. Sensitivity and specificity of the DSCAM-AS1 expression for diagnosing breast cancer was obtained by the receiver operating characteristic (ROC) curve analysis.
RESULTS:
Our results showed that the expression of DSCAM-AS1 was significantly up-regulated in breast cancerous tissues compared with the matched adjacent non-neoplastic tissues (P < 0.05). Furthermore, we observed a significant association between the DSCAM-AS1 expression and lymph node metastasis (P = 0.011) but no other clinicopathological characteristics (P > 0.05). ROC curve analysis resulted in an area under the curve (AUC) of 0.67 and showed that the DSCAM-AS1 expression level may discriminate cancerous and non-cancerous tissues with 68% sensitivity and 76% specificity.
CONCLUSION:
This study provides further evidence that the expression of DSCAM-AS1 is deregulated in breast cancer and highlights its potential in breast cancer development.
Introduction
Breast cancer is the most frequently diagnosed life-threatening malignancy in women worldwide, and associates with nearly 12.3 % lifetime risk of developing the disease and 80–90% five-year relative survival rate [1–3]. Despite remarkable advances in our understanding of the essence of cancer, early diagnosis and the treatment strategies over the past decade, breast cancer is still the second leading cause of cancer-related deaths among women in developed countries [1,2]. In 2019, an estimated 268,600 new cases of breast cancer were expected to be diagnosed, and 41,760 deaths to be occurred from the disease in the United State, according to the annual statistics reporting from the American Cancer Society [4]. It is a complicated disease caused by interplay between a multitude of genetic, lifestyle and environmental risk factors [5,6]. Although a large body of studies has been previously conducted to elucidate the underlying molecular mechanisms involved in etiopathogenesis of breast cancer, many questions still remain unanswered.
Long non-coding RNAs (lncRNAs), a novel and highly diverse class of cellular regulatory RNAs, are arbitrarily defined by two key characteristics: length longer than 200 nucleotides and lacking protein coding potential [7,8]. The human genome consists of more than 15,000 lncRNA genes based on GENCODE reference human genome annotation and more and more new lncRNAs are coming into the light [9]. They are involved in a wide range of biological processes such as cell signaling pathways, cell growth and proliferation, regulation of cell cycle, cell differentiation and development, programmed cell death and DNA repair, as well as about all processes of gene expression, ranging from transcription to post-translation through interacting with DNA, RNA, and proteins [10–17]. Not surprisingly, dysregulated expression of lncRNAs may contribute to development and progression of human malignancies. Down Syndrome Cell Adhesion Molecule-antisense RNA 1 (DSCAM-AS1), an estrogen receptor 𝛼 (ER𝛼)-dependent lncRNA, lies in chromosome 21q22.2, is transcribed from the antisense strand of DSCAM gene belonging to the immunoglobulin superfamily of cell adhesion molecules [18,19]. Recently, accumulating shreds of evidence suggest that dysregulated expression of DSCAM-AS1 can be involved in migration, invasion, metastasis and angiogenesis of various types of cancers including ovarian cancer [20], colorectal cancer [21,22], cervical cancer [23], melanoma [24], non-small cell lung cancer [25], hepatocellular carcinoma [26] and breast cancer [19,27,28]. However, the mechanism by which DSCAM-AS1 mediates development and progression of cancer is not yet completely understood. This opens a new avenue for future research to dwell into using DSCAM-AS1 as a potential therapeutic target.
Here, we posed a question as to whether the expression of DSCAM-AS1 is dysregulated in patients with breast cancer. To this end, we evaluated the expression of DSCAM-AS1 in breast cancerous and matched adjacent non-neoplastic tissue samples using quantitative real-time polymerase chain reaction (qPCR) technique. Also, the possible association between expression of DSCAM-AS1 and clinicopathological characteristics was assessed.
Materials and methods
Sample preparation
In the current study, paired cancerous tissue and adjacent non-neoplastic samples were obtained from fifty patients with breast cancer (mean age ± SD = 47.83 ± 8.14 year, range 34–71) who underwent surgical resection at the Nour-nejat hospital, Tabriz, Iran. All tissue samples were immediately frozen in liquid nitrogen after excision during the surgical procedure and stored in −80 °C until RNA extraction. To determine histological grade, stage of disease, tumor size and lymph node metastasis of all resected samples, histopathological tissue analysis was performed by pathologist. Patients who received any neoadjuvant therapy such as chemotherapy and/or radiotherapy before surgery were excluded from the study. The present study was approved by the Ethics and Human Rights Committee of the University of Tabriz and informed consent was obtained from all participants before enrolment.
RNA extraction and cDNA synthesis
The total RNA was extracted from breast cancerous and matched adjacent non-neoplastic tissue samples using Tri-Pure® Isolation Reagent kit (Cat No. 11667165001, Roche Applied Science, Germany) according to the manufacturer’s instructions. To eliminate DNA contamination, the isolated total RNA was treated with RNase-free DNase I (Cat No; EN0521, Thermo Fisher Scientific, USA) according to the manufacturer’s protocol. The quantity and integrity of extracted RNA were evaluated using NanoDrop® ND-1000 Spectrophotometer (Thermo Fisher Scientific, USA) and agarose gel electrophoresis, respectively. Subsequently, 1 μL of total RNA was reverse-transcribed into cDNA using PrimeScript RT reagent Kit (Takara Bio Inc., Shiga, Japan) with random hexamer primer following the manufacturer’s protocol.
Quantitative real-time polymerase chain reaction (qPCR)
To evaluate the relative expression of DSCAM-AS1 gene at mRNA level, we performed qPCR on StepOnePlusTM Real-Time PCR System (Applied Biosystems). QPCR amplification was carried out in triplicate format using Syber Green-I dye (AccuPower® 2X GreenStarTM qPCR Master Mix, Bioneer, South Korea) and gene-specific primers including F: 5 ′ -CGAAACCCCATCTCAAAACC-3 ′ and R: 5′-GTATTCAAGGGACAGTCACAG-3 ′ for DSCAM-AS1, and, F: 5 ′ -GAGAAGTATGACAACAGCCTC-3 ′ and R: 5 ′ -TGAGTCCTTCCACGATACC-3 ′ for GAPDH gene as an internal control. A 20 μL PCR reaction mixture contained 100 ng cDNA, master mix 2X, ROX dye 50X, and 10 pmol of each primer pairs for DSCAM-AS1 and GAPDH genes. Thermocycling program consisted of initial denaturation step at 95 °C for 15 min, followed by 40 cycles of denaturation at 95 °C for 15 sec, annealing at 61 °C for 15 sec and extension at 72 °C for 20 sec. No template control (NTC) was used as negative control. To normalize the variability of the relative expression of DSCAM-AS1 gene, the expression of the housekeeping gene (GAPDH) was used.
Statistical analysis
Statistical analysis was performed with SPSS Version 19.0 statistic software package (SPSS Inc., Chicago, IL, USA). The continuous variables were expressed as mean ± standard deviation (SD). The expression of DSCAM-AS1 gene between breast cancerous and matched adjacent non-neoplastic tissue samples was compared by student’s t-test. The association between DSCAM-AS1 gene expression and clinicopathological characteristics was evaluated using Fisher’s exact test. Receiver operating characteristic (ROC) curve analysis was used to evaluate the biomarker potency of the DSCAM-AS1 expression for breast cancer diagnosis. P values less than 0.05 was considered statistically significant.
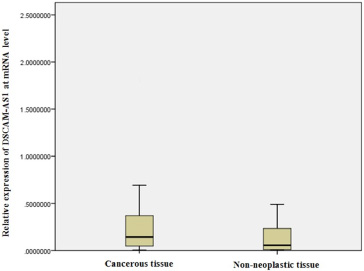
Box plot depicting the relative expression of DSCAM-AS1 in the breast cancerous and the matched adjacent non-neoplastic tissue samples.
The clinicopathological characteristics of the patients with breast cancer and their association with the expression of lncRNA DSCAM-AS1
Expression of DSCAM-AS1 in breast cancerous and matched adjacent non-neoplastic tissue samples
The expression of DSCAM-AS1 gene in breast cancerous and matched adjacent non-neoplastic tissue samples was quantified. Our results showed that the lncRNA ASCAM-AS1 expression level was significantly increased in the cancerous tissues compared with the matched adjacent non-neoplastic tissues (P = 0.029) (Fig. 1).
Association between DSCAM-AS1 expression and clinicopathological parameters
The clinicopathological characteristics of the patients were represented in the Table 1. We evaluated the association between these variables and the lncRNA DSCAM-AS1 expression level. There was no association between the DSCAM-AS1 expression and the tumor size, histological stages, as well as the age of the patients (Table 1). However, the mean of DSCAM-AS1 gene expression was higher in the patients with histological grade III compared with the patients having histological grade I and II, but the difference was not statistically significant (P > 0.05) (Table 1). Also, we found that the mean of DSCAM-AS1 expression in the patient with lymph node metastasis was significantly higher than in the patients without lymph node metastasis (P = 0.011) (Table 1). Considering the molecular subtypes, 94% of the samples were luminal A/luminal B, so in the present study we report association of the expression level of DSCAM-AS1 with the luminal type breast cancer.
Biomarker potency of the lncRNA DSCAM-AS1
To understand whether the lncRNA DSCAM-AS1 expression level may play as a biomarker for the breast cancer development, we implemented ROC curve analysis. ROC curve analysis resulted in an area under the curve (AUC) of 0.67. Moreover, it discriminated the breast cancerous and non-neoplastic tissues from each other with a sensitivity of 68% and a specificity of 76% (95% CI, 0.8120 to 0.9717, P < 0.0001) (Fig. 2).
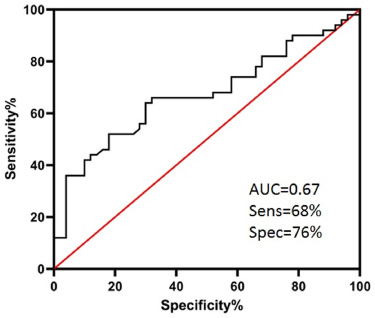
Receiver operating characteristic (ROC) curve analysis of lncRNA DSCAM-AS1 expression between breast cancerous and matched adjacent non-neoplastic tissue samples. AUC; area under the curve.
An increasing number of papers in the literature have reported that dysregulated expression of DSCAM-AS1 have been implicated in various types of human malignancy [19–28]. In this study, we investigated the expression pattern of DSCAM-AS1 in fifty breast cancerous and matched adjacent non-neoplastic tissue samples and evaluated the association between its expression and clinicopathological characteristics. Our findings demonstrate that DSCAM-AS1 was significantly overexpressed in breast cancerous tissues compared with the adjacent non-tumor tissues (P < 0.05) (Fig. 1). The mRNA level of DSCAM-AS1 was higher in the high-grade and high-stage cancerous tissues than in low-grade and low-stage specimens, but the differences were not statistically significant (P > 0.05), however, we found a significant increase of its expression in the metastatic tissues compared with the non-metastatic specimens (Table 1). The ROC curve analysis showed that the lncRNA DSCAM-AS1 expression level may discriminate cancerous and non-cancerous samples, implying its potential as a biomarker for the breast cancer development (Fig. 2). In line with our findings, more recently, Khorshidi et al. reported that the expression of DSCAM-AS1 was significantly up-regulated in invasive ductal breast carcinoma tissues compared with the matched adjacent non-neoplastic tissues [27]. In contrast to our results, they observed that the expression of DSCAM-AS1 was significantly higher in patients aged less than 55 compared with older patients, but, they did not observe any association with other clinicopathological characteristics [27]. Liang et al. reported its up-regulation in the breast cancer tissues [19]. Liang et al. showed that lncRNA DSCAM-AS1 plays a fundamental role in proliferation and invasion of breast cancer by reducing miR-204-5p and enhancing ribonucleotide reductase M2 (RRM2) expression [19]. Sun et al. showed that lncRNA DSCAM-AS1 was highly expressed in luminal and Her-2 overexpressing breast cancers, and its knockdown in two luminal breast cancer cell lines (MCF-7 and T47D) using siRNA, significantly inhibited cell proliferation and colony formation in both cell lines [29]. They demonstrated that lncRNA DSCAM-AS1 induces cell proliferation and colony formation in MCF-7 and T47D cell lines through promoting the G1/S transition [29]. They observed no association between clinicopathological characteristics and the DSCAM-AS1 expression level, but, most importantly, in a retrospectively registered clinical analysis they identified that the high expression of DSCAM-AS1 was a poor prognostic factor in patients with the luminal breast cancer treated with endocrine therapy [29]. Ma et al. reported that DSCAM-AS1 acts as a competing endogenous RNA of miR-137 and regulates epidermal growth factor receptor pathway substrate 8 (EPS8) to promote cell reproduction and suppresses cell apoptosis in Tamoxifen resistant breast cancer [30]. Xu et al. observed that knockdown of DSCAM-AS1 may result in inhibited cell proliferation, increased apoptosis and inhibited cell cycle progression [31]. Considering DSCAM-AS1 expression level in 42 breast tumor samples, Miano et al. reported a significant association with luminal breast cancer but no significant association with the other clinicopathological data [32]. To some extent, our results confirmed these findings, because, the significant higher expression level that we obtained here was correlated with the luminal A and luminal B molecular subtypes, though the lower numbers of HER-2 (n = 2) and triple negative (n = 1) samples limited a conclusive finding.
Dysregulation of DSCAM-AS1 has been reported in different cancer types, including colorectal cancer, non-small cell lung carcinoma, ovarian cancer, melanoma and hepatocellular carcinoma [21–26]. For example, Liang et al. showed that it was highly expressed in cervical cancer tissues, and associated with enhanced cell proliferation, migration and invasion [23]. They clarified that DSCAM-AS1, as an oncogenic lncRNA, is implicated in tumorigenesis of cervical cancer by targeting miR-877-5p/ATXN7L3 axis [23]. The study of Pei demonstrated DSCAM was significantly up-regulated in CRC samples and promoted the cell proliferation and metastasis by reducing the expression level of miR-144-5p in CRC [21]. It therefore appears that up-regulated expression of DSCAM-AS1 might promote proliferation and thus contribute to the transformed phenotype.
In conclusion, our study provides further evidence that the lncRNA DSCAM-AS1 is up-regulated in breast cancer and may contribute to the pathobiology of breast cancer. This work supports the results of others and highlights the potential of DSCAM as a biomarker for cancer development. Analysis of its plasma levels for establishing as a breast cancer biomarker is suggested as future direction.
Footnotes
Acknowledgements
This study was supported by the research and technology deputy of the University of Tabriz. Authors would like to express their sincerest appreciation to all subjects for participating in this study.
Conflict of interest
There are no potential conflicts of interest.
