Abstract
Endogenous and exogenous somatostatin receptors are commonly targeted for imaging using radiopharmaceutical analogues of somatostatin. Ligand binding activates receptor-mediated signaling. We assessed whether somatostatin receptor type 2A (SSTR2A) imaging can be uncoupled from signal transduction. In both human fibrosarcoma (HT1080) and human embryonic kidney (HEK293) cells, reverse transcriptase–polymerase chain reaction and enzyme-linked immunosorbent assay found similar levels of expression of hemagglutinin A tagged SSTR2A (HA-SSTR2A) or the same fusion protein with a deletion of the C-terminus beyond amino acid 314 (HA-SSTR2Δ314). Scatchard analysis demonstrated similar degrees of ligand binding by the wild-type or mutant receptor to 111In-octreotide in both cell pairs. Cyclic guanosine monophosphate (cGMP) production and inhibition of forskolin-induced cylic adenosine monophosphate (cAMP) production were evaluated at the signaling level, and growth inhibition was evaluated at the cellular level before and after stimulation. Unlike wild-type receptor, HA-SSTR2Δ314 was deficient in inhibiting forskolin-induced cAMP production (p < .05) and in inciting cGMP (p < .05) production; furthermore, at the cellular level, HA-SSTR2Δ314 was deficient in inhibiting cell proliferation (p < .05). Yet tumors expressing HA-SSTR2Δ314 could be imaged in vivo. Thus, in vivo imaging of SSTR2 can be uncoupled from cAMP and cGMP signaling as well as growth inhibition.
For somatostatin, there are six receptor types: 1, 2A, 2B, 3, 4, and 5. They belong to the family of G protein–associated receptors that contain seven alpha-helical transmembrane domains. Types 2A and 2B are alternate splice variants that are identical, except for the intracytoplasmic C-terminus. 8 On activation, somatostatin receptor type 2 (SSTR2) regulates signaling such as cyclic adenosine monophosphate (cAMP) 9 and cyclic guanosine monophosphate (cGMP) production. The latter appears to regulate cell proliferation. 10
Among the receptor subtypes, human type 211–15 has the highest affinity for the most common clinically used somatostatin imaging analogue, 111indium-labeled octreotide. This radiopharmaceutical and 99mtechnetium-labeled analogues are used in clinical practice to detect tumors that endogenously overexpress somatostatin receptors 1 and have been used in animal models to image tumors that express exogenously introduced SSTR2, for example, as a reporter gene. In many potential clinical applications of reporter imaging, signaling is not desired because it may negatively impact cellular function and/or interfere with a linked therapeutic gene.
There are two clinically approved somatostatin receptor type (SSTR) imaging agents. Expression of SSTR2 can be localized and quantified in vivo using 111In-labeled octreotide. 16 Clinically, the 111In-labeled analogue better distinguishes neuroendocrine tumors, especially in the liver, than the 99mTc-labeled agent 17 and has more favorable biodistribution for imaging in mice. 18 The 99mTc-labeled agent has more equal affinity for SSTR3, −5 and −2 19 compared with the 111In-labeled analogue. Alterations in imaging agent design and labeling can have major effects on receptor specificity and on the routes and rates of elimination,20,21 which may confound interpretation of downstream signaling.
Instead of using various ligands, for evaluating signaling, the receptor itself may be mutated. For somatostatin receptors, signaling may be mediated through the C-terminus of the receptor.9,22 In this study, strategically, we created cell-line pairs expressing similar amounts of wild-type or mutant receptors and tested the ability of the mutant receptor to elicit signaling on ligand binding. We tested the competency of the signaling deficient receptor for binding 111In-octreotide in vitro and for imaging in vivo to assess whether somatostatin receptor type 2A (SSTR2A) imaging can be uncoupled from signal transduction.
Materials and Methods
Cloning
Plasmids for SSTR1, −4, and −n5 were obtained from the Donald Guthrie Foundation for Education and Research Inc. (Guthrie cDNA Resource Center, Sayre, PA) and for SSTR3 from the American Type Culture Collection (ATCC, Rockville, MD). Human SSTR2A and the deletion mutant (SSTR2Δ314) were constructed from a phage (ATCC) containing the SSTR2A insert by polymerase chain reaction using the following oligonucleotide primers: SSTR2 FL-F (TCC CCG CGG CAT GGA CAT GGC GGATGA), SSTR2 FL-R (AAT CTG CAG CTG TCA GAT ACT GGT TTG GAG), and SSTR2 T71B-R (AGA AGG CAT ATA GGA TAG GGT TGG CAC AGC). The full-length SSTR2A and SSTR2Δ314 inserts were ligated into the Sac II and Pst I inserts in pDisplay vector (Invitrogen, Carlsbad, CA) separately. The inserts were placed downstream of the membrane localization sequence (5' Igκ leader) and the sequence for the hemagglutinin A (HA) epitope tag. The inserts were confirmed by sequencing.
Cell Lines
HT1080 (human fibrosarcoma, ATCC) and HEK293 (human embryonic kidney, ATCC) cells were maintained in Dulbecco's Modified Eagle's Medium (DMEM) supplemented with 10% fetal bovine serum and 1× penicillin-streptomycin mixture. Adherent cultures were incubated at 37°C in 5% CO2. For transfection of HEK293 cells, 2 μg deoxyribonucleic acid (DNA) mixed with 3 μL FuGENE 6 (InvivoGen, San Diego, CA) transfection reagent in 50 μL DMEM was added to 1 × 105 cells in 24-well plates per the manufacturer's instructions. For HT1080 cells, lipofectin (Invitrogen) was used per the manufacturer's instructions. 23 Cells were selected with 1 mg/mL G418.
Enzyme-Linked Immunosorbent Assay and Immunofluorescence
Enzyme-linked immunosorbent assay (ELISA) and immunofluorescence were performed as previously described. 23 Prospective colonies expressing fusion protein were screened by ELISA, and expression in selected clones was quantified using a quantitative ELISA; 25 mU/mL horseradish peroxidase conjugated rat anti-HA antibody (clone 3F10 from Roche, Indianapolis, IN) was used for ELISA. Indirect immunofluorescence employed an anti-HA primary antibody (HA-11, Babco, Richmond, CA).
Reverse Transcriptase–Polymerase Chain Reaction
Total ribonucleic acid (RNA) was extracted with TRIzol Reagent (Invitrogen), and 4 μg was used in each reverse transcriptase–polyerase chain reaction (RT-PCR) using a Superscript One-Step RT-PCR kit (Invitrogen) in a thermal cycler (Eppendorf, Westbury, NY). The conditions were one cycle each (30 minutes at 50°C and 2 minutes at 94°C) for complementary DNA synthesis and denaturation, 35 cycles (15 seconds at 94°C, 30 seconds at 60°C, and 1 minute at 72°C) for PCR amplification, and one cycle (5 minutes at 72°C) for final extension and included 0.2 μM primers and 2 units of taq in EcoTaq buffer. The primer pairs used were SSTR2-FL-F: ATGGACATGGCGGATGAGCCACTCAATGG with SSTR2-FL-R: TCAGATACTGGTTTGGAGGTCTCCATTGAG for the wild-type receptor transcript and SSTR2-Fl-F with SSTR2-Δ314-R: AGAAGGCATATAGGATAGGGTTGGACAGC for transcripts coding up to amino acid 314. Primers described previously for SSTR1, −3, −4, and −5, 24 as well as SSTR2-FL-F and SSTR2-Δ314-R, were used for RT-PCR of untransfected HEK293 and HT1080 cells and SSTR1 to SSTR5 plasmid controls. Primers for β-actin were used to confirm the quality of cellular complementary DNA. Ten microliters of each reaction underwent electrophoresis on a 1% agarose gel that was stained with ethidium bromide and visualized under ultraviolet light. Each group was repeated at least twice.
Binding
Cells were harvested in binding buffer (50 mM Tris-HCl, pH 7.8, containing 1 mM EGTA, 5 mM MgCl2, 10 μg/mL leupeptin, 10 μg/mL pepstatin, 200 μg/mL bacitracin, and 0.5 μg/mL aprotinin) and were centrifuged in a microfuge at 12,000 rpm for 10 minutes at 4°C. The pellet was homogenized with a Polytron (Omni International, Marietta, GA) at 50 Hz for 20 seconds in binding buffer. Five micrograms per well of the cell membrane preparation was used for the radioligand binding studies. 25 Glass fiber filter/B type (GF/B) multiscreen plates (Millipore, Bedford, MA) were pretreated with 100 μL of 0.1% polyethyleneimine per well for 2 hours. After removing the solution using a multiscreen vacuum manifold (Millipore), plates were washed once with 200 μL of 50 mM Tris-HCI (pH 7.8). Next, assay reagents were added to triplicate wells in the following order: 160 μL cell membrane and 40 μL (200 nM to 0.7 nM final concentration) 111In-octreotide (Octreoscan, Mallinckrodt, St. Louis, MO) or 160 μL cell membrane, 20 μL unlabeled ligand (1 μM final concentration of unlabeled somatostatin), 20 μL 111In-octreotide (200 nM to 0.7 nM final concentration). After a 40-minute incubation at room temperature, the wells were washed two times with 350 μL of unlabeled 50 mM Tris-HCl (pH 7.8). The radioactivity of each filter punched out from the plates was quantified with a gamma counter. Scatchard analysis was performed.
Internalization
For radioligand uptake, 30,000 cells/well were exposed to 10−7 M 111In-octreotide (Octreoscan) with or without 10−6 M somatostatin for varying amounts of time at 37°C. The cells were then washed three times with phosphate-buffered saline (PBS), and the radioactivity associated with the cells was measured by a gamma counter. The experiment was performed in triplicate.
cAMP and cGMP
Intracellular levels of cAMP or cGMP in response to somatostatin were measured in triplicate by modified ELISA methods. For measuring cAMP, the cells were incubated at 37°C for 30 minutes in 1 mL PBS with or without forskolin (10−7 M) or somatostatin (10−11–10−7M) plus forskolin (10−7 M). For cGMP detection, the cells were incubated with or without somatostatin (10−11–10−7 M). The levels of cAMP or cGMP in the cell lysates were detected using commercially available ELISA kits (Sigma-Aldrich, St. Louis, MO). The concentrations of cAMP or cGMP were calculated from standard curves.
Cell Growth
Cells (3,000 cells/well) were seeded overnight in 96-well plates in 5% serum-containing medium and then treated in the same medium with or without 10211 to 10 27 M octreotide acetate (Sandostatin, Novartis, East Hanover, NJ) in triplicate. Somatostatin is not stable in serum long term26,27; therefore, a stabilized analogue, Sandostatin,26,27 was used to maintain receptor activation. After 2 days of growth, to quantify viable cells, incubation for 1 hour at 37°C with 20 μL of 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT)28–31 (Promega, Madison, WI) followed, and the color reaction was quantified using an automatic plate reader (VERSAmax, Sunnyvale, CA) at 490 nm.
Imaging
The animal protocol was approved by the institutional animal care and use committee. Eight- to 10-week-old female nude mice bred within the institution were used. Each of the six mice received three subcutaneous inoculations of 5 × 106 cells: left thigh, HT1080 cells transfected with vector; left shoulder, HT1080 cells expressing wild-type fusion protein; and right shoulder, HT1080 cells expressing Δ314 fusion protein. After approximately 1.5 weeks, imaging and biodistribution analyses were performed as previously described using a combination of in vivo magnetic resonance and gamma-camera imaging. 16 The mice were injected intravenously via the tail vein with 300 μCi of 111In-octreotide (Octreoscan). The same day, the mice were imaged with a T2-weighted fast spin echo sequence in a 4.7 T small-animal magnetic resonance imaging magnet (MR) (Bruker, Billerica, MA) using a previously validated method to obtain tumor weight. 16 Twenty-four hours after radiopharmaceutical injection, the animals were imaged for 10 minutes using a gamma camera fitted with a medium-energy collimator (mCAM, Siemens Medical Solutions, Hoffman Estates, IL). Acquisition was with a 512 × 512 matrix that was compressed to a 256 × 256 matrix for viewing, and region of interest analysis for radiopharmaceutical uptake was used to calculate the percent injected dose using a previously validated technique. 16
Statistical Methods
For ELISA, receptor binding, radioligand uptake, cAMP, cGMP, and MTT assays, as well as biodistribution analysis, two-sided Student t-tests were performed using Excel (2000, Microsoft, Redmond, WA). Animal numbers (N = 6) provide a power of 0.96 with a two-sided α of 0.05 for separating a twofold difference in in vivo biodistribution. For all experiments, a p value of less than .05 was considered statistically significant.
Results
Creation of Cell Lines Expressing Similar Levels of Wild-Type or Mutant Receptors
Human embryonic kidney (HEK293) and human fibrosarcoma (HT1080) cells do not endogenously express somatostatin receptors. No SSTR subtype expression was found using RT-PCR of RNA extracts from either cell line (Figure 1A). RT-PCR of the same RNA extracts did result in a product for the ubiquitiously expressed β-actin, confirming the quality of the RNA. The functionality of the SSTR subtype primers was confirmed by PCR of SSTR subtype complementary DNA.
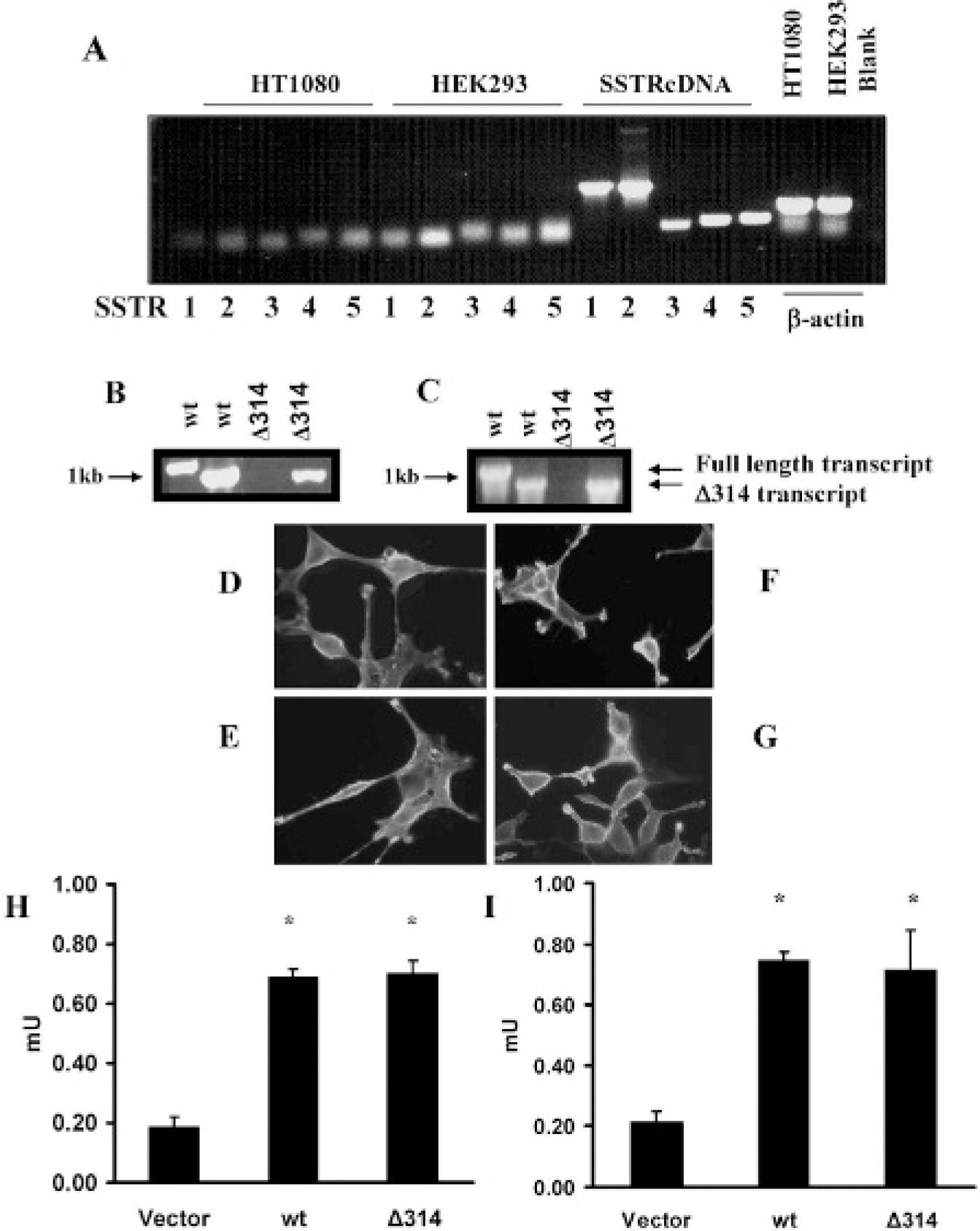
Cell lines transfected with hemagglutinin A wild type (wt) SSTR2 or HA-SSTR2Δ314 (Δ314) express appropriate transcripts and similar levels of fusion protein. Untransfected HT1080 or HEK293 cells do not express somatostatin receptors (A). No bands are seen in lanes SSTR1 to SSTR5 of HT1080 or HEK293 representing reverse transcriptase–polymerase chain reaction (RT-PCR) of cellular ribonucleic acid (RNA). The quality of the same RNA for RT-PCR is confirmed by bands in the β-actin lanes (blank, no RNA). Lanes SSTRcDNA 1 to 5 demonstrate PCR of the expected sized bands for SSTR subtypes 1 to 5 using SSTR subtypes 1, 2, 3, 4, and 5 cDNA as templates. Transcripts of the appropriate size are detected by RT-PCR in HT1080 cells (B) or HEK293 cells (C) transfected with HA-SSTR2 (HA-wt) or HASSTR2Δ314. Lane 1 and 3 primers for wild-type transcript; lane 2 and 4 primers for transcripts coding up to amino acid 314. Immunofluorescence targeting the HA domain demonstrates cell membrane localization of HA-wt or HA-SSTR2D314 fusion proteins; HT1080 clone expressing HA-wt (D) or HA-SSTR2Δ314 (E), HEK293 clone expressing HA-wt (F) or HA-SSTR2Δ314 (G). Equal protein expression is detected by enzyme-linked immunosorbent assay targeting the HA domain in HT1080 (H) or HEK293 (I) cell-line pairs transfected with HA-wt or HA-SSTR2Δ314, whereas no expression is seen in cells transfected with vector. Error bars represent standard deviation of triplicate samples (*p < .05, vector vs HA-wt SSTR2 or HA-SSTR2Δ314).
To inhibit signaling by the SSTR2A, the intracytoplasmic portion of the C-terminus was deleted beyond amino acid 314 (Δ314). Gene chimeras consisting of an Igκ membrane localization sequence, HA epitope tag, and full-length (wild type) or Δ314 SSTR2 were constructed. These were used to clone stable transfectants in both HEK293 and HT1080 cells, and RT-PCR of cellular RNA extracts confirmed expression of appropriate transcripts (Figure 1, B and C). Primers for either wild-type receptor or the Δ314 mutation resulted in appropriately sized products when RNA from either HEK293 or HT1080 cells transfected with wild-type receptor was used. In comparison, only primers for Δ314 resulted in appropriately sized products when RNA from HEK293 or HT1080 cells transfected with the Δ314 receptor was used, whereas no product was seen when primers for the full-length receptor were used.
To assess expression at the protein level, a common epitope tag was targeted instead of sites in the receptors to avoid confounding results that may arise from potential changes in ligand or antibody affinity owing to receptor mutation. Immunofluorescence demonstrated expression of wild-type or Δ314 SSTR2 fusion protein on the cell membranes of transfected HT1080 or HEK293 clones (Figure 1, D-G) and background signal in vector transfected cells. Quantitative ELISA demonstrated similar levels of fusion protein expression in cells transfected with wild-type receptor or the Δ314 deletion mutant for pairs of either HT1080 (Figure 1H) or HEK293 (Figure 1I) cells. Background signal was seen in cells expressing vector. Thus, HEK293 or HT1080 clonal pairs express similar levels of wild-type or Δ314 fusion protein.
Receptor-Ligand Binding
To test the hypotheses that the cytoplasmic C-terminus beyond amino acid 314 of human SSTR2 is not essential for ligand binding and is not essential for detecting the receptor in vivo, 111In-octreotide was used in binding experiments to evaluate a ligand relevant to in vivo imaging. Scatchard analysis demonstrated similar degrees of binding to membranes from clonal pairs expressing either full-length or Δ314 SSTR2 receptors (Figure 2). Similar results were seen whether HEK293 or HT1080 cell transfectants were evaluated; thus, binding to the imaging agent 111In-octreotide is conserved in the HA-SSTR2Δ314 (hemagglutinin A tagged SSTR2A with deletion of the C-terminus beyond amino acid 314) receptor.
No significant difference in uptake of 111In-octreotide was appreciated between HT1080 or HEK293 clonal cell pairs expressing wild-type or Δ314 SSTR2 receptors (see Figure 2E). Unlabeled somatostatin inhibited uptake, as expected (Figure 2E).
Biochemical Activity and Signaling
To assess receptor function, two separate signaling pathways were evaluated. We first assessed the ability of the human Δ314 SSTR2 mutant to inhibit cAMP production (Figure 3). No difference in baseline cAMP was noted in the cell-line pairs expressing wild-type or Δ314 fusion protein. Forskolin was used to activate adenylate cyclase. The addition of somatostatin inhibited forskolin-induced production of cAMP in either cell line expressing full-length receptors. In comparison, somatostatin did not inhibit forskolin-induced production of cAMP in HEK293 or HT1080 cells expressing the Δ314 mutant.
We then assessed the ability of the human Δ314 SSTR2 mutant to inhibit another signaling pathway, incitement of cGMP production (Figure 4). No difference in baseline cGMP was noted in the cell-line pairs expressing wild-type or Δ314 fusion protein. The addition of somatostatin to HEK293 or HT1080 cells expressing wild-type receptors resulted in increased cGMP production. In comparison, somatostatin did not cause cGMP production in HEK293 or HT1080 cells expressing the Δ314 mutant. Thus, biochemical signaling pathways regulating two different downstream mediators, cAMP or cGMP, are deficient in the Δ314-SSTR2 mutant.
Cellular Activity and Mitosis
The function of the Δ314 mutant was further assessed at the cellular level. Whether deleting amino acids distal to 314 affects the capacity of the SSTR2A to inhibit proliferation was tested in HEK293 and HT1080 cells (Figure 5). As predicted, the somatostatin analogue, Sandostatin, inhibited serum-induced proliferation in HEK293 cells or HT1080 cells expressing full-length receptors but not those expressing the Δ314 mutants. Thus, the Δ314 SSTR2 mutant is deficient in inhibiting cellular proliferation.
In Vivo Imaging
Next, we evaluated whether in vivo imaging of SSTR2 can be uncoupled from cAMP- and cGMP-mediated signal transduction as well as growth inhibition by using the SSTR2Δ314 mutant. On planar imaging of nude mice, both tumors derived from cells expressing Δ314 (Figure 6A, right shoulder) and tumors derived from cells expressing wild-type receptor (see Figure 6A, left shoulder) were clearly visible, whereas tumors derived from cells transfected with vector were not (see Figure 6A, left thigh). For 111In-octreotide, the normal route of excretion is primarily renal, with a secondary route via the hepatobiliary system. As expected, physiologic excretion of the radiopharmaceutical resulted in prominence of the kidneys.

HA-SSTR2 and HASSTR2Δ314 demonstrated similar binding to 111In-octreotide. Representative Scatchard (left) and saturation (right) plots of 111In-octreotide binding to membranes from HT1080 cells transfected with HA-wt SSTR2 (A) or HA-SSTR2Δ314 (B) and of HEK293 cells transfected with HA-wt SSTR2 (C) or HA-SSTR2Δ314 (D). Dissociation constant (Kd) and maximum specific binding (Bmax) values are the average and standard deviation of triplicate samples. Uptake of 111In-octreotide by cells in vitro in the presence or absence of unlabeled somatostatin by HT1080 (E, left) or HEK293 (E, right) clones. ss = somatostatin; V = vector; wt = wild type.
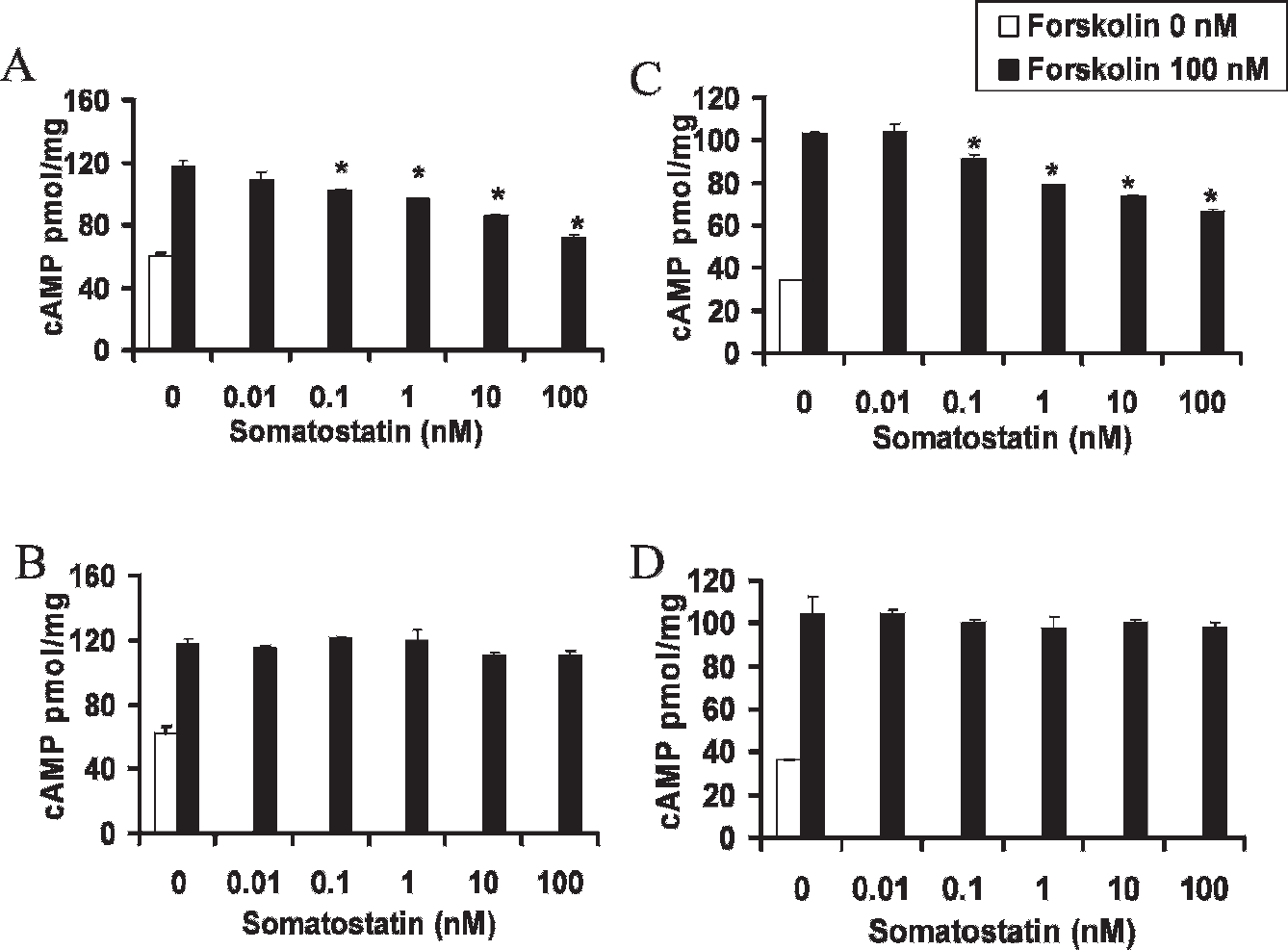
HA-SSTR2Δ314 is signaling deficient for the cyclic adenosine monophosphate (cAMP) pathway. Upon ligand binding, HA-wt SSTR2 decreases forskolin-induced cAMP production (A, C), whereas HA-SSTR2Δ314 (B, D) does not when expressed in HT1080 cells (A, B) or in HEK293 cells (C, D) (*p < .05, forskolin vs forskolin + somatostatin).
Radiopharmaceutical uptake by tumors was quantified by in vivo biodistribution analysis. 16 To control for tumor weight, magnetic resonance imaging was performed. Tumors expressing either wild-type receptor or the Δ314 mutant had greater uptake of 111In-octreotide than tumors derived from cells transfected with vector (p < .05, N = 6; see Figure 5B). No significant difference in uptake was seen in tumors expressing either wild-type receptor or the Δ314 mutant (p > .05, N = 6; see Figure 5B). The biodistribution analysis confirmed the imaging findings that the SSTR2Δ314 receptor is amenable to imaging in vivo.

HA-SSTR2Δ314 is signaling deficient for the cyclic guanosine monophosphate (cGMP) pathway. Upon ligand binding, HA-wt SSTR2 incites cGMP (A, C) production, whereas HA-SSTR2Δ314 (B, D) does not when expressed in HT1080 cells (A, B) or in HEK293 cells (C, D) (*p < .05, phosphate-buffered saline vs somatostatin).

HA-SSTR2Δ314 is deficient in inhibiting cell growth. Upon ligand binding, HA-wt SSTR2 decreases proliferation induced by serum (A, C), whereas HA-SSTR2Δ314 (B, D) does not when expressed in HT1080 cells (A, B) or in HEK293 cells (C, D). (*p < .05). Values on the abscissa are expressed as a percentage of the color product of cells treated with Sandostatin compared with the same cells not exposed to Sandostatin.
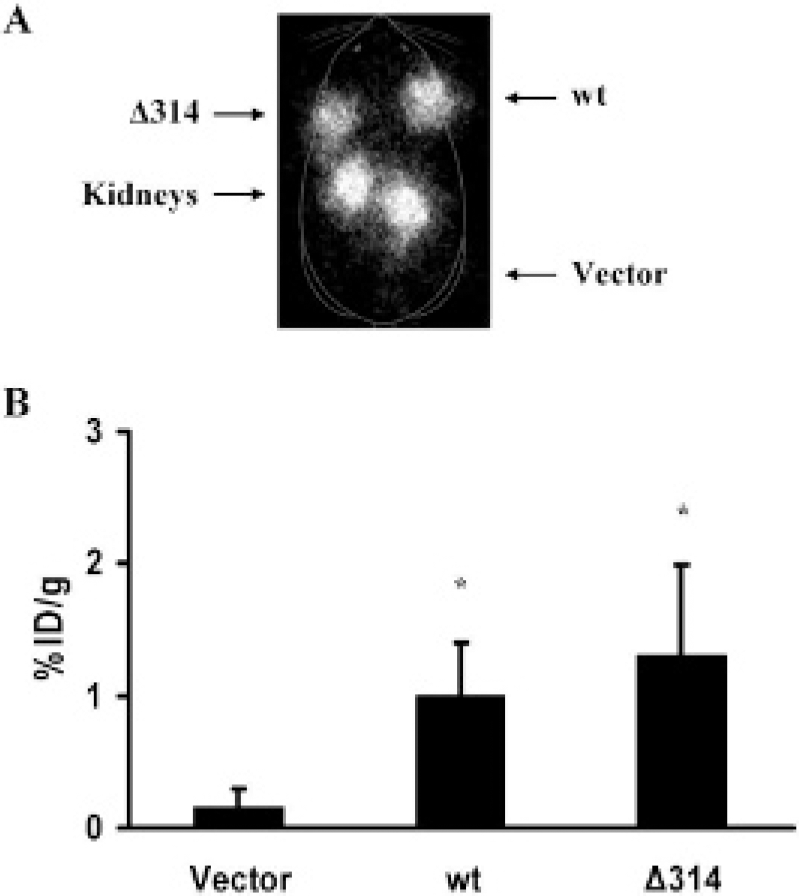
HA-wt SSTR2 and HA-SSTR2Δ314 are competent for imaging in vivo. Representative planar gamma-camera image of a nude mouse (A) demonstrates that tumors derived from HT1080 cells expressing HA-wt SSTR2 (left shoulder) or expressing HA-SSTR2Δ314 (right shoulder) are visible, but the tumor derived from HT1080 cells transfected with vector is not visible (left thigh). Nude mice bearing subcutaneous tumors (in both shoulders and the right thigh) were injected intravenously with 111In-octreotide (13 MBq) and imaged 24 hours later. Greater in vivo biodistribution (B) of the radiopharmaceutical is seen in tumors derived from HT1080 cells expressing HA-wt SSTR2 or HA-SSTR2Δ314 cells than in tumors derived from cells transfected with vector. Error bars represent standard deviation (N = 6 mice; *p < .05, vector vs HA-wt SSTR2 or HA-SSTR2Δ314).
Discussion
The findings suggest that in vivo imaging of SSTR2 can be uncoupled from cAMP- and cGMP-mediated signal transduction as well as cellular growth inhibition. We predicted that SSTR2 with deletion of amino acids beyond 314 would be signaling deficient yet amenable to imaging. The common HA tag in the wild-type and mutant fusion proteins was used to assess expression in stable cell clones. The tag provided a target for quantifying expression independent of binding sites that in the receptor may be altered by the receptor mutation. Limited signaling and growth inhibition by SSTR2Δ314 were verified in two different human cell lines, yet binding to 111In-octreotide was maintained. Furthermore, this mutant receptor was amenable to imaging in vivo.
Previous work with rat SSTR2 9 or human SSTR522,32 suggested that the C-terminus may influence but may not be necessary for binding somatostatin; however, the C-terminus did appear to regulate cAMP.9,22 Unlike most G protein–coupled transmembrane receptors that induce cAMP production upon ligand binding, somatostatin receptors usually inhibit cAMP production.9,22,33 Through cAMP, somatostatin can regulate secretion34–36 and chemotaxis. 37 A dopamine 2 receptor with a point mutation resulting in reduced ability to regulate cAMP has been imaged. 38 Consistent with the hypothesis that the C-terminus of SSTR2 regulates the cAMP pathway, we found that in HEK293 and HT1080 cells, SSTR2Δ314 is deficient in inhibiting forskolin-induced cAMP production.
SSTR2 also activates signaling cascades involved in cell division that appear to be separate from those regulated via cAMP. Instead, inhibition of proliferation by SSTR2 involves other downstream mediators,39–41 including cGMP. 10 We found that in HEK293 and HT1080 cells, SSTR2A can regulate the cGMP pathway and that this requires the C-terminus of the receptor. In both cell types, SSTR2Δ314 was deficient in eliciting cGMP production.
Further, we assessed SSTR2Δ314 function at the cellular level. Somatostatin binding to SSTR2 has been shown to cause cell-cycle arrest and inhibition of cell proliferation. 42 In both HEK293 and HT1080 cells, activation of wild-type SSTR2 inhibited serum-induced proliferation, indicating that the receptor not only alters signaling but cell function as well. In comparison, SSTR2Δ314 did not significantly inhibit serum-induced cell proliferation in either cell line. Thus, amino acids beyond 314 in the cytoplasmic C-terminus of SSTR2 regulate cAMP and cGMP signaling as well as serum-induced cell proliferation but are not necessary for binding the imaging agent 111In-octreotide. The corollary is that the SSTR2Δ314 mutant is deficient at the biochemical level in altering at least two signaling pathways and at the cellular level in inhibiting serum-induced proliferation, yet SSTR2Δ314 is competent for binding 111In-octreotide in vitro and for imaging in vivo.
One implication of the data is that in vivo imaging agents need not activate receptor-mediated cellular signaling. For producing new imaging agents, this suggests a hypothesis that may be tested in the future that instead of measuring downstream changes such as alterations in signaling, binding may be a better primary screen for selecting lead receptor imaging agents. Unlike therapeutic agents that seek to alter cellular function to treat disease, phenotypic change is not the goal of imaging and has the potential to cause toxicity. Consideration of receptor activation or deactivation is becoming more important as agents for radiologic modalities are being developed because they often require larger doses and longer circulation times. For example, unlike usually subpharmacologic (subnanomolar) concentrations often used for nuclear medicine imaging, agent concentrations in the millimolar range or higher 43 and circulation times of 1 to 2 days are often needed for magnetic resonance imaging. 44
In this study, stable cell clones were used to have a relatively homogeneous population of cells and tumors expressing approximately equivalent amounts of wild-type or mutant SSTR2 fusion protein to allow comparison. Data imply that a truncated receptor can retain binding for an imaging agent while being muted in eliciting signaling events. SSTR2 belongs to the large family of G protein–coupled receptors with seven transmembrane domains. The particular strategy of intracytoplasmic C-terminus deletion may be applicable to this family of receptors to create signaling deficient receptors, some of which may be good candidates for serving as reporters of gene expression.
Reporters can be introduced into cells in vitro, such as by transfection or infection, or in vivo. They usually consist of enzymes,43,44 transporters, 45 or receptors23,46 whose actions can alter physiology in target and nontarget cells or interfere with linked genes of interest. Reporters can be activated by exogenous and endogenous substrates or ligands. Through its receptors, somatostatin initiates signaling that affects cellular events such as secretion, chemotaxis, and growth suppression. The latter can sometimes be beneficial in the setting of cancer but may not be desirable for cellular therapies, where cell growth, function, and/or differentiation may be important; for evaluating gene therapy of diseases such as diabetes, where cell preservation and insulin secretion may be a goal; or for evaluating the effects of a linked gene of interest without confounding effects by the reporter. SSTR2 is one of the few reporters that can be imaged by US Food and Drug Administration–approved radiopharmaceuticals, such as 111In-octreotide, significantly increasing its potential for clinical translation. The same radiopharmaceutical was used in this study to image the signaling-deficient mutant. This suggests that SSTR2Δ314 potentially may serve as a reporter for exogenous gene expression imaging that has the desirable features of being limited in eliciting signaling and cellular responses.
In summary, the C-terminus of SSTR2 regulates signaling via cAMP and cGMP, as well as cellular responses such as growth. An SSTR2 mutant receptor deficient in eliciting such signaling and cellular responses can be imaged in a mouse model. Thus, in vivo imaging of SSTR2 can be uncoupled from signal transduction.
