Abstract
Phosphatidylserine (PS) is a well-characterized biomarker for apoptosis. Ligands that bind to PS can be used for noninvasive imaging of therapy-induced cell death, particularly apoptosis. In this study, we screened a random 12-mer peptide phage library on liposomes prepared from PS. One clone displaying the peptide SVSVGMKPSPRP (designated as PS3-10) bound to PS approximately 4-fold better than its binding to phosphatidylcholine and 18-fold better than to bovine serum albumin in a solid-phase binding assay. In addition, the binding of the corresponding PS3-10 peptide to PS was significantly higher than that of a scrambled peptide. PS3-10 phages, but not a control 4-2-2 phage, bound to aged red blood cells that had PS exposed on their surface. Binding of PS3-10 phages and PS3-10 peptide to TRAIL-induced apoptotic DLD1 cells was 3.2 and 5.4 times higher than their binding to untreated viable cells, respectively. Significantly, immunohistochemical staining confirmed selective binding of PS3-10 phages to apoptotic cells. Our data suggest that panning of phage display libraries may allow the selection of suitable peptide ligands for apoptotic cells and that PS3-10 peptide may serve as a template for further development of molecular probes for in vitro and in vivo imaging of apoptosis.
Phosphatidylserine (PS) is a well-characterized biomarker for apoptosis. PS normally resides in the internal leaflet of plasma membranes in healthy cells. 4 When cells undergo apoptosis, PS flips to the external leaflet of the plasma membrane.5,6 PS on the surface of apoptotic cells is associated with bridging molecules that are recognized by the receptors on macrophages. Exposure of PS on the cell surface triggers the macrophages to engulf the apoptotic cells.7,8 Using anti-PS monoclonal antibody as a probe, Ran and Thorpe showed that PS is also a marker of tumor vasculature and a potential target for cancer therapy. 9
Annexin V, a calcium-dependent phospholipidbinding protein, binds with high affinity to PS. For this reason, radiolabeled annexin V has been evaluated as an imaging agent for the noninvasive detection of therapy-induced apoptosis.10–12 However, the labeled protein is rapidly cleared from the body with a predominant renal excretion and a very short blood half-life (< 7 minutes).11,12 On the other hand, complete annexin V binding requires times up to 1 hour and approximately 2.5 mM of extracellular Ca2+. 13,14 Given its large molecular weight (35 kDa) and slower diffusion rate compared with low-molecular-weight compounds, there is concern whether optimal imaging of apoptotic cells in solid tumors is achievable with annexin V directly labeled with radio-isotopes. The clinical feasibility of 99mTc-labeled annexin V demonstrated mixed results. In cancer patients, 99mTc-labeled annexin V was able to localize at sites presenting with histologically proven spontaneous apoptosis or drug-induced apoptosis. However, the contrast of images in terms of signal to background ratio was clearly not optimal. 11 An alternative solution to optimize the imaging properties of radiolabeled annexin V is to use a long-circulating PEGylated annexin V.15,16 Although improved visualization of apoptotic response in solid tumors was observed, a significant amount of the injected dose of annexin V was distributed to the liver and the kidneys. Other promising approaches toward optimizing biodistribution of annexin V for imaging applications include the use of mutant forms of annexin V with endogenous sites for the (99mTc) labeling, 17 site-specific labeling, 18 and labeling with 18F-fluorodeoxyglucose. 19
Small-molecular-weight peptides are a potential alternative to proteins for diagnostic applications. Such agents are often less immunogenic and rapidly binding and possess advantageous pharmacokinetic properties. 20 The purpose of this study is to identify peptides that can selectively bind to PS on apoptotic cells independent of Ca2+ by panning of a 12-mer peptide phage library. Screening of random peptide phage libraries has been used to identify novel types of target-binding ligands.21–23 Several peptides identified using phage display technology have been evaluated for molecular imaging applications. For example, Koivunen and colleagues and Arap and colleagues identified a cyclic peptide, RGD-4C, containing an Arg-Gly-Asp motif that selectively binds to integrin αvβ3.24,25 99mTc-labeled RGD-4C was subsequently evaluated for imaging αvβ3-integrin-expressing tumors. 26 DUP-1, a peptide with high and specific binding to prostate cancer cells, also was identified using phage display techniques. The peptide showed properties that are potentially useful for the diagnosis and treatment of prostate cancer. 27 Here we report a phage clone and its corresponding peptide that selectively bind to PS and apoptotic cells. Our data suggest the use of both the phage particles and the peptide for molecular imaging of apoptosis.
Materials and Methods
Materials
1,2-Dioleoyl-sn-glycero-3-phosphatidylserine (PS) and 1,2-dioleoyl-sn-glycero-3-phosphatidylcholine (PC) were purchased from Avanti Polar Lipids, Inc. (Alabaster, AL). All Nα-Fmoc-amino acids, 1-hydroxybenzotriazole (HOBt), O-benzotriazole-N,N,N′,N′-tetramethyl-uronium-hexafluorophosphate (HBTU), and Fmoc-Rink linker were purchased from Novabiochem (San Diego, CA). N,N-Diisopropylethylamine (DIPEA), trifluoroacetic acid (TFA), triethylsilane, bovine serum albumin (BSA), rabbit anti-M13 phage antibody, propidium iodide, and 2′2′-azino-bis-3-ethylbenzothiazoline-6-sulfonic acid diammonium salt (ABTS) substrate were purchased from Sigma-Aldrich (St. Louis, MO). Mouse anti-M13 phage antibody was obtained from New England Biolab (NEB, Beverly, MA). Rabbit anti–annexin V antibody was purchased from Hyphen Biomed (Maon, OH). Texas red-labeled goat antirabbit antibody was purchased from Molecular Probe (Pitchford, OR). Annexin V–fluorescein isothiocyanate (FITC) was purchased from BD Pharmingen (San Diego, CA). Avidin and biotinylated horseradish peroxidase (HRP) complex (ABC) reagent was obtained from Vector Laboratories (Burlingame, CA). 3,3′,5,5′-Tetramentylbenzidine (TMB) was obtained from Pierce (Rockford, IL). Human recombinant tumor necrosis factor–related apoptosis-inducing ligand (TRAIL) was purchased from Chemicon (Temecula, CA). Isopropyl-1-thio-β-
Buffer Solutions
The following buffers were used throughout the experiment: phosphate-buffered saline (PBS; pH 7.4); PBS with 0.05% Tween 20 (PBST); 50 mM Tris-HCl buffer with 150 mM NaCl (TBS; pH 7.5); TBS with 0.05% Tween 20 (TBST); and 50 mM Hepes buffer with 100 mM NaCl (pH 7.4).
Analytic Methods
Peptides were analyzed by positive ion electrospray mass spectrometry on an Agilent 1100 Series LC/MSD instrument (Santa Clara, CA). Analytic high-performance liquid chromatography (HPLC) was carried out on an Agilent 1100 system (Wilmington, DE) equipped with a Vydac peptide and protein analytic C-18 column (10 × 250 mm, 10 μm, 300 Å) (Anaheim, CA). Samples were eluted with water and 10 to 80% acetonitrile containing 0.1% TFA over 30 minutes. Fluorescence intensity was measured using a Spex Fluorolog spectrofluorimeter (Jobin Yvon, Inc., Edison, NJ). Particle sizes of the PS and PC liposomes were analyzed by a Submicron Particle Sizer (Model 370, NICOM Particle Sizing Systems, Santa Barbara, CA).
Preparation of the Phospholipid-Coated Surface
The method of Kastl and colleagues 28 was adapted to coat microwell plates with PS. Briefly, lipid film was prepared by drying the lipid dissolved in chloroform under a stream of nitrogen, followed by 2 hours under a vacuum. Vesicles were prepared by first dispersing the lipid film in Hepes buffer (10 mg/mL) and then vortexing the solution for 2 minutes. Vesicles were formed by forcing PS solution through a polycarbonate membrane (pore size 50 nm) 50 times using a mini-extruder (Avanti Polar Lipids, Inc.), resulting in particles with an average diameter of 126 nm. To coat the microtiter wells with PS, diluted lipid vesicle solution (50 μL, 240 μg/mL) was added to each well, and the well was air-dried at 4°C. The unbound vesicles were removed by flushing with PBS, and the wells were treated with BSA (200 μL, 20 mg/mL PBS) at 4°C for 2 hours to block nonspecific binding. Wells were washed three times with PBS after blocking.
Enzyme-linked immunosorbent assay (ELISA) was used to verify the presence of PS on the surface of microtiter plates using annexin V as a probe. Briefly, PS-coated plates were incubated sequentially with annexin V (0.5 μg/mL), rabbit anti–annexin V antibody (final concentration, 1 μg/mL; Hyphen Biomed, Maon, OH), and HRP-conjugated goat antirabbit immunoglobulin G (IgG) (1:5,000 dilution). A signal was developed by the addition of ABTS substrate. BSA (0.5 μg/mL) was used as a control in place of annexin V. The optical density at 450 nm was 0.8 and 0.05 (arbitrary units) when PS-coated plates were probed with annexin V and BSA, respectively. Additionally, the optical density values increased with increasing dose of annexin V but not with increasing dose of BSA, indicating successful coating of PS to the surface of microtiter plates.
Phage Panning with PS
The Ph.D.-12 12-mer peptide phage library (NEB, Beverly, MA) has 1.5 × 1013 transducing units (TU)/mL and a diversity of 2.7 × 109 recombinants. The randomized peptide sequences are expressed at the N-terminus of the minor coat protein pIII, resulting in five copies of the displayed peptide per phage. Phage panning was performed according to manufacturer-provided procedures. The phage library was first diluted (1:100) with TBS buffer containing 1.5% BSA (w/v) in the absence of Ca2+. Aliquots of diluted phage library solution were then added to PS-coated microtiter wells (100 μL/well), and the plates were incubated for 2 hours at room temperature with gentle shaking. After thorough washing with TBST buffer, plate-bound phages were eluted with 100 μL of 0.2 M glycine-HCl (pH 2.2) containing BSA (1 mg/mL). Elutions were immediately neutralized with 1 M Tris-HCl (pH 9.1). Eluted phages were amplified in Escherichia coli ER2738 culture and titrated as described in the NEB Ph.D.-12 kit manual. The amplified phages were then subjected to the next round of panning. After six rounds of selection, individual clones were picked at random and their deoxyribonucleic acid (DNA) was sequenced at the DNA code facility of The University of Texas M. D. Anderson Cancer Center using the primer sequence 5′-CCCTCATAGTTAGCGTAACG-3′ provided by NEB.
Peptide Synthesis
PS3-10 peptide SVSVGMKPSPRP-NH2 and its corresponding scrambled peptide PSGVMPSRVKPS-NH2 were synthesized using standard Fmoc chemistry and purified by reverse-phase HPLC. Biotin (3 Eq, 0.3 M) was conjugated to the N-terminus of the peptide on solid support in dimethylformamide (1:1) using HBTU/HOBt (3 Eq/3 Eq) as the coupling reagents and DIPEA as the base (6 Eq). Side chain deprotection with concomitant cleavage of the product from the resin was achieved by treatment with TFA:H2O:TES (95: 2.5: 2.5, v/v/v) for 3 hours at room temperature. The products were purified by HPLC and were analyzed by LC/MSD.
Phage Binding to PS
PS-, PC-, or BSA-coated microtiter plates were filled with blocking buffer (0.1 M NaHCO3, 5 mg/mL BSA, 0.02% NaN3; pH 8.6) for 1 hour at 4°C and incubated at room temperature for 2 hours with 4 × 1010 TU/well of PS3-10 phages in 50 μL TBS containing 1.5% BSA. Wells were washed 12 times with TBST, followed by incubation with 100 μL host bacteria at room temperature for 1 hour. The phage-bacteria solution was then transferred to 10 mL LB medium (1% Bacto Tryptone, 0.5% yeast extract, 0.5% NaCl) containing 50 μg/mL tetracycline and allowed to stand at room temperature for 20 minutes. For colony titration, the phage-bacteria-containing LB medium (100 μL) was mixed with 3 mL top agar (0.7% in LB medium) at 45°C, and the mixture was spread on the top of an agar plate containing IPTG and Xgal. The plates were incubated at 37°C overnight. The binding of phages was quantified by counting blue colonies from aliquots of phage-infected bacteria in LB medium.
Enzyme-Linked Immunosorbent Assay
For phage ELISA assay, microtiter plates were coated with PS or PC as described before. Plates coated with either PC or BSA were used as controls. For BSA coating, aliquots of BSA in 0.1 M NaHCO3 buffer (pH 8.6, 10 μg/100 μL) were added to microtiter wells, and the plates were incubated at 4°C overnight. The PS- and PC-coated wells were blocked with 200 μL 3% BSA (w/v) in PBS at room temperature for 1 hour and then washed with PBST buffer. Phages (4 × 1010 TU/well in 50 μL TBS buffer containing 1.5% BSA) were added to the plates and incubated for 1 hour at room temperature. Wells were then washed three times with PBST buffer, incubated with 50 μL of HRP-conjugated anti-M13 phage monoclonal antibody (1:4,000 dilution in PBS with 3% BSA; NEB) for 1 hour at room temperature, and washed again. The plates were developed with 100 μL ABTS (0.022% in 50 mM sodium citrate). In a separate experiment, the binding of PS3-10 phages to PS was compared with that of a control phage, 4-2-2. The 4-2-2 phage has the same transfection capability as PS3-10, but its pIII coating protein does not express a peptide insert at the N-terminus.
For biotinylated peptide ELISA, PS-coated plates were incubated with 50 μL of biotin-PS3-10 peptide solution or biotin-scrambled peptide conjugate (50 μM) for 1 hour at room temperature, incubated with 50 μL ABC complex (Vector Laboratories) for 30 minutes at room temperature, and washed three times with PBST. The plates were developed with 100 μL TMB for ABC-mediated color reaction. The absorbance values were measured at 405 and 450 nm for ABTS and TMB reactions, respectively, using a Tecan microtiter plate reader (Research Triangle Park, NC). The experiment was performed three times. The resulting data were evaluated using a two-tailed Student t-test. Values were considered significantly different when p < .05.
For blocking experiment, unlabeled PS3-10 peptide (500 μM) or PS3-10 phages (8 × 1010 TU) were incubated with PS-coated plates for 1 hour. After this preincubation, the plates were incubated with biotin-PS3-10 peptide (50 μL, 50 μM) for 1 hour at room temperature. After washing three times with 200 μL PBST buffer, bound biotin-PS3-10 was detected by ELISA as described above.
Induction of Apoptosis
Human colon cancer DLD1 cells (American Type Culture Collection, Manassas, VA) were cultured in RPMI 1640 medium supplemented with 10% fetal bovine serum (FBS), penicillin, and streptomycin. When the cells grew to 80% confluence, they were split (1:4) and cultured overnight. The culture medium was changed to serum-free medium containing 250 ng/mL TRAIL, and the cells were cultured continuously for 2.5 hours. The floating cells and attached cells, which were detached by treatment with PBS buffer with 10 mM ethylenediaminetetraacetic acid, were collected together. Cells were washed once with PBS, fixed with 70% ice-cold ethanol, and stored at −20°C overnight. The fixed cells were stained with propidium iodide, and apoptotic cells were identified by fluorescence-activated cell sorting analysis, as described previously. 29
Phage Binding to Red Blood Cells and Apoptotic DLD1 cells
Stabilized human red blood cells (Beckman Coulter Inc., Fullerton, CA) were stored at 4°C. The presence of PS in the red blood cells was confirmed using 99mTc-labeled annexin V according to the method described by Tait and Gibson. 30 The red blood cells contained approximately 500,000 annexin V binding sites per cell.
Phage ELISA was used to study selective binding of phage particles to PS on the cell surface. Cells (8 × 107 red blood cells; 2 × 106 apoptotic DLD1 cells) were washed two times with 1 mL PBS in a microcentrifuge tube. The cell pellet was suspended in 200 μL PBS with 3% BSA and incubated for 1 hour at 4°C. After washing with PBS once, the cells were suspended in phage solution (4 × 1010 TU/tube, 200 μmL in TBS with 1.5% BSA) and incubated for 2 hours at 4°C. Cells were washed six times with TBST, incubated with 100 μL bacteria medium for 1 hour at room temperature, and then transferred to 10 mL LB medium containing 50 μg/mL tetracycline. After the medium was allowed to stand for 20 minutes at room temperature, a mixture of the medium (100 μL) with 3 mL top agar (0.3% in LB) was spread on the top of an agar plate containing IPTG and Xgal. The plates were incubated at 37°C overnight. Blue colonies were counted as TUs.
PS3-10 Peptide Binding to Apoptotic Cells
Viable and TRAIL-induced apoptotic DLD1 cells were collected and incubated with 5% rabbit serum in PBS for 1 hour at room temperature. The cells (5 × 105) were transferred to Eppendorf tube and incubated with 100 μL of biotinylated PS3-10 peptide (50 μM), biotinylated scrambled peptide (50 μM), or PBS for 30 minutes. After washing with PBS three times, the cells were treated with 100 μL ABC complex for 30 minutes. The signal was developed with 100 μL TMB substrate, and the optical density signal was measured at 450 nm.
Immunohistochemical Staining and Imaging
The viable DLD1 cells were cultured on chamber slides, and TRAIL-induced apoptotic cells were mounted on the slides by cytospin. Cells were fixed with 4% paraformaldehyde at room temperature for 30 minutes. For immunohistochemical staining and microscopic examinations, the slides were blocked with PBS containing 3% FBS for 1 hour at room temperature and incubated with 100 μL phages (4 × 1010 TU in TBS containing 1.5% BSA) at 4°C overnight. The slides were washed three times with PBST, incubated with mouse anti-M13 phage antibody (1:50 diluted) for 1 hour at room temperature, and washed three times, followed by incubation with HRP-conjugated rabbit antimouse secondary antibody. The slides were then incubated with 3,3′-diaminobenzidine substrate for color reaction. Slides were counterstained with Mayer hematoxylin for 1 minute. After being rinsed two times with deionized water for 5 minutes each, the slides were dehydrated in 95% ethanol, 100% ethanol, and xylene. Cells were covered with coverglass after dehydration with xylene and examined with a Leica microscope (Bannockburn, IL) with X200 magnification.
To examine the colocalization of phages with annexin V, TRAIL-induced apoptotic DLD1 cells were smeared on the slides and fixed with 4% paraformaldehyde. The slides were blocked with 3% FBS and incubated sequentially with PS3-10 phages or 4-2-2 control phages, rabbit anti-M13 phage antibody, and goat anti-rabbit IgG–Texas Red conjugate (Molecular Probe). Cells were then incubated with annexin V–FITC according to the manufacturer's instructions (BD Pharmingen, San Diego, CA). Cells were examined with a Leica microscope with FITC and Texas Red filters at X630 magnification.
Results
Selection of Phage Clone with PS-Binding Activity
In vitro selection of PS-specific phage clones was performed using a phage library displaying random 12-mer peptides on PS-coated plates in the absence of Ca2+. After six rounds of panning, 12 phage clones were picked randomly for sequencing. Of the 12 clones sequenced, 2 clones (16.7%) showed the same peptide sequence (SVSVGMKPSPRP) (Figure 1), which did not match that of any natural protein in the database searched (ie, National Center for Biotechnology Information, NCBI Blast nr database). Four other peptides shared at least 3 amino acids, suggesting a possible motif. The phage clone with the SVSVGMKPSPRP sequence, designated PS3-10, was selected for further evaluation.

Amino acid sequences of phage clones enriched through selection rounds and identified by sequencing. Amino acid residuals shared with PS3-10 are highlighted in bold.
Binding of PS3-10 Phages to the PS-Coated Surface
The selectivity of PS3-10 phages was first evaluated by comparing their binding to PS-, PC-, and BSA-coated surfaces. Figure 2A shows the number of phages recovered after incubation of PS3-10 phages and extensive washing steps. PS3-10 phages (expressed as TUs) recovered from the PS-coated surface were 4.2-fold higher than those recovered from the PC-coated surface and 17.8-fold higher than those recovered from the BSA-coated surface (see Figure 2A). Moreover, ELISA confirmed that PS3-10 phages bound to PS in a dose-dependent manner and at significantly higher levels than did insertless 4-2-2 control phages (Figure 2B).
Binding of PS3-10 Peptide to the PS-Coated Surface
Binding of biotinylated PS3-10 peptide and a scrambled peptide to PS was examined by ELISA. PS3-10 peptide showed 1.8-fold higher binding to PS than did the corresponding scrambled peptide control (Figure 3). Preincubation with soluble PS3-10 peptide and PS3-10 phages blocked 32% and 37% of the binding of biotinylated PS3-10 peptide to PS, respectively. On the other hand, the level of binding of the scrambled peptide to PS was not affected by incubation with either synthetic peptide or PS3-10 phages (see Figure 3).

Binding specificity of PS3-10 phages to phosphatidylserine (PS). A, PS, phosphatidylcholine (PC), or bovine serum albumin (BSA) was coated onto microtiter wells. PS3-10 phages (4 × 1010 TU in 50 μL TBS buffer) were added to each well. After incubation and washing, the bound phages were determined by counting the transduction unit (TU) number of recovered phages. B, Comparison of binding to PS between PS3-10 phages (■) and control 4-2-2 phages (□) by enzyme-linked immunosorbent assay using a horseradish peroxidase–conjugated anti-M13 monoclonal antibody. The data in B are presented as mean ± standard deviation from triplicate experiments. Binding to PS was significantly higher with PS3-10 than with 4-2-2 phages at doses tested (8 × 1010 TU and 16 × 1010) (p < .005). Study with the control phage (4-2-2 phages) was not performed at the dose of 4 × 1010 TU.
Binding of PS3-10 Phages and PS3-10 Peptide to PS on Aged Red Blood Cells and Apoptotic Tumor Cells
PS exposure on the outer leaflet of the plasma membrane is associated with aged red blood cells. 31 As shown in Figure 4, substantial phage clones were recovered from aged red blood cells that had been incubated with PS3-10 phages. In contrast, no clones were recovered from red blood cells incubated with 4-2-2 control phages.
To further test whether PS3-10 phages selectively bind to apoptotic cells, human colon cancer DLD1 cells were treated with TRAIL to induce apoptosis. 32 Over 90% of the TRAIL-treated cells were apoptotic, as determined by flow cytometry analysis (Figure 5A). ELISA revealed that TRAIL-treated cells exhibited more than threefold higher optical density value than did untreated viable DLD1 cells when the cells were incubated with PS3-10 phage (Figure 5B). In addition, the binding of PS3-10 phages to apoptotic DLD1 cells was about fivefold higher than that of 4-2-2 phages (data not shown).
The binding of PS3-10 peptide to apoptotic cells was assayed using ELISA. Apoptotic DLD1 cells incubated with PS3-10 peptide had greater than sixfold higher optical density value than those incubated with the scrambled peptide. Moreover, binding of PS3-10 peptide to apoptotic DLD1 cells was 5.4-fold greater than its binding to viable DLD1 cells. No difference in signal intensity was found between apoptotic and viable cells when cells were incubated with the scrambled peptide (Figure 5C).
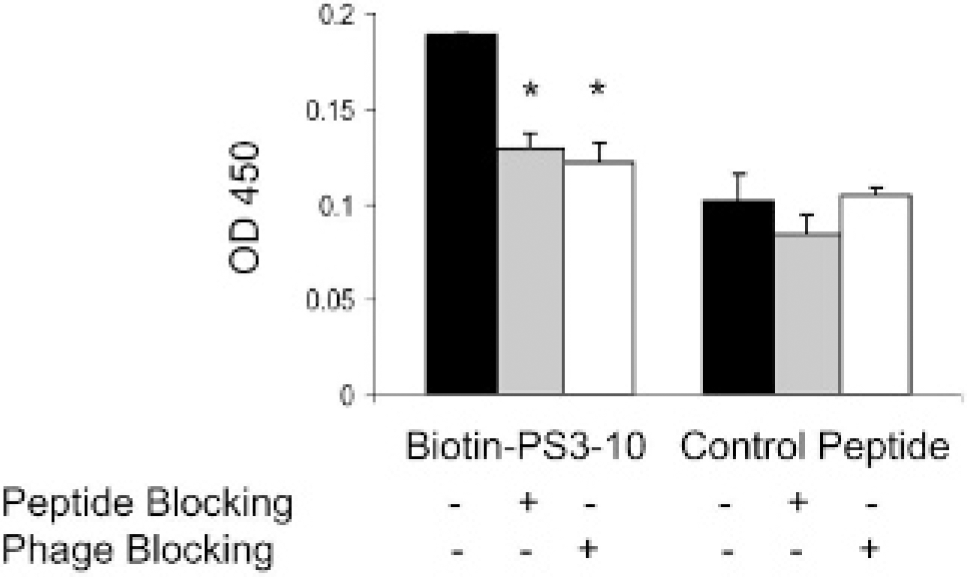
Binding of biotin-SVSVGMKPSPRP peptide to the phosphatidylserine (PS)-coated surface detected by enzyme-linked immunosorbent assay. Blocking experiments were performed in the presence of either SVSVGMKPSPRP (500 μM) or PS3-10 phages (8 × 108 transduction units [TU]). The PS3-10 peptide exhibited approximately twofold higher binding to PS than did a scrambled control peptide. Data are expressed as mean ± standard deviation of triplicate experiments. *Significant difference compared with the unblocked group with p < .005.

Binding of PS3-10 phages to aged red blood cells by counting transduction units (TU). A, Representative photograph of Xgal-stained blue phage colonies recovered from aged red blood cells. B, Quantification of TU. Red blood cells (8 × 107 cells) were incubated with PS3-10 phages or control 4-2-2 phages (4 × 1010 TU/well) in 50 μL TBS buffer. The phages bound to the red blood cells were recovered by infecting host bacteria and quantified by counting TU as described in Materials and Methods. The data represent mean ± standard deviation from duplicate wells of duplicate experiments.

Binding of PS3-10 phage and PS3-10 peptide to TRAIL-induced apoptotic cells. A, Flow cytometry of DLD1 cells. Treatment of DLD1 cells with TRAIL at a dose of 250 ng/mL for 150 minutes induced 91.6% apoptosis (M1 or sub-G1 population). B, Comparison of binding of PS3-10 phage to apoptotic and viable DLD1 cells. Viable or TRAIL-treated DLD1 cells (2 × 106) were incubated with PS3-10 phage (4 × 1010 transduction units [TU]), and bound phages were assayed by enzyme-linked immunosorbent assay (ELISA). C, Comparison of binding of PS3-10 phage to apoptotic and viable DLD1 cells. The binding of PS3-10 peptide to TRAIL-treated or viable DLD1 cells was determined by ELISA as described in Materials and Methods. The data represent mean ± standard deviation from duplicate wells of duplicate tests in experiment B and triplicate samples in experiment C. PBS = phosphate-buffered saline.
Imaging Apoptotic Cells with PS3-10 Phage
Neither untreated viable cells incubated with PS3-10 nor TRAIL-treated cells incubated with 4-2-2 control phage stained positive with anti-M13 phage antibody. Positive staining was observed only in TRAIL-treated DLD1 cells incubated with PS3-10 phages (Figure 6). The absence of normal nuclear staining and of the morphology integrity in the PS3-10 phage-stained cells suggests that these cells underwent apoptotic response. Interestingly, anti-M13 phage antibody colocalized with annexin V in TRAIL-treated DLD1 cells incubated with PS3-10 phages but not in apoptotic DLD1 cells incubated with 4-2-2 control phages (Figure 7).

Imaging of apoptotic cells using PS3-10 phages. TRAIL-induced apoptotic DLD1 cells and untreated viable DLD1 cells were stained for the presence of phages with horseradish peroxidase–conjugated anti-M13 monoclonal antibody and DAB substrate, as described in Materials and Methods. Arrows indicate cells stained positive for phage particles. Original magnification: X200.
Discussion
Using a bacteria phage display system, we have identified a new phage clone, PS3-10, and its corresponding 12-mer peptide, SVSVGMKPSPRP, that selectively bind to PS. The number of PS3-10 phages bound to PS was 4-fold to 18fold higher than phages bound to PC and BSA, respectively. PS3-10 phages also exhibited significantly higher binding to PS than did 4-2-3 control phages at all doses tested (see Figure 2). Moreover, PS3-10 phages, but not insertless control 4-2-2 phages, bound to aged red blood cells having their PS flipped to the outer surface of the plasma membrane (see Figure 4). PS3-10 phages bound to apoptotic DLD1 cells at a more than threefold higher level than their binding to untreated viable DLD1 cells (see Figure 5). Because PS3-10 phages showed selective binding to PS and PS on apoptotic cells, we further evaluated whether PS3-10 phages could be used to image apoptotic cells in vitro. Positive staining was visualized in TRAIL-treated DLD1 cells incubated with PS3-10 phages but was not observed in viable DLD1 cells incubated with PS3-10 phages or TRAIL-treated DLD1 cells incubated with 4-2-2 phages (see Figure 6), indicating specific binding of PS3-10 phages to apoptotic cells. Dual-color fluorescence labeling with annexin V and antiphage antibody revealed that only cells labeled by annexin V were also labeled by PS3-10 phages (see Figure 7A). The control 4-2-2 phages did not bind to apoptotic DLD1 cells that were positively labeled with annexin V (see Figure 7B). Taken together, these data indicate that PS3-10 phages bind specifically to PS on the surface of apoptotic cells.
In comparison with strong selective binding of PS3-10 phages toward PS and apoptotic cells, the corresponding PS3-10 peptide showed moderate selectivity to PS-coated surface. The binding of biotin-labeled SVSVGMKPSPRP peptide to PS was 1.8-fold higher than the binding of a scrambled peptide. Whereas the binding of PS3-10 peptide to PS could be partially blocked (up to 37%) by the corresponding soluble peptide and by PS3-10 phages (see Figure 3), the binding of PS3-10 phages to PS could not be blocked with PS3-10 peptide (data not shown), suggesting that PS3-10 phages had higher binding affinity to PS than did the PS3-10 peptide. The higher binding affinity with PS3-10 phages may be attributed to the multivalency effect associated with phage particles that contained five copies of the same peptide fused to the pIII protein. Of note, peptides with high binding affinity and selectivity can be accessed through peptidomimetic and multimeric design.33–35 Clearly, further studies are needed to increase the binding affinity and selectivity of PS3-10 peptide to PS before it can be used for in vivo imaging applications.

Representative immunofluorescent images of apoptotic DLD1 cells induced with TRAIL. Cells were stained with annexin V–FITC (green) and for phages (red), as described in Materials and Methods. Differential interference contrast image (DIC) and overlay (Merge) images are also presented. Only cells incubated with PS3-10 phages (A) but not with 4-2-2 control phages (B) showed colocalization with annexin V. Original magnification: X630.
The relatively low binding selectivity of PS3-10 peptide to purified PS may also be attributed to a high nonspecific binding of PS3-10 peptide to PS-coated surface. To assess the selective binding to PS exposed on apoptotic cells, both viable and apoptotic DLD1 cells were exposed to biotinylated PS3-10 peptide and the scrambled control peptide. As shown in Figure 5C, PS3-10 peptide demonstrated 5.4-fold higher binding to apoptotic cells than to viable cells. When compared with its scrambled peptide, PS3-10 peptide showed sixfold higher binding to apoptotic DLD1 cells (see Figure 5C). These results suggest that PS3-10 peptide exhibited much higher binding specificity in a cell-based assay than in an assay based on purified PS.
Previous studies have revealed the presence of primary peptide sequences in several PS-binding proteins that are responsible for their binding to PS. These proteins include Raf-1 translocated to membrane, 36 cell surface receptor Gas6, 37 and transmembrane glycoprotein CD36 38 ; the latter two are presumably involved in phagocyte recognition of cell debris. Another study reported PS-binding peptides identified by panning a 15-mer phage library. 39 In most reported sequences, the clusters of the basic amino acids arginine and lysine appeared to be critical for interaction with PS, and ionic interaction was suggested.36,38,39 PS3-10 also possesses two basic amino acids, a lysine and an arginine cluster, suggesting that its binding to negatively charged PS may be mediated through ionic interaction. However, since the scrambled peptide containing two Arg and two Lys units had significantly lower binding to PS than did the PS3-10 peptide, it is likely that in addition to ionic interaction, peptide sequence is another important factor determining the binding activity of PS3-10. Recently, Laumonier and colleagues reported PS-binding phages through using ex vivo biopanning of a hexapeptide M13 phage library in perfused apoptotic mice livers. 40 One of the lead phage clones exhibiting selective binding to PS carried the neutral TLVSSL peptide. These results support the notion that the presence of charge is not an absolute requirement for selective binding of peptides to PS.
In summary, we have demonstrated that phages and peptides selected by biopanning phage library against PS have the potential to be used for imaging apoptotic cells. Studies toward peptidomimetics suitable for in vivo imaging applications designed on the basis of PS3-10 peptide are in progress.
Footnotes
Acknowledgment
We thank Dawn Chalaire for editing the manuscript.
