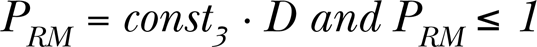Abstract
A stochastic model based on the Markov Chain Monte Carlo process is used to describe responses to ionizing radiation in a group of cells. The results show that where multiple relationships linearly depending on the dose are introduced, the overall reaction shows a threshold, and, generally, a non-linear response. Such phenomena have been observed and reported in a number of papers. The present model permits the inclusion of adaptive responses and bystander effects that can lead to hormetic effects. In addition, the model allows for incorporating various time-dependent phenomena. Essentially, all known biological effects can be reproduced using the proposed model.
INTRODUCTION
To date, a number of models of cellular responses to ionizing radiation have been published (UNSCEAR 1986; Calabrese and Baldwin 2003; Feinendegen 2005; Scott et al. 2007; Leonard 2008). Although the notion of probability of a given process is often used, most of the models demonstrate a rather deterministic approach resulting in more or less complicated mathematical formulas describing the biology of the cell and the physics of the radiation. Usually, the analytical formulas intend to show specific biological reactions of a cell culture or an organism in terms of probabilities and their distribution. However, at the very end, an analytical formula is used and the stochasticity of the processes is not obvious.
A relevant mathematical model should account for all variants of biological phenomena such as bystander effects (Prise et al. 2003; Leonard 2008), adaptive responses (Feinendegen et al. 2000), radiosensitivity, probability of natural death, cellular multiplication and mutation, and many others (ICRU 1983; Feinendegen and Pollycove 2001; Pollycove and Feinendegen 2003; Feinendegen and Neuman 2005; Lehnert 2007; Leonard 2008). This paper displays the results of simulations of possible outcomes of the exposure of cells to ionizing radiation. The simulations have been carried out within the framework of the Markov process.
METHODOLOGY
A stochastic Markov process, or memorylessness, is one for which “the present state of the system is independent of its previous states” (Encyclopaedia Britannica 2011). Hence, the conditional probability distribution of the future states depends solely upon the present state (Booth 1967; Podgórska et al. 2002; Trivedi 2002).
The reaction of a simulated simple organism (i.e., a group of cells) to ionizing radiation is analyzed in two numerical loops (see Fig. 1). Every step within the first loop represents the actual state of the cells. One can view the steps in this loop as the units of time. At each step, the cells may be growing old exhibiting the possibly altered states or those newly formed by the division of one or more progenitors. All these states are created in the second loop which inspects and acts on the set of cells delivered by the previous step. The second loop takes the cell one by one and uses a stochastic tree of probabilities which alter the state of the cell depending on whether it has been irradiated or not, has become mutated, cancerous, dead, etc. In this way, a new state of cells is created and the process reiterates in the next step, i.e. returns to the first loop.

General idea of the model: two loops over the steps and the cells.
The presented numerical loops contain the trees of probabilities, so purely mathematical values of these probabilities are intended to describe the biology of the cells (Lehnert 2007; ICRU 1983). An example of how the developed algorithm works is shown in Fig. 1 where the arrows displayed are just randomly selected for demonstration. A detailed presentation of the probability tree can be found in other figures. Since the primary aim of this paper is to show how such a type of the model works, some advanced biological mechanisms have not been considered here – they will be dealt with in the future calculations.
All the described rules work for the determined and constant value of dose per step: D. A change in D means that the calculations must start from the beginning. The total absorbed dose is a product of D multiplied by the number of steps K. Consistently, the dose D shown in all the figures is the dose per unit time and not the total dose which has led to the observed effect.
Construction of the stochastic tree of probabilities is the most important part of the model's algorithm (Booth 1967). The tree consists of a dozen of input parameters, e.g. the values of probabilities or the typically assumed constants.
The model presented in this paper shows a new method of simulation of a cellular response to an absorbed dose of radiation. The results demonstrate that, by using a given set of parameters, all the known reactions can be modeled. Currently, however, it is inherently difficult to find values of the probabilities which are closest to reality.
THE ALGORITHM
The numerical algorithm begins with setting a population of N cells. At first, all of the cells are “healthy“. At each (time) step, a stochastic tree of possibilities is applied to every i-th cell (see Fig. 2). Generally, the cell can be healthy, mutated, or cancerous. By definition, a healthy cell does not contain mutations and a mutated cell contains one or more mutations. However, not every cell that has been hit by radiation will develop a mutation. This process is regulated by the appropriate probability of mutation. A cancer cell is one transformed under certain conditions from a mutated cell, assuming that at least three genetic mutations have occurred (Hahn et al. 1999).
Every healthy, mutated, or cancerous cell can be irradiated (hit) with the probability Phit (Bryszewska and Leyko 1997; Simmons and Watt 1999):
where D denotes, as explained earlier, a dose per single cell in one simulation step, and const1 is a scaling constant. In other words, Phit is the probability of hitting a cell by radiation. The formula (1) is chosen just to describe a typical linear or quadratic dependence in the low-dose region and to make sure that the cell is hit at a very high dose per step. Obviously, one could use e.g. a stretched exponential or many other formulas as well. The choice of (1) is probably the simplest one (Bryszewska and Leyko 1997; Simmons and Watt 1999). For applications such as the one used herein, one is practically operating in the low dose region where eq. (1) is well approximated by a linear function of the dose. This equation can reflect the notion of cross section used, e.g. in particle physics.

The algorithm works based on the random numbers generator 1 which determines all the probabilities. Each probability is drawn and is assigned a random value from 0 to 1. Depending on the generated value, a given branch of the tree is chosen.
Within the first part of the stochastic tree (see Fig. 2) six scenarios are possible, each of them serving as a starting point for the following stages, as shown in Figures 3 through 8.
1. A healthy cell has been hit by radiation (Fig. 3).
There are seven possible consequences of hitting a healthy cell (Fig. 3), that is, the cell can:

A portion of the stochastic tree from Fig. 2 – a healthy cell has been hit.
naturally die (probability PD );
be killed by a single hit (probability PRD );
spontaneously mutate, irrespectively of the hit (probability PM );
naturally multiply (probability PS );
generate bystander signals to the nearby cells (probability PB );
become mutated by radiation (probability PRM );
stay intact (probability 1-PD-PRD-PS-PM-PB-PRM ).
The term “probability of natural cell death” (PD ), as used throughout, is a rather broad one. Death can be executed by two major mechanisms: apoptosis or necrosis. In order to keep the algorithm as simple as possible, we have used a single constant value of probability PD.
Probability of the immediate cell death as a result of a precise, single hit by radiation, PRD , is assumed to be linearly dependent upon dose per step D with a scaling factor const2 :
Eq. (2) must be constrained by the natural condition valid for all the probabilities, namely: PRD ≤ 1. When a cell becomes mutated, either spontaneously (PM ) or by radiation (PRM ), its status changes from “healthy” to “mutated“. In this case the number of mutations in the cell is one. Generally, cells can harbor more than one mutation. The value of PRM is again assumed to be a linear function of the dose per step, D:
where const3 is yet another scaling constant. The linear relationships (2) and (3) are selected for simplicity only, and can, if necessary, be replaced by any other form.
It has been shown in experimental kinetic analyses of human cell cultures that four to seven rate-limiting stochastic events – thought to be distinct somatic mutations and the triggered cellular signaling pathways thereof – are required for the formation of a tumorigenic cell (Renan 1993; Hahn et al. 1999; Hahn and Weinberg 2002). Based on this assumption, one can model the probability of transformation of a mutated cell into a cancerous cell using e.g. the following set of equations:
where Q is the number of mutations in the investigated cell. The value of 1-PRC describes the probability that the mutations generated in the cell will not result in tumorigenic transformation. Model (4) can be replaced by a continuous one once biological mechanisms of such a transformation are better understood.
One possible cause of mutations in cells that are in the vicinity of a hit cell is a bystander effect (with the probability PB ). The algorithm selects randomly one nearby cell from the investigated i'th cell. When the neighboring cell is healthy its status will change from “healthy” to “mutated” (assuming one mutation in the cell). The number of mutations in this nearby cell increases by 1 if the cell has already contained mutations. The investigated i'th cell preserves its status (and the number of mutations). The probability of the bystander effect, PB , is given by any probability distribution function with the maximum value at low doses (Feinendegen 2005). Four examples of such possible distributions (described by appropriate polynomials) are shown in Fig. 9.
Notably, in this approach we have assumed that the bystander effect causes only adverse outcomes and its possible beneficial effects (Prise et al. 2003; Leonard 2008; Mothersill and Seymour 2006) have been ignored. Within the present context of the algorithm a positive result would enhance the branch of the adaptive response. Again, for the sake of simplicity, such a possibility as well as an adaptive response limiting the number of non-radiogenic mutations has been excluded from the modeling.
2. A healthy cell has not been hit (Fig. 4).
There are four possible consequences of the case when a healthy cell has not been hit by radiation (Fig. 4):
the cell naturally dies (probability PD );
the cell spontaneously mutates (probability PM );
the cell naturally multiplies (probability PS );
nothing happens (probability 1-PD-PS-PM ).
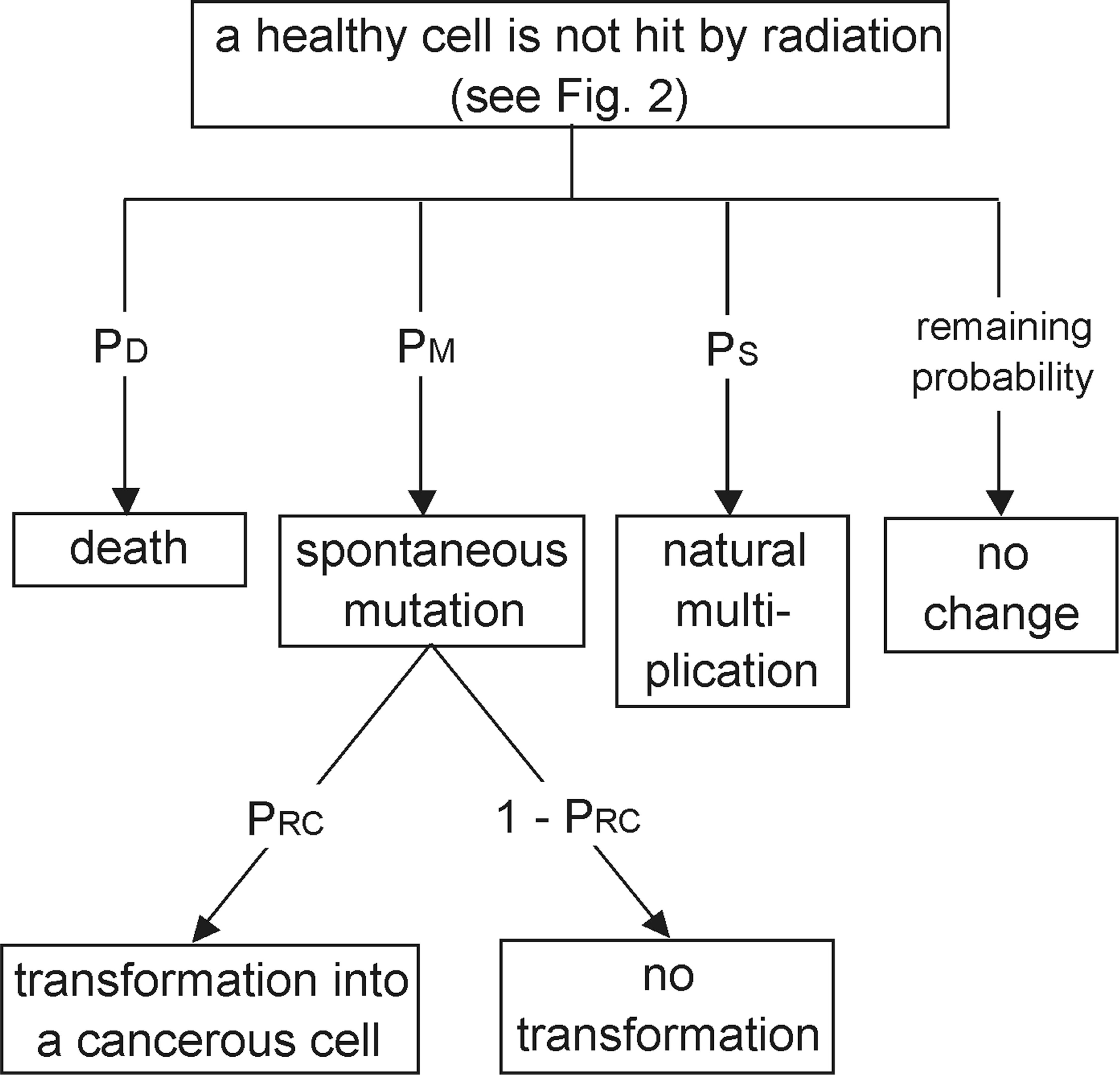
A portion of the stochastic tree from Fig. 2 – a healthy cell has not been hit.
As in section (1), the mutated cell can change into a cancer cell with the probability PRC
In this case, bystander effects may also be produced by non-hit cells which exhibit, for example, enhanced oxidative metabolism. As mentioned above, such effects have not been accounted for in the present algorithm.
3. A mutated cell has been hit (Fig. 5).
In this case the mutated cell can behave in one of the following ways (Fig. 5):
it can naturally die (probability PMD );
it can be killed by one precise hit of radiation (probability PRD );
it can spontaneously mutate one more time (probability PM );
it can naturally multiply (probability PMS );
an adaptive response can be generated (probability PA );
it can generate a bystander effect (probability PB );
it can spontaneously repair one of the existing mutations (probability PR );
it can be mutated by radiation (probability PRM );
nothing happens (probability: 1-PMD-PRD-PMS-PM-PA-PB-PR-PRM ).
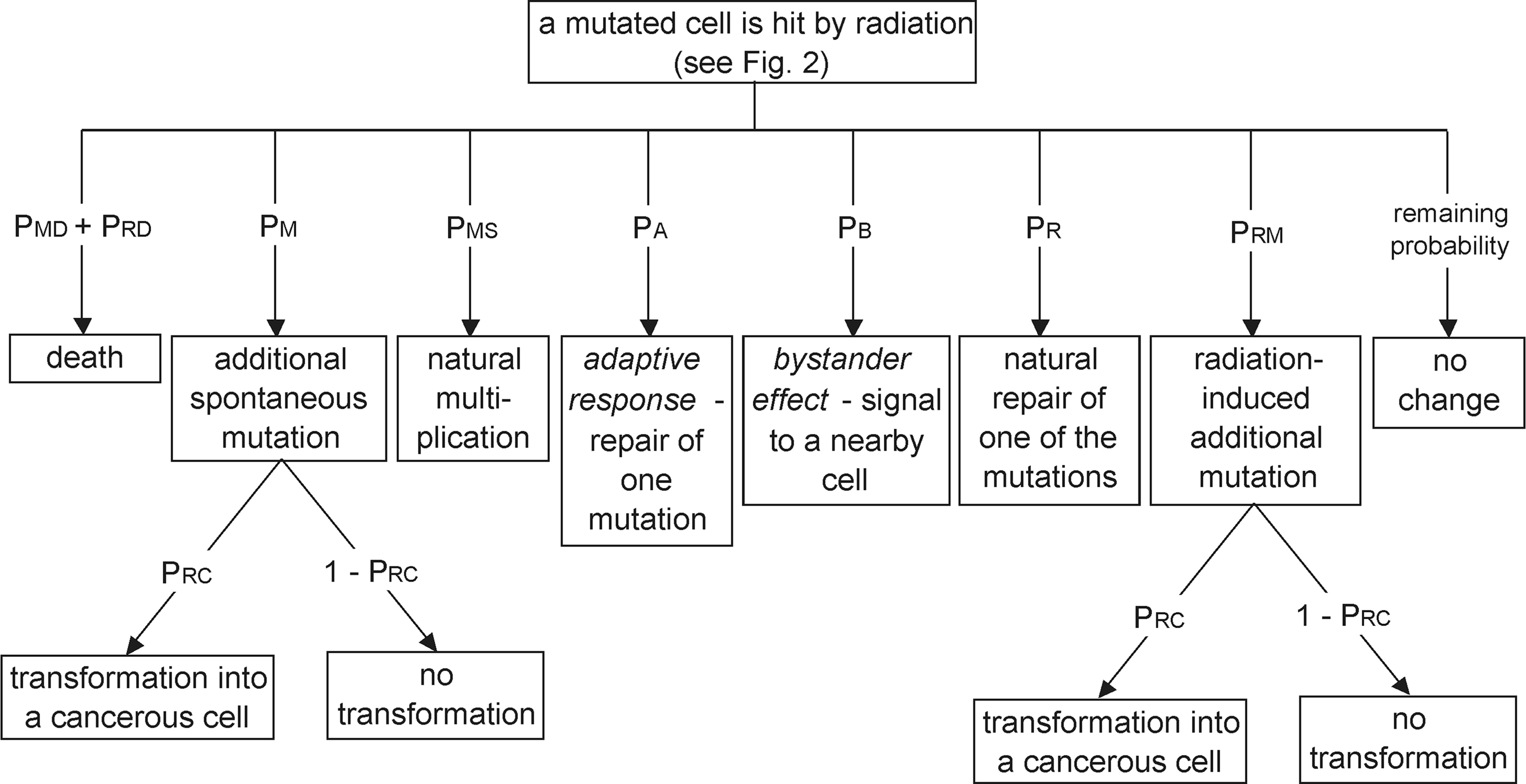
A portion of the stochastic tree from Fig. 2 – a mutated cell has been hit.
Generally, the probabilities PMD and PMS for mutated cells differ from the respective probabilities for healthy cells (PD and PS ). However, PMS may create a progeny (daughter cell) with the same number of mutations.
The new probability PA describes the adaptive response which is active only in mutated cells and reduces the number of mutations in the cell by 1. If the cell has only one mutation, its status will change from “mutated” to “healthy“. Probability of the adaptive response PA is given by any probability distribution function with the maximum value at low doses (Feinendegen 2005). Four examples of such distributions (described by appropriate polynomials) are shown in Fig. 10.
In a manner comparable to those described in the previous sections the mutated cell, having sustained mutations with probabilities PM and PRM , can be transformed into a cancer cell with the probability PRC.
4. A mutated cell has not been hit (Fig. 6).
There are six possible outcomes for a mutated cell that has not been hit (Fig. 6):
the cell naturally dies (probability PMD );
the cell spontaneously mutates one more time (probability PM );
the cell naturally multiplies (probability PMS );
an adaptive response develops (probability PA );
the cell naturally repairs one mutation (probability PR );
there is no effect (probability: 1-PMD-PMS-PM-PA-PR ).

A portion of the stochastic tree from Fig. 2 – a mutated cell has not been hit.
PMS creates a progeny (daughter cell) with the same number of mutations. As indicated earlier, the cell mutated with the probability PM can change into a cancer cell with the probability PRC.
5. A cancer cell has been hit (Fig. 7).
When a cancer cell has been hit by radiation there are six possible consequences (Fig. 7), i.e.:
the cell naturally dies (probability PCD );
the cell is killed by one precise-hit of radiation (probability PRD );
the cell naturally multiplies (probability PCS );
the cell dies because of the radiation-induced additional damage (probability PCRD );
the cell generates a bystander effect (probability PB );
nothing happens (probability: 1-PCD-PRD-PCS-PCRD-PB ).
The probabilities PCD and PCS should be significantly different from PMD and PMS because of the different biology of the cancer cell. In order to make the model as simple as possible, it assumed that the probability that the cancer cell will die because of an extra hit (due to its radiosensitivity) changes linearly with the dose per step:

A portion of the stochastic tree from Fig. 2 – a cancer cell has been hit.

A portion of the stochastic tree from Fig. 2 – a cancer cell has not been hit.
6. A cancer cell has not been hit (Fig. 8).
When a cancer cell has not been hit, one of the following three situations (Fig. 8) can be expected:
the cell naturally dies (probability PCD );
the cell naturally multiplies (probability PCS );
nothing happens (probability: 1-PCD-PCS ).
SUMMARY OF THE INPUT PARAMETERS
The described algorithm contains a stochastic tree and dozens of input parameters – probabilities and constant factors. All of them, listed in Table 1, can be selected subjectively and their suggested values are important for the final result. These parameters describe, rather intuitively, biological and physical effects of the interaction of ionizing radiation with cells. In particular, one considers a special form (1) of the probability distribution for a cell being irradiated (Phit ). Other probabilities, such as a cell developing a mutation (PRM ) or getting killed because of its radiosensitivity or the precise hit (PCRD and PRD ), or even the probability that a mutated cell is transformed into a cancerous one (PRC ) are assumed to vary linearly with the dose per step. There are also probabilities given by one simple constant value such as the probability that a healthy/mutated/cancer cell will naturally die (PD / PMD / PCD ) or multiply (PS / PMS / PCS ). The probability of a spontaneous mutation (PM ) and a natural repair (PR ) are also simple constants. In contrast, the bystander effect (PB ) or adaptive response to radiation (PA ) are given by certain dose dependent functions.
To cut off one of the branches in the stochastic tree (Figs 2–8) one can reduce the respective probability value to zero. It is also possible to modify the model by adding more branches of the tree representing the more detailed biophysical effects.
RESULTS
The present model can work with all values of the input parameters (Tab. 1), either intuitively assumed or precisely measured. Usually, it is difficult to obtain the precise data, because, in reality, there are many types of cells with different properties, age, etc. Sometimes, however, it is possible to draw more or less precise values of the parameters listed in Table 1. For example, in case of the parameter PRD one can use const2 ≈ 0.00006 (Feinendegen et al. 2010). Also, probability functions of the bystander effect (PB ) and adaptive response (PA ) can be taken as B1 and A1 (see Figs 9 and 10), respectively. Their forms correspond qualitatively with the ones found in the reference literature (Feinendegen 2005; Feinendegen et al. 2010).
One suggested collection of the parameters’ values is presented in Table 2. Results of the model based on the Table are shown in Figures 11 through 13.
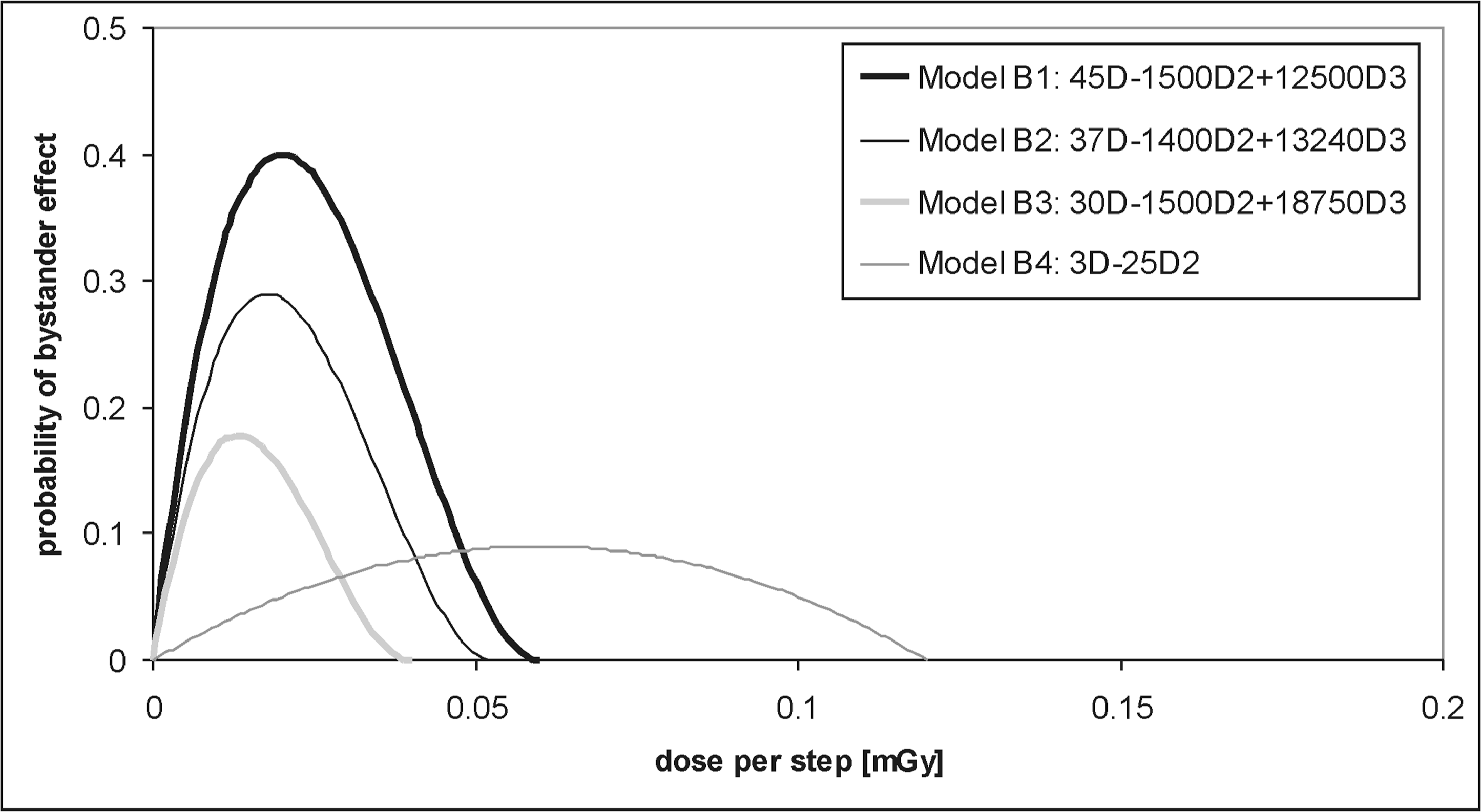
Four exemplary models (B1, B2, B3, B4) of the probability distribution of the bystander effect (PB) described by up to the third-order polynomials. The labels “D2” and “D3” in the figure's legend stand for “D2” and “D3“, respectively, where D is the dose per one state (step) of cells
In Fig. 11 the relative ratios of the mutated (solid line) and cancer cells (dotted line) to all the cells are shown in cases when the adaptive response and bystander effect are not taken into account. As shown, the model assumes that approx. 3% of the cells are naturally mutated at the beginning, i.e., independently of any radiation (Fig. 11). The straight line can easily be drawn through most of the data showing the number of the mutated cells. In the case of cancer cells, however, one can observe clear threshold at the dose per step of approx. 0.2 mGy. Cancer cells are also treated as mutated ones. In Fig. 12 the same data obtained for the lowest doses per step are shown (D < 0.4 mGy): a threshold for the formation of cancer cells at about 0.2 mGy per step and a nearly constant value of the mutated cells below the dose of 0.04 mGy per step are clearly indicated. At the dose of about 0.7 mGy per step practically all the cells are mutated and neoplastically transformed. Fig. 13 shows that at this dose per step the total number of cancer cells increases substantially – it is more than twice as high as the initial number of the “healthy” cells. Notably, the average number of mutations per each cell approximates six. With the still increasing dose the probability of the cell's death overcomes the probability of its multiplication; therefore the number of mutations per cancer cell is decreasing (Fig. 13).

Four exemplary models (A1, A2, A3, A4) of the probability distribution of the adaptive response (PA), described by up to the third-order polynomials. “D2” and “D3” in the figure's legend stand for “D2” and “D3“, respectively, where D is the dose per one state (step) of cells.
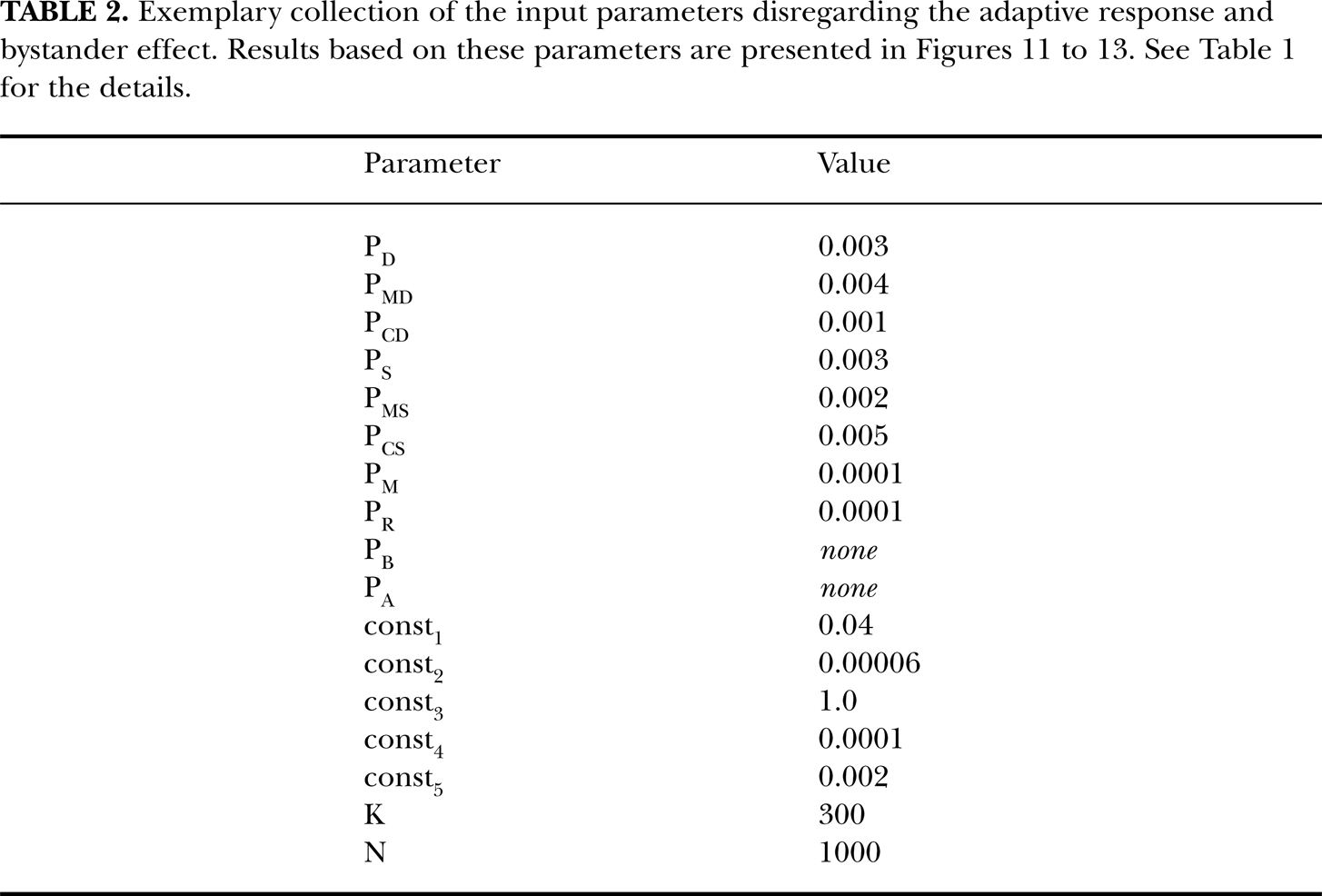
Exemplary collection of the input parameters disregarding the adaptive response and bystander effect. Results based on these parameters are presented in Figures 11 to 13. See Table 1 for the details.

Results obtained with use of the set of parameters from Tab. 2. The fraction is shown of mutated (solid line) and cancer cells (dotted line).

Results obtained with use of the set of parameters from Tab. 2. The fraction is shown of mutated (solid line) and cancer cells (dotted line). The dose is limited to 0.4 mGy per step.
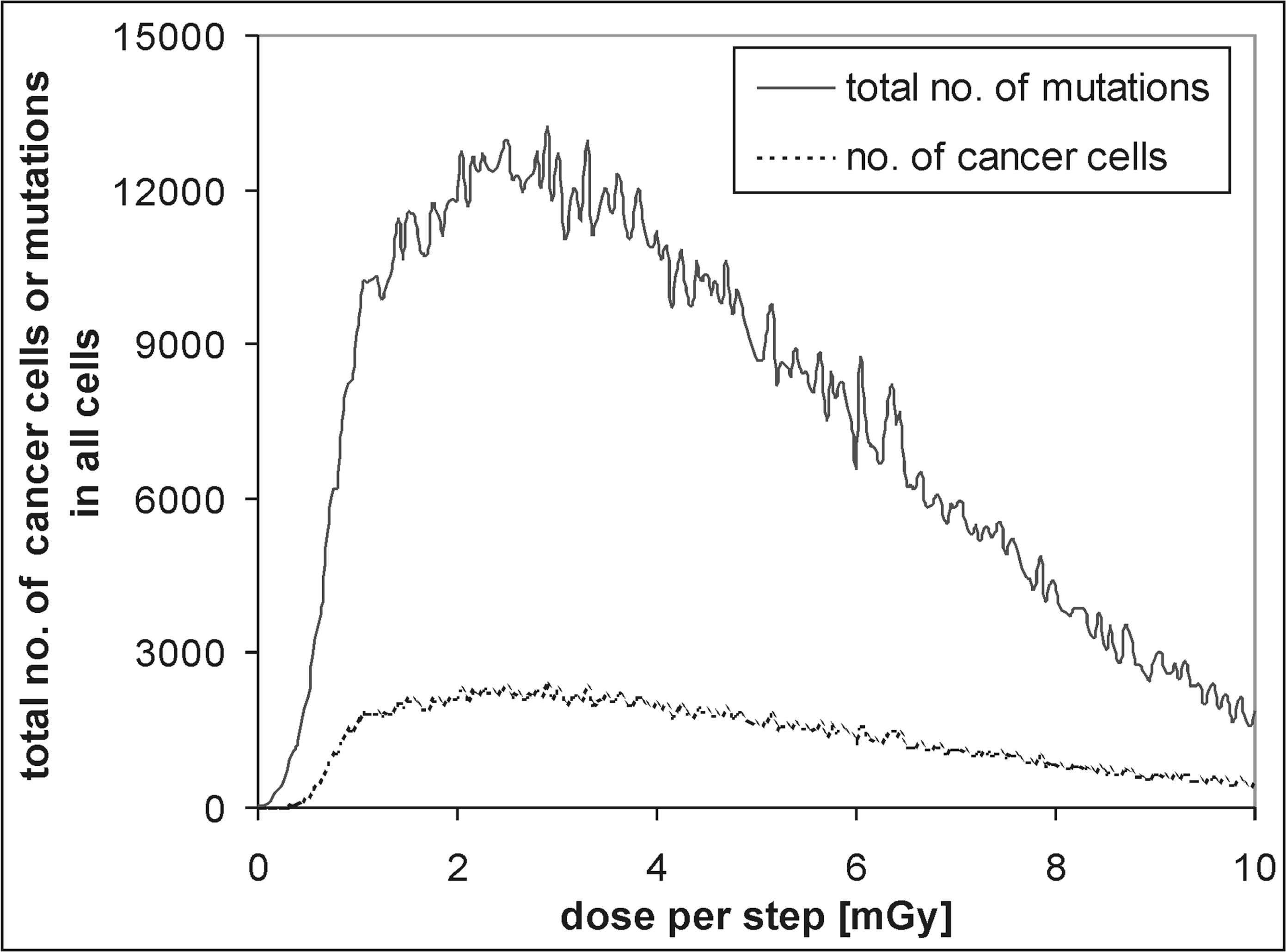
Results obtained with use of the set of parameters from Tab. 2. The total number is shown of mutations in all cells (solid line), along with the number of cancer cells (dotted line).
The results presented in Figures 11 through 13 do not include the adaptive response or the bystander effect. Figures 14 and 15 show the same results, but in this case PA is taken as a model A1 = D – 5D 2 + 6D 3 and PB as a model B1 = 45D – 1500D 2 + 12500D 3 (see Figures 9 and 10 for details). By comparing Fig. 15 with Fig. 12 one can note that when both the adaptive response and bystander effect are present a significant hormetic effect is manifested, especially below 0.15 mGy per step.
DISCUSSION AND CONCLUSIONS
The stochastic model used in the present paper most probably shows what can actually happen in a population of cells exposed to ionizing radiation. Such a stochastic approach seems to be better than many deterministic models because it takes into consideration some individual susceptibilities and probabilities rather than many phenomenological factors which, although called “probabilities” are not, in fact, used as such. In the present model, every run yields somewhat different results only because of the use of the truly probabilistic approach.
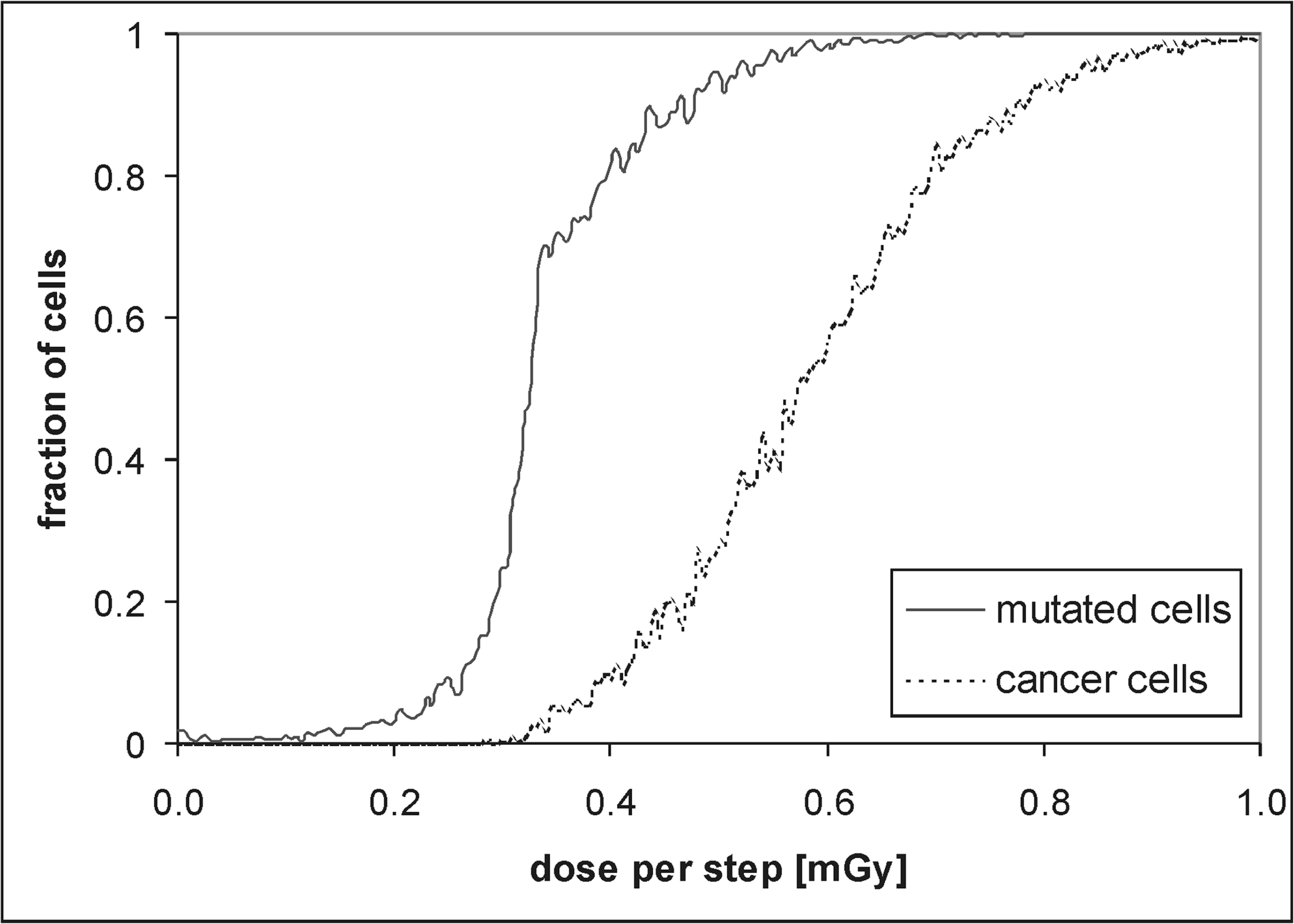
Results obtained with use of the set of parameters from Tab. 2 including the adaptive response and bystander effect. The fraction is shown of the mutated (solid line) and cancer cells (dotted line).

Results obtained with use of the set of parameters from Tab. 2 including the adaptive response and bystander effect. The fraction is shown of the mutated (solid line) and cancer cells (dotted line).
One of the advantages of using of a probability tree is a possibility of easily adding or cutting off some of its branches. The present algorithm has been significantly simplified in terms of its biological background, but it can certainly be further developed and made more complex in the future.
Based on the results presented in Figures 11 through 15, a number of conclusions can be drawn. Firstly, the observed fluctuations simply reflect the stochastic nature of the processes. Secondly, although most of the obtained relationships between the dose per step and the given outcome vary linearly with the dose (see Table 1), the final result is conditional upon the dose in a much more complicated way. Indeed, a clear-cut threshold for the development of cancerous cells can be seen regardless of whether or not any bystander effect or adaptive response has been accounted for (Figs 11 and 14). Thirdly, there is a diminution in the total number of mutations after absorption of a certain dose per step (Fig. 13), apparently, as a result of death of the cells exposed to a strong radiation field. Finally, when the bystander effect and adaptive response are taken into account, a significant hormetic effect can be seen in terms of the diminished number of the mutated cells (Fig. 15 versus Fig. 12) and a shift in the threshold for the appearance of cancer cells (Fig. 15).
In the present simulation an undefined cell culture with no association to any specific tissue has been used. Indeed, the present model has no ambition to develop the algorithm for more complex tissue reactions. However, a potential user of the model can take their own values of input parameters specifying the type of cells or the tissue they may want to investigate.
The obtained results are consistent with many epidemiological and experimental data demonstrating or implicating a threshold (Ullrich and Storer 1979; Miller et al. 1989; UNSCEAR 1986, 1994, 2000; Hosoi and Sakamoto 1993; Liu et al. 1996; Mitchel et al. 1999, 2003; Redpath et al. 2001, 2003; Ko et al. 2004, 2006; Elmore et al. 2005; Redpath and Elmore 2007; Nowosielska et al. 2008; Ulsh 2010). In particular, the specific shape of the dose-response relationship presented in Fig. 11 was recently demonstrated in the studies of death rates of the exposed laboratory rats and mice and the Chernobyl workers (Allison 2009; Henriksen and Maillie 2003; Zyuzikov et al. 2011). A similar shape is also presented in (Anderson and Storm 1992; UNSCEAR 1994), demonstrating a cumulative incidence of primary liver tumors among the Danish patients.
In turn, the shape of curves shown in Fig. 13 closely resembles the final cumulative rate of leukemia among mice (Di Majo et al. 1986; Robinson and Upton 1978; UNSCEAR 1986) and the absolute risk of breast cancer in women treated with radiotherapy (Shore et al. 1986; UNSCEAR 1994). Same shape of dose-effect relationship is seen in figure VI in (UNSCEAR 1994), where excess relative risk per sievert for mortality from leukemia of the life span study is presented. Finally, similar results were obtained by Shuryak et al. (2009) in their simulations of biological processes.
It must be stressed that the present model was not biased in any way towards a possibility of a threshold or hormesis. These effects seem to appear very naturally and, notably, the latter effect could be demonstrated only after taking the adaptive response into account. Obviously, the present model contains too many parameters to be readily adaptable to a particular situation. Nevertheless, one can easily control the influence of any of these parameters (processes) on the final result. The model can also be easily expanded to more sophisticated forms of the relationship between probabilities and doses.
The present approach may provide clues for modeling the impact of low doses on organisms with use of the Markov chain Monte Carlo techniques. A prospective user of the model can add new branches to the probability tree and/or can select a completely different set of the input probabilities. The values of such parameters listed in Table 2 are just an example of how the model can work. In order to describe a real biological system, the model will be further developed and elaborated in the future.
Footnotes
ACKNOWLEDGEMENTS
The authors wish to express their gratitude to Professor Zbigniew Jaworowski for his careful review of this paper and comments provided, as well as to Drs. Aneta Cheda, Ewa Nowosielska and Jolanta Wrembel-Wargocka for their valuable inputs.