Abstract
Enumeration of bacteria is a large component in standard testing of antimicrobial textiles; however, there are multiple plating methods used that are tedious and labor intensive to perform. TEMPO is an instrument that automates much of the enumeration process using the most probable number (MPN) technique. Selective reagents for Staphylococcus aureus (salt) and Pseudomonas aeruginosa (cetrimide) were used to expand TEMPO capability. It was demonstrated that TEMPO had either the same or smaller variance than the traditional spread plating for cells in solution and on textiles tested using AATCC Test Method (TM) 100. TEMPO is a faster and more cost-effective method, and with expanded selective capabilities, can be used to improve enumeration efficiency.
Introduction
Soldiers are exposed to a wide range of environments in the field including heat and moisture, parameters that promote bacterial growth. A significant degree of antimicrobial functionality on soldier textiles is of high importance to inhibit bacterial growth. Pseudomonas aeruginosa and Staphylococcus aureus are two bacteria known to be associated with skin infections and odor.
The industry standard for evaluating such textile finishes is the American Association of Textile Chemists and Colorists (AATCC) Test Method (TM) 100. 1 Additionally, the spread, spiral, or pour plate methods used for enumeration in TM 100 require labor intensive preparation, consumption of large quantities of supplies, and lengthy evaluation of results (counted manually or in a quicker, yet still tedious method, counted by automated colony counter). Many antimicrobial textiles cannot be sterilized due to inactivation of the antimicrobial technology, and the material may possess contamination from non-targeted bacteria arising from handling or storage. A selective enumeration method is of interest as it would eliminate contaminating microorganisms from potentially giving inaccurate results.
Selective plating media have been used for enumeration of microorganisms in a variety of environments including food safety, soil and water analysis, and clinical sample testing. The TEMPO (bioMérieux) automated system is a relatively new method for enumerating bacterial samples. Originally made for food sample quality indicators, TEMPO has the potential to be applied to a wider range of applications, such as enumeration of soil, skin, or textile bacterial samples. The automated TEMPO most probable number (MPN) enumeration method has been validated with soy food products, 2 cheese, 3 spices and food additives, 4 as well as food and dairy products. 5 TEMPO possesses selective capabilities to enumerate Staphylococcus spp., Escherichia coli, Bacillus spp., lactic acid bacteria, yeasts, fungi, molds, and a general aerobic card for non-selective enumeration of microbes. One limitation is that selectivity is not available for all microbes of interest. The bioMérieux STA card is selective for Staphylococci, however, is incompatible with the S. aureus strain American Type Culture Collection (AATC) 6538 required for textile testing using TM 100 (data not shown). During previous studies, about a quarter of data collected with S. aureus 6538 was not able to be used. To be compliant with TM 100, an alternative selective enumeration was needed for S. aureus 6538.
TEMPO works by enumerating with a statistical method based on the MPN method. TEMPO uses plastic cards that have three rows of 16-column wells of three volumes that result in dilutions covering three orders of magnitude. The cards are filled with the contents from the single use, TEMPO vials containing powdered culture medium and a fluorescent indicator to indicate growth. After the cards are inoculated and filled with the TEMPO filler, the bacteria are grown overnight (or longer depending on the organism). The TEMPO reader utilizes a statistical method to analyze the signal from the fluorescent indicator to calculate the number of microorganisms present in the sample. This method is less labor intensive and can be conducted more quickly than traditional plating techniques.
The purpose of this study is to overcome the selectivity limitation of the TEMPO instrument in enumerating P. aeruginosa and S. aureus, specifically, selective automated enumeration to improve speed and efficiency. Modifications were made to the pre-existing TEMPO AC cards by using known selective agents, specifically cetrimide for P. aeruginosa 6 and high salt for S. aureus. 7 Here we demonstrate incorporation of these selective agents for the enumeration of P. aeruginosa and an additional strain of S. aureus, expanding the capability of the MPN method for enumeration of bacteria both in solution and from textiles. Selective MPN correlated well with non-selective MPN enumeration and was confirmed as a suitable substitute for selective spread plating.
Experimental
Reagents, Bacterial Strains, and Growth Conditions
Chemicals were purchased from Sigma Chemical Co. unless otherwise indicated. Phosphate buffered saline (PBS) pH 7.2 consisted of 137 mM NaCl, 2.7 mM KCl, 4.4 mM Na2HPO4, and 1.4 mM KH2PO4. Cetrimide agar and cetyl trimethyl ammonium bromide (CN supplement) were purchased from Oxoid and reconstituted to 200 mg/L cetrimide and 15 μg/mL nalidixic acid in sterile water. Mannitol salt agar and Dey-Engley (D-E) neutralizing broth was purchased from Becton Dickinson (BD). Bacterial strains used for this study— S. aureus aATCC 6538 and P. aeruginosa ATCC 9027, 15692, and 27317—were purchased from the American Type Culture Collection (ATCC) and rehydrated according to manufacturer's instructions. Cells were grown overnight in 3 mL nutrient broth (BD Difco) with agitation at 37 °C, then diluted to OD600 0.5-1 (equivalent of 108 CFU/mL) for analysis.
Pseudomonas Selectivity
TEMPO AC reagent vials were filled with 3.9 mL rehydration solution consisting of 2-fold dilutions of CN supplement ranging from 200 mg/L centrimide and 15 μg/mL nalidixic acid to 12.5 mg/L centrimide and 0.94 μg/mL nalidixic acid. For ease of discussion, CN dilutions will be referred to using their cetrimide concentrations 200, 100, 50, 25, and 12.5 mg/L. Cells only (0 mg/L CN) was run as a control. P. aeruginosa and S. aureus cultures were serially diluted 10-fold in PBS and each concentration run in triplicate, and 0.1 mL of each dilution was added to the vials. TEMPO AC cards for P. aeruginosa and S. aureus were then filled with the cetrimide rehydration solutions from the vials using the provided TEMPO filler instrument instructions, incubated at 37 °C for 24 h, and read by the TEMPO reader to determine bacterial concentrations.
To determine the CN dose response against other strains in addition to P. aeruginosa 9027, kinetic microplate growth studies were conducted based on Arcidiacono et al. 8 CN solution was serially diluted 2-fold to replicate concentrations of the TEMPO study and incubated with P. aeruginosa strains ATCC 15692 and 27317. Strain 9027 was also run to compare with TEMPO enumeration. Briefly, 1 × 105 CFU/mL cells were added to 200, 100, 50, 25, and 12.5 mg/L CN in TEMPO AC growth medium in triplicate in a 96-well microplate. Control wells with cells were inoculated without CN supplement present. Microplate incubation was overnight at 37 °C and the optical density at 595 nm was determined every 30 min using a GENios microplate reader (TECAN Austria GmbH).
Staphylococcus Selectivity
S. aureus was serially diluted 10-fold and 0.1 mL cells tested in triplicate under selective (AC vials filled with 3.9 mL of 75 g/L NaCl) and non-selective (3.9 mL of sterile water) conditions. The vials were filled following the instructions provided by bioMérieux. A control containing 75 g/L NaCl without cells was run to determine background or interference in the AC card from addition of salt. To validate salt selectivity, S. aureus and P. aeruginosa were grown and prepared separately, and each tested in triplicate in 75 g/L NaCl or sterile water. TEMPO reagent vials were then prepared and read as described previously.
Selective Spread Plating versus Selective TEMPO
Overnight cultures of S. aureus and P. aeruginosa were prepared and serially diluted 10-fold in PBS for enumeration. Each dilution (0.1 mL) was spread plated in triplicate on mannitol salts and cetrimide agar for S. aureus and P. aeruginosa respectively. These dilutions were also enumerated by TEMPO. Vials were filled with 3.9 mL of NaCl or CN selective rehydration solution for S. aureus and P. aeruginosa respectively. Each dilution (0.1 mL) was added to respective vials for a final volume of 4 mL. The vials were processed as previously described.
TM 100 Selective versus Non-Selective TEMPO
TM 100 was used to evaluate if selective enumeration from textiles was possible using TEMPO. Two textiles were evaluated: Fabric 1 was untreated and Fabric 2 contained an antimicrobial treatment. Each test consisted of S. aureus and P. aeruginosa on three replicates of single textile swatches; tests were conducted in triplicate. Cells were grown and prepared as described previously for a target concentration of ∼108 CFU/mL. TM 100 was conducted with the following modifications: 1–2 × 107 CFU/mL cells were washed in 0.125× nutrient broth with 0.15% (w/v) Triton X-100 and applied to single textile swatches, using a volume to wet but not overly saturate the material. After 0 h and 24 h contact times, bacteria were recovered by placing the swatches in D-E broth at 100× the inoculum volume. D-E with cells was serially diluted 10-fold in PBS. Dilutions for each swatch replicate and inoculum were enumerated in duplicate by the TEMPO instrument using the General AC kit (non-selective) and AC cards with 75 g/L NaCl or 50 mg/L CN for selective enumeration of S. aureus and P. aeruginosa respectively.
For all plating studies, due to the dilutions plated, a cell concentration of 1 × 102 (2.00 log10) CFU/mL was the limit of detection.
Results and Discussion
There were two motivations for this study; the first was the lack of a specific medium for automated enumeration of Pseudomonas selectively, and second were issues involving using the TEMPO STA card enumeration of S. aureus 6538, an organism required by TM 100. When the STA card was initially used, ∼20–25% of data for each experiment resulted in read errors and invalid test results (data not shown). A second S. aureus strain, ATCC 27217, did not exhibit faulty readings (data not shown). The cause for this could not be identified by our own studies nor in discussions with bioMérieux.
Cetrimide Selectivity
A dose response study was conducted in TEMPO with CN to determine the CN concentration needed to create a selective media to enumerate P. aeruginosa. The selectivity of CN for P. aeruginosa was tested against S. aureus, each at a target concentration of 107 CFU/mL. In Fig. 1, CN dilutions with 50 to 12.5 mg/L cetrimide showed no reduction of P. aeruginosa, a slight reduction occurred at 100 mg/L, and a ∼1.5 log reduction was observed at 200 mg/L. The negative control of 0 mg/L CN showed full growth of both P. aeruginosa and S. aureus. CN demonstrated selectivity against S. aureus, with a significant reduction at 12.5 mg/L and complete reduction at and above 25 mg/L cetrimide. A CN solution of 25–50 mg/L cetrimide was sufficient to use as a selective agent, allowing for full growth of P. aeruginosa while completely inhibiting S. aureus. The concentration of CN chosen for further experiments was 50 mg/L, or 0.005%.
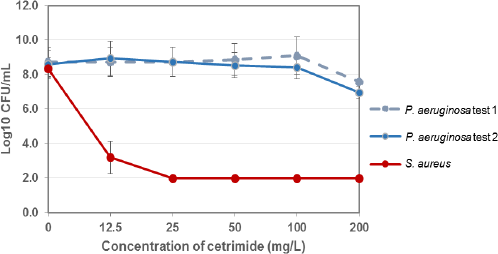
Determination of selective concentration of cetrimide in CN supplement. For each test, n = 3. Two separate tests of P. aeruginosa are shown. A concentration of 25-50 mg/L is selective for P. aeruginosa and inhibits S. aureus growth.
Cetrimide is a known selective agent for P. aeruginosa, and is used in numerous applications such as examination of cosmetics, 9 clinical specimens, 10 and evaluating disinfectant efficacy. 11 The standard concentration of cetrimide in selective medium was previously explored and determined to be optimal at a concentration of 0.03%. 6 Our results show that 25-50 mg/L (0.0025-0.005%) of CN was effective, which is significantly less than the 0.03% reported in literature when used in agar media for spread plating. There are two possible reasons for this finding: first, the nalidixic acid present in the CN supplement reduced the amount of cetrimide needed to be effective for selectivity of Pseudomonas, 12 and second, CN was more effective in the liquid TEMPO hydration solution than on a solid growth substrate. Since the recommended concentration of 200 mg/L CN in TEMPO resulted in a reduction of P. aeruginosa 9027 growth, kinetic microplate growth studies using 2-fold dilutions of CN were conducted to determine if this also occurred with other strains. P. aeruginosa strains 15692 and 27317 showed similar results, with growth unaffected at less than or equal to 50 mg/L CN, but slight and greater reductions occurred at 100 and 200 mg/L respectively (Fig. 2). P. aeruginosa 9027 showed the same growth patterns, which correlates to the results seen using TEMPO.
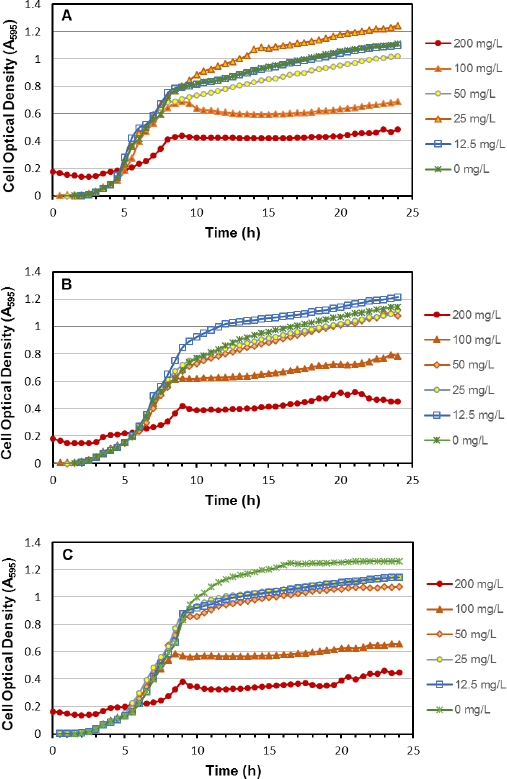
Kinetic microplate assays to determine effect of CN supplement concentration for P. aeruginosa strains: A. ATCC 9027, B. ATCC 15692, C. ATCC 27317. For all strains, growth at 50 mg/L and below of CN supplement was similar to cells only. Growth reduction was observed at 100 and 200 mg/L. Concentration effects on the 9027 growth curves correlate with results from enumeration using TEMPO.
Sodium Chloride Selectivity
Salt is the selective component of mannitol salts agar recommended for isolating pathogenic Staphylococci from clinical specimens, 13 cosmetics, 14 and microbial limit tests. 15 A study was done to determine if the 75 g/L NaCl used in selective plating media would be effective as a selective reagent for S. aureus when using the TEMPO. Both bacteria were enumerated twice, once with pure water as the TEMPO rehydration solution and once with 75 g/L NaCl as the rehydration solution. The S. aureus concentration was statistically the same in water and NaCl (data not shown). The selectivity of salt for S. aureus was tested against P. aeruginosa, each at a target concentration of 107 CFU/mL. Fig. 3 shows that when NaCl is present, P. aeruginosa was not detectable. A control without cells was run to determine if the addition of NaCl to the TEMPO hydration solution caused signal interference or additional problems in the enumeration step—no interference from NaCl was observed. The selectivity of NaCl using TEMPO concur with the findings by Chapman 7 who reported that 75 g/L (7.5%) NaCl is an effective selective agent for pathogenic and nonpathogenic Staphylococci.
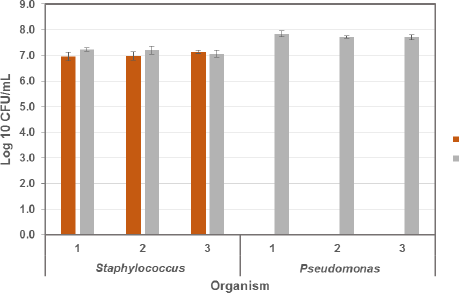
Log10 CFU/mL counts for both S. aureus and P. aeruginosa when enumerated with TEMPO in a selective solution of 75 g/L NaCl versus a non-selective solution of sterile water. Each organism was tested in triplicate. Because Pseudomonas was not detected in TEMPO under selective conditions, the salt solution was effective for S. aureus selective enumeration.
Selective TEMPO versus Selective Spread Plating
A study was done to confirm that the use of selective media with the TEMPO automated enumeration MPN instrument is a valid method to enumerate bacteria compared to spread plating. Statistical analysis was done to compare the variance for S. aureus and P. aeruginosa between plate count and TEMPO and is shown in a box and whisker plot (Fig. 4). For S. aureus, the mean log10 CFU/mL counts and variations were statistically equal for both methods. For P. aeruginosa, the variance was significantly smaller for TEMPO, indicating that TEMPO is a suitable method to replace spread plating.
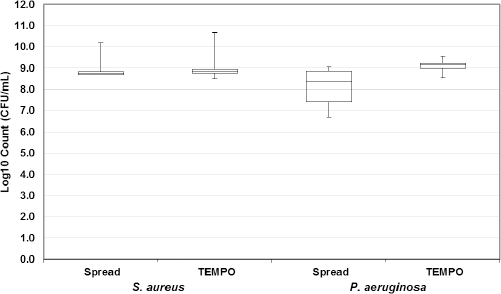
Bacterial counts for selective spread plate and TEMPO solutions. Cultures of S. aureus and P. aeruginosa were grown overnight and enumerated by spread plating and TEMPO (n = 4). Both enumeration methods used selective agents, mannitol salt agars and cetrimide agar plates for spread plating and 75 g/L NaCl and 50 mg/mL CN supplement for TEMPO. Enumeration by spread plating and TEMPO for both organisms produced similar results and were of equal variance.
TM 100 Enumeration by TEMPO with Selective versus Non-Selective Media
Testing was done to validate the developed selective media TEMPO method against the general bioMérieux AC cards in a full TM 100 test method environment. Samples were enumerated by both selective and non-selective TEMPO methods and statistical analysis was run to compare the variance between the results.
Fig. 5 compares the log reductions within a fabric enumerated by selective and non-selective TEMPO. Tests were run in triplicate for both bacteria, which were then averaged. The untreated Fabric 1 behaved as expected, showing essentially no reduction for S. aureus and growth for P. aeruginosa.
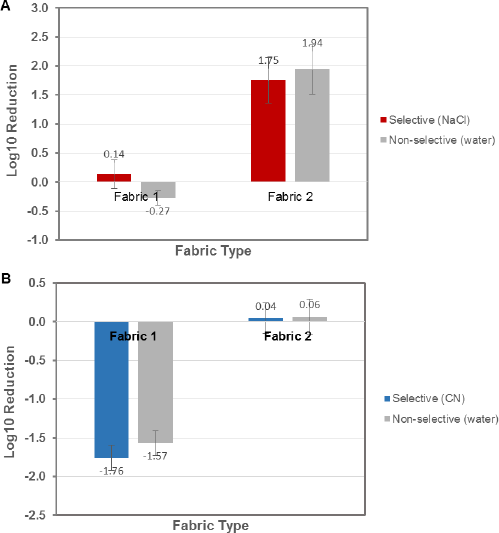
Selective and non-selective enumeration by TEMPO: A. S. aureus, B.P. aeruginosa, isolated from textiles using TM 100. Log reductions were determined from Fabric 1 (an untreated control) and Fabric 2 (treated with an antimicrobial technology). The selective media that were developed produced similar results to the non-selective media enumeration (n = 3).
Treated Fabric 2 resulted in a ∼2-log reduction for S. aureus, while P. aeruginosa growth was inhibited. For both fabrics, the enumerated cell counts were statistically similar using non-selective and selective conditions, with less than 0.25 log CFU/mL variance between tests. These results indicate that the developed selective method produced results similar to the pre-existing non-selective method and that the new selective method can be substituted in place of spread plating.
Use of the selectivity approach developed in this study allows for confidence in enumeration of the target bacteria in solution and on swatches, assuring the accurate effectiveness assessment of an antimicrobial applied to textiles. When added to TEMPO, agents CN and NaCl performed as intended, exhibiting selectivity for the appropriate target microorganisms. These results support the hypothesis of incorporating selective agents into an existing automated MPN method that can also translate to use with other non-sterile samples.
Use of plating for enumeration of large numbers of samples is labor and reagent intensive. Automated enumeration reduces the time needed for plate preparation, plating, and counting. Automated colony counters are helpful, but in some cases, their use may be difficult if there is insufficient contrast between the bacterial colonies and the growth medium, as has been observed with P. aeruginosa on cetrimide agar plates (data not shown). Enumeration is a major part of TM 100; incorporation of automated enumeration resulted in a 25% reduction in the overall time to conduct TM 100 relative to traditional methods.
Conclusions
In this work, we demonstrated that selective TEMPO enumeration was comparable to the traditional spread plating on selective media, and by extension, would also be comparable to spiral plating and pour plating. Automated selective enumeration was demonstrated with the organisms used in solution and with AATCC TM 100.
While selectivity against other strains was not determined, selectivity of cetrimide and NaCl is widely accepted and would be expected to work for other strains of P. aeruginosa and S. aureus. TEMPO was also shown to be a less labor-intensive method than spread plating, improving testing efficiency.
This study shows that not only can these selective agents for P. aeruginosa and S. aureus be used in the automated enumeration TEMPO instrument, but it also shows the potential for further manipulation of selectivity in other types of bacterial enumeration methods. For example, a previous study made modifications to a pre-established MPN enumeration method to separate aromatic and aliphatic hydrocarbon degrading bacteria in petroleum samples. 16 An additional study miniaturized a preexisting MPN method for oil-degrading microorganisms to enumerate microorganisms while at sea. 17 Fungal enumeration with TEMPO has also been expanded from yeast and molds to include filamentous fungi. 18
In the future, our modifications could potentially be applied to other skin microorganisms such as Micrococcus or Corynebacterium species. FTO agar (containing furoxone: 1-N-(5-nitro-2-furfuryli-dene)-3-amino-2-oxazolidene) was developed as the selective agent for Micrococci against Staphylococci; it can also be selective for Corynebacteria over Staphylococci, although to a lesser extent. 19 CBU medium was also shown to be selective for Corynebacterium species while inhibiting many other organisms. 20 Adapting selective enumeration methods to automated MPN could have implications for other microorganisms of interest, improving enumeration efficiency for numerous applications.
