Abstract
Kell and XK are related because in red cells they exist as a disulfide-bonded complex. Kell is an endothelin-3-converting enzyme, and XK is predicted to be a transporter. Absence of XK, which is accompanied by reduced Kell on red cells, results in acanthocytosis and late-onset forms of central nervous system and neuromuscular abnormalities that characterize the McLeod syndrome. In this study, expression of mouse XK, XPLAC, a homolog of XK, and Kell were compared by in situ hybridization histochemistry (ISHH) and RT-PCR. ISHH showed that Kell and XK are coexpressed in erythroid tissues. ISHH detected XK, but not Kell, mRNA in testis, but RT-PCR indicated that both Kell and XK are coexpressed. XK, but not Kell, was significantly expressed in brain, spinal cord, small intestine, heart, stomach, bladder, and kidney. ISHH did not detect XK in skeletal muscle but RT-PCR did. In brain, XK was predominantly expressed in neuronal rather than in supportive cells. By contrast, XPLAC was predominantly expressed in the thymus. Coexpression of Kell and XK in erythroid tissues and the different expressions in non-erythroid tissues suggest that XK may have a complementary hematological function with Kell and a separate role in other tissues.
XK
Available data on the expressions of Kell and XK were generated mainly by Northern blot and dot-blot analyses of different tissues (Ho et al. 1994; Russo et al. 2000; Camara-Clayette et al. 2001; Calenda et al. 20006). Expression of Kell was originally thought to be mainly in erythroid tissues (Lee et al. 1991), but later reports indicated that Kell was also present in testis and in small amounts in many other tissues (Russo et al. 2000; Camara-Clayette et al. 2001). Northern blots of human tissues indicated that XK is ubiquitously present in all tissues examined with highest levels in erythroid tissues and skeletal muscle. Because of the unequal expression of Kell with XK, it has been speculated that Kell and XK are not always covalently linked and that the functions of XK may be different, depending on whether it is complexed with Kell or if it exists by itself.
An immunohistochemical study of skeletal muscle from normal and McLeod patients showed that in normal tissues, but not in McLeod, XK is expressed in the sarcoplasmic reticulum of type II muscle cells but, unlike Kell, was not present in the sarcoplamic membrane of normal cells (Jung et al. 2001). Studies with transfected COS cells programmed to coexpress Kell and XK demonstrated that Kell and XK are covalently linked in the endoplasmic reticulum, and some Kell/XK complex travels to the plasma membrane (Russo et al. 1999). However, expression of XK and Kell by themselves also allowed them to be individually transported to the cell surface, indicating that linkage of Kell and XK is not obligatory for cell surface expression. The KEL promoter exhibits a stronger transcriptional activity in K562 cells of erythroid origin than in HeLa cells, indicating a preference for Kell expression in erythroid cells (Camara-Clayette et al. 2001).
Physiological functions of Kell and XK are not yet well understood. Kell is an endothelin-3-converting enzyme and its product, endothelin-3, is known to be involved in many biological processes, including regulation of blood pressure by affecting the contraction and proliferation of vascular smooth muscle, and developmental processes of enteric nervous system and melanocytes by affecting migration and differentiation of neural crest cells (Masaki 1995; McCallion and Chakravarti 2001; Motohashi et al. 2006; Nagy and Goldstein 2006). However, in humans, the rare Kell null phenotype that lacks Kell protein (Lee et al. 2001) does not express any obvious clinical disorders. By contrast, although the transport substrate for XK is unknown, absence of XK as occurs in the McLeod phenotype defines a set of clinical symptoms that invoke a role for XK in neuromuscular, central nervous system, and hematological functions (Danek et al. 2001a). unlike in humans, the pathogenicity caused by dysfunctional XK has not yet been established in murines.
McLeod phenotypes are derived from various XK gene mutations. To date, almost all the XK mutations in McLeod phenotypes result in either absence of XK or in truncated XK proteins caused by prematurely created stop codons (Danek et al. 2001a; Walker et al. 2007). All the truncations occur before cysteine (aa 347 of XK) that forms a complex with Kell. There are two reported missense mutations in XK and one single nucleotide mutation in an intron of XK near the splice junction that result in minimal expression of McLeod syndrome at ages that would be expected to develop clinical symptoms (Daniels et al. 1996; Russo et al. 2002; Jung et al. 2003; Walker et al. 2007).
A Caenorhabditis elegans protein, ced-8, which belongs to the outgroup cluster in the phylogenetic tree of XK-related proteins (Calenda et al. 2006), has been described. Ced-8 is involved in the timing of programmed cell death in C. elegans and is thought to function in downstream apoptotic processes (Ellis et al. 1991; Stanfield and Horvitz 2000). Whether the function of XK is similar to ced-8 remains to be shown. Homology of ced-8 with XK is weak, and thus XK may not share a similar function even though ced-8 shares the common motif, termed the ced-8 domain, which is present in all XK-related proteins.
Further information on the expression patterns of Kell, XK, and of an XK homolog, XpLAC, should prove useful in the eventual understanding of their physiological functions. To this end, we have performed a comparative in situ hybridization histochemistry (ISHH) study in mice, with RT-PCR when needed to verify ISHH, of these related and relevant mRNAs.
Materials and Methods
Animals
Brain, testis, and spleen tissues were obtained from adult male or female C57BL6 mice with a minimum age of 7-8 weeks. C57BL6, day 1, newborn mice were used for whole body sagittal sections. Animals were housed and tissues were collected according to the guidelines of the Institutional Animal Care and Use Committee (IACUC) of the New York Blood Center and the Ethical Committee of the Centre Hospitalier de l'Université de Montréal.
ISHH
ISHH was performed essentially as described previously (Young et al. 1986) with the following modifications for the detection of Kell, XK, and XPLAC transcripts.
Probe Preparations
mKell. For the preparation of antisense probe, a full-length mouse Kell cDNA (3′ to 5′ orientation) in pTAdv vector (Invitrogen; Carlsbad, CA) was cut with EcoRI (in the vector) and HindIII (Kell at nt 657), and the resulting Kell 1-657 cDNA fragment was subcloned in pBCSK(+) (Stratagene; La Jolla, CA), placing it in a 3′ to 5′ orientation for the T7 promoter. For the preparation of a sense probe, the same mouse Kell cDNA in pTAdv vector was cut with Xhol (in the vector) and HindIII (Kell at nt 657), and the resulting fragment was subcloned in pBCSK(+), placing it in a 5′ to 3′ orientation for the T7 promoter. A Smal site was used to linearize the plasmids for cRNA synthesis.
mXK. The following PCR primers were used in the amplification of a mouse XK cDNA fragment of 413 bp: mXK650F, 5′-GCTTCATCATGACCTTGTCCCTGCTGTC-3′ and mXK1035R, 5′-GAGCTCCGGATTGTCGATTTTCAGCTGA-3′.
The PCR product was subcloned in pCR4-TOPO vector (Invitrogen). An antisense clone and a sense clone (negative control) for the T7 promoter were used. Both plasmids were cut with PstI to linearize for cRNA synthesis.
mXPLAC. The following primers were used to amplify a 629-bp of mXPLAC cDNA: XPISHF, 5′-GAGCTCTGAACTCCAGACAGTAAGAGAG-3′ and XPISHR, 5′-TCTGCTGAGATCAGACTCACAT-3′.
The amplified cDNA fragment was subcloned in pCR4-TOPO vector in a 3′ to 5′ orientation of the vector. For anti-sense riboprobe synthesis, T3 promoter was utilized following cutting the plasmid with NotI. For sense probe, T7 promoter was utilized following cutting the plasmid with PstI.
All the cRNA transcripts were synthesized from the templates in vitro, according to the manufacturer's conditions (Ambion; Austin, TX) and labeled with 35S-UTP (>1000 Ci/mmol; Amersham, Pittsburgh, PA).
Tissue Preparation
Harvested and trimmed tissues were immediately placed in precooled 2-methylbutane at −35C for 5 to 7 min. Tissues were serially cryosectioned into 6- to 8-μm thickness, mounted on gelatin-coated microscope slides, and stored at −80C. Before ISHH, sections were fixed in 4% paraformaldehyde (Sigma-Aldrich; St Louis, MO) in PBS, treated with triethanolamine/acetic anhydride, washed, and dehydrated with a series of ethanols.
Hybridization
For the ISH experiments, the custom service from Phylogeny Inc. (Columbus, OH) was used (Lyons et al. 1990; Patapoutian et al. 1993). Briefly, tissue sections were hybridized overnight at 55C in 50% deionized formamide, 0.3 M NaCl, 20 mM Tris-HCl, pH 7.4, 5 mM EDTA, 10 mM NaH2PO4, 10% dextran sulfate, 1X Denhardt's, 50 μg/ml total yeast RNA, and 50-80,000 cpm/μl 35S-labeled cRNA probe. The tissue was subjected to stringent washing at 65C in 50% formamide, 2X SSC and 0.1X SSC for 10 min at 37C, the slides were dehydrated and dipped in NTB-2 nuclear track emulsion (Kodak; Rochester, NY), and exposed for 21 days. Two sets of slides were hybridized with antisense riboprobes, and one set of slides was hybridized with negative control sense riboprobes. Tissues were validated by ISHH to unrelated genes CD4, CD8, and CD38 as markers of spleen tissues (data not shown).
Imaging
Following ISH, gene expression patterns were analyzed by both x-ray film autoradiography for anatomical level analysis (5-day exposure time; MioMaxMR, Kodak) and emulsion autoradiography for cellular level analysis. Unstained sections or sections slightly stained with hematoxylin were viewed under both darkfield and lightfield illumination. Anatomic level data are shown at low magnification as bright labeling on dark field. Cellular level hybridization is shown at higher microscopical magnification as black labeling by silver grains on hematoxylin-stained background. In some cases, intermediate illumination that allows one to see hematoxylinstained blue nuclei as well as bright silver grains was used with combined optics that include a darkfield condenser and a brightfield lens.
Northern Blot
For XK Northern blot probe, a 600-bp cDNA fragment of XK cDNA cut with Pfl3MI (nt410-nt1027) was used. All probes were labeled with 32P using [α- 32P]dCTP (Amersham Biosciences) and used in hybridization of the human brain multiple tissue Northern blots (Clontech; Mountain View, CA). Hybridization was carried out overnight at 67C following prehybridization at 67C for 2 hr. Blots were washed once with 2X SSC containing 0.1% SDS at room temperature for 5 min, twice with 0.5X SSC containing 0.1% SDS at 67C for 20 min, and once with 0.1X SSC containing 0.1% SDS at 67C for 15-20 min.
Cell Culture
C2C12 cells (ATCC; Manassas, VA), a mouse myoblast cell line of skeletal muscle origin, were grown in monolayers in complete growth medium (DMEM with 4 mM
RT-PCR
Mouse tissues were removed, washed in PBS to remove blood, and homogenized in Trizol reagent (Invitrogen) using Power-Gen 125 (Fisher Scientific; Hampton, NH). Cultured C2C12 cells were directly lysed in Trizol reagent. RNA was extracted following the protocol provided by the manufacturer. Reverse transcription and DNA amplification were performed using GeneAmp RNA PCR kit (Applied Biosystems, Roche; Branchburg, NJ). Reverse transcription was carried out for 10 min at room temperature, 40 min at 42C, and 7 min at 99C. PCR amplification was performed for 32 cycles (94C for 30 sec, 62C for 30 sec, and 72C for 30 sec). Primers used for PCR are as follows: mXK (373 bp): MmXK-1F, 5′-CTGTCAGTCCGACCAGAATGAAGAACCT-3′; MmXK-1R, 5′-CCAGAGGAAGATACAGACATAAGCCAGG-3′; mXPLAC (456 bp): MmXP-1F, 5′-CTAACCCACGCTTTACCTTCCCGTTTAG-3′; MmXP-1R 5′-CCAGGAAGGCTTGAATCTGTGACATACG-3′; mKell (352 bp): MEX12F, 5′-AAGGATGCTGTCATCATACGCCTC-3′;MEX15R, 5′-GGTGGAAGAATGGAGGTTGGAGAA-3′; mGPA (235 bp): mGPA-F, 5′-TCCTGTGGTGGCTTCAACTGTAGGTAAC-3′;mGPA-R, 5′-GATAGTTCCGATAATCCCTGCCATCACG-3′; G3PDH (456 bp): G3PDH-F, 5′-GACCACAGTCCATGCCATCACTGC-3′; and G3PDH-R, 5′-AGGTCCACCACCCTGTTGCTGTA-3′.
The expected size of the PCR products is shown in parentheses following the identification of the genes as above. PCR products were analyzed by electrophoresis on 1% agarose gel.
Results
ISHH of mKell, mXK, and mXPLAC mRNAs
mKell. ISHH of a whole sagittal section of newborn mouse (Figure 1-I) showed that mKell mRNA is present in high amounts in erythroid tissues, spleen, bone marrow, and liver of a 1-day-old mouse but was not detected in brain or in testis. There was questionable labeling of mKell mRNA in the marginal zone of the cortex of the kidney. Microscopical analysis of the emulsion autoradiography of an adult spleen section showed that Kell is expressed in the red pulp, which is peripherally located, but not in the white pulp (Figures 1-IIA and IIB).
mXK. A sagittal section of a newborn mouse (Figures 2-I-2-III) showed mXK mRNA expression in the erythroid tissues, spleen, bone marrow, and liver and also in most other tissues including brain, spinal cord, layers of epithelial cells of small intestine, heart, stomach, transitional epithelium, and smooth muscle of the bladder and cortex of kidney. mXK mRNA in testis was detected in the cells of the seminiferous tubules (Figure 2-IV). Expression pattern of mXK mRNA in the adult spleen (Figure 2-V) was similar to the one shown for mKell and was present mostly in the red pulp.
mXK mRNA expression in adult brain was seen in many different regions of the brain (Figure 3-I), notably in the Purkinje cells of the cerebellum (Figure 3-II), magnocellular neurons of pons (Figure 3-III), mitral cell layer of olfactory lobe (Figure 3-IV), the cornu ammonis of hippocampus and dentate gyrus, and at low to moderate expression in cerebral cortex, caudate putamen, and hypothalamus (Figure 3-V). Brain gray matter supportive (glial) cells were not labeled. White matter oligodendroglial cells in the corpus callosum were also not labeled (Figure 3-I). Together the pattern of expression in the brain leads to the conclusion that XK expression is predominantly in neuronal rather than in supportive cells of the brain.
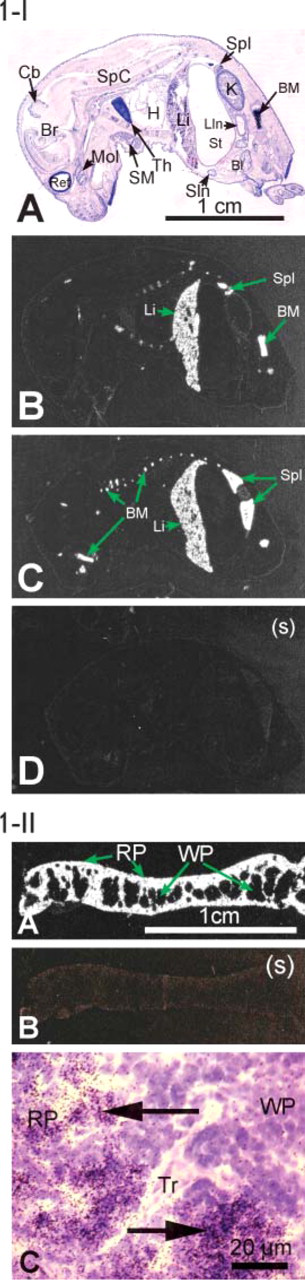
Expression of mKell mRNA. (
Expression of XK mRNA in human brain studied by Northern blots (Figure 3-VI) supports the ISHH results of mXK mRNA expression in mouse brain with the exception that, by Northern blot, a weak band was detected in human corpus callosum. Notably, in the XK Northern blots of human brain, the putamen region contains a second, slightly smaller minor transcript than the normal transcript. The original Northern blot of brain XK reported by Ho et al. (1994) also detected the presence of a minor XK transcript.
mXPLAC. In the newborn mouse (1-day old), a sagittal ISHH section showed expression ofmXPLAC mRNA in the thymus, skin, and stomach wall (Figure 4-I). An adult mouse sagittal section showed that mXPLAC mRNA is present in moderate levels in spleen (data not shown). Microscopical analysis showed that most labelings are seen in the cortex of the thymus (Figure 4-II) and in the squamous cornified epithelium of skin (Figure 4-III). mXPLAC mRNA was also present in the hair follicles of the dermis, mostly in hair bulb (Figure 4-IV) and within the external root sheaths and within the inner root sheaths of whiskers (Figure 4-V).
RT-PCR
Although ISHH did not detect mouse Kell mRNA in testis, it was shown to be present by RT-PCR (Figure 5A). The 352-bp band in Figure 5A, Lane 1 is the mKell-amplified product. Two light bands of ∼800 and 900 bp are due to PCR artifacts, and the 1010-bp band is the amplified product of the Kell gene from genomic DNA. mXPLAC mRNA is also present in a small amount in testis (Figure 5A, Lane 2) as is XK mRNA (Figure 5A, Lane 3).
XK transcript, but not Kell or XPLAC (Figures 5B and 5C), was present as measured by RT-PCR in mouse skeletal muscle and in C2C12 cells, a mouse myoblast cell line. The 1010-bp band in mKell lane (Figure 5B, Lane 2) is the amplified product of the Kell gene from genomic DNA. An 800-bp light band (Figure 5B, Lane 2) observed for mKell and three light bands in mXK lane (Figure 5B, Lane 4) are PCR artifacts. A small amount of mGPA, an erythrocyte marker, was amplified in skeletal muscle tissue (Figure 5B, Lane 6), indicating slight contamination of blood with the skeletal muscle mRNA preparation.
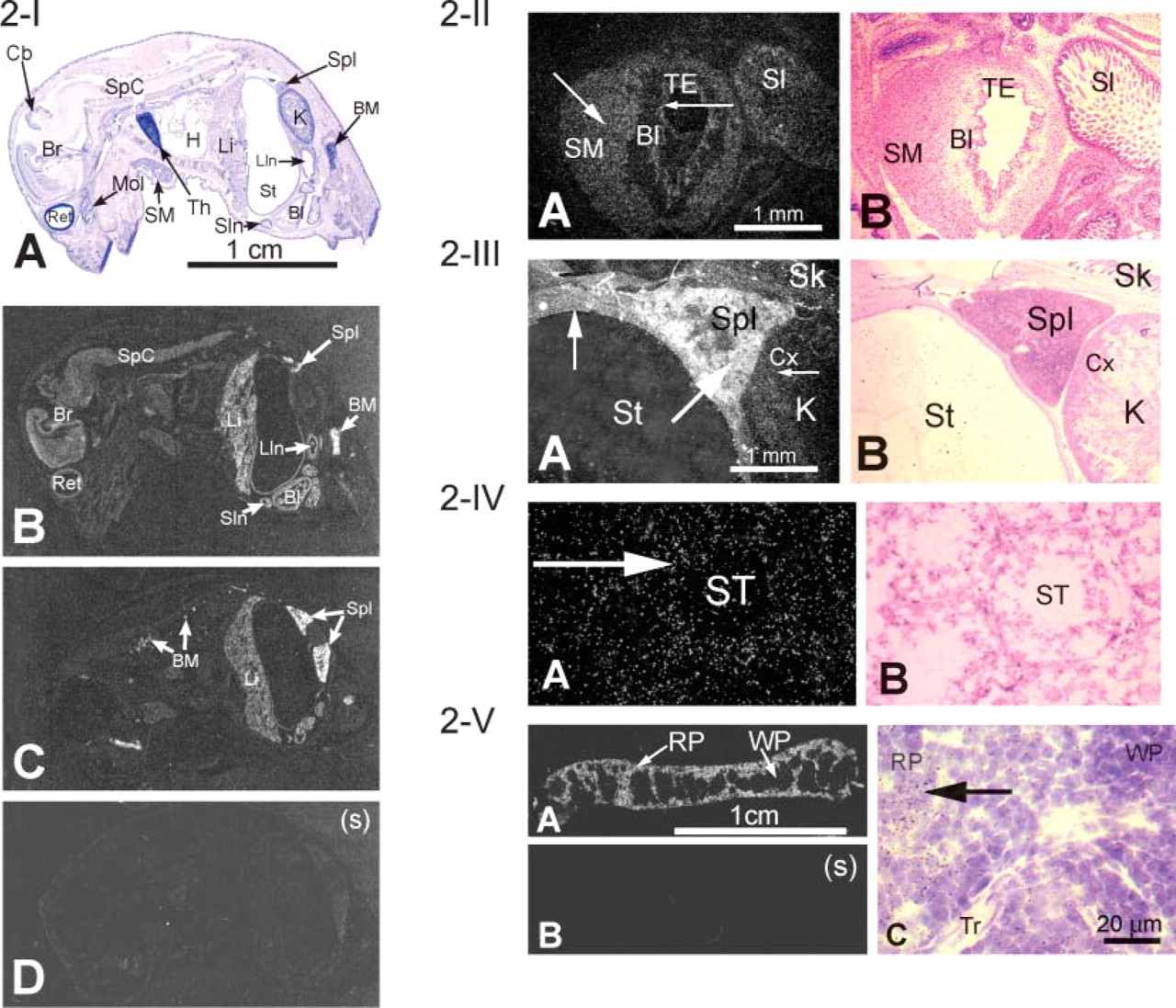
mXK mRNA expression profile in different tissues of the newborn mouse. (
ISHH did not detect mKell mRNA on mouse cerebellum, but RT-PCR showed a very weak 352-bp mKell band (Figure 5D, Lane 2, arrow). However, an mGPA band (235 bp) was also present (Figure 5D, Lane 6), indicating some erythroid cell contamination. mXPLAC was also positive by RT-PCR (456 bp), indicating that mXPLAC may be expressed in small amounts in brain (Figure 5D, Lane 3).
Discussion
Kell expression appears to be restricted to erythroid tissues and testis. The GATA family of transcriptional factors is not only specific in erythroid tissues but is also important in the expression of testis-specific proteins (Hales 2001). The Kell promoter contains multiple conserved GATA-1 binding sites and is expected to regulate late expression in both erythroid tissues and testis (Lee et al. 1995). We did not detect mKell mRNA in testis by ISHH but detected mKell mRNA by RT-PCR. The discrepancy may arise from the sensitivity of the two techniques. Human testis may have greater amounts of mKell than mouse testis. Northern and dot-blot analyses of human testis showed strong signals for Kell (Russo et al. 2000; Camara-Clayette et al. 2001), suggesting the possibility of species difference.
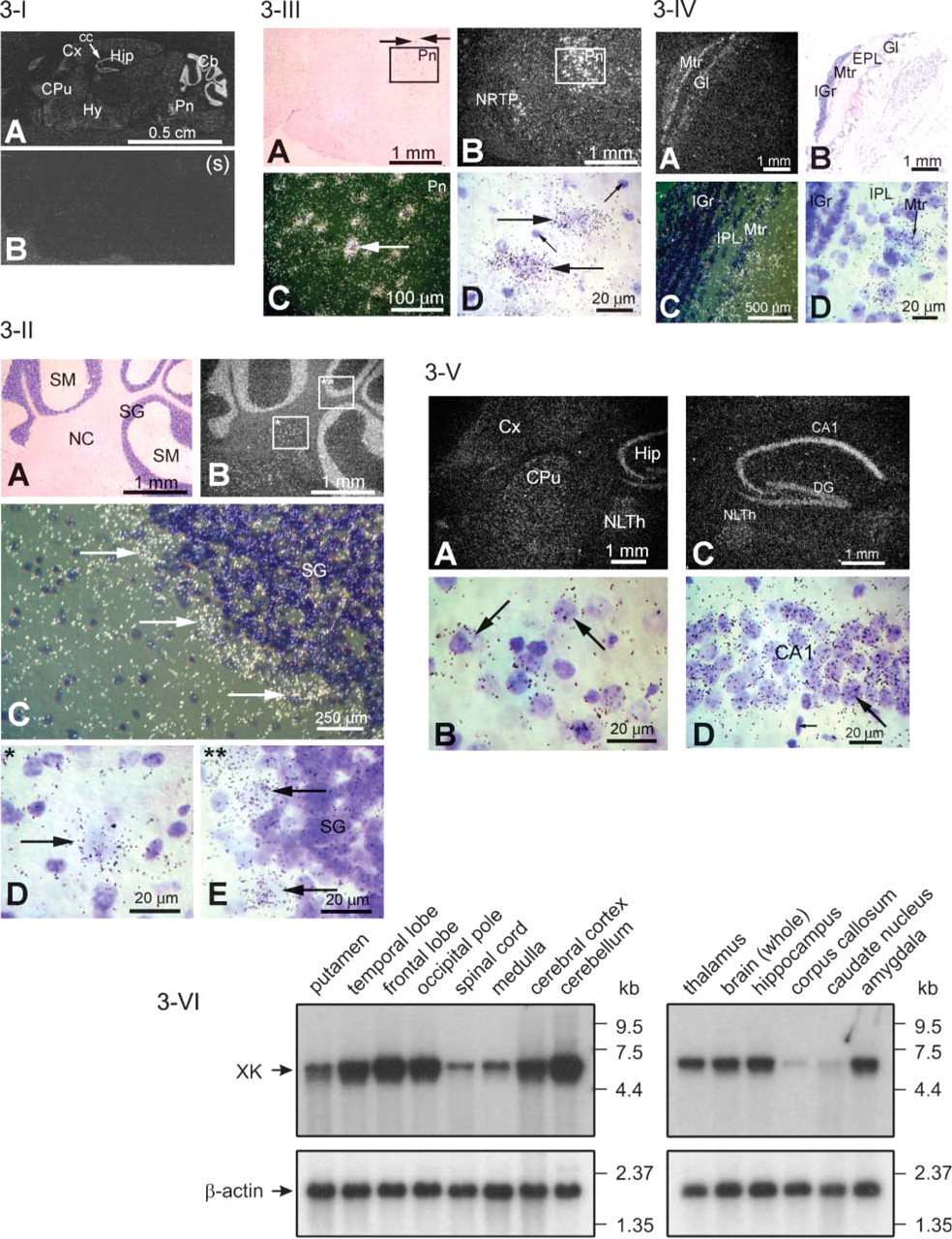
Expression of mXK mRNA in brain. (
Expression of Kell in non-erythroid tissue other than testis is very much in question, especially in skeletal muscle and brain where XK is present in large amounts. Our RT-PCR failed to detect mKell mRNA in mouse skeletal muscle and in C2C12 cells, which is a mouse myoblast cell line originated from mouse skeletal muscle. Earlier results showing Kell and XK complex in human skeletal muscle (Russo et al. 2000) could be due to erythroid cell contamination or to the possibility that the level of expression of Kell in skeletal muscle is different in human and mouse. It should be noted that Kell protein was detected by immunohistochemistry in the sarcoplasmic membrane of normal, but not in McLeod, skeletal muscle (Jung et al. 2001).
RT-PCR of mXK detected mXK mRNA both in skeletal muscle and C2C12 cells but ISHH of 1-day-old whole mouse body sagittal section did not detect mXK mRNA. Discrepancy in the results obtained by the two techniques may be due to differences in the detection sensitivity or there may be less XK in mouse skeletal muscle than in human skeletal muscle tissues where XK mRNA expression was shown in a large amount (Ho et al. 1994; Calenda et al. 2006).
Both ISHH and RT-PCR of mouse cerebellum support the view that there is little or no Kell expression. The dot-blot (Russo et al. 2000; Camara-Clayette et al. 2001) results of Kell mRNA expression in brain may have been due to contamination of the tissues with erythroid cells or to a nonspecific signal. Or there may be species differences. Our human brain Northern blots with Kell cDNA probes show hybridization with a higher molecular size band that is not the major Kell mRNA size (2.5 kb) (data not shown). We conclude that Kell and XK are coexpressed in erythroid tissues and testis, but in other tissues XK may be expressed by itself. If Kell is expressed in brain, it could be localized in a specific cell type that has not been identified by ISHH or Northern blots.
The Kell and XK disulfide-bonded complex is assembled in the ER, but they do not require each other to travel to the cell surface as evidenced by transfection experiments in COS cells (Russo et al. 1999). However, the facts that Kell null red cells that lack Kell have reduced amounts of XK, and McLeod red cells that lack XK have greatly reduced amounts of Kell (Lee et al. 2000b) indicate that coexpression of Kell and XK in erythroid cells is beneficial for protein stability and perhaps for efficient transport to the surface membrane.
The fact that both mXK and mKell are coexpressed in the spleen, liver (in newborn mouse), and bone marrow cells and that they are covalently linked in red cells suggests that they may play complementary roles in a hematopoietic function. Clearly, however, in non-erythroid tissues such as in brain neuronal cells and in bladder epithelial cells where Kell is not expressed, XK may have a separate neuronal or epithelial function.
Although Kell and XK are both expressed in testis, there is no evidence that they form a similar disulfide-linked complex as occurs on red cells. In tissues in which XK but not Kell is expressed, or where markedly less Kell than XK is expressed, there is also the possibility that XK may be linked either to itself or to another protein. However, at present there is no evidence to support this view. We have tried, using Western blot, to characterize XK in non-erythroid tissues of mouse, such as testis and cerebellum, to determine whether XK forms a complex with other proteins or with itself. However, the amount of XK in these tissues was insufficient to be determined by Western blot.
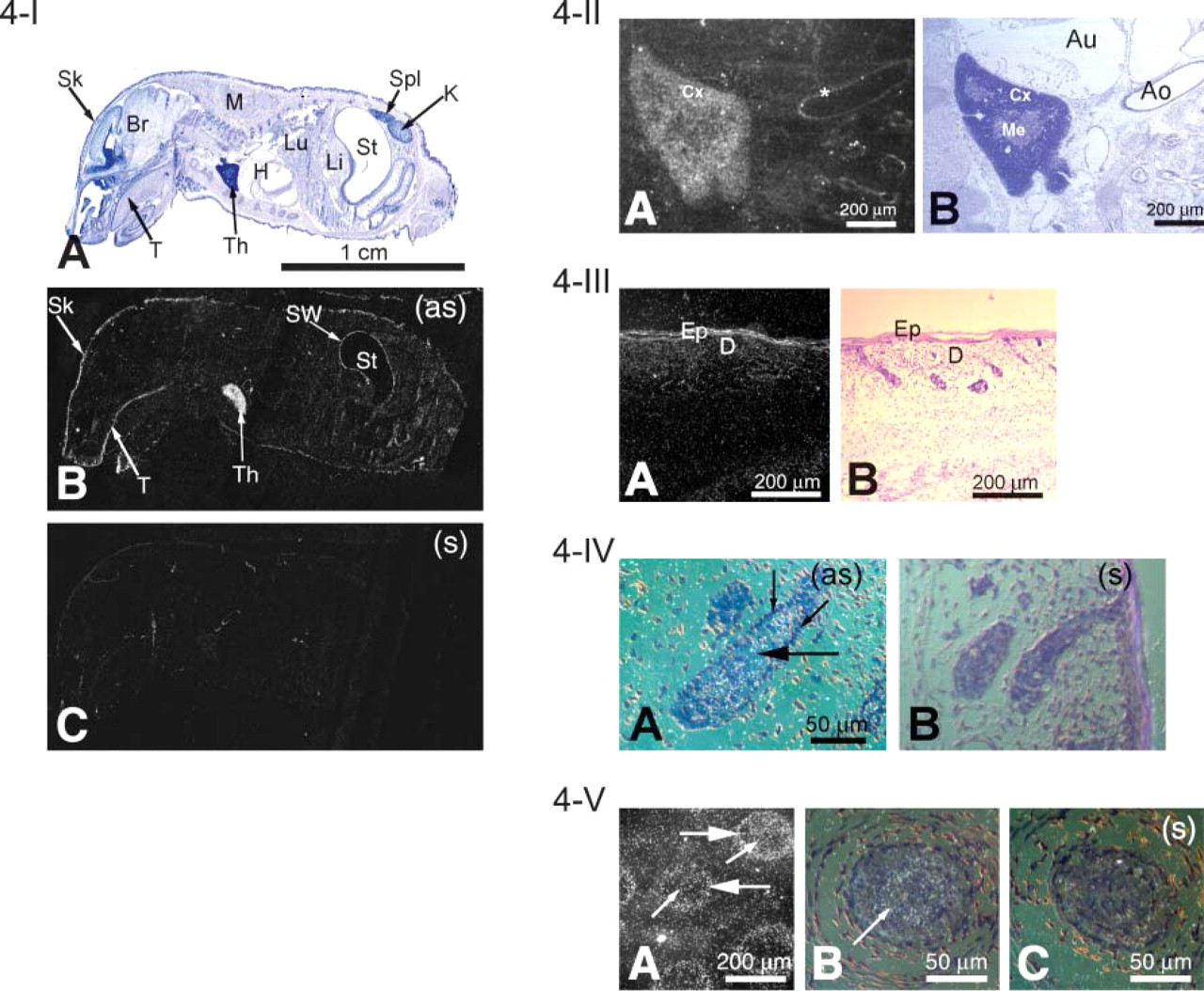
mXPLAC expression profile in different tissues of the newborn mouse. (
XPLAC (Calenda et al. 2006) is highly expressed in thymus and at moderate levels in skin and spleen. On the other hand, XK is expressed highly in spleen and only at a low level in thymus. Thus, expression of XPLAC is not like that of XK, indicating that the neuromuscular function of XK is likely not to be compensated by XPLAC when XK is absent. XPLAC may have a very different function from XK, and its presence in thymus suggests an immune response involvement. However, low levels of mXPLAC mRNA in testis and cerebellum were detected by RT-PCR, and the possibility of increased XPLAC expression when XK is absent, as in the case of McLeod phenotype, to compensate XK function cannot be ruled out.
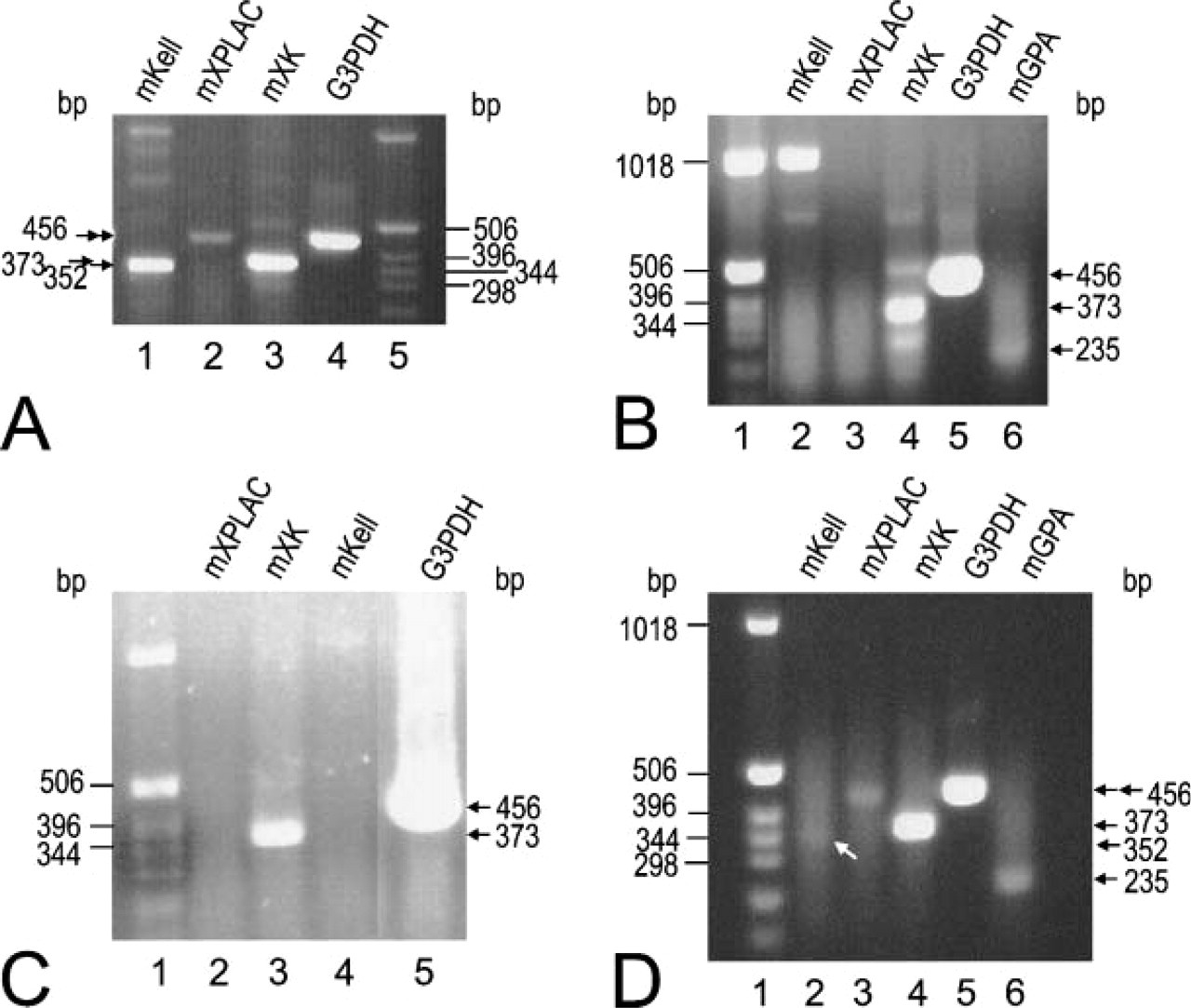
Detection of mKell, mXK, and mXPLAC mRNA by RT-PCR. Expected size of PCR products: mKell, 352 bp; mXPLAC, 456 bp; mXK, 373 bp; G3PDH, 456 bp; mGPA, 235 bp. G3PDH was used as a positive control. The amount of template cDNA for amplification of G3PDH for
Brain imaging of McLeod patients often shows atrophy of the caudate nucleus (Danek et al. 1994,2001a,b; Malandrini et al. 1994) suggesting that XK may be highly expressed in this region. However, our study shows that XK expression is pan-neuronal with highest expression levels in the pontine region where magnocellular neurons are located. There is also moderate to low expression of XK throughout several regions of the brain. This indicates that the late-onset form of neurological symptoms exhibited in McLeod syndrome may derive from a composite of prolonged abnormal neuronal function that results from absence of XK in many different types of neurons, but not particularly neurons of the basal ganglia. However, the possibility of the existence of age-related differential expressions of Kell, XK, and XPLAC that may contribute to the late onset of McLeod syndrome cannot yet be ruled out.
Footnotes
Acknowledgements
The study was supported in part by a National Institutes of Health (NIH) Specialized Center of Research (SCOR) Grant in Transfusion Biology and Medicine (HL-54459) and by an NIH Grant (RO1 HL-075716).
We thank Dr. Colvin M. Redman, Emeritus member of the New York Blood Center, for valuable discussions in preparation of the manuscript, Dr. Congrong Liu of the New York Blood Center, and Dr. Krista LaPerle of Research Animal Resource Center of the Weill Medical College of Cornell University, New York for examining the ISHH results. We are also grateful to Dr. Ruth H. Walker, James J. Peters Veterans Affairs Medical Center, Bronx, NY and Mount Sinai School of Medicine, New York, NY, whose expertise in the McLeod syndrome guided our selection of brain sections for the ISHH study. We thank the staff of the Nucleic Acid Analysis Laboratory of the New York Blood Center for DNA sequencing.
