Abstract
The greenbug Schizaphis graminum, is a major pest of wheat worldwide. Biotype screening of this pest is essential to develop pest management programs. In this research, eight greenbug clones, collected on wheat in the cereal-growing region of Béja (north Tunisia), were used to determine their damage on six reference wheat cultivars. All tested clones shared a unique biotypic profile, similar to biotype C. Moreover, DNA from the tested clones and that from seven reference clones of biotypes C, E, F, G, H, I and K, was analyzed, using 5 RAPD-PCR primers. The UPGMA method clustered samples into two distinct clades: a first one (I) included clones from north Tunisia, which were clearly associated to agricultural biotypes C, E, I and K, while a second clade (II) included non agricultural biotypes F, G and H. Results reported in this paper suggest that resistance genes Gb2, Gb3, Gb4, Gb5 and Gb6 in wheat would be the most efficient if used in wheat improvement programs for resistance against greenbug in Tunisia.
Introduction
The greenbug, Schizaphis graminum Rondani (Homoptera: Aphididae) is an aphid pest of several graminaceous crops worldwide. It causes severe injuries to host plants in all growth stages and often kills the entire cereal plant. It is prejudicial to the host, either due to the large quantity of sap it extracts, causing water and nutrients depletion (Cruz et al. 1998) or due to vectoring Barley Yellow Dwarf Virus (BYDV, Gray et al. 2007), Mosaic Maize Dwarf Virus (MMDV, Nault and Bradley 1969) or Sugarcane Mosaic Virus (SCMV, Ingram and Summers 1938). Over twenty greenbug biotypes have been recognized (Porter et al. 1997; Shufran et al. 2000; Burd and Porter 2006); nearly all of them (except D, which has been identified on the basis of insecticide resistance) have been characterized based on the preference for a host plant species and/or ability to damage specific cultivars of a defined species.
The term "biotype" usually designates an infraspecific group of organisms that are not morphologically distinguishable, but differing by a biological function (Eastop 1973). Within this definition, S. graminum biotypes could be described as a case of host race. In fact, host plant response remains the main criterion for recognition of greenbug biotypes. However, determination of host plant response is often laborious and time-consuming. Therefore, other methods based on morphological characters (Starks and Burton 1977), isozymes (Abid et al. 1989) and mitochondrial DNA (Shufran et al. 2000), have been used to assess genetic relationships among biotypes or to develop alternative identification procedures. These methods, however, have not fully distinguished all biotypes. Random Amplified Polymorphic DNA-Polymerase Chain Reaction (RAPD-PCR) has been successfully applied to reveal distinctive patterns among some greenbug biotypes (Black et al. 1992; Aikhionbare et al. 1998; Lopes-Da-Silva et al. 2004). Because biotype identification of S. graminum should be the first step for any cereal-breeding program aiming to obtain resistant cultivars to this pest, the main objective of this work was to characterize the biotype(s) of this insect in a wheat-growing area in the north of Tunisia.
Materials and Methods
Reactions of cereal differential cultivars to infestation by S. graminum clones from north Tunisia.
S: Susceptible, R: Resistant.
Results and Discussion
Biotype screening on a differential set of indicator cultivars showed that the studied greenbug clones from northern Tunisia exhibited profile similar to biotype C (Table 1). All DNA amplification products obtained were reproducible and 60 different markers were scored with the 5 primers used, ranging from 150 to 1100 bp in size. The percentage of polymorphic RAPD fragments generated by each primer alone varied from 60% with primer OP-C09 to 88.88% with primer OP-G18, with an overall percentage of 74.05%. The range of genetic distance was high, varying from 0.040 between clones Sg11 and Sg15, to 1.021 between biotype E and clone Sg11. The dendrogram yielded by the UPGMA method (Fig. 1) showed that the studied clones clustered into 2 distinct clades: a first clade (I), included S. graminum clones from north Tunisia, which were clearly associated to agricultural biotypes C, E, I and K (Shufran et al. 2000); while a second clade (II) included non agricultural biotypes F, G and H. These results do not contrast with the infestation assay; thereby tending to confirm the assignment of Tunisian clones to biotype C. Although the limited set of primers used in this study did not result in clear discrimination between the reference biotypes, the RAPD analysis was useful to distinguish between agricultural biotypes (C, E, I and K) and non-agricultural ones (F, G and H). In similar studies, Aikhonbare et al. (1998) with USA populations, and Lopes-Da-Silva et al. (2004) with Brazilian populations, could not distinguish biotypes C and E, by RAPD markers.
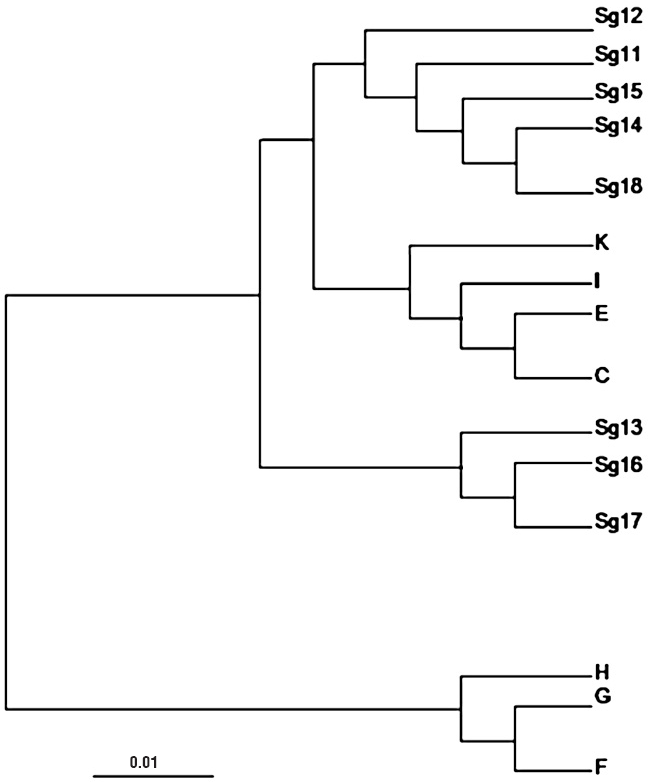
UPGMA dendrogram showing genetic relationships between 8 S. graminum clones from north Tunisia (Sg11-Sg18) and biotypes C, E, F, G, H, I and K, based on 60 RAPD markers generated by 5 primers.
Although greenbug clones from Tunisia, used in this study, showed a homogeneous behaviour at the physiological level, when used for infesting wheat cultivars; they were heterogeneous in molecular analysis, as they were relatively discarded in the dendrogram generated based on RAPD-PCR. This observation is in agreement with several reports where genetic differences were detectable between and within greenbug biotypes (Black et al. 1992; Shufran et al. 1992; Black 1993; Anstead et al. 2002). While some authors (Black et al. 1993) have treated biotypes as evolutionary lineages/units, Anstead et al. (2002) suggested that a conceptual distinction should be made between three terms, which are "genotype", "biotype" and "host race". In other words, a biotype may be characterized by its pattern of virulence to resistance genes or associated with a particular host plant species; yet, it should not be treated as an evolutionary unit or be given taxonomic status and, therefore, it could potentially display genotypic diversity. For example, Shufran et al. (1992) found that biotype E in the field was genetically heterogeneous and concluded that it was comprised of many clonal lineages. In order to distinguish normal variation "between clones within a biotype" from differences "between biotypes", Anstead et al. (2002) suggest that phylogeographical surveys should include large samples and integrate data from different genomic and mitochondrial markers. Anyway, biotype designation remains very useful to researchers, and especially to breeders, as it enables the identification of efficient resistant genes. Biotype C, characterized in this study, implies that resistance genes Gb2, Gb3, Gb4, Gb5 and Gb6 in wheat would be efficient if integrated in wheat improvement programs in Tunisia. The transfer of these genes into commercial wheat cultivars in Tunisia could be facilitated by a marker-assisted selection (MAS), as several molecular markers associated with most genes have been characterized (Weng et al. 2005; Lu et al. 2010).
For a better efficiency of cereal breeding programs, the findings reported here should be continued by extensive surveys, to identify greenbug biotypes in Tunisia, at a greater scale encompassing the South of the country where barley and sorghum are usually grown. Besides, regular biotype surveys, through field and molecular assays, should be conducted, as biotype shifts may occur, rendering previously efficient genes, susceptible to the new biotypes.
Footnotes
Acknowledgements
The authors thank the USDA-ARS (Stillwater, Oklahoma-USA) for providing plant and insect material. Special thanks are addressed to the referees who have reviewed this work for "Revista Colombiana de Entomologia". This work was funded by a "PRF" project between the Ministry of Higher Education (Tunisia) and the Ministry of Agriculture (Tunisia).
