Abstract
ATP-sensitive K+ (K
ATP-
Functional K
K
To date, little is known about the cellular localization, the role, and the regulation of K
With the questions noted above in mind, the present study attempted to clarify the expression and localization of the K
Materials and Methods
Animal and Tissue Preparation
Male Wistar rats (8–10 weeks) were used (Japan SLC; Hamamatsu, Japan). The protocols for animal experimentation described here were previously approved by the Animal Research Committee, Akita University. All subsequent animal experiments completely followed the University guidelines for animal experimentation.
Rats were deeply anesthetized by diethyl ether inhalation. Specimens for RT-PCR were quickly put into liquid nitrogen and kept at −80C until use. Specimens for immunoblot were cut into pieces, homogenized in a buffer as previously described (Zhou et al. 2005), and stored at −80C until use. Specimens for immunohistochemistry were fixed by transcardial perfusion with cold physiological saline, then further fixed with Zamboni fixative (100 ml 2% paraformaldehyde with 15 ml of saturated picric acid in 0.1 M PBS), pH 7.4. After perfusion, specimens were quickly dissected out, immersed in the same fixative for 6 hr at 4C, and subsequently transferred into 30% sucrose in PBS. Cryosections were cut at a thickness of 8 to 10 μm and thaw-mounted on MAS-coated glass slides (Matsunami Glass Industries; Kishiwada, Japan).
RT-PCR
RT-PCR was carried out as previously described (Zhou et al. 2008). Briefly, total RNA was extracted from SMG or skeletal muscle using the RNeasy Mini Kit (QIAGEN, GmbH; Hilden, Germany) according to the manufacturer's instructions. First-strand cDNA was synthesized by using the total RNA as a template with oligo(dT)12–18 primer and superscript II reverse transcriptase (Invitrogen; Tokyo, Japan) according to the manufacturer's instructions.
PCR was performed as previously described (Zhou et al. 2008) with primers for rat Kir6.x and SURs (see below) as Kir6.1 (GenBank accession no. D42145), Kir6.2 (GenBank accession no. D86039), SUR1 (GenBank accession no. L40624), SUR2A (GenBank accession no. D83598), and SUR2B (GenBank accession no. AF019628), or the housekeeping gene glyceraldehyde-3-phosphate dehydrogenase (G3PDH, GenBank accession no. AF106860). PCR reactions (50 μl volume) contained 5 μl of 10 × buffer, 3 μl of 25 mM MgCl2, 4 μl of dNTP mixture (2.5 mM each), and 3 U Taq DNA polymerase (Biotech International; Tokyo, Japan). The specimens were heated to 94C for 1 min, then subjected to 30 cycles of denaturation (94C, 30 min), annealing (30 min; 55C for Kir6.1, SUR2A, and G3PDH; 60C for SUR1; 58C for Kir6.2 and SUR2B), and extension (72C, 30 min). A final extension phase (72C, 3–5 min) was included for all samples. PCR products were separated on 1.5% agarose gels and visualized by ethidium bromide staining under ultraviolet light. To test for any residual contamination of the RNA samples by genomic DNA, control RT-PCR reactions were performed in a manner identical to that described above, but with the omission of the reverse transcriptase enzyme. All PCR products were extracted from the gel and purified with a gel extraction kit (Omega Bio-Tek; Doraville, GA), then labeled with BigDye Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems; Foster City, CA) and analyzed by sequencing test with an ABI Prism 3100-Avant Genetic Analyzer (Applied Biosystems; Hitachi, Tokyo, Japan) to confirm correct amplification.
K
atp
Channel Primers
Primers for all K
Western Blot Analysis
SDS-PAGE was carried out as previously described (Zhou et al. 2007) using 8% or 10% polyacrylamide gels. Proteins extracted from whole SMG were denatured in a modified sample buffer (125 mM Tris-HCl buffer, pH 6.8, 2% SDS, 25% glycerol, 0.01% bromophenol blue, and 10% 2-mercaptoethanol) and electrophoresed (10 μg per lane). After electrophoresis, proteins were electrophoretically transferred to a polyvinylidene difluoride membrane (Bio-Rad; Hercules, CA) by use of a semi-dry transfer unit (Hoefer TE70 series; Amersham Pharmacia Biotechnology, Buckinghamshire, UK). After blocking with 5% (w/v) BLOT-QickBlocker reagent (Chemicon International; Temecula, CA) in PBS-T (PBS containing 0.1% Tween-20), the membranes were incubated with goat anti-human Kir6.1 (Sc-11224; Santa Cruz Biotechnology, Inc., Santa Cruz, CA) (Zhou et al. 2005), or goat anti-human Kir6.2 (Sc-11228; Santa Cruz Biotechnology, Inc.) (Zhou et al. 2005), or rabbit anti-rat SUR2A (Zhou et al. 2007), and/or rabbit anti-rat SUR2B (Zhou et al. 2007), diluted 1:1000 for 60 min. After rinsing with PBS-T, the membranes were exposed to horseradish peroxidase (HRP)-conjugated donkey anti-goat IgG (AP180P; Chemicon International), or HRP-conjugated donkey anti-rabbit IgG (NA9340; Amersham Pharmacia Biotechnology), diluted 1:5000 for 30 min. The antigen–antibody complexes were visualized with chemiluminescence detection reagents (Amersham Pharmacia Biotechnology), according to the manufacturer's instructions.
Immunohistochemistry
Cryosections of SMG were kept in PBS containing 0.3% Tween-20 for 45 min. Prior to the incubation with first antibodies, sections were treated with a 0.3% solution of H2O2 in methanol and the ABC Blocking Kit (Vector Laboratories, Inc.; Burlingame, CA) to reduce the endogenous peroxidase reaction as well as nonspecific binding with avidin–biotin complex. After incubation with 5% normal goat serum or 5% normal rabbit serum for 1 hr, the sections were reacted with antibodies of goat anti-human Kir6.1, goat anti-human Kir6.2, rabbit anti-rat SUR2A, or rabbit anti-rat SUR2B, at a dilution of 1:200 to 1:500 for 12 hr at room temperature. After thorough rinsing with PBS containing 0.05% Tween-20, the sections were exposed to biotinylated rabbit anti-goat IgG (BA-5000; Vector Laboratories, Inc.) or biotinylated goat anti-rabbit IgG (BA-1000; Vector Laboratories, Inc.), diluted 1:200 for 30 min, and then with ABC complex (Vectastain ABC Kit; Vector Laboratories, Inc.) according to the manufacturer's instructions. Reaction sites were visualized by incubating the sections in 0.001–0.005% DAB (3,3'-diaminobenzidine tetrahydrochloride) reaction with 0.003% H2O2, and counterstaining with methyl green. Negative control was carried out either by omitting the first antibody or by adding corresponding immunizing peptide antigen against which the antibody was raised.
Immunofluorescence Double Staining
Double staining was performed as previously described (Zhou et al. 2005), although with some modification in relation to double labeling. After preincubation with 5% normal donkey serum in PBS for 1 hr, the sections for double labeling with Kir6.1 and SUR2A were incubated with goat anti-human Kir6.1 antibody (1:200) and rabbit anti-rat SUR2A antibody (1:500) diluted together in PBS. Those for Kir6.2 and SUR2A were incubated with goat anti-human Kir6.2 antibody (1:200) and rabbit anti-rat SUR2A antibody (1:500) diluted together in PBS. Those for double labeling with Kir6.1 and SUR2B were incubated with goat anti-human Kir6.1 antibody (1:200) and rabbit anti-rat SUR2B antibody (1:500) diluted together in PBS, and those for Kir6.2 and SUR2B were incubated with goat anti-human Kir6.2 antibody (1:200) and rabbit anti-rat SUR2B antibody (1:500) diluted together in PBS for 12 hr at room temperature. After rinsing with PBS, the sections were reacted with Alexa 488–conjugated donkey anti-goat IgG (A11055; Molecular Probes, Inc., Eugene, OR) and Alexa 594–conjugated donkey anti-rabbit IgG (A21207; Molecular Probes, Inc.), diluted together at 1:500 in PBS for 30 min. The sections were then coverslipped with PermaFluor aqueous mounting medium (Thermo; Pittsburgh, PA) after counterstaining with 4',6-diamidino-2-phenylindole, dihydrochloride (DAPI). Fluorescence immunolabeling signals were detected by a laser-scanning microscope (LSM510; Carl Zeiss, Oberkochen, Germany).
Results
RT-PCR
During the course of the studies, reactions were replicated at least twice. The PCR products were sequenced, and positive control (skeletal muscle) was also used. RT-PCR of rat SMG RNAs using the primers for Kir6.1 generated a 212-bp product specific for Kir6.1; using the primers for Kir6.2 generated a 297-bp product specific for Kir6.2; using primers for SUR1 generated a 558-bp product specific for SUR1 in skeletal muscle, but no product for SMG; using the primers for SUR2A generated a 155-bp product specific for SUR2A; and using the primers for SUR2B generated a 152-bp product specific for SUR2B and a 328-bp product specific for SUR2A, respectively (Figure 1). It is suggested that rat SMG contains all K
Western Blot Analysis
In SMG extractions, the anti-Kir6.1 antibody recognized a prominent ×50-kDa band, with a greater band of ×100 kDa detected as a dimer (Figure 2, Lane Kir6.1), the anti-Kir6.2 antibody recognized a prominent ×48-kDa band (Figure 2, Lane Kir6.2), the anti-SUR2A antibody recognized a prominent ×140-kDa band (Figure 2, Lane SUR2A), and the anti-SUR2B antibody recognized a prominent ×120-kDa band (Figure 2, Lane SUR2B), respectively.
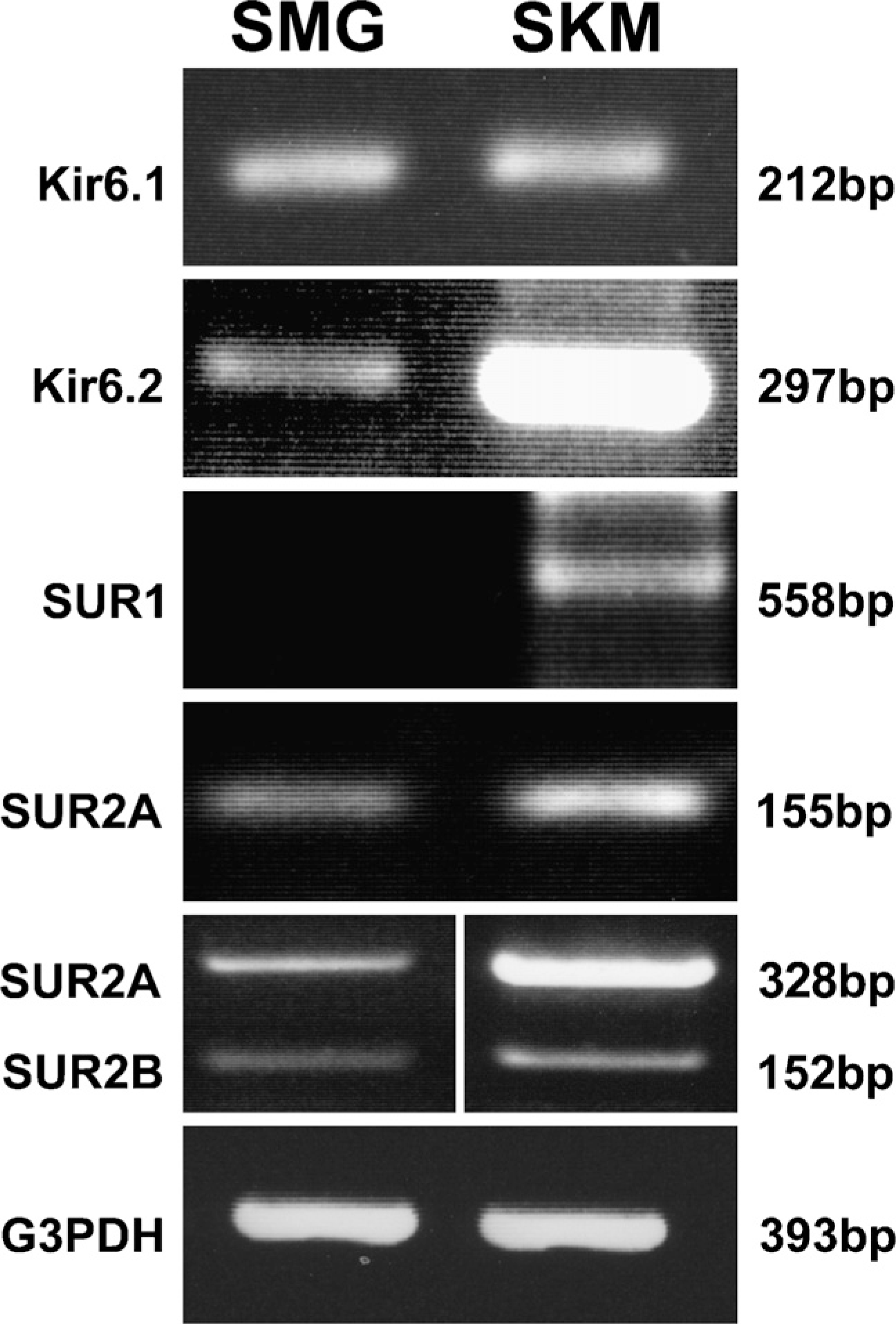
RT-PCR of rat submandibular gland (SMG) and skeletal muscle (SKM) total RNA. The 1.5% agarose gels show bands of DNA fragments amplified with primer pairs specific for Kir6.1, Kir6.2, SUR1, SUR2A, and SUR2B, as well as glyceraldehyde-3-phosphate dehydrogenase (G3PDH). Primers were tested to amplify fragments of 212 bp for Kir6.1, 297 bp for Kir6.2, 558 bp for SUR1, 155 bp for SUR2A (primers for SUR2A only), 328 bp for SUR2A, 152 bp for SUR2B (primers for both SUR2A and SUR2B), and 393 bp for G3PDH. The result shows that all ATP-sensitive K+ channel subunits, but not SUR1, are expressed in the SMG, compared with the positive control of the SKM RNA.
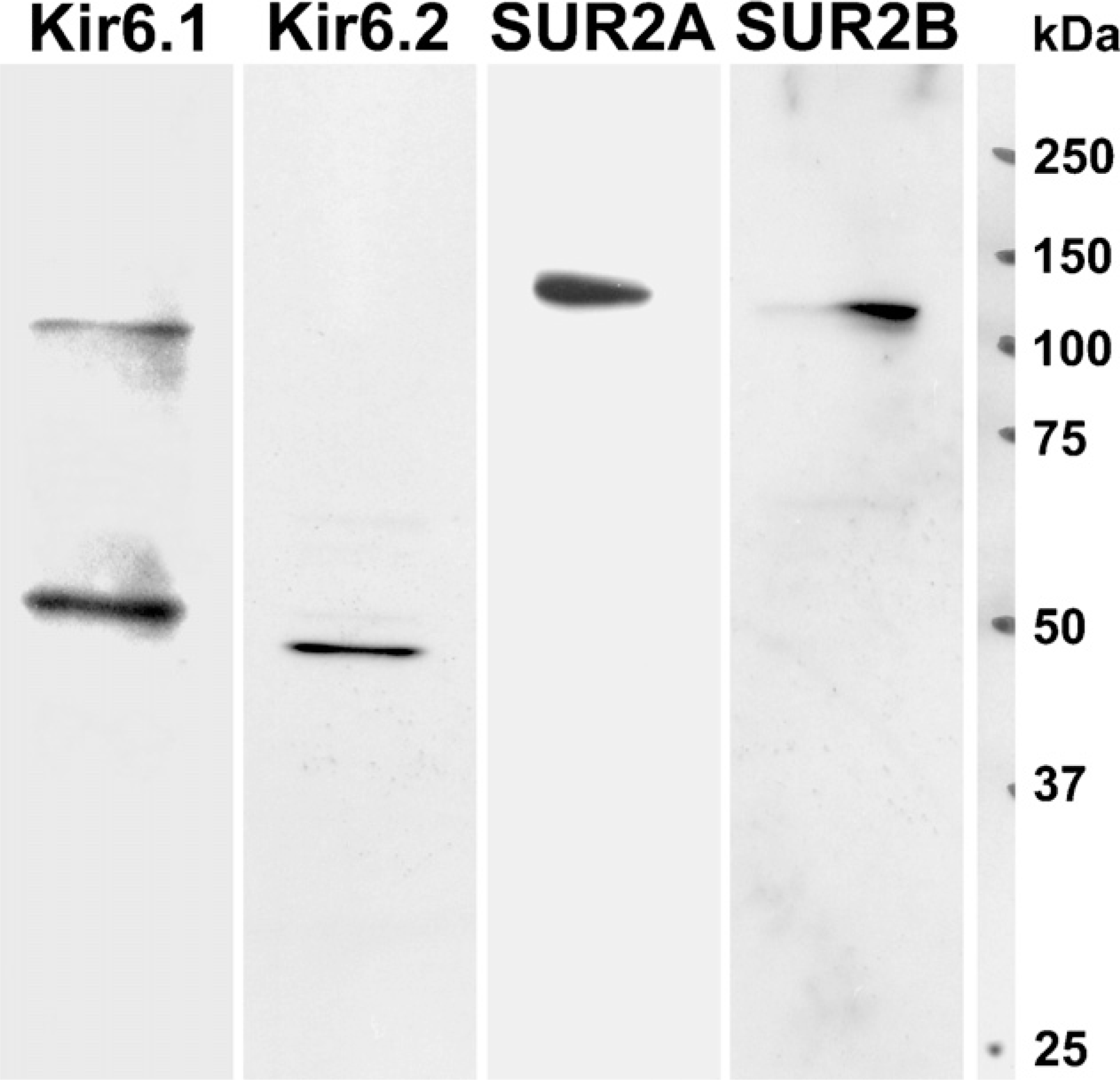
Immunoblot analysis of Kir6.1, Kir6.2, SUR2A, and SUR2B from rat SMG. Anti-Kir6.1 antibody recognized a prominent polypeptide of ×50 kDa with a dimer of ×100 kDa (Lane Kir6.1), and anti-Kir6.2 antibody recognized a ×48-kDa polypeptides (Lane Kir6.2) in the SMG. Anti-SUR2A antibody recognized a remarkable polypeptide of ×140 kDa (Lane SUR2A), and anti-SUR2B antibody revealed a polypeptide of ×120 kDa (Lane SUR2B) in the SMG.
Immunohistochemistry
In the acinus, immunoreactivity with anti-Kir6.1, anti-Kir6.2, or anti-SUR2A and/or SUR2B antibody was not observed (Figures 3A–3D and 4A–4C). In the myoepithelial cells, faint immunoreactivity with those antibodies of K
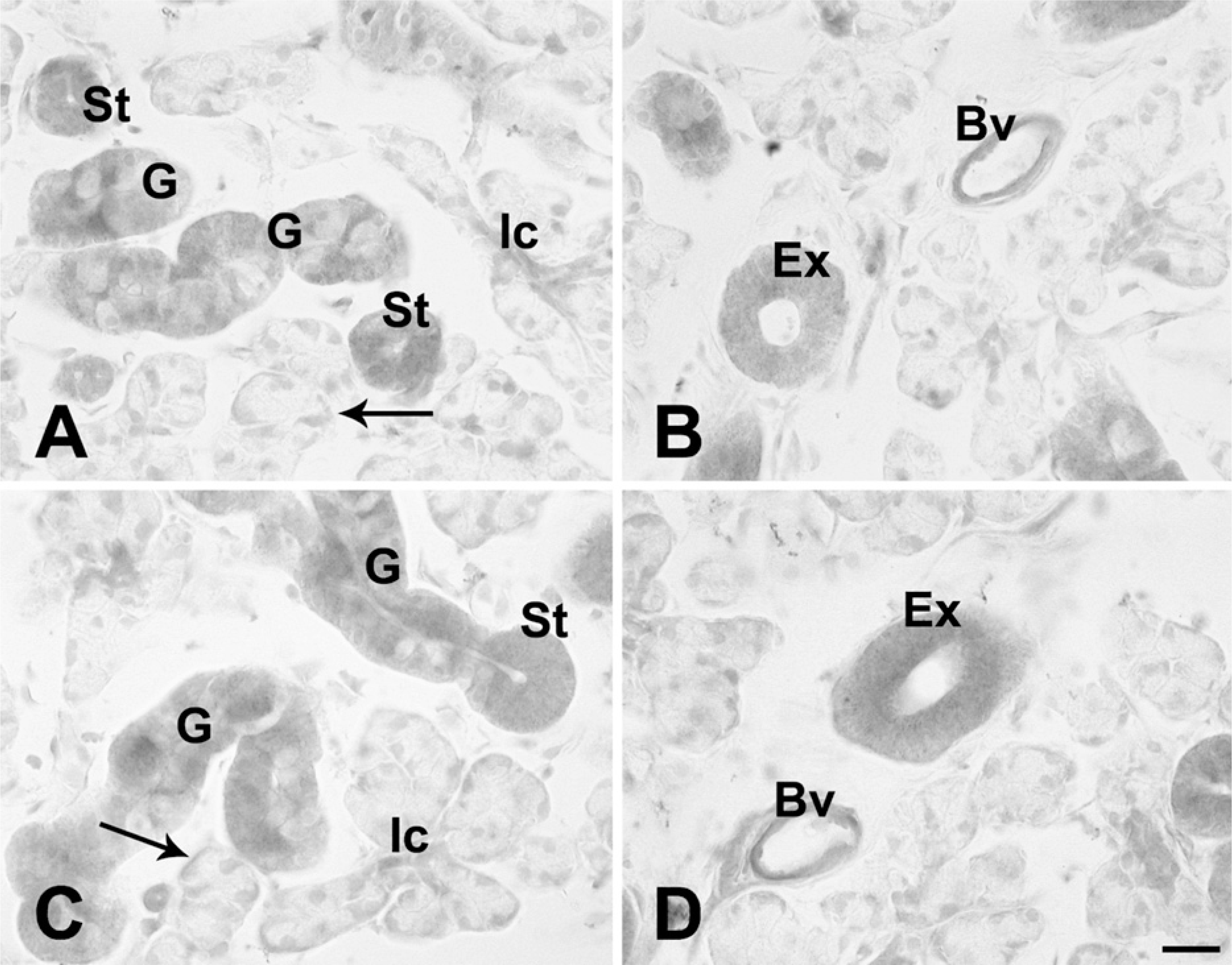
(
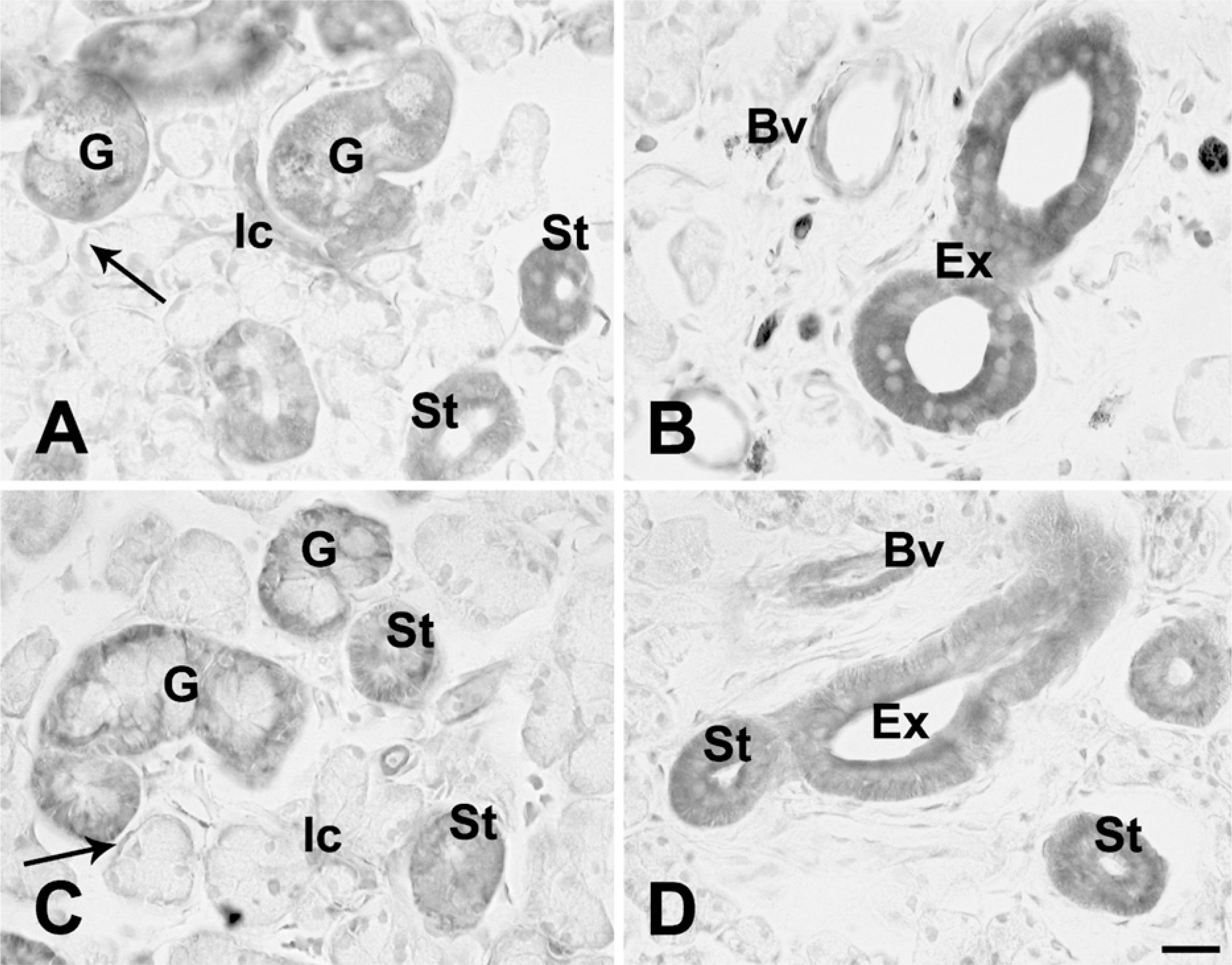
(
Localization of K
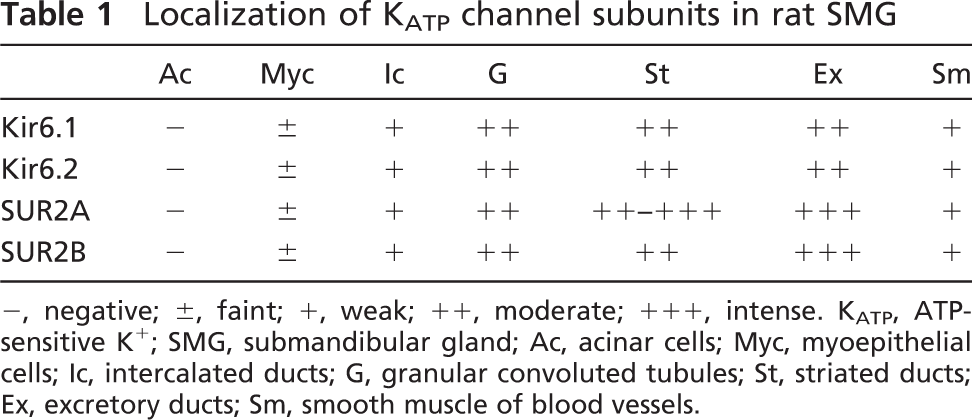
-, negative; ±, faint; +, weak; ++, moderate; +++, intense. K
Immunofluorescence Double Staining
To determine whether the Kir6.1 or Kir6.2 is colocalized with SUR2A or SUR2B in the epithelial cells of the rat SMG, immunofluorescence double staining was performed. Immunoreactivity with Kir6.1 or Kir6.2 antibody was detected as green fluorescence (Alexa 488) (Figures 5A, 5D, 5G, and 5J), whereas immunoreactivity with SUR2A or SUR2B antibody was detected as red fluorescence (Alexa 594) (Figures 5B, 5E, 5H, and 5K). When the images were merged, yellow fluorescence was detected in the cytoplasm (Figures 5C, 5F, 5I, and 5L).
Results showed that Kir6.1 overlapped with SUR2A in the granular convoluted tubules around the nucleus, at both the basal and apical portions. In the striated and excretory ducts, the Kir6.1 and SUR2A were widely colocalized. Colocalization with Kir6.1 and SUR2A was also observed in myoepithelial cells and small blood vessels (Figure 5C). Kir6.2 and SUR2A are widely distributed in myoepithelial cells, striated and excretory ducts, and blood vessels (Figures 5D and 5E). In the granular convoluted tubules, Kir6.2 was mainly localized in the apical portion and slightly localized in the basal portion (Figure 5D), whereas SUR2A was mainly localized in the basal portion rather than the apical portion (Figures 5B and 5E). Thus, colocalization with Kir6.2 and SUR2A was observed mainly in the basal portion and to some extent in the apical portion (Figure 5F). The colocalization with Kir6.2 and SUR2A was also observed in the striated and excretory ducts, in the myoepithelial cells at the base of the acinus, and in blood vessels (Figure 5F). Kir6.1 and SUR2B overlapped in both the basal and the apical portion in the granular convoluted tubules. In the striated and excretory ducts, as well as blood vessels, colocalization with Kir6.1 and SUR2B was also observed (Figure 5I). Kir6.2 overlapped with SUR2B mainly in the apical portion and to some extent in the basal portion in the granular convoluted tubules (Figure 5L). In the striated ducts and smooth muscle of the blood vessels, colocalization with Kir6.2 and SUR2B was also observed (Figure 5L). There was no clear colocalization of Kir6.1 and SUR2B or Kir6.2 and SUR2B in the myoepithelial cells observed.
Discussion
To our knowledge, the present observation provides the first evidence for the localization of K
ATP-activated Ca2+-dependent K+ channel (K
As in other exocrine secretion, the saliva secretion proceeds in two steps; the acinar cell produces primary isotonic, NaCl-rich saliva, which is then modified during its transportation through the ductal system, changes in electrolyte composition, and to some extent, the osmolarity of the primary fluid, by absorbing the NaCl and secreting KHCO3 − to the mouth as hypotonic final saliva (Zeng et al. 1997; Chaib et al. 1999). During passage through the excretory ducts of the salivary glands, the electrogenic component of duct transport consists of Na+ influx through the Na+ channel, Cl− flux through the Cl− channels, and K+ flux through the K+ channels (Chaturapanich et al. 1997). In general, the concentration of inorganic ions, such as Na+ and Cl− in the saliva, changes from the acinar cavity to the duct cavity, because they are reabsorbed during their passage through the duct system. The K+ ion is secreted from the duct cells into the saliva when it passes through these ducts (Nakamoto et al. 2008).
Containing K
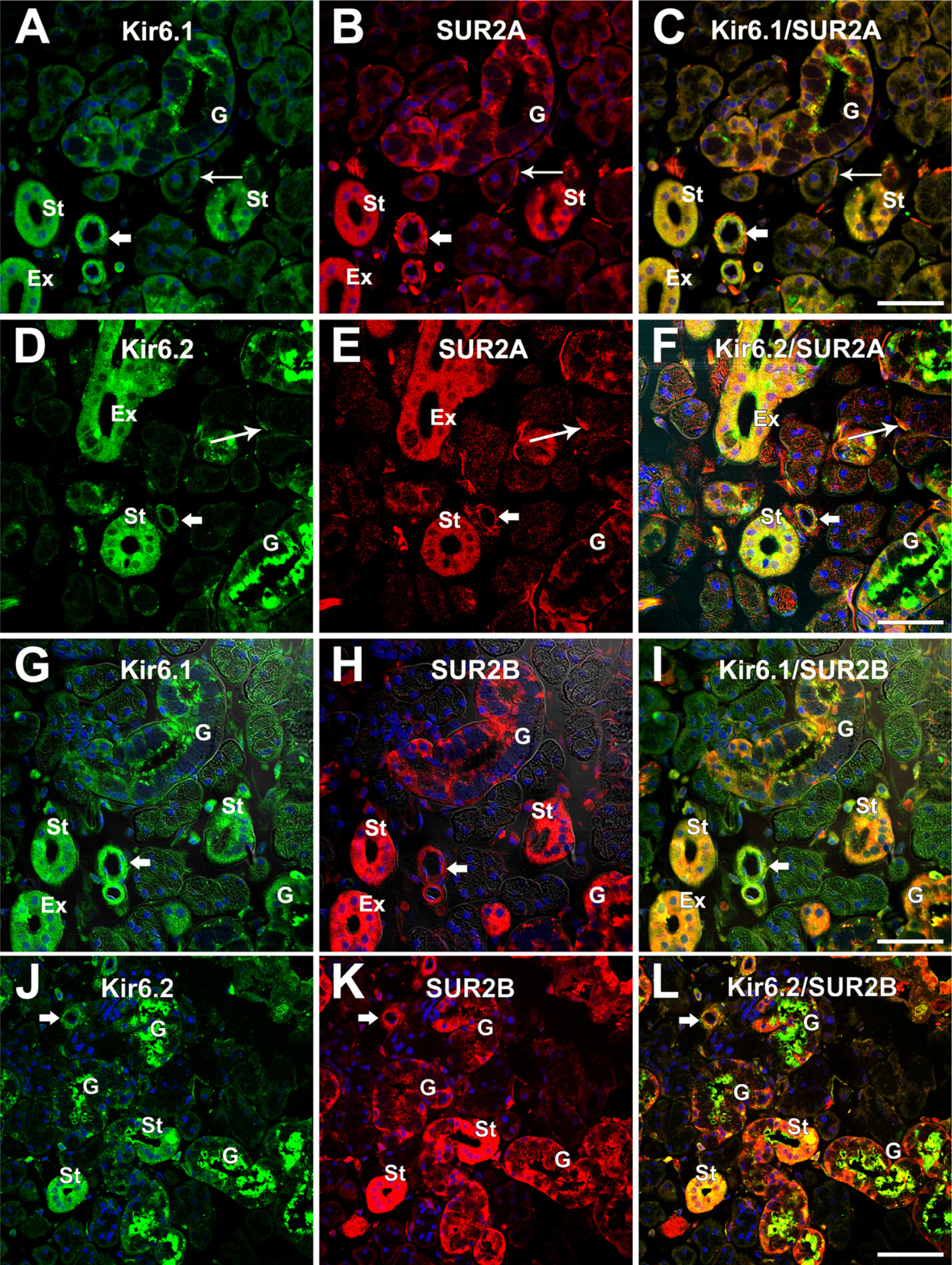
Immunofluorescence double staining shows the expression of Kir6.1 and Kir6.2 in the SMG as green (Alexa 488,
It is well known that there are different types of K
Thus, the present study revealed that four subunits of the K
In conclusion, K
Footnotes
Acknowledgements
This work was supported in part by research grants from the Akita University Graduate School of Medicine and Faculty of Medicine (to HA) and from Kitasato University School of Medicine (to KK).
The authors thank Professor H. Kondo, Tohoku Bunka Gakuen University Faculty of Medical Science and Welfare, for reviewing this manuscript, and the staff of Bioscience Research Education Center of Akita University Graduate School of Medicine and Faculty of Medicine for their help.
