Abstract
Keywords
Schizophrenia occurs in individuals with a genetic predisposition who encounter environmental factors that trigger the onset of the disorder [1,2]. It has been proposed that gene and environment interactions causes changes in central nervous system (CNS) gene expression, resulting in the underlying pathophysiology [3]. Thus, such changes can only be investigated in tissue from individuals diagnosed as having schizophrenia. This notion has resulted in numerous studies of the human transcriptome in post-mortem CNS tissue from subjects with schizophrenia; these have shown that schizophrenia is a complex disease affecting the expression of genes involved in glutamatergic neurotransmission [4–6], protein turnover [7], regulation of presynaptic function [4,8], G-protein signalling [4,9], GABAergic function [10], metabolism [11], mitochondrial function [7,12], myelination [13,14], as well as neurite outgrowth, cytoskeletal proteins and synapse plasticity [7].
Whilst existing microarray studies have significantly advanced our understanding of the pathophysiology of schizophrenia, they have not accommodated data from neuroimaging and clinical studies suggesting schizophrenia is a progressive disorder. We postulated there could be temporally related changes in gene expression in the CNS of subjects with schizophrenia, and this was supported by our findings on two genes, muscleblind protein 1 and protocadherin 17, where increased levels of mRNA for these two genes were present in the pre-frontal cortex from subjects with schizophrenia with a short (< 7 years) but not long (> 22 years) duration of illness (DOI) [14]. To further explore the hypothesis, we recently performed a genome-wide microarray study in the dorsolateral prefrontal cortex (Brodmann's area 46; BA 46) from subjects with schizophrenia with varying durations of illness (DOI) [15]. This study showed differences in gene expression profiles at different illness stages, with the greatest changes occurring in the short DOI subjects compared to control. Similarly, in a study looking at genes associated with glycobiology, the highest number of differentially expressed genes occurred in short DOI subjects [16]. Significantly, one transcript shown to be decreased in short, but not intermediate or long, DOI schizophrenia was phospholipase C β1 (PLCβ1) [15]. PLCβ1 is the rate-limiting enzyme in the generation of inositol 1,4,5-trisphosphate (IP3) and diacylglycerol (DAG) from phosphatidylinositol 4,5-bisphosphate (IP2) and is important in postnatal-cortical development and plasticity [17–20]. Therefore changes in the enzyme could have profound effects on CNS function. PLCβ1 is also a crucial step in the signalling of several neurotransmitters implicated in schizophrenia [18,21–28]. Thus, changes in PLCβ1 expression could be involved in the genesis of some symptoms of schizophrenia. To begin addressing this proposal we extended our microarray study, using focused methodologies (real-time PCR and western blotting), to measure PLCβ1 mRNA and protein in BA 46 from larger cohorts of subjects with schizophrenia with different durations of illness.
Materials and methods
Human post-mortem tissue collection
Consent was obtained from the Ethics Committee of the Victorian Institute of Forensic Medicine and the North Western Mental Health Program Behavioural and Psychiatric Research and Ethics Committee. Tissue was obtained from the Victorian branch of the National Neural Tissue Resource Centre. Psychiatric diagnoses were made according to DSM-IV criteria [29] by consensus after extensive case history reviews using the Diagnostic Instrument of Brain Studies (DIBS) [30].
Tissue was excised from BA 46, according to Brodmann's criteria, of the left hemisphere of 15 subjects with schizophrenia with short DOI (SDOI), < 7 years, and 15 subjects with long DOI (LDOI), > 22 years, along with 15 non-psychiatric controls for both SDOI and LDOI groups matched as closely as possible for age, sex, post-mortem interval (PMI) and brain pH (see Table 1).
Demographic data for cohorts
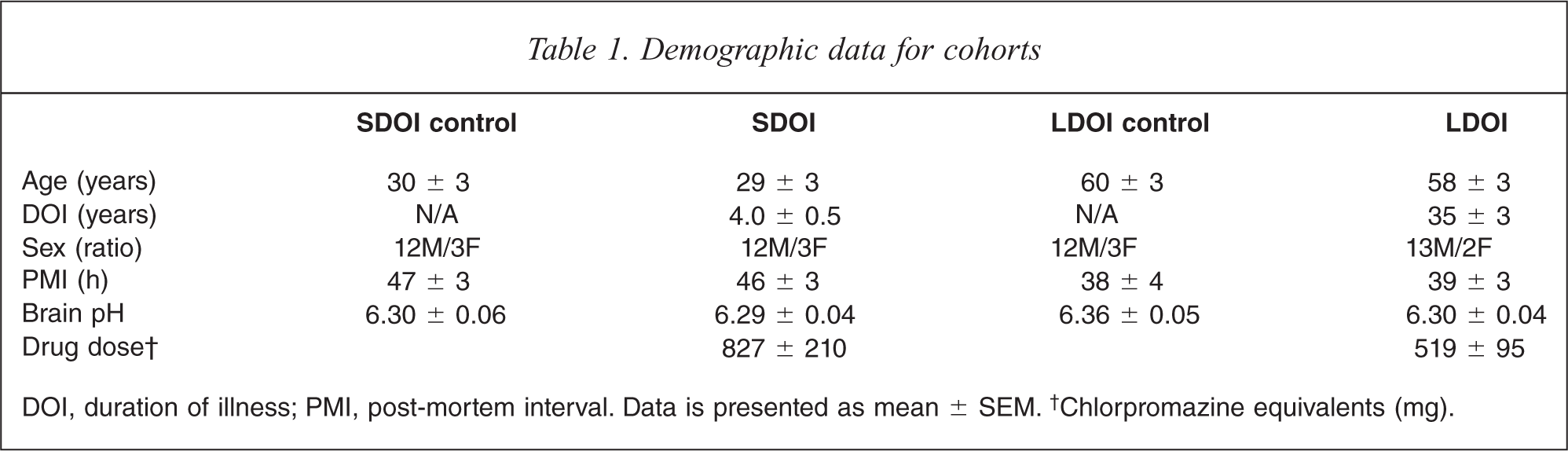

DOI, duration of illness; PMI, post-mortem interval. Data is presented as mean ± SEM.† Chlorpromazine equivalents (mg).
RNA purification and first-strand cDNA synthesis
Total RNA was isolated from 100 mg frozen tissue using 1 mL TRIzol® reagent (Invitrogen, Carlsbad, CA), according to the manufacturer's instructions. The RNA was treated with DNAse I (Applied Biosystems, Scoresby, Victoria) at 37°C for 25–30 min, then purified by phenol/chloroform extraction and stored at −80°C. RNA quantity and quality were determined by spectrophotometer readings and RNA integrity numbers (RINs). DNA contamination was checked by PCR using primers specific for genomic DNA.
First strand cDNA was synthesized from 2 μg RNA using 100 units M-MLV-RT (Applied Biosystems) with 2.5 μM random decamers and 2.5 μM oligo dT primers (Applied Biosystems), 0.5 mM of each dNTP and 20 units RNase inhibitor in 1× RT Buffer (50 mM Tris-HCL, pH 8.3, 75 mM KCl, 3 mM MgCl2, 5 mM DTT) in a final volume of 20 μL. The reaction was incubated at 44°C for 1 h then inactivated at 92°C and the product stored at −20°C.
Real-time PCR assay
cDNA was used as a template for real-time PCR performed with SYBR green detection using a Bio-Rad iQ5 Real-Time PCR Detection System (Bio-Rad, Hercules, CA). Reactions were performed in 25 μL volume containing cDNA diluted 1:125, 0.4 nM primers (Table 2) and 1× IQ SYBR green supermix (Bio-Rad), with cycling conditions of 95°C for 3 min, 40 cycles of 30s each at 95°C, 57°C and 72°C, followed by a melt curve. Amplification levels of target gene were normalized to the reference gene Cyclophilin, which did not differ between subjects with schizophrenia and matched controls (4.99 e−6 versus 4.51 e−6, P = 0.75 for SDOI and 1.69 e−6 versus 1.48 e−6, P = 0.71 for LDOI). qPCR data was acquired using IQ5 optical system 2.0 software (Bio-Rad). The identities of the amplicons were confirmed by sequencing. Relative expression was determined using the Pfaffl method [31]. Reaction efficiencies were calculated from standard curves run on each plate, constructed from a 10-fold dilution series of the purified amplicon of each primer pair, and found to be between 85–110%.
Primer sequences for real-time PCR

F, forward primer; R, reverse primer.
Western blotting
Homogenates were prepared from BA 46 at 5% w/v in 10 mM Tris, (pH 7.4), containing 1 % w/v sodium dodecyl sulphate and 1 mM fresh sodium orthovanadate (Sigma-Aldrich, St Louis, MO). Five μg of total protein, determined by colorimetric protein assay, was loaded per lane and separated using polyacrylamide gel electrophoresis (4% stacking gel, 6% running gel; 150 V constant), then transferred (100 V constant, 1 h) onto Hybond-ECL nitrocellulose membrane (GE Health Life Sciences, North Shore, NSW). Nitrocelluloses were blocked in Tris-buffered saline with 0.1% Tween-20 (TTBS) for 1 h at room temperature, then incubated with mouse anti-PLCβ1 antibody (BD Biosciences, 1/2500 in TTBS), for 1 h in an orbital incubator at 30 rpm, 26°C. Equal protein loading and transfer was confirmed by staining with 0.1% ponceau S in 3% trichloroacetic acid. Nitrocelluloses were washed and incubated with HRP conjugated goat anti-mouse IgG (BD Biosciences; 1/4000 in TTBS), for 1 h at 30 rpm, 26°C, before final washes in TTBS. Antigenic bands were visualized using SuperSignal West Pico Chemiluminescent Substrate (Thermo Scientific, San Francisco, CA) and the image captured with a Kodak 440 CF imaging system. To control for inter-blot variation, an internal control (IC) membrane preparation was run in 12 wells on two gels to establish both intra- and inter-blot variation for the measurement of PLCβ1 protein. This internal control was subsequently run (in duplicate) on every gel. Gels were imaged so that the optical density of the internal control fell within the range established by the mean ± 1 SD of the initial measurements. These internal control values were used to standardize the data by expressing it as ratio to internal control.
The bands of interest were identified by molecular weight and their presence/absence in cortical samples from SV129/C57Bl6 (wild type) and PLCβ1 KO mice [21] (see Figure 1).
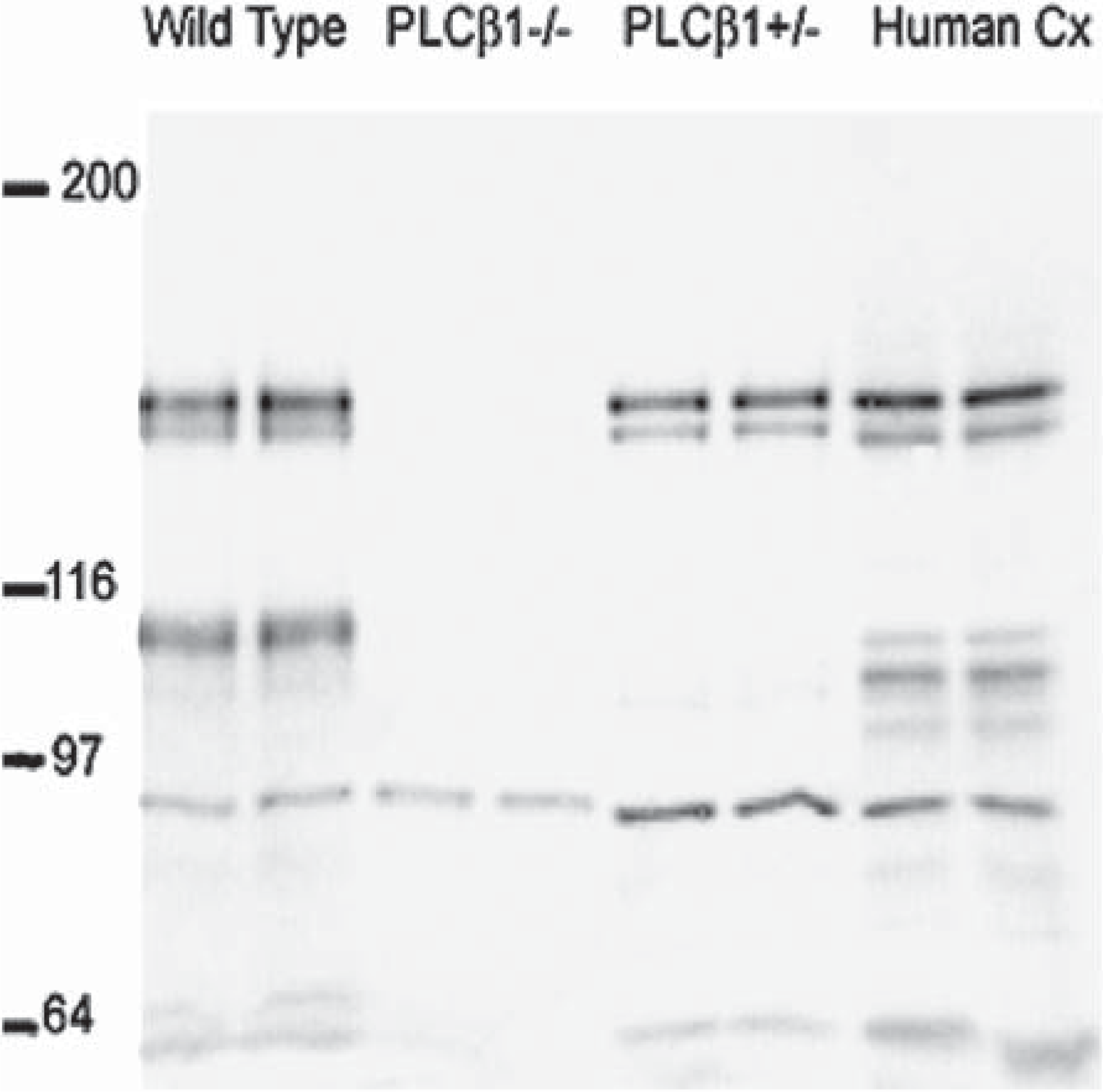
Western blot generated using the anti-PLCβ1 antibody. Five μg of cortical brain homogenates were run in duplicate, as follows, from the left, SV129/C57Bl6 (wild type, WT), PLCβ1 knockout mice, PLCβ1 heterozygous mice and human cortex.
Statistical analysis
Statistical outliers were identified using Grubb's test and the normality of the data distribution assessed with D'Agostino and Pearson omnibus normality tests. Two-way ANOVA with Bonferroni post-tests were employed to identify variance in levels of PLCβ1 protein and mRNA associated with diagnosis and DOI. Correlations between expression levels and potential confounds (age, PMI, brain pH and chlorpromazine equivalents) were assessed by Pearson correlation, with an r2 value of over 0.5 indicating a significant correlation [32]. All analyses were performed using GraphPad Prism version 5.01 (GraphPad Software, San Diego, CA).
Results
Demographics
Two-way ANOVAs showed no significant variance associated with brain pH (F1,56 = 0.04, P = 0.84) or DOI (F1,56 = 1.42, P = 0.24) across the diagnostic groups (Table 1). There was a variation in PMI with DOI (F1,56 = 5.07, P = 0.03) due to SDOI subjects and their matched controls having a higher PMI (46.55 ± 2.2 h) than LDOI subjects and their matched controls (38.52 ± 2.7 h), but there was no variance with diagnosis (F1,56 = 0.01, P = 0.93). As expected from the selection criteria, there was variation in age with DOI (F1,56 = 97.14, P < 0.0001); subjects with SDOI and their matched controls were significantly younger (30 ± 2 years) than those with LDOI and their controls (59 ± 2 years), although there was no variance with diagnosis (F1,56 = 0.11, P = 0.74). Final recorded drug doses (chlorpromazine equivalents) were not different between subjects with SDOI (827 ± 210) and LDOI (519 ± 95, P = 0.193).
Real-time PCR
Grubb's tests on the normalized levels of PLCβ1 mRNA revealed a number of outliers: PLCβ1a expression; subject 9 (SDOI control) and subject 14 (LDOI schizophrenia), PLCβ1b expression; subject 13 (SDOI schizophrenia) and subject 14 (LDOI schizophrenia). Analyses were performed with and without the outliers; graphs are presented minus outliers. Relative PLCβ1a and b expression in LDOI schizophrenia did not meet the criteria for the D'Agostino and Pearson omnibus normality test, however following exclusion of the outliers all data sets met normality criteria.
PLCβ1a
Including outliers, PLCβ1a expression varied with diagnosis (F1,56 = 7.01, P = 0.011) and DOI (F1,56 = 10.87, P = 0.002), with no interaction between variables (F1,56 = 0.55, P = 0.46). The variance with diagnosis was due to significantly decreased levels of mRNA in tissue from subjects with SDOI schizophrenia (0.022 ± 0.003 ratio to CYPA) compared to their controls (0.036 ± 0.015 ratio to CYPA, P = 0.011). PLCβ1a mRNA levels were also lower in SDOI (0.022 ± 0.003) than LDOI schizophrenia (0.038 ± 0.005, P = 0.012).
Without outliers there was still variance with diagnosis (F1,54 = 10.65, P = 0.002) and DOI (F1,54 = 14.26, P = 0.0004; Figure 2A), but no interaction (F1,54 = 0.01, P = 0.93). PLCβ1a mRNA levels in SDOI schizophrenia were decreased compared to matched controls (0.022 ± 0.003 versus 0.033 ± 0.003, P = 0.02; Figure 2A) and compared to LDOI schizophrenia (0.022 ± 0.003 versus 0.035 ± 0.004, P = 0.02). PLCβ1a expression was also decreased in LDOI schizophrenia compared to matched controls (0.035 ± 0.004 versus 0.046 ± 0.003, P = 0.04; Figure 2B), as well as in SDOI controls compared to LDOI controls (0.033 ± 0.003 versus 0.046 ± 0.003, P < 0.01).
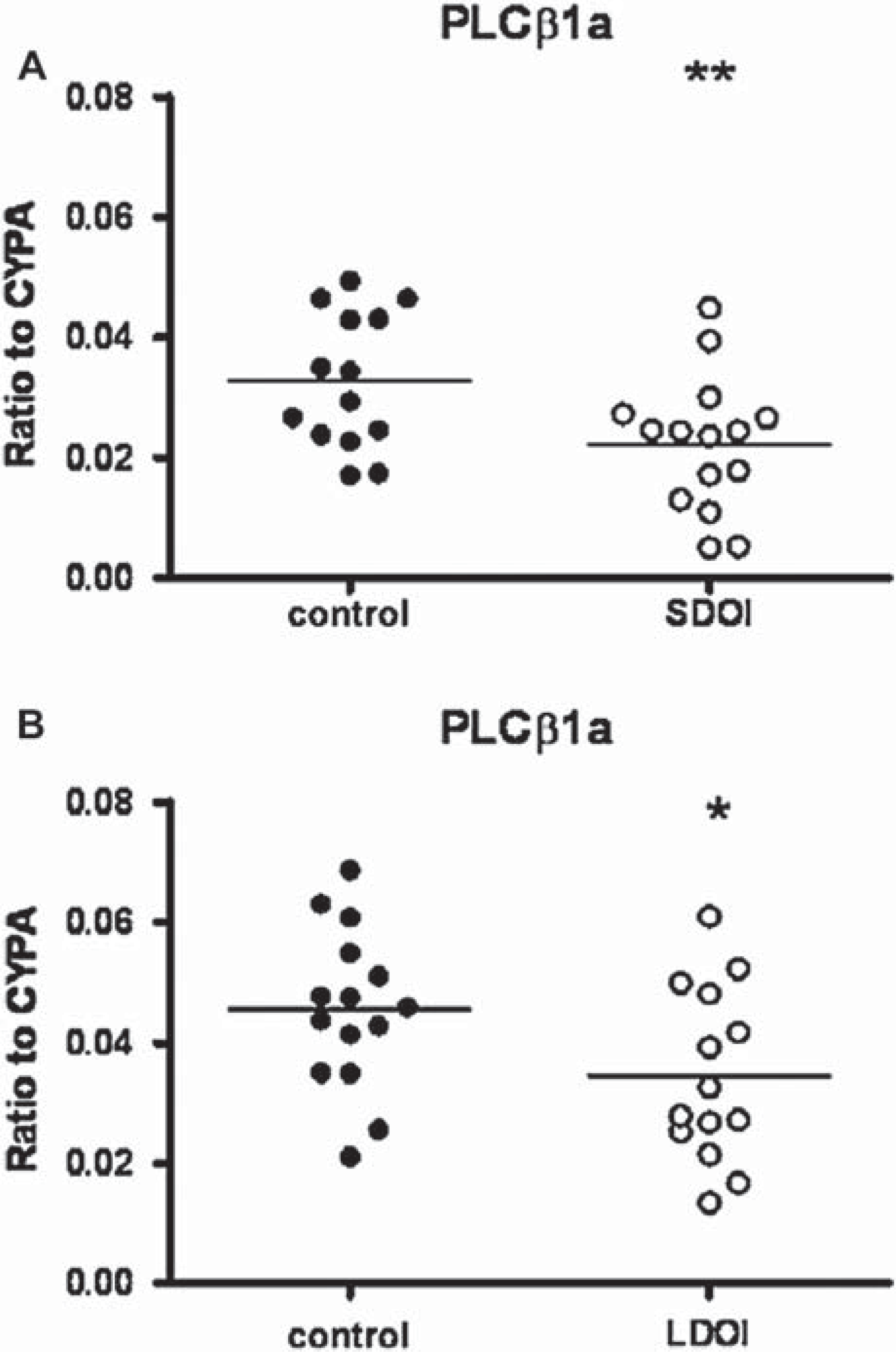
Relative mRNA expression of PLCβ1a in BA 46 from 15 subjects with schizophrenia (•) of (A) short duration of illness (SDOI) or (B) long DOI (LDOI) and 15 age/sex matched control subjects (○), normalized to house-keeping gene cyclophilin A (CYPA). Outliers were removed from the graph. ∗∗P < 0.01, ∗P < 0.05 compared to control subjects.
No strong correlations were found between PLCβ1a expression levels and age, PMI, brain pH or chlorpromazine equivalent (r2 ranged from 0.01 to 0.12). The effect of gender was not assessed due to the low number of female subjects.
PLCβ1b
PLCβ1b mRNA expression varied with DOI (F1,56 = 4.04, P = 0.049) but not diagnosis (F1,56 = 0.0 P = 0.97), with no interaction between variables (F1,54 = 1.12. P = 0.3). This variation was due to decreased PLCβ1b mRNA expression in SDOI subjects (0.0017 ± 0.0002) compared to their controls (0.003 ± 0.0003, P = 0.003).
With the outliers removed, PLCβ1b mRNA expression showed variation with diagnosis (F1,54 = 9.32, P = 0.0035) as well as DOI (F1,54 = 12.43, P = 0.0009), with no interaction between the variables (F = 0.28. P = 0.6; Figure 3A). The variation with diagnosis was due to decreased PLCβ1b mRNA in SDOI schizophrenia (0.0015 ± 0.0002) compared to matched controls (0.003 ± 0.0003, P = 0.0008). There was no difference in levels of mRNA in LDOI schizophrenia compared to their controls (0.0032 ± 0.0005 versus 0.0043 ± 0.0005, P = 0.163; Figure 3B). The variation with DOI was because of decreased mRNA levels in SDOI compared to LDOI schizophrenia (0.0015 ± 0.0002 versus 0.0032 ± 0.0005, P = 0.0083) and in SDOI controls compared to LDOI controls (0.003 ± 0.0003 versus 0.0043 ± 0.0005, P = 0.042).
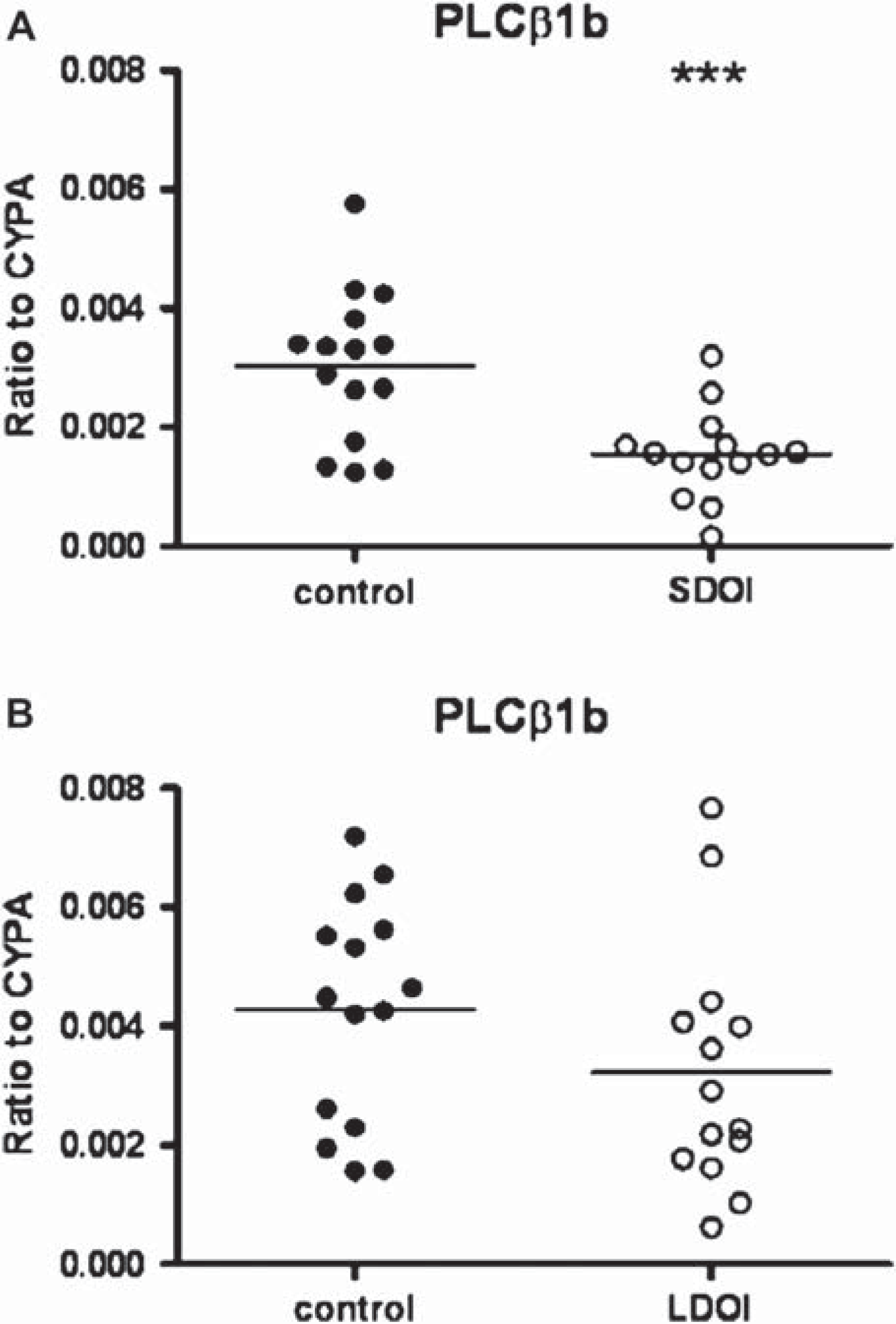
Relative mRNA expression of PLCβ1b in BA 46 from 15 subjects with schizophrenia (•) of (A) short duration of illness (SDOI) or (B) long DOI (LDOI) and 15 age/sex matched control subjects (○), normalized to house-keeping gene cyclophilin A (CYPA). Outliers were removed from the graph. ∗∗∗P < 0.001 compared to control subjects.
PLCβ1b expression levels showed no strong correlation with PMI, brain pH or chlorpromazine equivalent (r2 from 0.01 to 0.08) but showed a weak correlation with age (r2 = 0.56, P = < 0.0001; Figure 4).
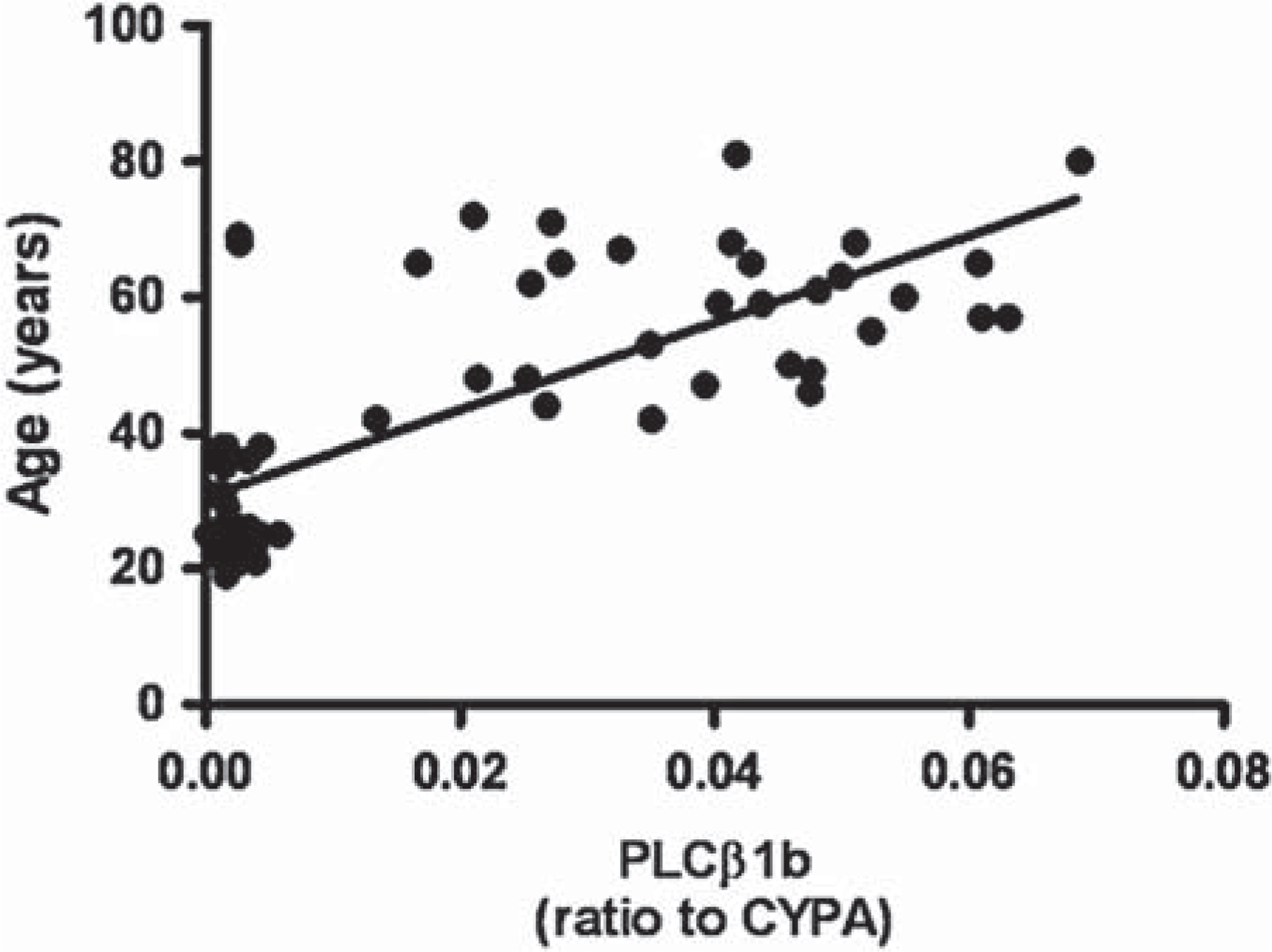
Correlation of relative PLCβ1b mRNA expression normalized to house-keeping gene cyclophilin A (CYPA) with age of the subject.
Western blot analysis
Two bands detected by western blot were selected for analysis of PLCβ1 expression. The 150kDa band corresponded to the PLCβ1a variant while the 140kDa band corresponded to PLCβ1b. Grubb's test on the data revealed subject 2 of SDOI schizophrenia as an outlier for PLCβ1a expression. Data was analysed with and without the outlier and is represented in the graph without the outlier. Removal of this data point resulted in all data sets meeting the criteria for the D'Agostino and Pearson omnibus normality test.
No variance was seen in PLCβ1a protein levels with diagnosis (F1,56 = 0.04, P = 0.84) or DOI (F1,56 = 0.02, P = 0.89), with no interaction between the variables (F1,56 = 0.31, P = 0.58; Figure 5). Without the outlier there was no variance with diagnosis (F1,55 = 0.41, P = 0.52) or DOI (F1,5 5= 0.07, P = 0.79), and no interaction (F1,5 = 0.03, P = 0.86; See Figure 5A).
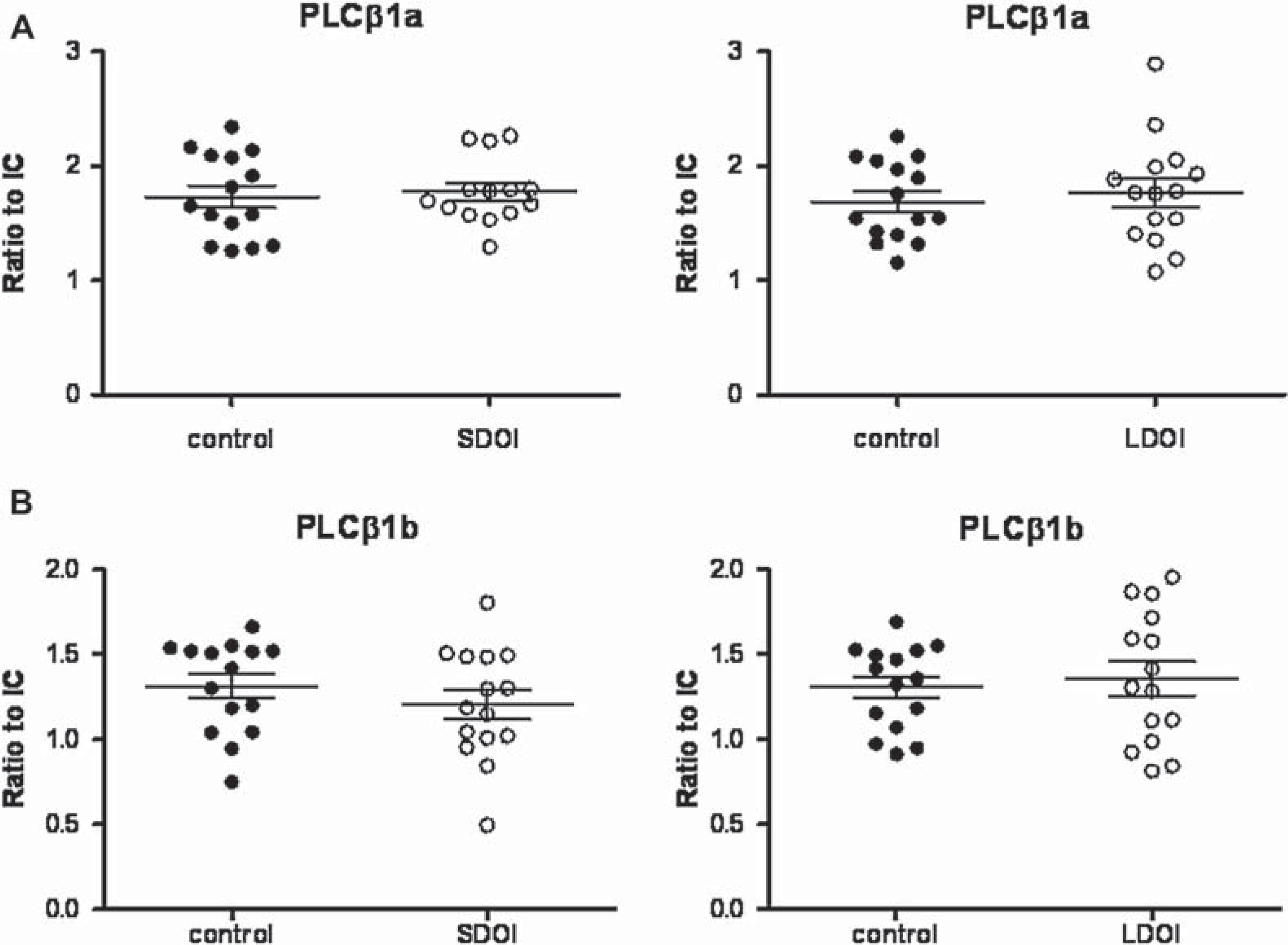
Expression of PLCβ1a (A) and PLCβ1b (B) in BA 46 from 15 subjects with schizophrenia (•) of short duration of illness (DOI; left) or long DOI schizophrenia (right) and 15 age/sex matched control subjects (○), expressed as ratio to internal control (IC). Outlier was removed from the graph.
Similarly there was no significant variation in PLCβ1b protein levels with diagnosis (F1,56 = 0.12, P = 0.73) or DOI (F1,56 = 0.82, P = 0.37) and no interaction between the variables (F1,56 = 0.95, P = 0.33; Figure 5B). There were no strong relationships between levels of PLCβ1a or b expression and possible confounding factors (r2 from 0.00 to 0.11).
Discussion
As in our previous microarray study [15], this study shows a significant decrease in levels of PLCβ1 mRNA expression in BA 46 from subjects with schizophrenia and SDOI. However, this study is the first to measure PLCβ1 variants individually; reporting a 39% decrease in PLCβ1a mRNA in subjects with SDOI and a 24% decrease in subjects with LDOI. By contrast, the level of PLCβ1b mRNA was decreased in subjects with SDOI by 50% but not changed in subjects with LDOI.
We failed to find changes in protein levels of either PLCβ1 variant in BA46 with schizophrenia, highlighting problems in interpreting changes at the level of the transcriptome in isolation. PLCβ1 is involved in the signalling of a number of G-protein coupled receptors, regulating cardiovascular function [33,34] as well as development of neuronal plasticity [35], and thus is tightly regulated by several feedback mechanisms [36]. Therefore, regulation of the signalling in the CNS may be occurring at the level of the receptor rather than effector, with the lack of change in PLCβ1 protein reflecting compensation for changes in the signalling pathway, an ability absent in single cells in culture [37] and possibly rodent CNS [37–39] for this protein. Such mechanisms could protect cells in the human CNS from the potential adverse effects of prolonged changes in PLCβ1 activity, as this protein is involved in various pathways. Previous investigations of signalling proteins in post-mortem CNS tissue from patients with schizophrenia showed no change in protein Kinase C levels in the temporal lobe [40], despite a decrease in [3H]phorbol ester binding specific to this region [41], whilst changes in protein kinase glycogen synthase kinse-3 (GSK-3) were shown to be region and hemisphere specific [42–44], suggesting changes in signalling pathways in schizophrenia to be complex.
Although there was no change in protein levels, it is possible the change in mRNA expression has a functional effect. For example, protein synthesis is dependent upon mRNA availability, therefore, a change in mRNA levels could affect a neuron's ability to rapidly respond to stimuli, particularly if the mRNA is localized to cellular compartments such as in the dendrites, where proteins synthesis has been shown to occur within minutes of synaptic activation [45]. Whilst the detected difference between the expression of the two transcripts could reflect differences in their stability due to varying length and sequence, another possibility is differences in the mechanisms controlling the alternate splicing at the 3-prime end of the PLCβ1 gene that causes the two variants [46], which could be affected over the course of the illness. In cardiomyocytes, the splice variants play differing roles in signal transduction [47] and thus may have different roles in the CNS. Given the changing symptom profile of schizophrenia, it might be argued that PLCβ1a, which is altered in LDOI, is more associated with negative symptoms, which are more prominent later in the illness.
One limitation in this study is that we measured levels of PLCβ1 proteins in homogenates; a proportion of PLCβ1 is found in the nucleus, where it has been linked to cell cycle regulation [36]. Hence, it is possible that, while there is no global change in expression of the protein, there may be changes restricted to certain cellular compartments. This could explain the differences between this and an earlier study showing PLCβ1 immunoreactivity was increased in the particulate fraction (nuclear and membrane proteins) of BA9, decreased in the particulate fraction of the left superior temporal cortex from patients with schizophrenia and unchanged in the soluble fractions (cytosolic protein) from these regions [48]. The variants are also differentially expressed between the cell membrane, cytosol and nucleus [47,49] and thus may be differentially regulated in each compartment. Therefore, measuring PLCβ1 protein levels in different cell fractions may provide insight into how a decrease in PLCβ1 mRNA may affect either receptor signalling or cell proliferation and differentiation.
As with all post-mortem CNS studies in schizophrenia, treatment before death is a potential confound in our study. Significantly, the doses of antipsychotic drugs prescribed to subjects with SDOI and LDOI schizophrenia do not differ (see Table 1), therefore differences in expression with DOI would not seem to be due to drug treatment
The use of post-mortem tissue poses further restrictions, one of which is that phosphorylated proteins are often rapidly dephosphorylated [50,51]. Thus, we may not be measuring an accurate representation of levels of proteins at the time of death. An alternative would be to measure levels of PLCβ1 activity, the ultimate measure of function, as it dictates the level of signalling. PLCβ1 hydrolyses phosphoinositide (PI), and PI hydrolysis is increased following GTPγS stimulation in the prefrontal cortex (BA8/9) from patients with schizophrenia [52,53]. Moreover, this change in activity occurred in the absence of changes in PLCβ1 protein levels [52]. This further supports the possibility of decreased PLCβ1 mRNA having a functional effect despite no observed change in protein expression.
Our data suggests that changes in PLCβ1 gene expression do not result in a change in PLCβ1 protein levels and highlights the importance of expanding studies of transcriptome expression to determine if the gene product is changed. However, with our increasing understanding of the importance of levels of various RNAs in cell function, it may be premature to conclude that changes in PLCβ1 mRNA would have no physiological consequences for CNS function.
Footnotes
Acknowledgements
We thank Geoff Pavey and Susan Juzva for technical assistance.
