Abstract
This review article describes our simplified biophysical model for the response of a group of cells to ionizing radiation. The model, which is a product of 10 years of studies, acts as (a) a comprehensive stochastic approach based on the Monte Carlo simulation with a probability tree and (b) the thereof derived detailed deterministic models describing the selected biophysical and radiobiological phenomena in an analytical manner. Specifically, the presented model describes effects such as the risk of neoplastic transformation of cells relative to the absorbed radiation dose, the dynamics of tumor development, the priming dose effect (also called the Raper–Yonezawa effect) based on the introduced adaptive response approach, and the bystander effect. The model is also modifiable depending on users’ potential needs.
Introduction
From a purely physical perspective, every living organism is a complex system. Such systems are characterized by a very unpredictable behavior, as many different influencing factors must be taken into account. Therefore, it is very difficult, if not impossible, to describe the response of a complex system to a given factor in a strict mathematical formalism. However, using appropriate numerical methods, one can create a model which provides a good approximation of reality within an acceptable range. Such a model can successfully describe the behavior of an organism exposed to one specific external stressor. One of the most important effects of stressor interaction is cancer induction, which is of crucial interest to the presented paper.
Cancer is a genetic disease—it is caused by stable changes in DNA which controls the normal functioning of a cell, created as a result of error or damage. In 2020, 19.3 million estimated worldwide cancer cases were reported, of which almost 10.0 million were fatal. 1 Exposure to external agents may potentially result in induction of a neoplasm in the human body. According to the mutational theories, more than one oncogenic mutation is responsible for initial steps in the neoplastic process, until the cells eventually sustain enough genetic alteration to become autonomous, which results in cancer. It is conservatively estimated that 5% of human cancers are caused by viruses, 5% by radiation, and the remaining 90% by chemicals. 2
In the context of the presented paper, ionizing radiation is a special case of an external factor (the stressor) that affects the complex system of a living organism. The subject of the general impact of radiation on health is very wide and will not be discussed here—this article focuses on the description of various attempts to model the impact of ionizing radiation on carcinogenesis initiation and progression, methods of model’s programming and statistical description, as well as on the description of selected biophysical and radiobiological phenomena, which contribute significantly to the complexity of its response to radiation.
Comparison of different computational stochastic models describing ionizing radiation interaction with the organism on the cellular level.

This review article shows our own approach which presents selected methods of computer modeling of the effects of ionizing radiation on the body at the cellular level. The authors present a stochastic model using the Monte Carlo technique based on a Markov probability tree with memory approach, which can be successfully used for computer simulation of various types of radiation phenomena. A deterministic approach is also described, which focuses on specific processes, such as tumor growth, or the phenomenon of radiation adaptive response.
Monte Carlo Stochastic Modeling of Irradiated Cells
The Monte Carlo modeling technique has been implemented in various types of biophysical simulations for many years. The development of the model discussed in this chapter started 10 years ago 19 when the first simple Monte Carlo Markov Chain (MCMC) with probability tree model of cellular radiation response was published. Over the years, this model has been refined, bringing its results closer to reality.20–25 Details of the current version of the model are described below.
Cell Arrangement
The model can simulate the behavior of any number of cells (N), limited only by computer capabilities. All cells are organized into a three-dimensional matrix that simulates real tissue (see Figure 1 and 2A and B). This matrix may undergo changes (e.g., cells may die, multiply, and mutate) with successive discrete time steps. Moving from one time step to the next means that each cell has passed through the probability tree (described in detail later). At each time step, each cell has one of the following states: (a) healthy, (b) damaged, (c) mutated, (d) cancerous, and (e) dead cell/lack of cell (empty). Of course, all those states are on the general level of biological processes. Cell’s numerical organization in the three-dimensional matrix which geometry is presented in Figure 2. Two possibilities of geometrical cell arrangements in the 3D matrix: (a) the cube or (b) sphere, where 3 axes represent geometric coordinates. The plot (c) shows the schematic possibility of radiation beam implementation.
25


Damaged cells are those whose DNA has been successfully hit by ionizing radiation: the act of ionization in the space of DNA has occurred (irrespective of the type of lesion or radiation). A mutant cell, on the other hand, is a cell that has transformed from a damaged cell because of damage that was not repaired or was repaired incorrectly (via all potential ways). A cancerous cell is a neoplastically transformed mutant cell, that is, that has undergone the process of cytogenetic transformation due to the accumulation of an appropriate number of oncogenic mutations in its DNA. 26 More details about cell statuses and their transitions with time (number of time steps) will be described later.
Dosimetry
In the simplest variant, all cells are pre-assigned with a certain dose of ionizing radiation. This may be a single dose value (D) at a specific fixed time step (k) or a fixed dose value in all subsequent time steps which corresponds to a constant dose-rate. The two scenarios mentioned are the most commonly used, although it is possible to enter an arbitrarily variable dose-rate (dose per step) as well as to differentiate the dose depending on the cells’ matrix region. In particular, by applying microdosimetric methods, it is possible to simulate any dose distribution based on the adopted external radiation field. For example, a dose distribution for tumor cells was simulated for the interaction of alpha radiation (Bragg curve) in a specific direction of the cell matrix. 27 Another example is a possibility of cell irradiation by a specific beam with its dedicated dimensions and angle 25 (see Figure 1C).
Probability Tree
At each successive time step, each cell is tested against a probability tree (the so-called Bernoulli tree). The iterative algorithm in three loops passes through each i-th cell in turn and first checks its status (healthy/damaged/mutated/cancerous), which determines further treatment. Then, the algorithm uses Monte Carlo methods to check the appropriate probability functions, such as whether the cell was hit by radiation, whether the cell died naturally, whether the cell divided (creating a daughter cell), or if any further biophysical effects occurred. The diagram of the probability tree used in the model is presented in Figures 3-7. A description of all probability functions is presented in Table 2. More details are presented in References.20,24,25 However, one has to note that all biophysical processes are implemented on a rather general level, where many details (e.g., chromosomal aberrations) are just a part of single phenomenon (e.g., mutation creation). Part of a probability tree from Figure 3 concerning healthy cell which was (a) hit and (b) not hit by ionizing radiation. Part of a probability tree from Figure 3 concerning damaged cell which was (a) hit and (b) not hit by ionizing radiation. Part of a probability tree from Figure 3 concerning mutated cell which was (a) hit and (b) not hit by ionizing radiation. Part of a probability tree from Figure 3 concerning cancerous cell which was (a) hit and (b) not hit by ionizing radiation.




It should be emphasized that the passage of all cells in the matrix through the probability tree (each cell individually) equals one time step. In the following time step, the situation repeats itself and again all cells go through the probability tree according to the same pattern (see Figure 8). However, due to the random nature of probability, the results will be different each time. This approach allows for an evolutionary change of the entire cell layout depending on time, dose, and other model input parameters.
Probability Functions
The probability tree described in the previous section is based on a number of probability functions that describe biophysical processes in a cell. Some of them are simple functions, described by constant values, but most of them are complex functions of many variables, which include, among others, time, age of the i-th cell, dose in a given time step, radiation doses in previous time steps, number of lesions, or number of mutations. These variables are accompanied by appropriate constants, the so-called model parameters, the determination of which is non-trivial. These parameters can be different for various types of cells, different tissues, or organisms. Therefore, one of the basic activities is to calibrate the model based on the existing experimental data, allowing determination of the values of parameter inputs to the model for specific conditions. Such calibration is difficult and of high complexity but is also necessary for the correct quantitative operation of the model. Of course, arbitrary values of the input parameters could be assumed; however, the results of such simulations would be only qualitative and would not relate to any particular cell type or experimental case.
As mentioned earlier, the model contains many probability functions that comprehensively, albeit on a general level, describe the biophysics of irradiated cells. A detailed description of these functions, along with their mathematical formulae, is given in Table 2. Some of them describe quite obvious biological processes, such as death, multiplication, or radiation damage repair, but some of them address less known phenomena, called non-targeted effects, which include the radiation adaptive response and the bystander effect. Some of them are described below in a more precise way.
Radiation Adaptive Response
The phenomenon of radiation adaptive response includes stimulating the body with low doses of radiation in order to stimulate its repair mechanisms.31-35 This phenomenon is very subtle and does not always occur; 33 hence, there is a strong necessity to describe it using stochastic methods. If it does occur, it plays an essential role in the cell response to ionizing radiation in the low-dose area and is one of the factors that generate the so-called non-linear response.21,36
In the presented model, an equation derived from other studies is used, describing the probability of an adaptive response for the dose D received k time steps ago:20,21,37 Exemplary hunchbacked shape of equation (1) representing the radiation adaptive response probability function, after single constant dose irradiation.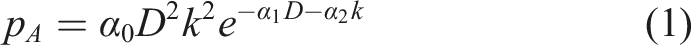

A special case of the radiation adaptive response is the so-called priming dose effect, also known as the Raper–Yonezawa effect. 38 It describes the response to a situation where a given organism is first irradiated with a low priming dose and after some time with a high challenging dose. Due to this effect, the negative health effects on the body are smaller than if it was exposed to only a high dose. More about the Raper–Yonezawa effect will be provided later.
Bystander Effect
The bystander effect is a classic example of the non-targeted effect of the interaction of ionizing radiation. It describes the phenomenon where the non-irradiated cell adjacent to the one irradiated behaves as if it were irradiated itself. This is due to the transmission (via diffusion) of molecular signals from a cell hit by radiation to a neighboring cell. Firstly, the probability of the bystander effect occurring depends on the dose-related function, which is saturated for medium and high doses:20,30,39

The presented description is not the only approach to the bystander effect. A good alternative is based on the diffusion equation with 2 ways of the signal transmission, analogically as in Japanese model,
29
which can be directly implemented in the presented model
30
using the simplified equation:
Estimated values of input parameters for probability functions (see Table 2) of Monte Carlo stochastic model, dedicated for human lymphocytes and their response to ionizing radiation (X-rays). The parameter estimation was obtained by partial model calibration on experimental data for dedicated processes.3,29,39–51

Neoplastic transformation
The neoplastic transformation (cancer transformation) is a key element of the presented model, as it was its original purpose. Due to this, it is possible to evaluate the so-called radiation risk, that is, the overall probability of cancer formation for a specific radiation dose. 37
The described model uses the sigmoid function (the same as for the phase transition dynamics) to describe the probability of neoplastic transformation depending on the number of M mutations with critical index n:
The above formula is known as the Avrami equation and shows that only a certain accumulated number of M mutations determines the highly non-linear probability that a mutated cell changes its status to a cancerous one. The derivation of this equation is based on the theory of nucleation and growth of crystals, as an analogue to cancer. 28
Applicability
The presented model has wide applications in simulating various cellular processes, but its main goal is to simulate the processes of radiation carcinogenesis. This means that the model can be used to support radiation risk analyses and can be used, for example, for strictly radiobiological purposes, as well as auxiliary in radiological protection or probabilistic safety analyses (PSA) of the Level 3 in a nuclear power plant and its emergency planning.
Within the presented model, any initial conditions can be set (e.g., all damaged or healthy cells, a specific number of cells and their spatial distribution, and specific dose fractionation). Therefore, it is possible to simulate, for example, radiotherapeutic processes, where cells with a set status of “cancer” are exposed to high doses of radiation, and neighboring healthy cells may be affected by, for example, the bystander effect. This is one of many examples of how the model can be used.
In this context, an interesting example of the results that can be achieved with the model is the simulation of the cell survival curve for a group of cells with “cancer” status. Based on data from the in vitro experiment for the U251 human glioblastoma and D384 medulloblastoma cell lines, 52 the model allows for a quantitative assessment of the effect of cancer cell killing. 25 Additionally, the application of the ionizing radiation beam (Figure 2C) enables the simulation of a radiotherapy treatment process by directing the beam to a specified location in the simulation space with an appropriately entered angle as an additional input parameter of the simulation.
What’s more, additional work is ongoing to develop the model’s capabilities and thus increase its applicability for potential users.
Exemplary Calibration
Calibration of the model requires determining the values of parameters used in the probability functions, so that the simulation results match experimental data for exact cell type. In particular, this chapter focuses on the exemplary calibration process on human lymphocytes as the most frequently used type of cells. Parameters from each function were determined individually, without including other mechanisms in the model, unless otherwise stated. Subsequent cellular effects were then included in the simulation to determine values more accurately. 24
There were different ways in which the calibration was done depending on the function type. In most cases, the calculated parameter was a mean value that characterizes the entire population, for example, mutation rate, division rate, or a number of lesions per cell. In some cases, two functions were related. For example, the number of damages in a cell depends on the probability that spontaneous damage occurs in a cell and on the probability of natural repair of one of the lesions in a cell. In those cases, parameter values were initially estimated for each individual function and then changed with each subsequent simulation. This allowed the observation of how parameter alterations influenced the obtained results and to identify which changes obtained results similar to those presented in experimental studies. 50
To determine the parameters describing the probability of cell multiplication and cell death, the mean frequency of cell division in a stable colony (where all the elements of the array are occupied by cells) was used, which was determined as an average for different types of lymphocytes. 45
The probability of cell death from radiation depends on whether it was hit or not. That is why the parameter used for calculation of the probability of death directly from radiation (a RD ) was determined simultaneously with the parameter used for finding the probability of a cell being hit (a hit ). Survival curves were generated for subsequent values of the parameter a hit and aRD, and a chi-square test was performed to see which curve was the best fit to the curve presented in the experimental study. 44
Parameters associated with the occurrence of mutations in a cell were determined for non-irradiated cells. 43 The constants for determining the probability of radiation damage were then adjusted to generate a correspondingly higher number of mutations after irradiation. 41 Then, because the frequency of occurrence of mutated cells is low, parameters for determining the probability of subsequent mutation 43 and malignant transformation 48 were determined using only the mutated cell population.
In the determination of parameters describing tumor growth, 49 a higher density of cancerous tumors was included by reducing the radius (r) around the mother cell (where a new cell can be located) by a value of 1. The value of the parameter used to determine the probability of death of a cancer cell due to its radiosensitivity was determined by finding the best fit of the survival curve—obtained in the model—to the experimental curve 40 while considering cancer cells that died due to irradiation.
Due to limitations in the available experimental data, calibration of {β} parameters (equations (2) and (3)) related to the bystander effect was carried out for fibroblasts irradiated with alpha particles. The number of damages induced by the bystander effect dominates for doses <0.2 Gy; therefore, the maximum value of the bystander effect probability was determined as the intersection point with the function describing the probability of radiation damage for a dose equal to 0.2 Gy. The parameter determining the reach of the bystander effect was chosen so that for a colony of 100 cells approximately 10% of them were damaged. 39
Parallelly, the values of the parameters used to describe the bystander effect according to the Japanese model (equation (4)) were taken directly from a paper describing the original model 29 (see Table 2). Analogically, parameters {α} describing the adaptive response (see equation (1) and Table 2) were taken directly from Reference 38.
Estimated values of all parameters (related to human lymphocytes) are presented in Table 3. These are calculated values based on the experimental data from specific papers and, due to large discrepancies in available experimental data, do not guarantee the consistency of simulation results with data from other articles, especially for different types of cells.
Deterministic Modeling of Irradiated Cells
Monte Carlo modeling uses stochastic formalism as it is the only possible mathematical description of a complex process such as the response to the irradiation of group of cells. It is not possible to present an equivalent single analytical description of this entire process, only of a selected process; this can be extracted from the probability tree and described with one analytical deterministic mathematical formula. Selected examples of such deterministic approaches, which can be considered as separate models describing specific biophysical phenomena, are presented and discussed below.
“Lesion to Cancer” Model
Based on the previously presented Monte Carlo model, it is possible to analytically present a narrow path from a broad probability tree, starting from the factor inducing a single damage to the neoplastic transformation.
37
This approach is called the “Lesion to Cancer” model
53
—it covers all processes from initial lesion (damage), through repair (or lack thereof), the generation and accumulation of mutations and, consequently, neoplastic transformation. This model focuses on the biophysical impact of a DNA region that is being hit. Based on general knowledge, it was assumed that in order to initiate the carcinogenic process, it is crucial to cause a mutation in the DNA coding region, preferably either in the oncogene or tumor suppressor gene. This is, however, a simplification of this complex mechanism, but it allowed us to make the model more biologically accurate. Thanks to the applied approaches, it was possible to develop one mathematical formula describing the number of cells that underwent neoplastic transformation:
37

Tumor Growth Model
For longer periods of time (t→∞), equation (6) presented in the previous subchapter can be summed over all values of M to obtain the tumor growth function. 37 This has been discussed in a recent paper by Fornalski et al, 54 where the Gompertz function for the dynamics of tumor growth was precisely described based on biophysical grounds. Additionally, the herein presented calculations have been independently supported by the Monte Carlo simulations23,25 demonstrating that the Gompertz curve does describe cancer cells dynamics adequately, except at the very early phase where a parabolic function is more appropriate. These results are consistent with the previous reports by other authors.55,56
Tumor growth modeling is a topic which can be discussed in the context of the dose, time, or even dose-rate relationships. In this regard, the Gompertz curve is not the only model that can be considered: other sigmoidal functions can, to a certain extent, also properly describe the tumor dynamics. 36
The Priming Dose Effect Model
The recent example of the deterministic solution related to the adaptive response description only is the one dedicated to the priming dose effect (also called the Raper–Yonezawa effect, which was mentioned earlier). In this case, the radiation adaptive response was taken as a theoretical background to calculate the percentage decrease (δ) of post-radiation mutations after the irradiation scheme of priming + challenging doses (D
1
+D
2
) in relation to the isolated challenging dose (D
2
) case:
38

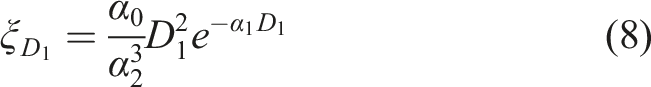
and {α} are mentioned earlier model’s free parameters (obtained via its calibration), while Δt corresponds to the time interval between small priming dose (D
1
) and the large challenging dose (D
2
). The diagram of the Raper–Yonezawa effect is presented in Figure 10. Diagram of the Raper–Yonezawa effect (also known as the priming dose effect): the single priming (small) dose (D1) generates much fewer negative effects than the single detrimental (high) dose (D2). However, when D2 follows D1 with some time distance between them (Δt), the overall biological effect for that D1+D2 total dose is lower than for single D2. The parameter δ is therefore showing the percentage difference between the effect (e.g., mutations or lesions) generated by the single dose D2 (without the priming dose) and the combination of D1+D2.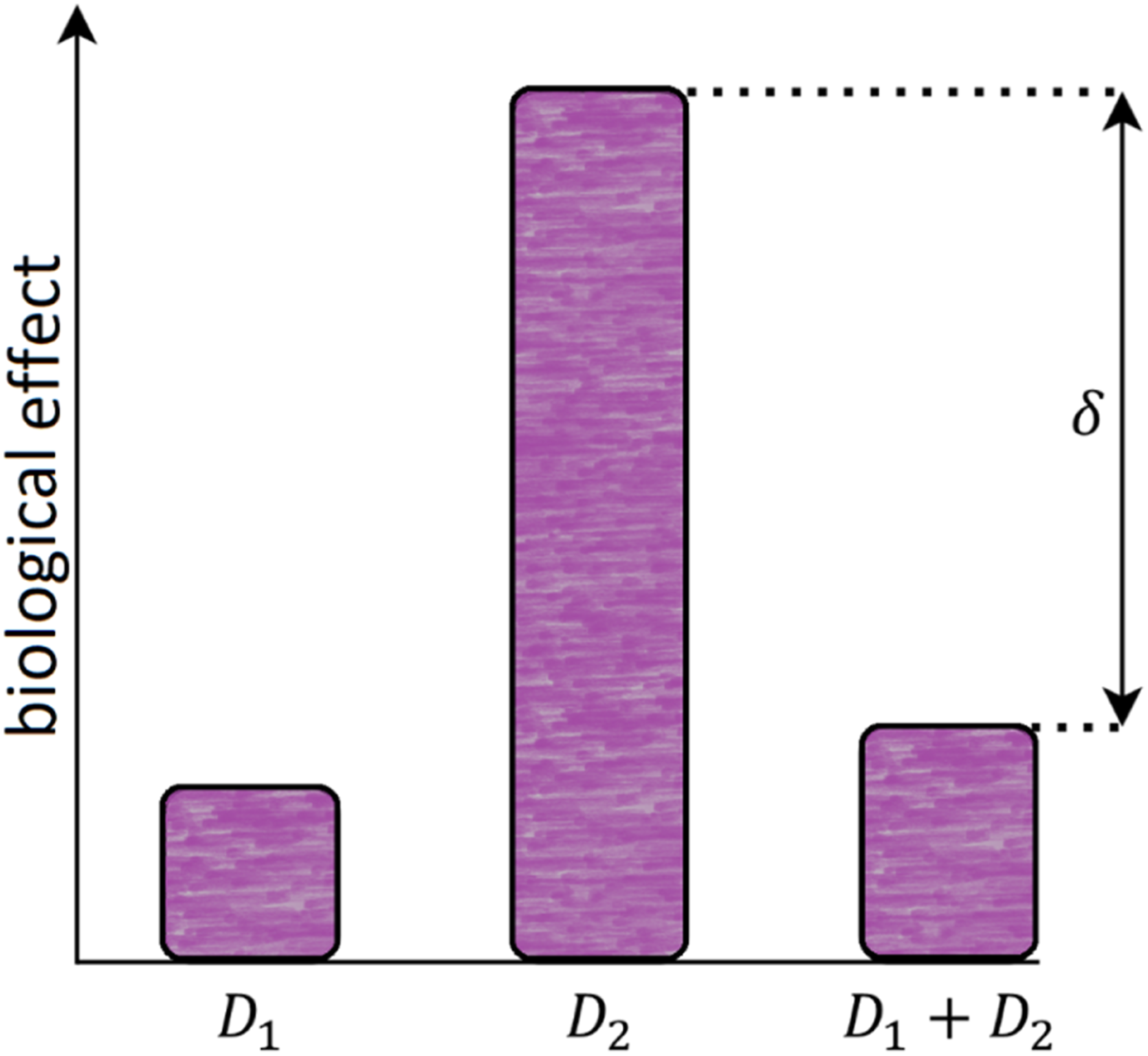
The presented model works well to describe the practical application of the phenomenon of radiation adaptive response. It was calibrated on various experimental data, such as lesions in human lymphocytes and chromosomal inversions in mice. 38
Constant High Background Radiation Model
Another popular aspect of the implementation of radiation adaptive response is the case of high background radiation areas (HBRAs), where some epidemiological studies show significant radioadaptation.57,58 However, the presented theoretical approach given by equation (1) is related to the single-dose pulse (D) while HBRA corresponds to the constant dose-rate (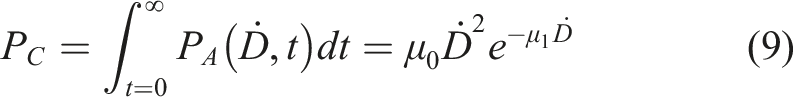
Discussion and Conclusions
This review article describes our proposed stochastic model of a cell group response to ionizing radiation using the Monte Carlo technique with a probability tree. This universal but simplified approach allows for modification and the creation of virtually any model: depending on the needs, the appropriate branches of the tree can be expanded; thus, enabling a more detailed simulation of the selected phenomenon. This model can be calibrated (i.e., its input parameters estimated) based on existing or dedicated experimental data describing a specific cell type, both human and animal.
Thanks to the applied stochastic approach, it is possible to treat a set of cells as a complex physical system, the precise analytical description of which seems to be currently impossible. However, using computer simulations, basically any simplified behavior of such cells can be modeled. An interesting case in particular is the determination of the probability of neoplastic transformation for a given dose, which can be used to determine the radiation risk (the risk of stochastic effects), especially cancer risk. For such an estimate to be reliable, the model takes into account many different types of biophysical phenomena, such as repair mechanisms, as well as the non-targeted effects, namely, the bystander effect and the adaptive response.
The presented model can be simplified and narrowed in certain ranges and, for example, only a selected phenomenon can be described, which can be demonstrated in the form of one narrow branch in a broad probability tree. Such a narrow branch, with the probability functions included within it, can therefore be written in an analytical form, creating de facto a new (secondary) deterministic model, which is, in fact, an independent fragment of the main model. So far, five such secondary deterministic models have been developed,36–38,54,59 but more are in preparation. This means that the presented approach is broad and universal which enables a relatively accurate biophysical description of various aspects of irradiated cells within natural model limits. Although each model is just an approximation of the reality, its proper implementation with validated input data allows for irradiated cellular studies, including cancer treatment, prognosis, or even prevention.
Footnotes
Acknowledgments
The authors would like to kindly thank Prof. Marek Janiak (Poland) and Dr Yehoshua Socol (Israel) for substantial help during the model development. Additional thanks to Prof. Marcin Kruszewski (Poland) and Dr Sylwester Sommer (Poland) for consultations in radiobiology and to Ms Charley Jeynes (University of Manchester) for linguistic corrections.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The research was funded by the National Centre for Nuclear Research (NCBJ), Poland.

