Abstract
Molecular imaging probes can target abnormal gene expression patterns in patients and allow early diagnosis of disease. For selecting a suitable imaging probe, the current Molecular Imaging and Contrast Agent Database (MICAD) provides descriptive and qualitative information on imaging probe characteristics and properties. However, MICAD does not support linkage with the expression profiles of target genes. The proposed Disease-specific Imaging Probe Profiling (DIPP) database quantitatively archives and presents the gene expression profiles of targets across different diseases, anatomic regions, and subcellular locations, providing an objective reference for selecting imaging probes. The DIPP database was validated with a clinical positron emission tomography (PET) study on lung cancer and an in vitro study on neuroendocrine cancer. The retrieved records show that choline kinase beta and glucose transporters were positively and significantly associated with lung cancer among the targets of 11C-choline and [18F]fluoro-2- deoxy-2-D-glucose (FDG), respectively. Their significant overexpressions corresponded to the findings that the uptake rate of FDG increased with tumor size but that of 11C-choline remained constant. Validated with the in vitro study, the expression profiles of disease-associated targets can indicate the eligibility of patients for clinical trials of the treatment probe. A Web search tool of the DIPP database is available at http://www.polyu.edu.hk/bmi/dipp/.
IT IS WIDELY ACCEPTED that genes code for proteins, which maintain cellular function and growth. Any pathologic changes in the gene expression pattern can lead to cellular malfunction, giving rise to disease. Thus, the observed change in the gene expression pattern can provide a sign for early diagnosis and theranosis.
Molecular imaging nowadays provides detailed information at the cellular and biochemical levels. 1 Such emerging technology offers biologists and clinicians new opportunities to noninvasively localize and quantify certain in vivo molecular events.2–4 Clinically, molecular imaging has been widely used as a tool for monitoring, evaluating, and optimizing treatment. 5 The existing molecular imaging techniques include magnetic resonance imaging (MRI), magnetic resonance spectroscopy (MRS), ultrasonography, single-photon emission computed tomography (SPECT), positron emission tomography (PET), and, specifically in the preclinical small-animal setting, optical bioluminescence and fluorescence imaging. 6 For research application, optical imaging has been widely used for detecting gene expression and protein-ligand interactions in animal studies. 1
Molecular imaging probes can interrogate the in vivo change in regional metabolism of specific pathways in the body, which are regulated by specific proteins owing to the fact that the cellular uptake concentration and/or spectral properties of imaging probes are altered by the specific biologic process, such as pathologic or physiologic change, under investigation. 7 Therefore, an imaging probe can respond to the abnormal gene expression pattern with signals in the patients, and the obtained images facilitate early diagnosis and more accurate characterization of disease. Conventional direct binding probes include positron-emitting analogues of dopamine (e.g., 3-(2-[18F] fluoroethyl)spiperone, [18F]FESP), which monitor the dopamine receptors of the striatum, and indirect imaging probes include hexokinase substrate 2-deoxy-2-[18F] fluoro-D-deoxyglucose (FDG), which monitors glucose metabolism. 7
Multimodality imaging, the combined application of multiple imaging techniques, has been launched to provide accurate anatomic and functional information simultaneously. The combinations include PET/computed tomography (CT), which was introduced commercially in 2001, followed by SPECT/CT in 2004 and PET/MRI in 2008. 2 PET/CT acts as an efficient tool for whole-body staging and restaging within one imaging modality. 8 As a result, multimodal imaging probes become more important for developing multimodality imaging techniques. 2 By administering new imaging probes with multimodality-imaging instruments, structural and functional data can be merged and allow physicians to perform multiple functional imaging assays simultaneously with anatomic analyses. 7
Other potential imaging probes are suggested. For example, Oxygen-15 was proposed for studying regional cerebral oxygen extraction and metabolism using PET. 9 Nanoparticles in nanoplatform-based molecular imaging, for example, iron oxide nanoparticles as MRI contrast agents, 10 are already available in clinical practice for tumor targeting and have become more popular due to their unique properties and multifunctionality in nanoplatforms. 5 Iron oxide PEGylated (IO-PEG) nanoparticles have been suggested to serve as an efficient magnetic resonance contrast agent compared to the commercial contrast agent Feridex. 10 Moreover, the bioconjugated near-infrared quantum dot probes are recognized as highly sensitive optical imaging tools for in vivo study. 5
Molecular diagnostics, including in vivo imaging of human bodies and in vitro laboratory test of clinical samples, involves the use of biologic markers to predict the risk of developing disease at an early stage. The molecular phenotype is essential for individualized therapy and therapeutic monitoring. As a rapidly growing area, genomics also plays an important role in defining new direct-imaging targets where gene expression profiling and proteomic analyses of human tumors have already been started. 8 The approach of reporter gene imaging has been changed from postmortem tissue sampling and processing to emphasize noninvasive imaging techniques involving live animals and human subjects. 11 Lifelong reporter gene imaging in the lungs of mice following polyethyleneimine-mediated sleeping-beauty transposon delivery is an example. 12
A number of choices of imaging probes with different specific molecular targets exist. Each imaging method or probe contributes a unique molecular target profile characterizing the biologic system. 13 Therefore, it is important to choose an imaging probe whose signal originates at abnormal cells, which can effectively reflect the pathologically altered gene expression profile. This goal can be achieved by integrating the information of the molecular imaging probes with the genomic and proteomic, morphologic (such as tissue banks), and functional data of their targets. 14 Such an effort facilitates disease-specific comparison of gene expression patterns among different imaging methods or probes.
The Molecular Imaging and Contrast Agent Database (MICAD) provides descriptive and qualitative information on the characteristics and properties of imaging probes, such as their molecular targets and the corresponding gene-disease association. 15 However, MICAD could not provide a systematic, structural, and quantitative data repository and presentation of the target gene expression changes associated with particular disease phenotypes. To address this issue, relevant research effort has been made in the identification of prognostic imaging biomarkers by leveraging public gene expression microarray data in nonsmall cell lung cancer (NSCLC). 16 Such an effort provides insight for profiling the probes with the molecular context of the cell. Supported by bioinformatics resources, such as MetaCore, the Kyoto Encyclopedia of Genes and Genomes (KEGG), and the Gene Ontology (GO), MICAD was further extended to establish the Researching Agents through Molecular Pathways (RAMP) database that integrates imaging probes with the signaling pathways of their target genes. 17 However, the RAMP database still lacks the support of the disease-specific quantitative data of target genes.
An initiative for the development of a systematic and structural database exists that quantifies the target gene-disease relationship of every imaging probe. In this study, it is expected to quantitatively archive and present the differential gene expression profiles of targets across different diseases, anatomic regions, and cellular components. Such an informatics resource provides a comparative target profiling of various imaging and treatment probes as an objective and quantitative reference for supporting the clinical decision and, if the probes have not yet been tested in humans, piloting the clinical trials.
Methods
Database Design and Construction
The database delivered in this work is called the Disease-specific Imaging Probe Profiling (DIPP) database. The DIPP database was constructed by three processes:
collection of the information of imaging probes and their corresponding targets; (2) retrieval and processing of the microarray data sets of specified diseases; and (3) data integration and identification of significant records. Processes 1 and 2 were performed in parallel followed by process 3. Figure 1 shows the flowchart of the database construction. A Web search tool of the DIPP database is available at http://www.polyu.edu.hk/bmi/dipp/ and is best viewed by Firefox or Safari.
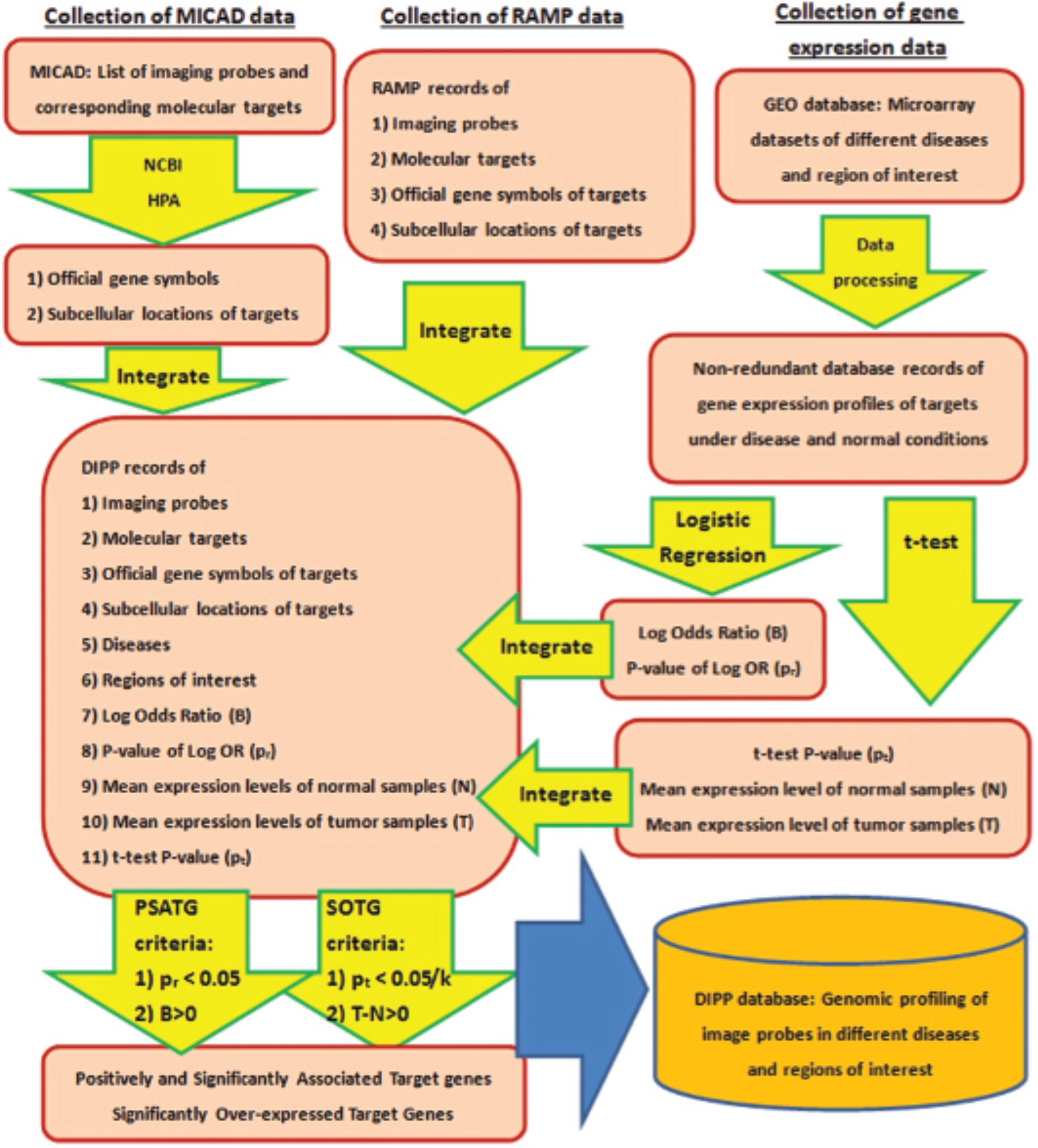
The schematic diagram of the Disease-specific Imaging Probe Profiling (DIPP) database construction: k is the number of targets of an imaging probe. Logistic regression and t-test were performed to identify disease-associated and differentially expressed targets. Bonferroni correction, which adjusts the significance level by dividing it by k, was applied to control the type I error of the multiple t-tests. GEO = Gene Expression Omnibus; HPA = Human Protein Atlas; MICAD = Molecular Imaging and Contrast Agent Database; NCBI = National Center for Biotechnology Information; PSATG = positively and significantly associated target gene; RAMP = Researching Agents through Molecular Pathways; SOTG = significantly overexpressed target gene.
Collection of Probe-Target Mappings
The mappings between imaging probes and their targets were obtained using the export function of a RAMP search tool (http://www.rampsearch.com/) and manually curated using MICAD (http://www.ncbi.nlm.nih.gov/books/NBK5330/).
Each record collected from RAMP provides four information items: imaging probes, target descriptions, target official gene symbols, and target subcellular locations. Although about 4,100 records were collected from RAMP, some commonly used imaging probes were not covered. To supplement the inadequacy of the RAMP database, manual curation of probe-target mappings, including those for FDG, 11C-choline, 68Ga-DOTATATE, and 177Lu-DOTATATE, was performed using MICAD. To identify the targets of those manually curated records consistently and uniquely, their official symbols and subcellular locations were obtained from the National Center for Biotechnology Information (NCBI) and the Human Protein Atlas (HPA), respectively.18,19
Collection and Processing of Gene Expression Data
In this work, eight diseases, including breast cancer, colon cancer, NSCLC, head and neck cancer, neuroendocrine tumor (NET), ovarian cancer, cervical cancer, and nasopharyngeal carcinoma (NPC), were considered. For each disease, the related studies were searched and the corresponding microarray data sets were collected from the Gene Expression Omnibus (GEO). 20 Information about the region of interest (ROI) was also provided in the study description in the GEO. The selection criterion of microarray data sets is the presence of both tumor and normal subject groups in the studies. Based on this criterion, microarray data sets of the experiment series, GSE12452 (NPC), GSE6791 (cervical cancer, head and neck cancer), GSE43458 (NSCLC), GSE42568 (breast cancer), GSE26712 (ovarian cancer), and GSE1037 (NET), were retrieved.21–26 All the data sets collected in the work have already been normalized. According to the microarray probeset design, a gene may be interrogated by more than one microarray probe. In this case, the expression level of the gene is represented by the average intensity value across the corresponding microarray probes. For each data set, nonredundant records of targets were maintained by averaging the multiple microarray probe intensities.
Multivariate statistical analysis should be applied to examine the complex interrelationships between the probes, targets, and diseases. Since a probe may target multiple genes whose expression levels jointly contribute to the occurrence of a disease, logistic regression analysis was performed to identify a linear equation whose output variable is the logarithm of the odds of a disease and input variables are the expression levels of target genes. The coefficient associated with each target gene represents the log odds ratio (log OR) of the disease with respect to the expression level of that target gene corrected for the influence of the others. The p value was also estimated to determine the significance of the log OR. The positive and significant log ORs indicate that the corresponding genes are positively and significantly associated with the disease and thus contribute to the disease-specific uptake of their targeting probe.
After the disease-associated genes were identified, their differential expressions between normal and disease groups were examined using the Student t-test. Mean expression levels over control samples (N) and disease samples (T) were calculated for each record of target. The difference between T and N was used to indicate whether the gene was overexpressed or underexpressed in the disease group.
Data Integration and Identification of Disease-Associated and Differentially Expressed Genes
Using the database table-joint method, the records of probe-target mappings and the differential gene expression parameters obtained from processes 1 and 2 were linked to form the disease-specific genomic profiling of imaging probes. The key for joining records is the official gene symbol. The integrated database records include 11 fields: imaging probe, target description, official gene symbol, subcellular locations, disease, ROI, log OR (B), p value of log OR (pr), mean expression level of normal samples (N), mean expression level of tumor samples (T), and t-test p value (pt).
For a particular disease and ROI, each imaging probe could have k distinct targets. The disease-associated genes were identified using the following logistic regression model:
where logit is the logarithm of the odds, X1, ·, Xk are the expression levels of targets and B1, ·, Bk are the corresponding log ORs. The disease-specific uptake of a probe can be predicted by its targets, whose expression levels are positively and significantly associated with the disease. These positively and significantly associated target genes (PSATGs) are defined as genes satisfying the criteria of pr < .05 and B > 0. For each combination of imaging probes, disease, and ROI, the PSATG records were identified based on the criteria.
The individual contributions of the PSATGs to the differential uptake of the probe were further examined using the Student t-test. To control the type I error of multiple t-tests of these k database records, Bonferroni correction was applied to adjust the significance level (a, originally 0.05) for differential gene expression. Significantly overexpressed target genes (SOTGs) are defined as genes whose expression profiles satisfy the criteria of pt < .05/k and T – N > 0. The SOTGs were identified based on the criteria to predict the differential uptake of the probe comparing the disease to normal cases.
Validation of Database
For a particular disease, we hypothesize that the PSATGs and SOTGs characterize, respectively, the disease-specific and the differential uptake properties of an imaging probe with respect to the disease and the anatomic ROI. To test this hypothesis, we queried the DIPP database with criteria corresponding to a clinical PET study. 27
In the clinical PET study, the changes in standardized uptake values (SUVs) against tumor size were compared between two imaging probes, FDG and 11C-choline, in lung cancer. The search criteria were “lung” as ROI, “NSCLC” as disease, and “FDG or 11C-choline” as imaging probes. Such validation mimics the scenario for comparing the uptake properties of the candidate imaging probes for molecular imaging. 27
As some probes are eligible for treatment, the DIPP database can also be applied to identify therapeutic radionuclide and predict its efficacy with respect to the disease and ROI. To identify potential treatment probes for NET, we searched for PSATG and SOTG records with the criteria “neuroendocrine cancer” as disease and “lung” as ROI. The retrieved record was validated by the findings of an in vitro study on pulmonary NET. 28
The Web search tool provides links to the PubMed webpages of the probes found in the DIPP database. By clicking on the tracer name of the retrieved records, the PubMed webpage of the selected probe will be brought out, showing the excerpt, the background, the synthesis approach, and the information about the related in vitro animal and human studies supported by references.
Results
Construction of the DIPP Database
To demonstrate the database construction process, the information collection from MICAD and HPA corresponding to two commonly used tracers, FDG and 11C- choline, is illustrated as follows. Table 1 provides the overview of target-disease relationships identified in MICAD and HPA. FDG potentially traces 18 targets and 11C-choline 3 targets. The targets include enzymes, trans-porters, and regulators of metabolic processes. Table 2 shows the detailed information of the relationships, such as the type, subcellular location, and chromosomal location of the target. Note that choline transporter was not defined as a target of 11C-choline at MICAD, but it is important for incorporating choline into the cell. As such, an extra record for choline transporter was added to the database.
Relationships of Target Genes of Two Imaging Probes with Cancers

FDG = [18F]fluoro-2-deoxy-2-D-glucose.
Relationship between Imaging Probes and Target and Related Information Found at MICAD, NCBI, and HPA
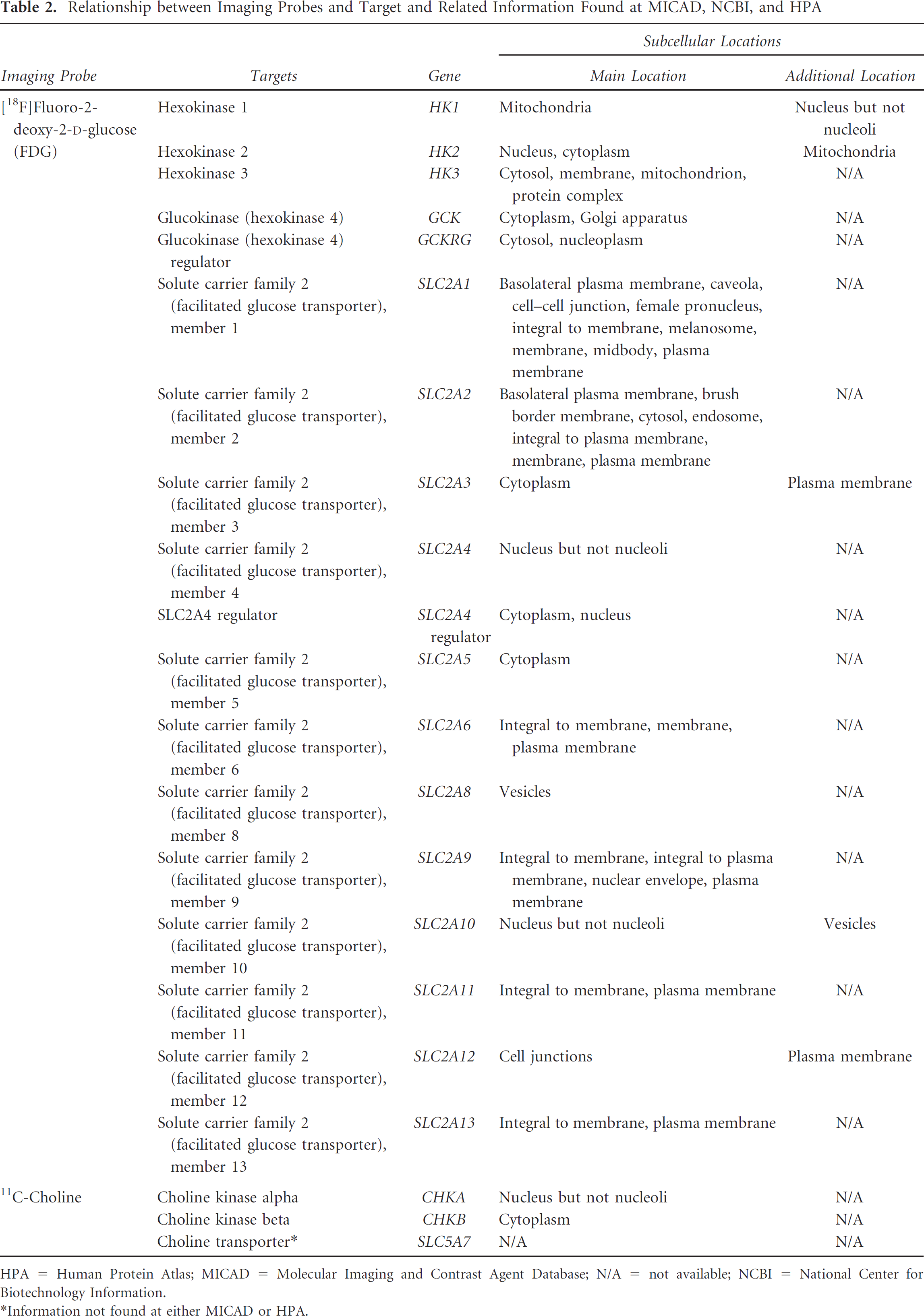
HPA = Human Protein Atlas; MICAD = Molecular Imaging and Contrast Agent Database; N/A = not available; NCBI = National Center for Biotechnology Information.
Information not found at either MICAD or HPA.
Validation Results
By setting “lung” as ROI, “NSCLC” as disease, and “FDG” and “[11C]choline” as tracers, we retrieved 13 and 3 target gene records from the DIPP database for FDG and 11C- choline, respectively. The records of PSATGs and SOTGs are shown in Table 3 and Table 4, respectively. Among 3 targets of 11C-choline, choline kinase beta (CHKB) is the only PSATG that was found to be positively and significantly associated with NSCLC (log OR = 3.67, pr = 4 × 10−4) and also the only SOTG that was found to be significantly overexpressed (pt = 4.13 × 10−5 < α/k = 0.0167) in the microarray data set (GSE43458). Among 13 targets of FDG, solute carrier family 2 member 8 (SLC2A8), which is one of the glucose transporters, and hexokinase 2 (HK2) were identified as the PSATGs (log OR = 9.46, pr = .015 and log OR = 2.72, pr = .013, respectively). Four glucose transporters and one hexokinase were significantly overexpressed among the 13 FDG targets (α/k=3.85 × 10−3). The SOTGs are SLC2A2 (p = 1.42 × 10−4), SLC2A4 (p = 4.40 × 10−6), SLC2A4RG (p = 4.77 × 10−6), SLC2A8 (p = 8.55 × 10−8), and GCK (p = 8.06 × 10−5). SLC2A8 is the only SOTG of the two
DIPP Database Records of PSATGs of 11C-Choline and FDG in NSCLC
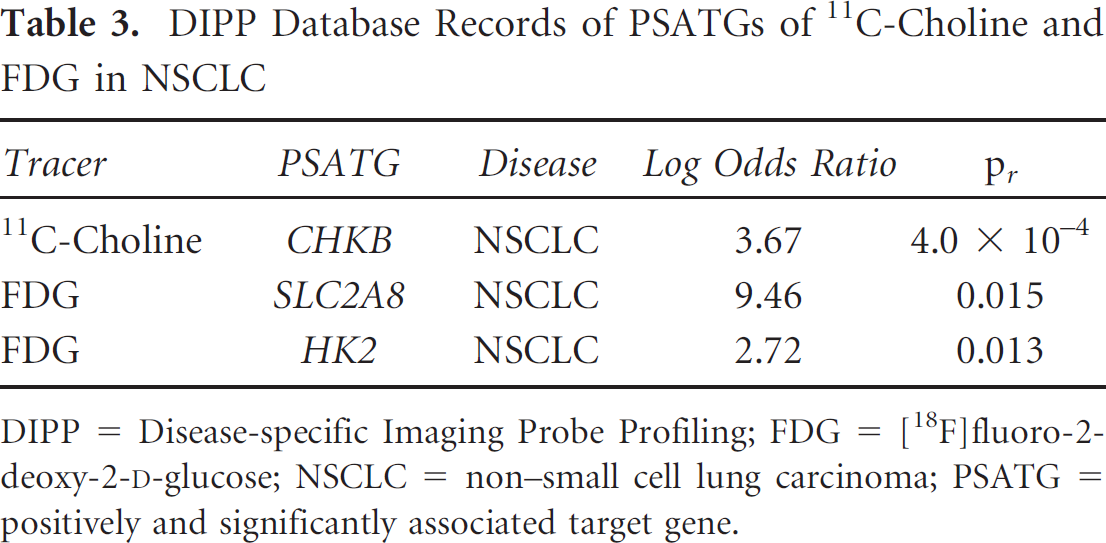
DIPP = Disease-specific Imaging Probe Profiling; FDG = [18F]fluoro-2- deoxy-2-D-glucose; NSCLC = non-small cell lung carcinoma; PSATG = positively and significantly associated target gene.
DIPP Database Records of SOTGs of 11C-Choline and FDG in NSCLC

DIPP = Disease-specific Imaging Probe Profiling; FDG = [18F]fluoro-2- deoxy-2-D-glucose; NSCLC = non-small cell lung carcinoma; SOTG = significantly overexpressed target gene.
PSATGs. For profiling the associations with disease, the targets were characterized by a panel of two features, y = log(OR) and × = log(a/pr). For profiling the differential expression, the targets were characterized by another panel of two features, y = (T – N) and × = log(α/(pt-k)). The scatter plots of targets in these two panels of features are shown in Figure 2 and Figure 3. The first quadrant of the scatter plots represents the region where the PSATGs and SOTGs are located.

The scatter plot of targets of 11C-choline and FDG with respect to the log odds ratio and its significance of association with non-small cell lung carcinoma. The points in the first quadrant represent the positively and significantly associated target genes, consisting of HK2, SLC2A8, and CHKB.
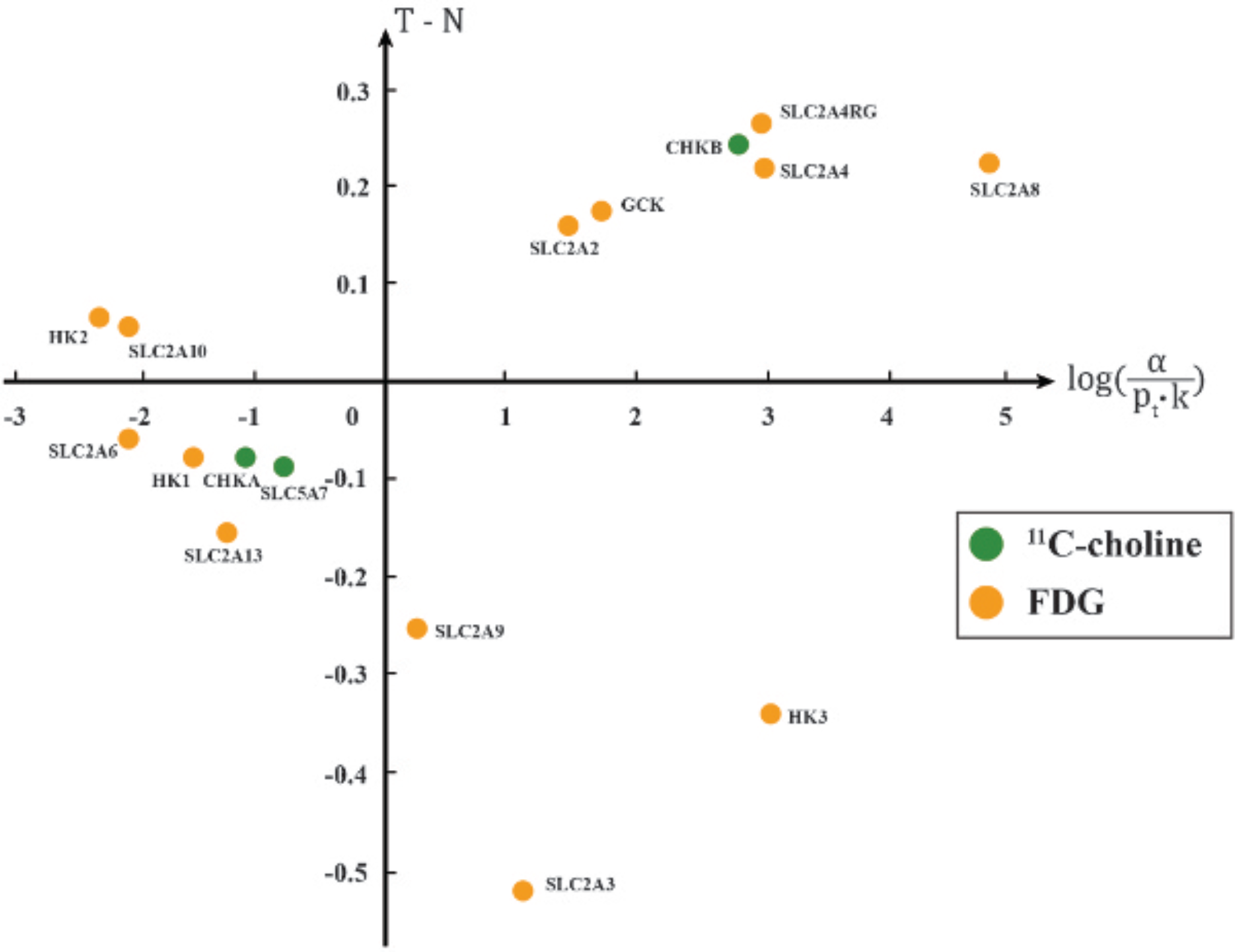
The scatter plot of targets of 11C-choline and FDG with respect to the mean expression level difference (T – N) and its significance of differential expression in non-small cell lung carcinoma. The points in the first quadrant represent significantly overexpressed target genes, consisting of GCK, SLC2A2, SLC2A4, SLC2A4RG, SLC2A8, and CHKB.
The identified PSATGs include two enzymes, HK2 and CHKB, which are targeted by FDG and 11C-choline, respectively. These two PSATG records justify the similar disease-associated increase in enzymatic activity and tumor cell proliferation in lung cancer and support the baseline disease-specific uptake of these two imaging probes in clinical PET images. 27 However, only SLC2A8 and CHKB were significantly overexpressed among the three PSATGs. The types of SOTGs targeted by FDG and 11C-choline were so different that the two tracers have different uptake properties in terms of tumor size. It was found in the clinical PET study that the SUV of FDG increased but that of 11C-choline remained unchanged when tumor size increased. Such a differential uptake property was characterized by the types of significantly overexpressed PSATGs that are an enzyme for 11C-choline and a transporter for FDG.
To prove the ability of the DIPP database to identify and evaluate treatment probes, it was queried for PSATGs and SOTGs by setting “neuroendocrine cancer” as disease, “lung” as ROI, and “yes” as significantly overexpressed. Among the 32 retrieved SOTG records, a gene, carbonic anhydrase IX (CA9), was targeted by a treatment probe, 177Lu-Bn-DTPA-SIP-A3, and its expression level was found to be associated with neuroendocrine cancer (log OR = 5.55, pr = .01). Labeled with the γ- and β-emitting isotope lutetium 177 with a 6.73-day half-life, the probe is a more efficient option in radionuclide therapy. 29 In the only SOTG record of a treatment probe for neuroendocrine cancer, the identified target is CA9. In the in vitro study, the carbonic anhydrase inhibitor was applied to the lung carcinoid cell lines, and a significant dose-dependent reduction in cell viability and colony formation was observed. 28 The finding suggests the application of 177Lu-Bn-DTPA-SIP-A3 to NET, where CA9 is overexpressed. By clicking on the tracer name “177Lu-Bn-DTPA-SIP-A3” of the retrieved records, its popup PubMed webpage shows a summary of an animal study, which investigated the tracer biodistribution in mice bearing colorectal tumors and thus provided important data supporting further clinical trials. 30
Discussion
According to the validation results, the PSATG records of the DIPP database are able to justify the baseline disease-specific tracer uptake in PET imaging of cancer. In tumor cells, an excessive amount of glucose is used to produce adenosine triphosphate (ATP) through anaerobic glycolysis. The glycolysis is promoted by the increased activity of glucose transporters and glycolytic enzymes that can be reflected by a high SUV of FDG. For tumor cell proliferation, choline is transported into the tumor cells and then metabolized to phosphatidylcholine, leading to the cell membrane synthesis. The increased activities of choline transporter and kinases resulting in the enhanced cell membrane synthesis can be reflected by a high SUV of 11 C-choline. However, the enzymatic activity of tumor cells is further affected by hypoxia. The blood supply to the tumor becomes poorer, leading to hypoxia when the tumor grows. The increase in tumor size further reduces the molecular oxygen supply and promotes the enzymatic activity. The significant overexpression of glucose transporters supports this neoplastic metabolic process and the tumor size-specific differential uptake property of FDG. 27 In contrast, the overexpression of choline kinase does not support any tumor size-dependent increase in 11C-choline uptake. 27
The validation demonstrated a proof of concept for using the database as a quantitative reference for predicting the performance of the imaging probes with respect to particular diseases. In the potential applications of the DIPP database in clinical practice across all diseases, the query using a disease and its ROI as the search criteria corresponds to a list of target gene records. The PSATGs and SOTGs of the retrieved records, their types, and involved metabolic processes facilitate the prediction and comparison of the uptake properties of different imaging probes under different pathologic conditions, such as tumor size. Such properties indicate if the imaging probes fulfill the specific diagnostic and prognostic needs. Objective selection of tracer is allowed as the PSATGs and SOTGs provide quantitative evidence.
Apart from the suspected diseases, tracer uptake depicted unexpectedly in some regions of the molecular image can be explained by the DIPP database search. By setting the ROI and imaging probes as the search criteria, the PSATGs can be identified accordingly, and the associated diseases provide clinicians with useful information for making a differential diagnosis.
The database may also help clinicians identify the physiologic uptake of the imaging probe at a region other than the primary region of disease of interest. By setting ROI and disease as the search criteria, the SOTG records about other regions, such as the lymph nodes, can provide useful information for clinicians to differentiate the physiologic uptakes in metastatic cases from those of the primary tumors. The current version of the DIPP database is limited to the ROI, which is the same as the disease origin. Such function extension will be implemented in the future version of the DIPP database.
The DIPP database further facilitates the selection of optimal imaging method among different imaging modalities. Using disease as a search criterion, the intermodality summary of SOTGs can be obtained so that a better choice of imaging modalities is suggested.
Using the DIPP database, the exploration of disease mechanism is possible. Since the DIPP database is able to store detailed information, such as gender, age, source of tissue, histology, histotype of cancer, type of tumor (benign, malignant, metastasis), and stage and grade of cancer in the PSATG records, the corresponding association of the gene expression pattern with the disease progression can be identified for suggesting further experimental studies on the PSATGs.
The DIPP database potentially explains the interrelationship of target proteins functioning in vivo by tracing the over- and underexpression patterns of the protein-coding genes over different disease states. A new hypothesis of a previously unexplained mechanism of proteins in cellular pathways may be facilitated, gaining momentum for the flourishing of related pathologic analysis and drug development.
However, there are a number of concerns regarding the application of the DIPP database. The MICAD collection of information of targets will not be further updated after June 30, 2013. As the imaging probes could target proteins apart from those listed in the MICAD, such new targets will be identified, validated, and reported in future publications. Depending on funding support, periodic updates of DIPP on the information of targets will be performed by the scientific editors and curators of our research team.
Not all of the probes in the MICAD collection have human studies available. However, the in vitro experiments and animal studies available at the PubMed webpage, in combination with the disease-specific gene expression profiling in the DIPP database, provide supportive data for suggesting further clinical trials of the probes. In contrast, some imaging probes, such as FDG and 11C-choline, have been widely used in clinical applications for the diagnosis, treatment monitoring, and staging of various diseases, including cancers and central nervous system disorders. In addition to the related human studies, the “Related Resource Links” under the background section of the PubMed webpage provides a link to ClinicalTrials.gov that lists all of the ongoing and completed clinical trials of these imaging probes. Although there are still many ongoing clinical trials of probes, such as 177Lu-DOTA-TATE (ClinicalTrials.gov Identifier: NCT01456078), which have not yet been referenced by MICAD, the scientific editors and curators of our research team will periodically review and incorporate such findings into the DIPP database, depending on funding support.
Furthermore, the expression profiling of imaging probes can be extended to genes coding proteins that participate in the downstream and upstream of the same pathways as targets do. The RAMP database does not cover many such genes involved in the downstream and upstream of target pathways. For example, the activation of the phosphatidylinositol 3′-kinase(PI3K)-Akt signaling pathway results in the translocation of glucose transporter to the cell membrane, facilitating glucose uptake. The genes involved in such a signaling cascade, such as PI3K and Akt, are the indirect targets of FDG, and their variants were found in various cancer types, including ovarian and colorectal cancers. 31 It was also found that the choline metabolic process interacts with the oncogenic pathways, such as the RAS pathway and the PI3K-Akt pathway, where RAS, RAF, and PI3K regulate the enzymatic activity of choline kinase, and dysregulation was found in breast and ovarian cancers. 32 Thus, RAS, RAF, and PI3K are indirectly targeted by 11C-choline. If the expression profiles of these indirect targets of imaging probes can be further integrated with the DIPP database, a deeper understanding of the uptake properties of imaging probes will be possible with the support of the cancer signaling pathways.
This work also suggests application of the DIPP database for the identification and evaluation of treatment probes in diseases, such as neuroendocrine cancer. A treatment probe, 177Lu-Bn-DTPA-SIP-A3, was identified through the retrieval of PSATG records in NET. To suggest the clinical trial of the personalized therapy, the efficacy of the treatment probe can be evaluated through the differential expression of PSATGs in the NET biopsy of the patients who are eligible for such a radionuclide therapy option.
In this work, the construction of DIPP database was achieved by data collection, manual curation, integration, and processing efforts, which were relatively time consuming and labor intensive. To continuously extend and update the database, data integration and update processes should be computerized and automated.
Conclusion
The DIPP database developed in this study was validated using a clinical PET study on lung cancer. The hot spots depicted on the images reflecting the tracer uptake were verified against the PSATGs and SOTGs in the target gene records. The tumor size–dependent differential uptake of FDG was further explained by the fact that the glucose transporters were overexpressed significantly. The validation demonstrated a proof of concept for using the database as a quantitative reference for predicting the performance of the imaging and treatment probes with respect to particular diseases.
Disclaimer
After June 30, 2013, the MICAD website will no longer provide the new and revised chapters of molecular imaging probes and contrast agents. The current chapters are still accessible online at the MICAD website. On the condition of sufficient future funding support, the research team will continue to maintain and update the content of the DIPP database periodically.
Footnotes
Acknowledgment
Financial disclosure of authors: This work was supported by an Internal Grant [G.55.09.YL61] and an FHSS Summer Research Studentship from The Hong Kong Polytechnic University. Financial disclosure of reviewers: None reported.

