Abstract
To improve spatial resolution in in vivo bioluminescence imaging, a photon scattering correction, image restoration method was tested. The chosen algorithm was tested on in vivo bioluminescent images acquired on three representative tumor models: subcutaneous, pulmonary, and disseminated peritoneal. Tumor size was chosen as a quantitative criterion, such that the tumor reference measurements (determined photographically or by computed tomography) were compared to those derived from bioluminescent images, before and after restoration. This technique allowed a significant reduction to be achieved in the relative error between reference measurements and dimensions derived from bioluminescent images. In addition, improved delineation of the tumor foci was achieved. The restoration method allows spatial resolution in bioluminescence imaging to be improved, with interesting perspectives in terms of staging and quantitation in experimental oncology.
BIOLUMINESCENCE IMAGING (BLI) is a real-time noninvasive technique that is used increasingly commonly in biomedical and pharmaceutical research for the monitoring of therapeutic response to novel drug candidates.1–3 BLI is based on the noninvasive detection by a charge-coupled device (CCD) sensor of photons emitted by the oxidation reaction of intraperitoneally administered luciferin, by luciferase-expressing cells in the living animal. Nevertheless, for routine applications of this technique, scattering and absorption of photons by the tissues lead to a global degradation of performance, in particular when quantitative imaging is required. Indeed, spatial resolution is poor as a consequence of tissue scattering, such that when the tumor foci are close to one another, the bioluminescent signals can merge and appear to originate from a single focus. In addition, a deeply located focus will appear larger than an equivalent one located just beneath the skin.
Several scattering correction strategies have been reported. One proposes to modify the skin's optical properties using hyperosmotic agents such as dimethyl sulfoxide. 4 However, this method has a significantly decreased sensitivity and suffers from obvious limitations in biologic studies. The other strategies are used to perform bioluminescence tomography. Some of them are based on the photon diffusion model.5,6 They require knowledge of the depth of each luminescent source inside the body (it can be determined with a multi-spectral approach 7 ) and the characterization of all tissues interacting with the photons between the tumor and the skin, but in two-dimensional BLI, these informations are not determined. The others use diffuse optical tomography, a tool to detect and reconstruct the optical properties inside highly scattering media. 8 The approach described in this article can potentially be applied on images acquired with any two-dimensional bioluminescent system.
At least two phenomena are responsible for the poor spatial resolution observed in BLI: tissue scattering and image degradation within the imaging system itself. Indeed, the overall image acquisition system can be characterized by a transfer function referred to as the point spread function (PSF), such that the acquired image generated by a luminous point source within the body is a blurred spot rather than a single point. Because any given optical system can be characterized by a representative PSF, it is possible for an estimation of the latter to be used to correct for this blurring phenomenon, thereby allowing sharper, more realistic images to be obtained. The raw image produced by the optical system can be expressed as the convolution of the nondegraded image with the PSF. As the inverse operation, that is, deconvolution of the recorded image, is difficult to achieve because both the PSF and the additive noise need to be determined, a different strategy was chosen. This consisted of the use of restoration algorithms to recover the original light sources. 9 An initial study was carried out to determine whether an image restoration process could be used to improve the spatial resolution. By comparing several restoration methods using synthetic images, the blind deconvolution method was found to be the most suitable for our application. 10
The aim of the present study was to apply this restoration method to in vivo acquisitions of bioluminescent tumors, to validate the algorithm using slices of computed tomographic (CT) images, or tumor photography, as reference images, and to evaluate the algorithm's effectiveness in restoring bioluminescent images. Four tumor models were used. First, because the subcutaneous tumor is the most widely used model, we selected a subcutaneously implanted glioma that had a delineated edge to verify that the corrected images corresponded to the reality. Lung metastasis is a common event during the progression of a variety of cancers, and it is of great interest to achieve early detection and staging inside such an organ, characterized by a high scattering coefficient in addition to respiratory motion. To model such a situation, a lung carcinoma was orthotopically implanted, the reference sizing method being CT. The third model was a peritoneal carcinoma with foci located on different organs at varying ranges beneath the skin, which is representative of normal tumor progression in the abdominal cavity. The last model was an ovarian adenocarcinoma with foci located mainly on intestines with ascites in the abdominal cavity.
Material and Method
Cell Lines and Culture Conditions
The U87-MG tumor derived from surgical explants of a human malignant glioma and the NIH:OVCAR-3 tumor derived from surgical explants of a human ovarian adenocarcinoma were procured from American Type Culture Collection (Manassas, VA). U87-MG and NIH:OVCAR-3 cells were stably transfected with the firefly luciferase reporter gene to generate the U87-Luc and the OVCAR-3-Luc cell line (Institut de Recherche Pierre Fabre). The human large cell lung carcinoma NCI-H460-Luc2 cell line and the human colorectal adenocarcinoma cell line were procured from Caliper Life Sciences (Roissy, France). U87-Luc, HCT116-Luc2, and OVCAR-3-Luc cells were grown in a minimum essential medium (Gibco, Cergy-Pontoise, France) supplemented with 10% fetal calf serum, glutamine, 100 U/mL of penicillin, and 100 U/mL of streptomycin. Immediately before implantation into nude mice, the cells were trypsinized, washed with phosphate-buffered saline (PBS), and adjusted to a concentration of 2 × 107 cells per milliliter in PBS for subcutaneous or intraperitoneal implantation. The NCI-H460-Luc2 cells were grown in an RPMI-1640 medium (Gibco, Cergy-Pontoise, France) supplemented with 10% fetal calf serum,
Mice
Homozygous female athymic nude mice of the Swiss nu/nu strain, BALB/c nu/nu strain (Charles River, Saint-Germain-sur-l'Arbresle, France), and female athymic nude mice (Harlan, Gannat, France) were used. The animals were handled and cared for in accordance with the Guide for the Care and Use of Laboratory Animals (National Research Council, 1996) and the European Directive EEC/86/609, under the supervision of authorized investigators.
Mouse Models
Swiss nude mice (female, 12 weeks, n = 7) were used for the subcutaneous implantation of U87-Luc; 4 × 106 cells in 200 µL of PBS were injected under the skin on the right flank. Athymic nude mice (female, 6 weeks, n = 20) were used for the intraperitoneal administration of 4 × 106 HCT116-Luc2 cells in 200 µL of PBS or 5 × 106 OVCAR-3-Luc cells in 250 µL of PBS. BALB/c nude mice (female, 6 weeks, n = 3) were used for the intrabronchial implantation of NCI-H460-Luc2; 2.5 × 105 cells in 25 µL of NaCl 0.9% were implanted at a given bronchial location, under interventional x-ray monitoring.
Micro-CT Imaging
The mice were anesthetized with a continuous flow of 2% isoflurane (Aerrane, Maurepas, France), and their thorax was imaged with respiratory gating using the GE eXplore Locus system (GE Healthcare, USA). The isotropic resolution was 90 µm. The x-ray tube settings were 80 kV and 70 mA. The resulting raw data were reduced, and the reconstructed slices were output in the CT manufacturer's raw format, without being scaled to Hounsfield units. The reconstructed images were viewed and analyzed using MicroView software (GE, USA). Size measurements were determined on axial and coronal slices where the tumor was the tallest.
Bioluminescence Imaging
The mice were intraperitoneally injected with 125 mg/kg of luciferin (Promega, Paris, France) before undergoing anesthesia with an intraperitoneal injection of 100 mg/kg of ketamine hydrochloride (Imalgene 1000, Merial, London, UK) and 10 mg/kg of xylazine hydrochloride (Rompun 2%, Bayer, Deerfield, IT). BLI was performed 10 minutes later using a high-sensitivity CCD camera, cooled to −70°C (ORCA II BT C4742-98-26 LW, Hamamatsu Photonics, Herrsching, Germany). The field of view and the binning (1 × 1) were kept constant, as opposed to the exposure times, which were adjusted according to the signal strength to optimize the signal to noise ratio. In addition, cosmic ray events recorded by the CCD camera were subtracted. The resulting images had a resolution of 512 × 512 pixels, coded onto 16 bits, with a spatial resolution of 300 µm.
Restoration of In Vivo Bioluminescent Images
On the basis of previous results,
10
the blind deconvolution method was used to restore the bioluminescent images. With this method, accurate a priori knowledge of the PSF is not necessary but allows the algorithm to converge more quickly. As the bioluminescent sources are always located below the skin, the PSF of the system was determined, including the skin. An image was acquired using a point light source, produced by a 200 µm diameter optical fiber (smaller than the pixel size), positioned under the skin of an anesthetized mouse. This optical fiber was connected to a xenon light source (ELS-2 xenon arc automatic light source, Solos Endoscopy, Boston, MA), using a customized bandpass filter to produce the same spectrum as that emitted by firefly luciferin (Figure 1).
Schematic drawing of the experimental setup used to determine the point spread function of the imaging system, including the mouse's skin. A bioluminescent point source was simulated using bandpass (BP) filtered light delivered via a 200 µm optical fiber. CCD 5 charge-coupled device camera; PC 5 personal computer.
Validation with an In Vivo Model
To verify the performance of the previously selected restoration algorithm with synthetic images, dimensional measurements were made on bioluminescent images (before and after correction) and were then compared to reference measurements. The latter were determined using either suitably scaled photographic images of HCT116-Luc2 or U87-Luc tumors after skin removal or CT images of a NCI-H460-Luc2 lung tumor. The relative errors were computed by comparing the real dimensions (determined on the tumor) with the measured values derived from the images, according to the following expression:
With a similar approach, surface area analysis of the subcutaneous U87-Luc acquisitions was carried out using an edge detection tool as described by Chan and Vese 11 on photographs and bioluminescent images. This method could not be applied to the other two models owing to their deeper location.
Results
Determination of the PSF
The PSF of the imaging system, including the skin, was determined with the experimental setup shown in Figure 1. The (200 µm) point source placed under the skin produced an image corresponding to a blurred spot, approximately 4 mm in diameter (Figure 2B) and larger than when the source was placed directly under the CCD camera (Figure 2A). The pixel matrix of the obtained image corresponded to the PSF and was included in the blind deconvolution algorithm used to restore the bioluminescent images.
Images acquired with the punctual light source placed on the stage (A) or under the skin (B) and profile of the recorded image (C).
In Vivo Bioluminescence Image Restoration
The restoration process was first tested on subcutaneous U87-Luc tumors. The original and restored bioluminescent images are shown in Figure 3. To validate the performance of the algorithm, several size determinations were made on the basis of photographic images of tumors and then compared to the bioluminescence images. The average relative error between the real value and the initial bioluminescent image was 0.37 ± 0.22, whereas the relative error achieved after image restoration decreased to 0.05 ± 0.03 (n = 17 measurements). Comparisons were also made between the surface area of the tumors, determined photographically (reference values), and the bioluminescent images, before and after restoration. The results are summarized in Table 1. This shows that a significant decrease in relative error is achieved after restoration (1.0 ± 0.3 vs 0.04 ± 0.02).
Photographs of subcutaneous U87-Luc tumors (used to determine tumor size) and in vivo bioluminescent images before and after restoration with the edges detected by the algorithm.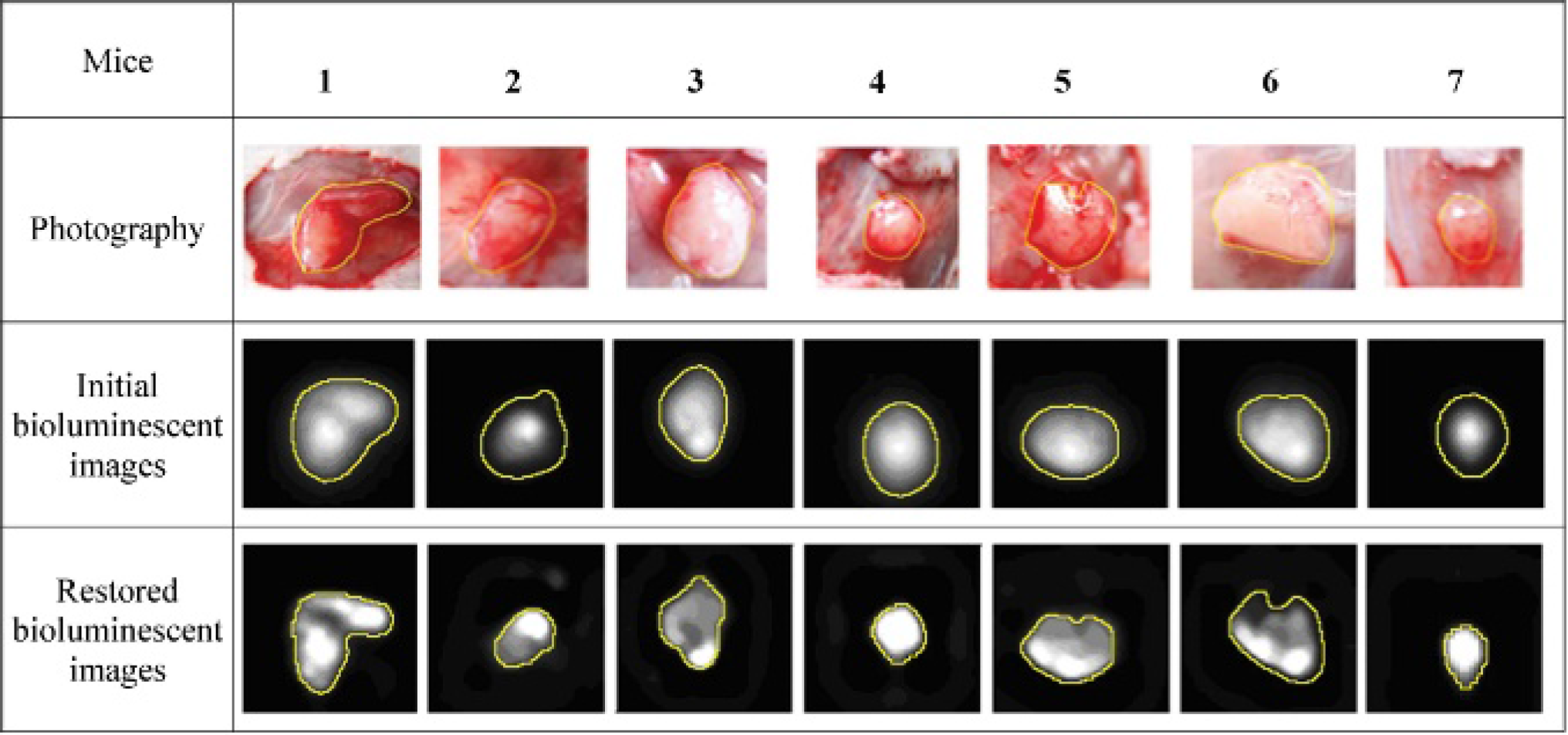
Discussion
In this study, we investigated the use of an image restoration method to improve the spatial resolution of in vivo BLI.
Surface Area Measurements of Subcutaneous U87-Luc Tumors

Actual surface area: area measured on a photograph of each tumor.
Surface areas 1 and 2: surface areas measured on the initial and restored bioluminescent images, respectively.
Relative errors 1 and 2: errors between the actual surface area and surface areas 1 and 2, respectively.
When applied to a NCI-H460-Luc2 intrabronchial tumor, the average relative error between the measurements taken from the CT images (Figure 4) and the initial bioluminescence images was 0.72 ± 0.33, whereas after restoration, the relative error decreased to only 0.12 ± 0.16 (n = 6 measurements). Figure 5 represents the bioluminescent images of an intrabronchial tumor before and after the restoration process.
Coronal CT slice of thorax allowing size determination of an intrabronchial NCI-H460-Luc2 tumor (indicated by the circle). Bioluminescent images of intrabronchial NCI-H460-Luc2 tumors before (A) and after (B) image restoration with the same intensity scale.

The method was applied to HCT116-Luc2 intraperitoneal tumors (Figure 6). It can be seen in this figure that the tissue scattering correction improves the sharpness of the imaged bioluminescent foci. In this model, as it was difficult to measure all of the tumor foci owing to their adherence to the skin, the relative errors were not computed. Nevertheless, when the bioluminescent images were compared to the measurements recorded during the autopsy, they were found to be significantly closer to the reality after the image restoration algorithm had been applied.
Bioluminescent images of intraperitoneal HCT116-Luc2 tumors before (A) and after (B) image restoration with the same intensity scale.
Finally, the method was applied to OVCAR-3-Luc tumors (Figure 7), and the restoration improved the visualization of the tumor foci. Before the restoration, it seemed that the signal was present in the entire abdominal cavity, whereas after the correction, it appeared that this was not the case. The restored signal was consistent with the observations performed during the autopsy.
Bioluminescent images of intraperitoneal OVCAR-3-Luc tumors before (A) and after (B) image restoration with the same intensity scale.
In addition, all images were quantitated before and after correction to assess the impact of the restoration on the signal intensity. We showed that the restoration had no significant impact on the quantitation (n = 15 measurements) according to the results obtained with synthetic images. 10
To assess the quality of the restored bioluminescent images, size measurements of tumor foci were used as quantitative criteria, as well as the areas, which are easily comparable on tumor photographs and bioluminescent images and not operator dependent thanks to an automatic edge detection tool. First, we applied the blind method without supplying the specific PSF of the system. The resolution was better, but this improvement was not sufficient even after 5,000 iterations. So, to improve the quality of the restoration, an experimental PSF including the detector and the skin was determined on nude mice but equally on BALB/c mice. In these conditions, this restoration method appeared to be equally applicable to mice with white fur. To measure experimental PSF, we used a system composed of a xenon lamp source emitting a filtered light and an optic fiber. This device is not usually available in the laboratory using BLI, so, to apply this approach, an alternative could be to use a set of theoretical PSFs corresponding to different gaussian curves. Each PSF would be tested on a representative situation (ie, a well-delineated subcutaneous tumor) and the best suited one used to restore all the images.
The results showed that measurements taken from restored images were closer to the reality than those based on raw images. This technique was found to be particularly efficient for subcutaneous tumors. It appeared to have a slightly decreased performance for the observation of intrabronchial tumors as a consequence of two factors known to complicate lung imaging in biophotonics: a high scattering coefficient together with lung blurring resulting from both respiratory motion and beating of the heart. The results would be better if we applied a specific PSF corresponding to the lung, but it is very difficult to determine a specific PSF for the lung owing to the risk of pneumothorax. Furthermore, it is not realistic to determine the PSF for each location in the mouse, and it would not be strictly applicable because in two-dimensional BLI, we don't know the exact location of the source. Even if this approach does not entirely resolve the problem of scattering, it allows obvious improvement of the resolution. Recording images with sharper edges, together with an improved resolution of the tumoral structure (see Figure 3, tumors 1, 2, and 6, and Figure 5), allows essential characteristics to be assessed, from which the tumor's sensitivity to a given treatment can be determined. As an example, this approach makes it possible to improve the visualization of tumor heterogeneities and areas of hypoxia (better delineation of hypoactivity area using conventional luciferin) or apoptosis (by using caspase 3–processed luciferin 12 ).
For lung cancer and a variety of other primary tumors, this technique could allow the detection of the presence of a ganglionic extension in the vicinity of the primary tumor, which is a crucial parameter in the staging of the disease. This improvement could be of great value for pharmacologic studies when it is necessary to constitute homogeneous groups of animals with the same pattern of tumor dissemination (lymph nodes, metastasis, or local extension of the primary tumor).
In the case of disseminated tumors (HCT116-Luc2), the objective of BLI is usually to identify the different foci and to assess the level of proliferative activity, based on the quantitation of emitted photons. In such cases, it is not relevant to quantify the global light emission within the mouse because the foci are located at different positions throughout the body, and the tissue–photon interactions are not similar. Accurate assessment requires the sum of the photons emitted by each individual focus to be computed, following correction for tissue absorption by spectral analysis. 13 The use of scattering corrections should improve the focus delineation process.
The size measurements realized on restored bioluminescent images are much closer to reality at the time this parameter is used to assess the accuracy of the restoration method. However, the goal of BLI is not to perform sizing but to determine proliferative activity, so better delineation could improve the global reliability. Furthermore, as for the OVCAR model without the scattering correction, it was not possible to determine where the bioluminescent cells were: in the entire abdomen (eg, in the ascites) or on the intestines. So this correction is important to study more precisely the proliferation without euthanizing the animals. In addition, if we acquire bioluminescent images with multiple views (eg, dorsal and lateral views) and correct them from the scattering, it will be possible to discriminate superimposition of multiple tumor foci to improve their location and their count, which constitutes an important parameter for assessing tumor progression.
Footnotes
Acknowledgment
Financial disclosure of authors and reviewers: None reported.

