Abstract
Plasmids tend to have much lower expression than viruses. Gene expression after systemic administration of plasmid vectors has not been assessed using somatostatin receptor type 2 (SSTR2)-based reporters. The purpose of this work was to identify gene expression in non–small cell lung cancer (NSCLC) after systemic liposomal nanoparticle delivery of plasmid containing SSTR2-based reporter gene. In vitro, Western blotting was performed after transient transfection with the plasmid cytomegalovirus (CMV)-SSTR2, CMV-TUSC2-IRES-SSTR2, or CMV-TUSC2. SSTR2 is the reporter gene, and TUSC2 is a therapeutic gene. Mice with A549 NSCLC lung tumors were injected intravenously with CMV-SSTR2, CMV-TUSC2-IRES-SSTR2, or CMV-TUSC2 plasmids in DOTAP:cholesterolliposomal nanoparticles. Two days later, mice were injected intravenously with 111In-octreotide. The next day, biodistribution was performed. The experiment was repeated including single-photon emission computed tomography/computed tomography (SPECT/CT). Immunohistochemistry was performed. In vitro, SSTR2 expression was similar in cells transfected with CMV-SSTR2 or CMV-TUSC2-IRES-SSTR2. TUSC2 expression was similar in cells transfected with CMV-TUSC2 or CMV-TUSC2-SSTR2. Biodistribution demonstrated significantly greater 111In-octreotide uptake in tumors from mice injected with CMV-TUSC2-IRES-SSTR2 or CMV-SSTR2 than the control plasmid, CMV-TUSC2 (p < .05). Gamma-camera and SPECT/CT imaging illustrated SSTR2 expression in tumors in mice injected with CMV-TUSC2-IRES-SSTR2 or CMV-SSTR2 versus background with control plasmid. Immunohistochemistry corresponded with imaging. SSTR2-based reporter imaging can visualize gene expression in lung tumors after systemic liposomal nanoparticle delivery of plasmid containing SSTR2-based reporter gene or SSTR2 linked to a second therapeutic gene, such as TUSC2.
LUNG CANCER accounts for the greatest number of deaths related to cancer in both men and women. There are an estimated 221,130 new cases of lung cancer and 156,940 deaths, accounting for approximately 27% of all cancer-related deaths in the United States, as expected in 2011. 1 The method of treatment is typically dependent on the type (small cell or non–small cell) and tumor stage. The great majority of lung cancers are of the non–small cell variety. Common treatment options include surgery, radiation therapy, chemotherapy, or multiple modalities. 2 However, the survival rate is often limited using these treatment options because the disease has usually spread by the time of diagnosis. Adjuvant chemotherapy prolongs overall survival by only 5% at 5 years. 3 Thus, new methods of treatment are currently being assessed. For example, gene therapy has the potential to improve the outcome of systemic therapy in patients. 3 However, it has been limited by a lack of clinically relevant techniques to assess gene expression in patients after gene transfer. Imaging-based reporter systems provide a means to overcome this limitation by evaluating gene expression at the target site.
Reporter systems commonly consist of a gene that encodes an enzyme, transporter, or receptor,4–11 which typically binds or entraps an imaging agent. Human somatostatin receptor type 2 (hSSTR2)-based reporters present several advantages.11–14 For example, hSSTR2 is of human origin, which should limit the immune response, and its level of expression in vivo can be quantified over time. 12 Its small size takes up little of the limited insert space of vectors. Moreover, a signaling-deficient variant has been created, 15 which should limit potential interference with the intracellular milieu and a linked gene of interest. Previously, our laboratory demonstrated imaging of hSSTR2 gene expression after intratumoral delivery of adenoviral vectors in mouse models using cognate human imaging machines and (111In)-octreotide,12–14 a radiolabeled somatostatin analogue that is approved by the Food and Drug Administration for use in the clinic. Plasmids present safety advantages compared to viruses, such as being less immunogenic and having a tendency to be nonintegrating. However, plasmids commonly have much lower expression than viruses. Assessing gene expression after systemic administration of plasmid vectors with or without a linked gene has not been assessed using SSTR2-based reporters. A therapeutic gene that has shown promise is the tumor suppressor candidate 2 TUSC2/FUS1 (TUSC2).16–18 It has been previously determined that the expression of TUSC2 protein is either absent or reduced in the majority of lung cancers and premalignant lung lesions. 16 TUSC2 gene expression has been found to significantly inhibit tumor cell proliferation of non–small cell lung cancer (NSCLC) by induction of apoptosis both in vitro and in vivo.16,19–21 In a recent phase I dose escalation clinical trial using DOTAP:cholesterol to deliver a plasmid containing a TUSC2 insert, TUSC2 expression and apoptosis were seen in biopsy specimens, and several patients achieved stable disease, with one demonstrating a metabolic response by 18 F-fluorodeoxyglucose positron emission tomography. 22 The formulation could be safely given, and the maximum tolerated dose was found to be 0.06 mg/kg. 22 The purpose of the present study was to identify gene expression in NSCLC tumor xenografts in mice after intravenous delivery of a plasmid containing a therapeutic gene linked to an SSTR2-based reporter gene.
Materials and Methods
Cell Culture, Western Blotting, and Immunofluorescence
A549-luc human NSCLC lines were obtained from Dr. Lin Ji of the Department of Cardiothoracic Surgery, The University of Texas MD Anderson Cancer Center, and were maintained in RPMI 1640 supplemented with 10% fetal bovine serum (FBS; Mediatech Inc., Manassas, VA), 1% L-glutamine, and 1% penicillin-streptomycin mixture. Adherent monolayer cultures were maintained on plastic flasks and incubated at 37°C in a mixture of 5% CO2 and 95% air. HT1080 cells (human fibrosarcoma; ATCC, Rockville, MD) were grown in Dulbecco's Modified Eagle's Medium (DMEM) containing 1× glutamine, penicillin, streptomycin, and 10% FBS.
Plasmid cytomegalovirus (CMV)-TUSC2 is as described previously.22,23 Using the same backbone, for CMV-SSTR2, SSTR2A gene 11 was placed in an expression cassette driven by a CMV promoter and ended with a bovine growth hormone polyadenylation (BGH-polyA) tail. For CMV-TUSC2-IRES-SSTR2A, the tumor suppressor gene TUSC2 and the reporter gene SSTR2A were separately coexpressed with an internal ribosome entry site (IRES) inserted between them and driven by the same CMV promoter and tailed by a BGH-polyA signal sequence. A kanamycin-resistant gene was used as a selective marker for plasmid amplification in the bacterial host.
For transfection, 1 μ DNA (CMV-TUSC2-IRES-SSTR2, CMV-SSTR2, or CMV-TUSC2) was added with Lipofectamine 2000 (Invitrogen, Carlsbad, CA) to 1 × 105 cells according to the manufacturer's instructions. After 48 hours, cells grown to near-confluence in six-well dishes were washed with phosphate-buffered saline (PBS), exposed to Tris/sodium dodecyl sulfate (SDS) lysis solution (1% SDS, 10 mM Tris [pH 7.4], 1 mM sodium orthovanadate). The lysate was boiled for 3 minutes and then cooled on ice. After a 10-minute centrifugation at 12,000 rpm, the supernatant was collected and its protein concentration was determined using the Bradford method (Bio-Rad Laboratories, Hercules, CA). Thirty micrograms of protein was loaded per lane on a 9% SDS gel. After transfer, the membrane was washed in PBS, blocked with 5% milk for 1 hour at room temperature, and washed in PBS with 0.1% Tween. Next, it was exposed to rabbit-anti-SSTR2A (Santa Cruz Biotechnology, Santa Cruz, CA) at 1:3,000 dilution or rabbit-anti-TUSC2 (obtained from Dr. Lin Ji) at 1:1,000 dilution at room temperature for 2 hours. The membrane was washed three times for 5 minutes each with PBS containing 0.1% Tween and exposed to secondary goat antirabbit antibody (Bio-Rad) at 1:4,000 dilution for 2 hours at room temperature. Following three 5-minute washes with PBS containing 0.1% Tween, the membrane was covered with a chemiluminescent horseradish peroxidase (HRP) substrate (PerklinElmer Life Sciences, Boston, MA) for filming.
For immunofluorescence, transfected cells were plated on coverslips overnight and then fixed with 10% formalin for 30 minutes, washed, and exposed to rabbit-anti-SSTR2A (1:2,000 dilution, Santa Cruz Biotechnology) or rabbit-anti-TUSC2 antibody (1:1,000 dilution) for 1 hour at room temperature. Three PBS washes for 5 minutes each followed. Next, the cells were exposed to diluted (1:250) fluorescein isothiocyanate (FITC)-goat-antirabbit antibody (Sigma, St. Louis, MO) or (1:250) rhodamine donkey-antirabbit antibody (Santa Cruz Biotechnology) for 1 hour at room temperature. After three washes for 5 minutes each, the coverslips were mounted on slides using Gel Mount (Biomeda, Foster City, CA).
Animal Implantation of Tumor Cells
The animal experiments were approved by the Institutional Animal Care and Use Committee at The University of Texas MD Anderson Cancer Center and performed in accordance with the guide for the care and the use of laboratory animals. Female athymic nude mice (8–10 weeks old) were purchased from Charles River (Wilmington, MA). Tumors were produced by harvesting A549-luc cells from subconfluent cultures and briefly exposing the cells to 0.25% trypsin and 0.02% ethylene diaminetetraacetic acid (EDTA). RPMI medium containing 10% FBS was used to stop trypsinization. Then the cells were washed once with PBS, resuspended, and counted using a cell counter. A549-luc cells in the amount of 2 × 106 in 100 μ PBS were injected via the tail vein of each animal.
In Vivo Plasmid Treatments, Animal Imaging, Biodistribution
Tumors were monitored by luciferase imaging and then confirmed with computed tomography (CT) approximately 2 weeks after the A549-luc cells were injected. After CT imaging, a total of six mice/per group were distributed into three different groups. Each group received one of the following plasmid DNAs via the tail vein: CMV-SSTR2, CMV-TUSC2-IRES-SSTR2, or CMV-TUSC2 (25 mg plasmid DNA: 40 nM DOTAP:cholesterol in 100 μL volume). 23 Two days later, the mice were injected intravenously via the tail vein with 11 MBq (300 μCi) of 111In-octreotide (Octreoscan, Mallinckrodt, St. Louis, MO). The next day, the mice were sacrificed; the organs and lung tumors of each animal were dissected and weighed. Associated radioactivity was determined using a gamma counter (Packard Instrument Company, Downers Grove, IL) to obtain the percentage of injected dose per gram (% ID/g) biodistribution. 11
The experiment was repeated with three animals per group and 24 hours after 111In-octreotide injection and gamma-camera and single-photon emission computed tomography (SPECT)/CT imaging were performed. Mice anesthetized with 2% isofluorane were imaged for 10 minutes using a gamma camera (mCAM, Siemens Medical Solutions, Hoffman Estates, IL) fitted with a medium-energy collimator. The same day, the anesthetized mice were imaged using a SPECT/CT scanner (Flex XO X-SPECT, Gamma Medica, Salem, NH) for 20 minutes. CT images were obtained at 75 kV and 310 μA at a scanning time of 2 minutes per animal. The images have an isotropic voxel size of 170 μm at 250 milliseconds per frame for a total of 512 images per animal. The Image Segmentation editor of the imaging software Amira 3.1 (VSG, Burlington, MA) was used to view and normalize the images. Lung nodule size by CT and number visible by CT versus SPECT were assessed. After imaging, the mice were sacrificed; the organs and lung tumors of each animal were dissected and weighed to perform ex vivo biodistribution analysis.
Immunohistochemistry
Mouse lung tumor sections from the three groups of mice underwent hematoxylin-eosin staining or immunohistochemistry. Tumor tissue was fixed in 10% formalin for 24 hours. Paraffin-embedded sections were processed for immunohistochemistry using the Dako LSAB system-HRP kit per the manufacturer's instructions (Cat# K0679, DakoCytomation Inc., Carpinteria, CA). The sections were probed with primary polyclonal rabbit-anti-SSTR2 antibodies (1:250 dilution, Santa Cruz Biotechnology) or rabbit-anti-TUSC2 antibodies (1:250) and stained with a 3,3′-diaminobenzidine (DAB) substrate kit. Caspase 3 (Cat# CP 229, Biocare Medical, LLC., Concord, CA) was used as a marker for apoptosis. The slides were counterstained with Meyer's hematoxylin stain (Sigma), washed with running water, and mounted on glass slides using Gel Mount (Biomeda).
Statistical Analysis
The results are presented as means ± standard deviation. For biodistribution analysis, a two-sided t-test was performed using Microsoft Excel 2007 (Microsoft Corp., Redmond, WA). For all results, p < .05 was considered statistically significant.
Results
SSTR2 and TUSC2 Expression Detection In Vitro
We assessed whether SSTR2 reporter and TUSC2 tumor suppressor gene expression could be detected in vitro upon plasmid-based delivery.
By Western blotting, similar SSTR2 expression was seen in HT1080 cells transfected with CMV-TUSC2-IRES-SSTR2 or CMV-SSTR2 plasmid DNA. In contrast, no expression was observed in cell lines transfected with CMV-TUSC2 plasmid DNA (Figure 1A). Similar TUSC2 expression was seen in HT1080 cell lines transfected with CMV-TUSC2-IRES-SSTR2 or CMV-TUSC2 plasmid DNA. In contrast, no expression was observed in HT1080 cells transfected with CMV-SSTR2 plasmid DNA (Figure 1B).

Western blot and immunofluorescence results demonstrating SSTR2 and TUSC2 expression after plasmid-based gene transfection. Western blot results in HT1080 cell lines transfected with CMV-TUSC2-IRES-SSTR2, CMV-SSTR2, and CMV-TUSC2 plasmid DNA with anti-SSTR2 antibody for the expressed SSTR2 protein (A). The blot was stripped and then probed with anti-TUSC2 antibody for the expressed TUSC2 protein (B). Immunofluorescence stain results showed the fluorescent labeling in HT1080 cell lines transfected with CMV-TUSC2-IRES-SSTR2, CMV-SSTR2, and CMV-TUSC2 plasmid DNA with anti-SSTR2 antibody for the expressed SSTR2 protein (C) and with anti-TUSC2 antibody for the expressed TUSC2 protein (D). Western blot results in A549-luc cell lines transfected with CMV-TUSC2-IRES-SSTR2, CMV-SSTR2, and CMV-TUSC2 plasmid DNA with anti-SSTR2 antibody for the expressed SSTR2 protein (E). The blot was stripped and then probed with anti-TUSC2 antibody for the expressed TUSC2 protein (F).
Immunofluorescence confirmed the above results. SSTR2 expression was seen in HT1080 cells transfected with CMV-TUSC2-IRES-SSTR2 or CMV-SSTR2 plasmid DNA (Figure 1C). In contrast, no expression was observed in cell lines transfected with CMV-TUSC2 plasmid DNA. Similar TUSC2 expression was seen in HT1080 cell lines transfected with CMV-TUSC2-IRES-SSTR2 or CMV-TUSC2 plasmid DNA (Figure 1D). In contrast, no expression was observed in HT1080 cells transfected with CMV-SSTR2 plasmid DNA.
As with HT1080 cells, Western blotting demonstrated similar SSTR2 expression in A549-luc cells transfected with CMV-TUSC2-IRES-SSTR2 or CMV-SSTR2 plasmid DNA. In contrast, no expression was observed in cell lines transfected with CMV-TUSC2 plasmid DNA (Figure 1E). Similar TUSC2 expression was seen in A549-luc cell lines transfected with CMV-TUSC2-IRES-SSTR2 or CMV-TUSC2 plasmid DNA. In contrast, no expression was observed in A549-luc cells transfected with CMV-SSTR2 plasmid DNA (Figure 1F).
Biodistribution in Lung Tumors
We assessed whether SSTR2 reporter expression after plasmid nanoparticle systemic delivery could be detected in lung tumors by biodistribution analysis. Animals with lung tumors were injected with plasmid DOTAP:cholesterol nanoparticle and then 111In-octreotide. Ex vivo biodistribution analysis illustrated significantly greater uptake in tumors from mice injected with CMV-TUSC2-IRES-SSTR2 (p < .001; Figure 2A) compared to control plasmid, CMV-TUSC2. Similarly, compared to the control, there was significantly greater uptake in tumors from mice injected intravenously with CMV-SSTR2 (p < .05). There was no significant difference between tumors from mice injected intravenously with CMV-SSTR2 and CMV-TUSC2-IRES-SSTR2. There was no statistically significant difference in the organs of test versus control groups of mice injected intravenously with the different plasmids (Figure 2B).

Biodistribution demonstrating SSTR2 expression in NSCLC lung tumor metastases after systemic plasmid SSTR2-based gene transfer. Biodistribution analysis illustrates significantly greater uptake of 111In-octreotide in lung metastases of mice injected intravenously with CMV-SSTR2 (*p < .05, n = 6) or CMV-TUSC2-IRES-SSTR2 (**p < .001, n = 6) plasmid DNA compared to control plasmid DNA, CMV-TUSC2. There was no significant difference between CMV-SSTR2 and CMV-TUSC2-IRES-SSTR2 (A). The graph including organs and tumors shows no significant difference in biodistribution among organs (B) of mice injected intravenously with CMV-SSTR2, CMV-TUSC2-IRES-SSTR2, or CMV-TUSC2 plasmid DNA, whereas differences among tumors are as above in A.
In Vivo Imaging
We assessed whether SSTR2 reporter expression after plasmid nanoparticle systemic delivery could be detected in lung tumors by in vivo imaging. Representative bioluminescent images of mice before they received plasmid nanoparticles (Figure 3, A–C) demonstrate lung tumors. Representative planar gamma-camera images demonstrate increased uptake in the lungs of mice injected intravenously with CMV-SSTR2 or CMV-TUSC2-IRES-SSTR2 plasmid DNA in DOTAP:cholesterol nanoparticles compared to CMV-TUSC2 plasmid DNA in DOTAP:cholesterol nanoparticles (Figure 3, D–F). SPECT/CT imaging of SSTR2 gene expression in representative mice (Figure 4) demonstrates expression localized in lung tumors in mice injected with CMV-TUSC2-IRES-SSTR2 and CMV-SSTR2 in DOTAP:cholesterol nanoparticles. Eighty percent of the lung nodules visible by CT in mice that received CMV-SSTR2 or CMV-SSTR2-TUSC2 were seen by SPECT imaging. Sizes ranged from 1 to 3 mm. As expected, in the CMV-TUSC2 group, lung tumors are seen, but no uptake was noted in the tumors. After imaging, the mice were sacrificed for ex vivo biodistribution analysis, and, again, greater expression was seen in tumors from mice injected with CMV-TUSC2-IRES-SSTR2 or CMV-SSTR2 compared to the control plasmid, CMV-TUSC2 (p < .001, n = 3; Figure 4D).

Gamma-camera images demonstrating SSTR2 expression in NSCLC lung tumor metastases after systemic SSTR2-based plasmid gene transfer. Representative bioluminescence luciferase imaging for monitoring tumor development demonstrated lung tumors in groups (A–C) that were then injected with plasmid nanoparticles and, subsequently, 111In-octreotide. Representative planar images demonstrate increased uptake (arrows) in the thorax of mice with NSCLC lung metastases injected systemically with CMV-SSTR2 (D) or CMV-TUSC2-IRES-SSTR2 (E) compared to control CMV-TUSC2 plasmid DNA (F). Mice bearing lung tumors and exposed to plasmid systemically 2 days earlier were injected intravenously with 111In-octreotide (11 MBq) and imaged 24 hours later.
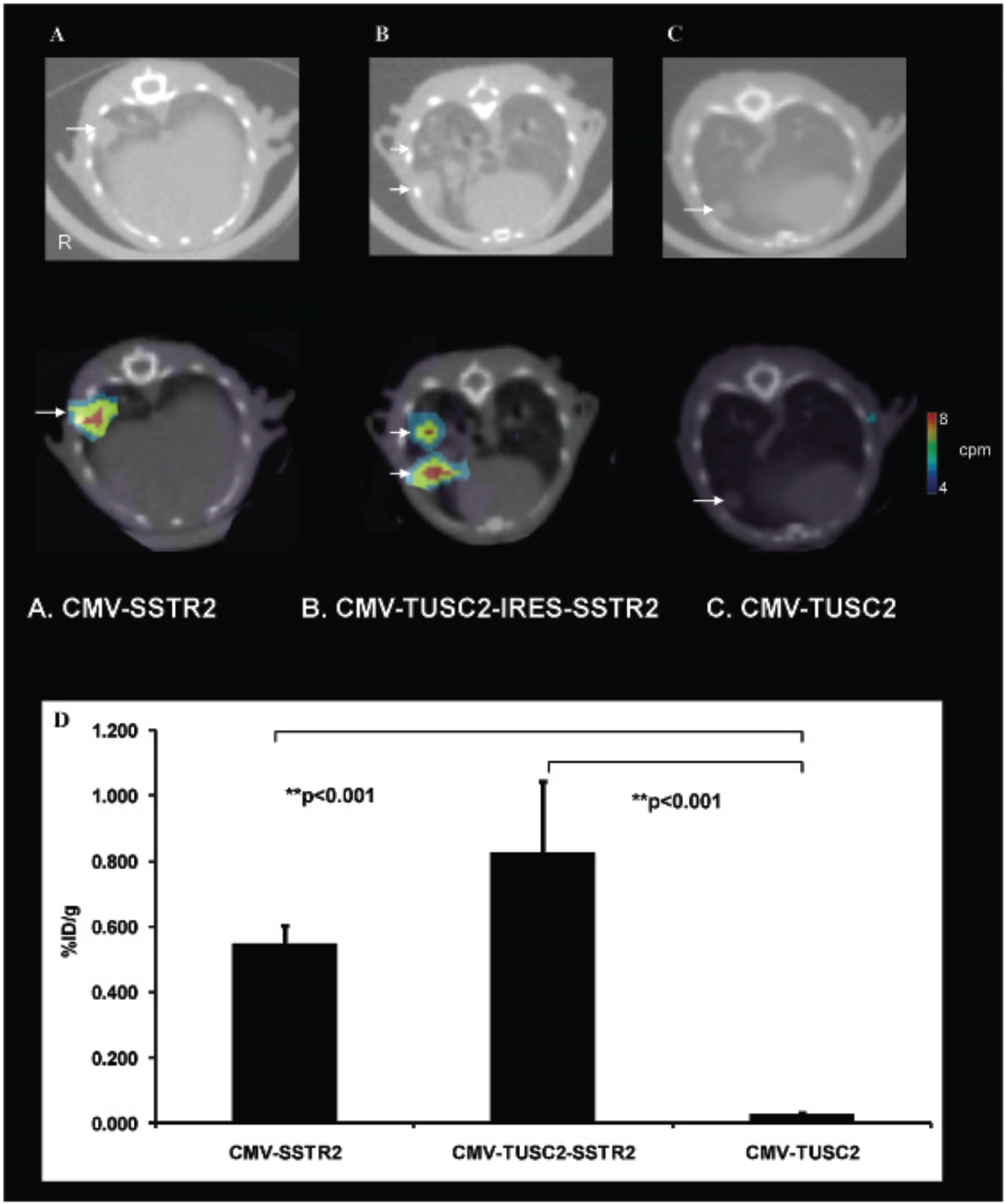
SPECT/CT images demonstrating SSTR2 expression in NSCLC lung tumor metastases after systemic SSTR2-based plasmid gene transfer. Representative slices of three-dimensional, tomographic SPECT/CT images demonstrate increased uptake in the lung tumors of mice with NSCLC lung metastases injected systemically with CMV-SSTR2 (A) or CMV-TUSC2-IRES-SSTR2 (B) compared to CMV-TUSC2 plasmid DNA (C). Top: Lung windows demonstrating lung tumors (arrows). Bottom: Soft tissue windows with SPECT overlay. Biodistribution analysis (D) illustrates significantly greater uptake of 111In-octreotide in lung metastases of mice injected intravenously with CMV-SSTR2 (**p < .001, n = 3) or CMV-TUSC2-IRES-SSTR2 (**p < .001, n = 3) plasmid DNA compared to control plasmid DNA, CMV-TUSC2. There was no significant difference between CMV-SSTR2 and CMV-TUSC2-IRES-SSTR2. Mice bearing lung tumors and exposed to plasmid systemically 2 days earlier were injected intravenously with 111In-octreotide (13 MBq) and imaged 24 hours later.
Immunohistochemical Staining
Immunohistochemical analysis confirmed SSTR2 expression in tumors from mice injected intravenously with CMV-SSTR2 plasmid and in tumors from mice injected intravenously with CMV-TUSC2-IRES-SSTR2 plasmid (Figure 5). In contrast, no expression was observed in tumors in mice injected intravenously with CMV-TUSC2 plasmid. Immunohistochemical evaluation with anti-TUSC2 antibody showed TUSC2 expression in tumors from mice injected with CMV-TUSC2 plasmid and in tumors from mice injected with CMV-TUSC2-IRES-SSTR2 plasmid. In contrast, no expression was observed in tumors in mice injected with CMV-SSTR2 plasmid (see Figure 5). Apoptosis staining followed TUSC2 staining (see Figure 5). Thus tumors from mice injected with CMV-TUSC2-IRES-SSTR2 plasmid expressed both TUSC2 and SSTR2.

Immunohistochemical staining demonstrating SSTR2 expression in NSCLC lung metastases after systemic SSTR2-based plasmid gene transfer. Representative images show expression of SSTR2 (brown staining) in NSCLC metastases from mice injected systemically with CMV-SSTR2 (A) or CMV-TUSC2-IRES-SSTR2 (B) but not control CMV-TUSC2 (C) plasmid DNA. Images do not show expression of TUSC2 (brown staining) in NSCLC metastases from mice injected systemically with CMV-SSTR2 (D) plasmid DNA as expected but do show TUSC2 expression in NSCLC metastases from mice injected systemically with CMV-TUSC2-IRES-SSTR2 (E) or CMV-TUSC2 (F) plasmid DNA. Images do not show expression of apoptosis (brown nuclear staining) in NSCLC metastases from mice injected systemically with CMV-SSTR2 (G) plasmid DNA as expected but do show TUSC2 expression in NSCLC metastases from mice injected systemically with CMV-TUSC2-IRES-SSTR2 (H) or CMV-TUSC2 (I) plasmid DNA (X200 original magnification).
Discussion
The data demonstrate that plasmid-based in vivo delivery to multiple lung metastases from NSCLC can be assessed using SSTR2-based reporters, including when such reporters are linked to a therapeutic gene, such as TUSC2. NSCLC is commonly metastatic at the time of detection. Thus, methods for systemic delivery to numerous tumors are needed. Among vectors, plasmids are advantageous in that they have a low rate of integration into chromosomal DNA,24,25 suggesting increased safety. Lipids, such as those that are DOTAP:cholesterol based have been used to increase delivery and expression. However, plasmids commonly result in relatively low levels of expression that may limit detection in vivo.
The amount of expression with plasmids is usually significantly lower than with most viral vectors,24,25 presenting a challenge to reporter systems. In addition, for eventual human use, reporter systems that can be employed for percutaneous imaging in patients are needed. Light-based methods are prone to scatter, limiting their use in such settings. Instead, reporters that can be imaged using translatable methods, such as by nuclear medicine, are required.
In our recently published study, we found that it is possible to visualize and noninvasively quantify SSTR2-based reporter gene transfer in NSCLCs after intratumoral adenovirus injection. 13 It had been thought that NSCLCs endogenously express SSTR2, which would confound reporter-based imaging with SSTR2-based reporters. However, the previous study found that SSTR2 was not or was minimally expressed in the great majority of the 70 human NSCLC tumor samples. 13 Furthermore, even in cell lines that endogenously expressed other SSTR subtypes, exogenous SSTR2 gene expression could be detected, including in vivo. 13 The previous study used adenoviral vectors and intratumoral injection. The current study demonstrates imaging in metastatic tumors in the lungs after systemic plasmid delivery using a SSTR2-based reporter system.
Reporter genes may be used alone to assess expression or may be linked to therapeutic genes. Clinically, therapeutic genes are necessary to generate desired phenotypic change. The linkage may inhibit expression, or the downstream phenotypic change has the potential to confound imaging of the reporter. Moreover, among methods for linking two genes,26–28 a common method for linking expression is placing an IRES29,30 between the gene of interest and the reporter. Although this results in a ratio of expression of the first and second gene, it often results in decreased expression of the second gene.31–33 Thus, it was of interest to investigate the effects of linking SSTR2 to a therapeutic gene to determine if SSTR2 expression continues to be visualized with an additional gene. Both the reporter gene and the therapeutic gene could be expressed and detected in vitro and ex vivo.
The therapeutic gene used in this study, TUSC2, is a novel tumor suppressor gene that was identified in the human chromosome 3p21.3 critical region, where allele losses and genetic modifications occur both early and frequently in many human cancers.17,19,34,35 Studies have suggested that TUSC2 protein expression is commonly diminished or absent in many types of cancer, including lung and breast cancers.17,19,36,37 It was previously found that when the function of TUSC2 in 3p21.3-deficient NSCLC cells was restored, tumor growth inhibition occurred as a result of apoptosis induction and cell cycle disruption.17,19,21 A problem with therapeutic genes is that they could induce inflammation and necrosis, which could potentially accrue the radiopharmaceutical and confound imaging. Significantly, although we used a therapeutic gene whose product induces apoptosis and placed this therapeutic gene first before the IRES and SSTR2 after the IRES, SSTR2 expression could be visualized on systemic plasmid-based gene transfer.
Plasmids tend to be nonintegrating, but their expression levels tend to be low and naked plasmids tend to express only in muscle. To increase plasmid expression in vitro, often lipid-based carriers are used. Such a strategy has shown success in vivo using DOTAP:cholesterol, including in a clinical trial for expressing the tumor suppressor gene TUSC2 that demonstrated low toxicity. 22 Compared to other lipids for gene delivery, DOTAP:cholesterol has less binding to serum proteins and less uptake by RES organs such as the liver and spleen.38,39 Moreover, SSTR2 expression was noted to occur more so in tumors than in normal surrounding lung, enabling imaging of tumors. Although greater expressions in living tumors than normal tissue may be due to greater susceptibility by the tumors, it is likely also due to particle formation by the DOTAP:cholesterol plasmid complex, resulting in the enhanced permeability and retention (EPR) effect. The EPR effect has been seen with multiple nanoparticles 40 and can be capitalized on to direct delivery to tumors. This has the potential for systemic delivery to multiple metastases, such as to the lung. Adenoviruses may prove more useful localized delivery since intravenous injection commonly results in expression in the liver.
The present study found SSTR2 expression in multiple metastatic NSCLC tumor xenografts using gamma-camera and SPECT/CT imaging after CMV-TUSC2-SSTR2 plasmid with DOTAP:cholesterol nanoparticles was given systemically (intravenously). Biodistribution analysis confirmed the imaging findings. Additionally, immunohistochemistry further illustrated SSTR2 and TUSC2 expression in the tumor metastases. Thus, plasmid-based in vivo delivery to multiple lung metastases of a NSCLC model can be visualized using SSTR2-based reporters, including when such reporters are linked to a therapeutic gene.
Footnotes
Acknowledgment
Financial disclosure of authors: This work was supported by National Institutes of Health/National Cancer Institute SBIR R41-CA 114924, RO1-CA116322, RCA129714, and CA016672. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. J.A.R. has been a consultant to, received grants from, and has stock in Genprex, Inc. Patents pending and issued: TUSC2/FUS1 NST: Novel DNA:liposome complexes for increased systemic delivery and gene expression. United States Letters Patent No. 6,413,544 B1 issued July 2, 2002. United States Patent No. 6,770,291 B2 issued August 3, 2004. International Publication Number: WO 98/07408. The institution holds patents regarding using SSTR2-based reporters.
Financial disclosure of reviewers: None reported.
