Abstract
To understand the possible functions and subcellular localizations of sulfonylurea receptors (SURs) in cardiac muscle, polyclonal anti-SUR2A and anti-SUR2B antisera were raised. Immunoblots revealed both SUR2A and SUR2B expression in mitochondrial fractions of rat heart and other cellular fractions such as microsomes and cell membranes. Immunostaining detected ubiquitous expression of both SUR2A and SUR2B in rat heart in the atria, ventricles, interatrial and interventricular septa, and smooth muscles and endothelia of the coronary arteries. Electron microscopy revealed SUR2A immunoreactivity in the cell membrane, endoplasmic reticulum (ER), and mitochondria. SUR2B immunoreactivity was mainly localized in the mitochondria as well as in the ER and cell membrane. Thus, SUR2A and SUR2B are not only the regulatory subunits of sarcolemmal KATP channels but may also function as regulatory subunits in mitochondrial KATP channels and play important roles in cardioprotection.
F
Sulfonylurea receptors (SURs) are transmembrane proteins of the ATP-binding cassette (ABC) superfamily (Aguilar-Bryan et al. 1998; Seino 1999). Two genes encoding SUR subunits SUR1, SUR2A, and SUR2B have been cloned (Inagaki et al. 1995, 1996; Chutkow et al. 1996; Isomoto et al. 1996; Seino and Miki 2003). SUR1 shares 67% identity with SUR2 subunits (Chutkow et al. 1996; Ashcroft and Gribble 2000; Tanemoto et al. 2000). Alternative splicing of the last exon of SUR2 gives rise to the isoforms SUR2A and SUR2B (Chutkow et al. 1996, 1999; Aguilar-Bryan et al. 1998), which differ only in their carboxy-terminal 42 amino acid residues (Isomoto et al. 1996; Shindo et al. 1998).
The functional diversity of KATP channels is determined largely by the SUR subunits. For example, the Kir6.2/SUR1 channel is activated by diazoxide but not by pinacidil (Inagaki et al. 1995), the Kir6.2/SUR2A channel is stimulated by pinacidil but not by diazoxide (Inagaki et al. 1996; Babenko et al. 1998b; Gribble et al. 2000), and the Kir6.2/SUR2B channel is stimulated by both diazoxide and pinacidil (Isomoto et al. 1996). The SUR2A-based channels are distinguished from SUR2B channels by their differential sensitivity to diazoxide (Babenko et al. 1998a), whereas nicorandil activates SUR2B/Kir6.2 more potently than SUR2A/Kir6.2 channels (Yamada and Kurachi 2004).
Although SUR2A and SUR2B differ only in their carboxy-terminal 42 amino acid residues, their distributions are quite different. SUR2A is predominantly expressed in heart and skeletal muscle (Inagaki et al. 1996), and SUR2B is ubiquitously expressed in many tissues including cardiomyocytes, as assessed by reverse transcription–polymerase chain reaction (RT-PCR) assay (Isomoto et al. 1996).
Cardiomyocyte mitochondria contain both Kir6.1 and Kir6.2 (Lacza et al. 2003; Singh et al. 2003; Zhou et al. 2005), although conflicting views exist (Seharaseyon et al. 2000; Kuniyasu et al. 2003). How do SURs cooperate with these pore-forming subunits in mitochondria? Singh et al. (2003) showed that SUR2A, but not SUR2B, is localized in the mitochondria, whereas Lacza et al. (2003) showed no SURs in mitochondria, except for a low molecular mass SUR2-like protein.
It is necessary to have antisera that can distinguish SUR2A from SUR2B to describe the differences in physiological significance, tissue distribution, and sub-cellular localization of these subunits. We determined the localization of SUR2A and SUR2B in rat cardiomyocytes and coronary vessels by immunoblotting and immunohistochemistry, using newly generated specific anti-SUR2A and anti-SUR2B antisera. These novel antibodies will open a new approach to elucidate the molecular mechanism of cardiomyocytes and their energy production in both normal and pathological conditions and will aid in the understanding of the possible functions of SUR2 in the regulation of cardiomyocytes.
Materials and Methods
Generation of Anti-SUR2A and/or Anti-SUR2B Antibodies
Rabbit anti-SUR2A antibody was raised against a synthetic 19-mer peptide, NH2-PNLLQHKNGLFSTLVMTNK(C)-OH (Biologica Co.; Nagoya, Japan), which corresponds to amino acid residues 1526–1544 of rat SUR2A (GenBank Accession No. D83598). Rabbit anti-SUR2B antibody was raised against a synthetic 14-mer peptide, NH2-(C)MKRGNI-LEYDTPES-OH (Hokkaido System Science Co.; Sapporo, Japan), which corresponds to amino acid residues 1514–1528 of rat SUR2B (GenBank Accession No. AF019628). In brief, the synthetic peptide representative of rat SUR2A or SUR2B was coupled to the carrier protein keyhole limpet hemocyanin (KLH) via the cysteine residue added to the C- or N-terminal. Japanese white rabbits (Japan SLC; Hamamatsu, Japan) weighing 2.5–3.0 kg were injected with ∼200 μg of peptide–KLH conjugate emulsified with an equal volume of Freund's complete adjuvant (Rockland Immunochemicals; Gilbertsville, PA) at multiple intradermal sites, followed by three boosters at 2-week intervals by injection with the same dosage of the peptide conjugate emulsified in Freund's incomplete adjuvant (Rockland Immunochemicals). Antisera were harvested 1 week after the final injection. Antisera were purified by peptide antigen immunoaffinity column chromatography (SUR2A) or protein A chromatography (SUR2B) before using for Western blot analysis and immunohistochemistry.
Transfection and Preparation of Cell Extracts
COS-7 cells were transfected with the mammalian expression vector pcDNA3 containing rat SUR1 (2 μg), pCMV6c containing SUR2A (2 μg), or pCMV6c vector containing SUR2B (2 μg) using FuGENE 6 Transfection Reagent (Roche Diagnostics; Indianapolis, IN) according to the manufacturer's instructions. Briefly, 24 hr prior to transfection, COS-7 cells were plated at a density of 2.0 × 106 cells/90-mm diameter cell-culture dish in DMEM (Sigma-Aldrich; St Louis, MO) supplemented with 10% fetal bovine serum (DMEM/FBS; Gemini Bioproducts, Woodland, CA). Cells were incubated at 37C in 5% CO2/95% air (as were all subsequent cell incubations). Transfection was performed with 2 μg of each plasmid vector using 6 μl of FuGENE 6 Transfection Reagent (Roche Diagnostics). At 5 hr following transfection, the medium was removed and replaced with fresh DMEM/FBS. Cells were harvested 48 or 72 hr after transfection, washed with PBS, suspended in a protein extraction buffer (10 mM Tris–HCl, pH 7.4, 0.25 M sucrose, 0.1 mM EDTA), and sonicated on ice with an Ultrasonic Processor (Sonics and Materials; Newtown, CT). Cell lysates were clarified by centrifugation and stored at −80C until use.
Animals
Four- to eight-week-old male Wistar rats were used. Protocols for animal experimentation described here were previously approved by the Animal Research Committee, Akita University. All subsequent animal experiments were conducted in accordance with the Guidelines for Animal Experimentation of Akita University.
The animals prepared for immunohistochemistry and immunoelectron microscopy were anesthetized by diethyl ether inhalation and then perfused through the left ventricle with 4% paraformaldehyde in 0.1 M PBS, pH 7.4. The hearts were quickly excised and placed in the same fixative for 6 hr at 4C, after which they were transferred to 30% sucrose in PBS. Eight- to 10-μm-thick cryosections were cut and thaw mounted on MAS-coated glass slides (Matsunami Glass Industries; Kishiwada, Japan).
Subcellular Fractionation of Rat Cardiomyocytes
All operations were carried out at 0–4C. Each rat was deeply anesthetized by inhalation of ethyl ether. The heart was removed immediately and quickly washed with 0.9% NaCl. The hearts were then cut into small pieces and homogenized in 0.25 M sucrose/50 mM Tris–HCl, pH 7.4, containing a protease inhibitor cocktail tablet (one tablet in 25-ml solution) (Roche Diagnostics GmbH; Mannheim, Germany). Subcellular fractions were extracted as described previously (Itoh et al. 2002). Briefly, after centrifugation at 600 × g for 10 min, the precipitate was discarded. The 600 × g supernatant was further centrifuged at 7000 × g for 10 min. The 7000 × g precipitate (P1) was re-dissolved in the buffer and further centrifuged at 5000 × g for 10 min. The 5000 × g precipitate was used as the mitochondrial fraction. The 7000 × g supernatant (S1) was centrifuged at 54,000 × g for 60 min, and the supernatant (S2) was further centrifuged at 105,000 × g for 60 min. The 54,000 × g precipitate was used as plasma membrane, the 105,000 × g precipitate as the microsomal fraction, and the 105,000 × g supernatant as cytoplasm.
Western Blot Analysis
SDS/PAGE was carried out according to the procedure of Laemmli (1970) using 8–10% polyacrylamide gels. Proteins extracted from cultured cells, heart tissue, brain, and kidney, and cellular fractions from rat hearts were electrophoresed (10 μg per lane). After electrophoresis, proteins were electrophoretically transferred to a polyvinylidene difluoride membrane (PVDF; Bio-Rad, Hercules, CA) by use of a semidry transfer unit (Hoefer TE70 series; Amersham Pharmacia Biotechnology, Buckinghamshire, UK). PVDF membranes were then blocked with 5% Blot-QuickBlocker (Chemicon International; Temecula, CA) in PBS overnight at room temperature. After washing with PBS-T (PBS containing 0.1% Tween-20), PVDF membranes were incubated in PBS-T containing rabbit anti-SUR2A (1.5 μg/ml), rabbit anti-SUR2B (4.3 μg/ml), rabbit anti-cytochrome c (1 μg/ml) (Sc-7159; Santa Cruz Biotechnology, Santa Cruz, CA), rabbit anti-calreticulin (NB600-101; Novus Biologicals, Littleton, CO) diluted 1:500, or rabbit anti-Na+/K+ ATPase β1 (1 μg/ml) (Sc-16053; Santa Cruz Biotechnology), for 60 min at room temperature. After rinsing with PBS-T, the membranes were exposed to horseradish peroxidase-conjugated donkey anti-rabbit IgG (NA9340; Amersham Pharmacia Biotechnology) diluted 1:3000, or donkey anti-goat IgG (AP180P; Chemicon International) diluted 1:2000 in PBS-T for 30 min. After three washes in PBS-T, signals were developed with ECL Western blotting detection reagents (Amersham Pharmacia Biotechnology) according to the manufacturer's instructions, and the membranes were exposed to X-OMAT film (Eastman Kodak; Rochester, NY). Negative control was carried out in adding peptide antigen to the corresponding antibody by which it was raised against. The ratio of antibody to antigen was 1:2 μg in reaction solutions. Because SUR2B shares 64% homology with SUR1 in the peptide amino acid sequences, to confirm the specificity of anti-SUR2B antibody, the peptide antigens of SUR2A, SUR2B, and SUR1 (Sc5789p; Santa Cruz Biotechnology) were used in preabsorption tests to check the specificity of anti-SUR2B against mitochondrial fractions.
Immunohistochemistry and Immunoelectron Microscopy
Cryosections of rat heart were kept in PBS containing 0.3% Tween-20 for 45 min. Prior to the incubation with normal serum and antibodies, sections were treated with a 0.3% solution of H2O2 in methanol for 10 min to reduce endogenous peroxidase reactions. After preincubation with 5% normal goat serum for 1 hr, sections were incubated with rabbit anti-SUR2A or anti-SUR2B antibody at a dilution of 1:200 to 1:500 for 12 hr at room temperature. After a thorough rinsing with PBS, the sections were exposed to biotinylated goat anti-rabbit IgG (BA-1000; Vector Laboratories, Burlingame, CA), diluted 1:200 for 30 min, and then with ABC complex (Vectastain ABC kit; Vector Laboratories) according to the manufacturer's instructions. Reaction sites were visualized by the DAB (3,3′-diaminobenzidine tetrahydrochloride) reaction and counterstained with methyl green. Negative control was carried out either by omitting the first antibody or by adding corresponding immunizing peptide antigen against which the antibody was raised.
Tissue sections showing immunopositive reactions to SUR2A or SUR2B were postfixed in 1% osmium tetroxide (OsO4) for 30 min, dehydrated in a graded ethanol series, and embedded in Quetol 812 (Nissin EM Ltd; Tokyo, Japan). Thin sections were cut and observed with an electron microscope without uranyl acetate and lead citrate staining.
Results
Western Blot Analysis
The anti-SUR2A antibody recognized a prominent polypeptide of 140 kDa in the extracts of COS-7 cells transfected with rat SUR2A cDNA, but did not react notably with extracts of cells transfected with SUR1 cDNA or SUR2B cDNA (Figure 1A, left three lanes). The anti-SUR2B antibody recognized a prominent polypeptide of 140 kDa in extracts of COS-7 cells transfected with rat SUR2B cDNA, with faint reaction to COS-7 cells transfected with rat SUR1 cDNA, but no notable signal was detected in extracts of COS-7 cells transfected with rat SUR2A cDNA (Figure 1A, right three lanes). Anti-SUR2A antibody recognized the same-size polypeptides of ∼140 kDa in the extract of COS-7 cells transfected with rat SUR2A cDNA and in the extracts of heart, brain, and kidney (Figure 1B). Anti-SUR2B antibody recognized a polypeptide of ∼140 kDa from the extract of COS-7 cells transfected with rat SUR2B cDNA, but different-size polypeptides of ∼120 kDa from theextracts of heart, brain, and kidney (Figure 1C). These data corroborated the specificities of the newly made antibodies. The purity of each cellular fraction was confirmed with respective marker antibodies. Anti-cytochrome c antibody (a mitochondria-specific antibody) was used to confirm the purity of the mitochondrial fraction. Only mitochondrial fractions (Lane mit), and neither membrane (Lane cm) nor microsome fractions (Lane ms), showed intense immunoreactivity to anti-cytochrome c antibody (Figure 1D). Anti-calreticulin antibody [an endoplasmic reticulum (ER)-specific antibody] was used to confirm the purity of the microsomal fraction (Figure 1E, left three lanes). It reacted only to the microsomal fraction (Lane ms) but not to the mitochondrial (Lane mit) and cell membrane fractions (Lane ms). The purity of the cell membrane fraction was confirmed by anti-Na+/K+ ATPase β1 subunit antibody (a membrane-specific antibody, Figure 1E, right three lanes), which reacted to the cell membrane fraction (Lane cm) with lower polypeptides in microsome fraction (Lane ms) but not to the mitochondrial fraction (Lane mit). In these cell fractions from rat heart, the anti-SUR2A antibody (Figure 1F, right three lanes) recognizes a prominent polypeptide of ∼140 kDa in the microsomal fraction (Lane ms), cell membrane fraction (Lane cm), and mitochondrial fraction (Lane mit). Preincubation of the antibody with the immunizing peptide prevented the staining of these polypeptides (Figure 1F, left three lanes, +peptide). Anti-SUR2B antibody (Figure 1G, right three lanes) detected a prominent polypeptide of ∼120 kDa in the microsomal (Lane ms) and cell membrane (Lane cm) fractions, whereas in the mitochondrial fraction, in addition to a weak band of ∼120 kDa, a prominent polypeptide of ∼50 kDa was detected (Lane mit). These bands were not detected when the antibody was first preincubated with the immunizing peptide (Figure 1G, left three lanes, +peptide). In mitochondrial fractions, anti-SUR2B antibody was only preabsorbed by SUR2B peptide (Lane 2B) but not preabsorbedby either SUR2A peptide (Lane2A) or SUR1 peptide (Lane1) (Figure 1H).
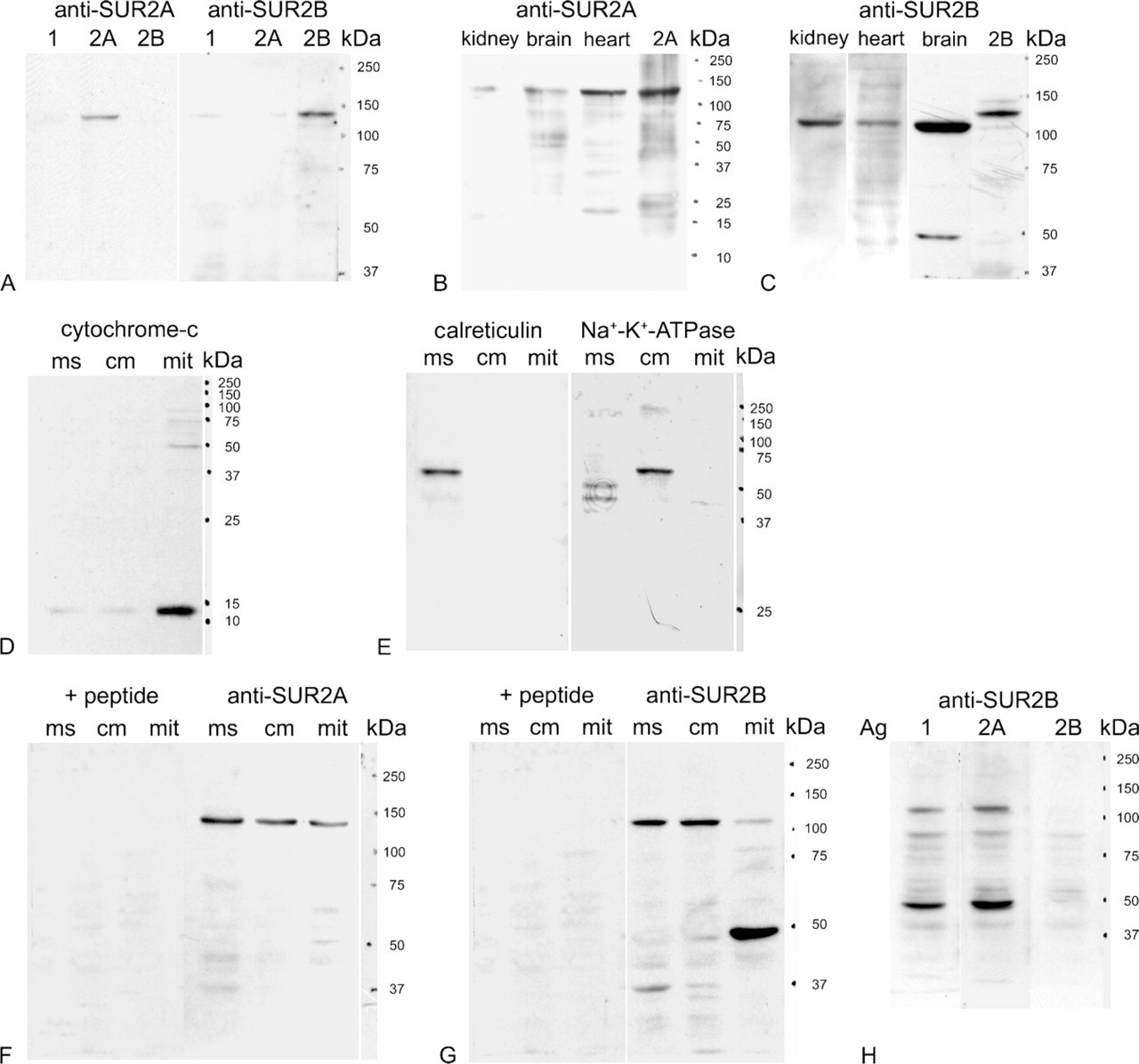
Immunoblot analysis of SUR2A and SUR2B from rat heart and preparation of cellular fractions. (
Expression of SUR2A and SUR2B
In the atria, immunoreactivity with anti-SUR2A antibody was observed in the atrial cardiomyocytes (Figure 2A). In the ventricles, weak to moderate immunoreactivity with SUR2A antibody was detected in both the longitudinal (Figure 2B) and transverse sections of cardiomyocytes (Figure 2C). It was also detected in the papillary muscles, endocardium, interatrial septum, and interventricular septum (not shown). The SUR2A protein was detected as punctate immunoreactive products in the cytoplasm (Figure 2A, inset; Figure 2C) and also showed immunoreactivity in the cell membrane as well as in the striated pattern (Figure 2B). Weak to moderate immunoreaction to SUR2A was also observed in the smooth muscles and endothelia of the middle- and large-size coronary arteries (Figure 2D). Immunoreactivity was absent after preabsorption with the immunizing peptide (Figure 2D, inset). Omission of the first antibody resulted in no immunoreaction (not shown).
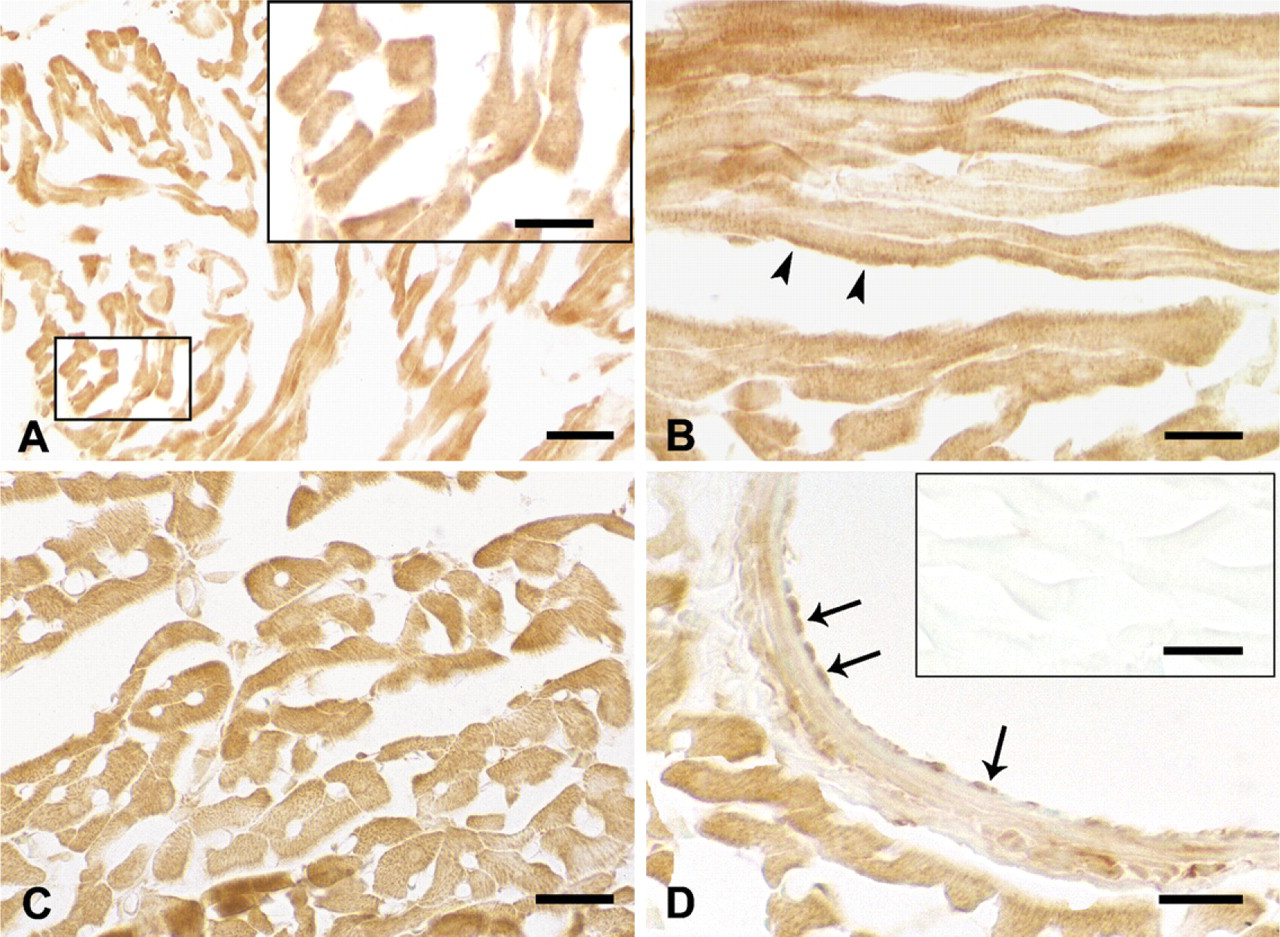
Immunohistochemistry showing expression of SUR2A protein in rat heart. Immunoreactivity with anti-rat SUR2A was seen in cardiomyocytes of the atrium (
Using the anti-SUR2B antibody, a weak immunohistochemical reaction was detected in the cytoplasm and cell membranes of cardiomyocytes in the atria (Figure 3A) and in the ventricles in both longitudinal and transverse sections (Figures 3B and 3C). Immunoreaction with anti-SUR2B was also detected in the papillary muscles, endocardium, interatrial septum, and interventricular septum (not shown). The SUR2B protein was detected as punctate staining in the cytoplasm (Figure 3C). In the longitudinal section, weak immunoreaction was also observed in a striated pattern consistent with localization in t-tubules and also in the cell membrane (Figures 3B, 3E, and 3F). Weak to moderate immunoreaction to SUR2B was also observed in the smooth muscles and endothelia of the middle- and large-size coronary arteries (Figure 3D). Immunoreactivity was absent after preabsorption with the immunizing peptide (Figure 3D, inset). Omission of the first antibody resulted in no immunoreaction (not shown). An antigenic absorption test showed that the immunoreaction was blocked only by SUR2B peptide antigen (Figure 3D, inset) but not by SUR2A or SUR1 peptide antigens (Figures 3E and 3F).
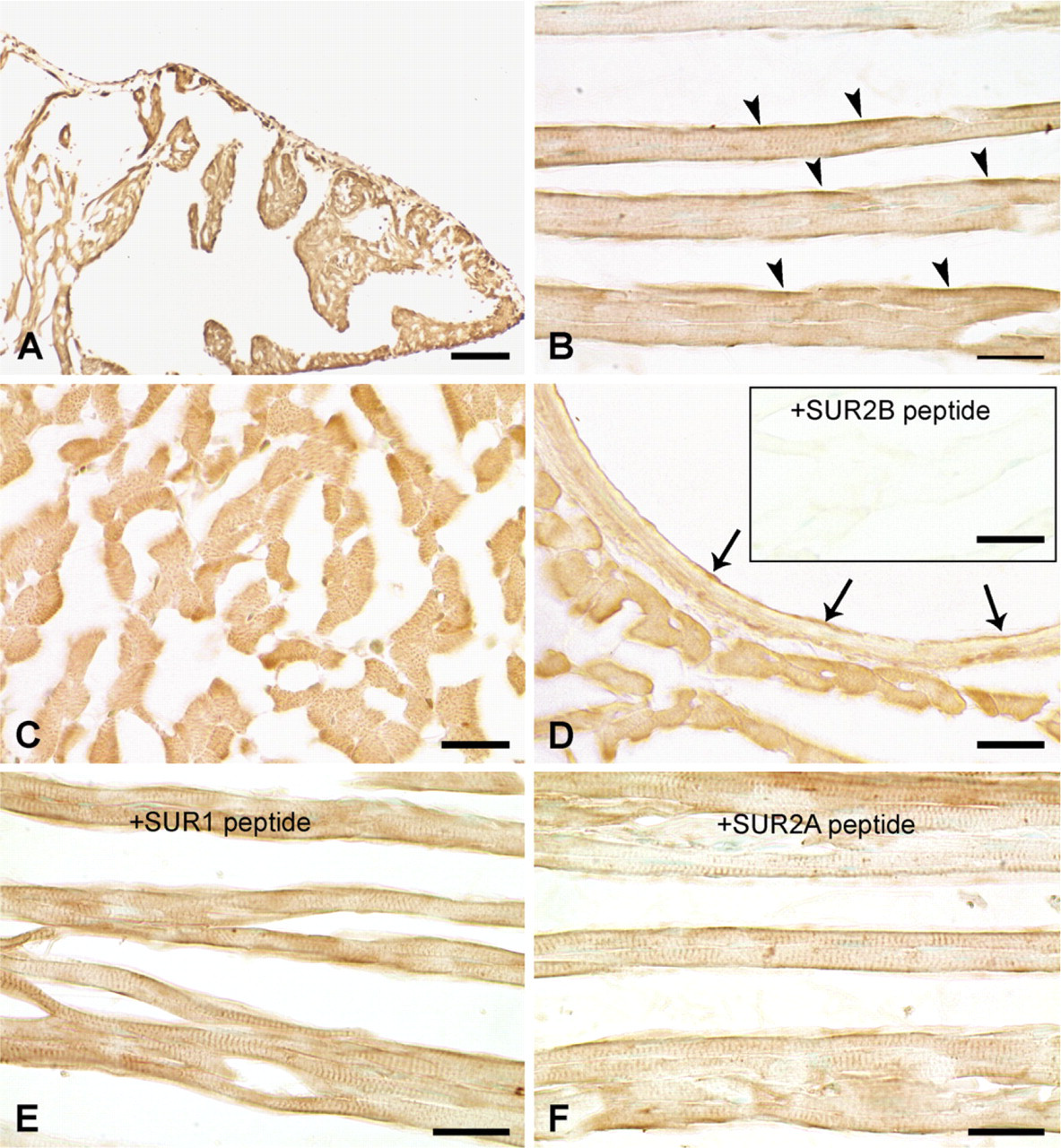
Immunohistochemistry showing expression of SUR2B protein in rat heart. Immunoreactivity with anti-SUR2B antibody was seen in cardiomyocytes in the atrium (
Under the electron microscope, SUR2A immunoreactivity, which was evident by light microscopy, was localized in the ER between or along the microfilaments, with additional signal detected in the mitochondria (Figure 4). Weak immunoreaction was also observed on the cell membranes of cardiomyocytes (Figure 4, inset). Some immunoreactivity was localized in t-tubules near the I bands of cardiac muscle fibers or in intracytoplasmic vesicles in the cardiomyocytes. As for SUR2B, the punctate immunoreaction products seen by light microscopy were mainly observed in the mitochondria among the myofilaments of cardiomyocytes when analyzed by immunoelectron microscopy (Figure 5). It is clear that immunoreactivity was also observed in the ER as well as in the cell membrane (Figure 5, inset).
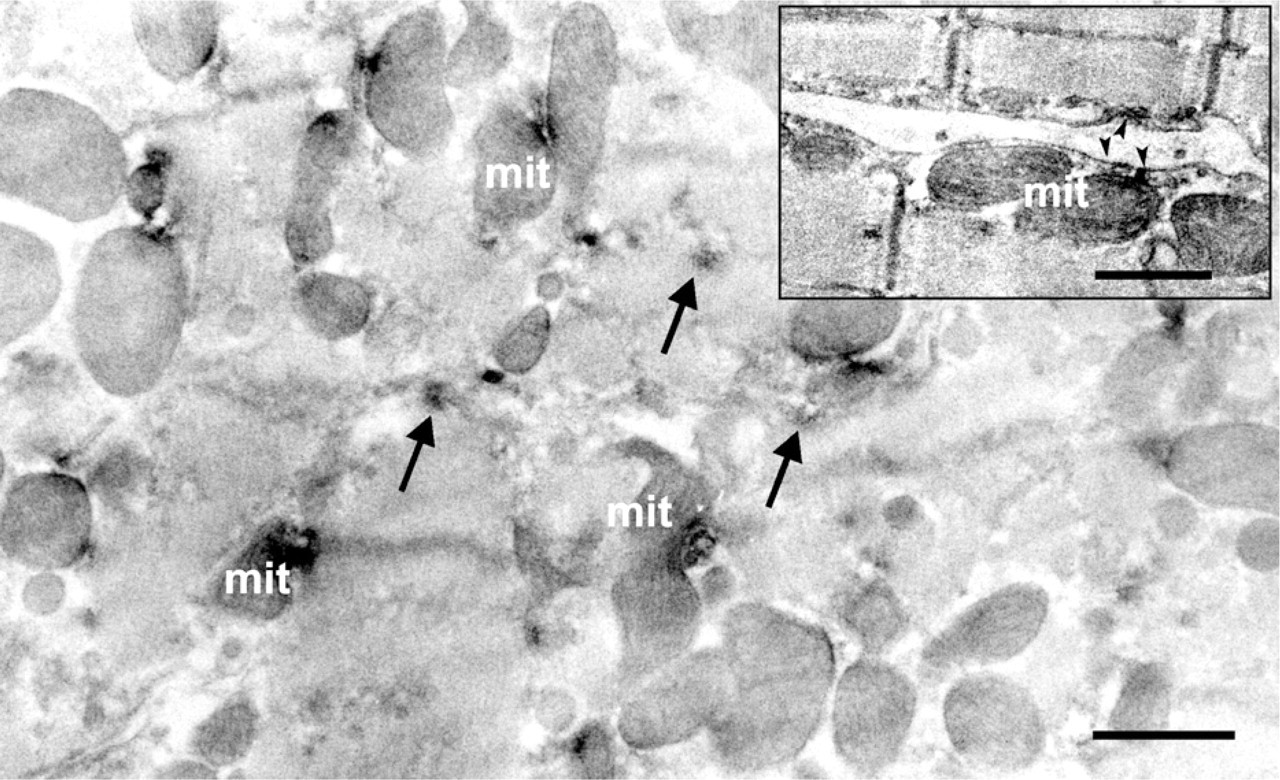
Immunoelectron micrograph of transverse section showing that SUR2A was expressed in the mitochondria (mit) and endoplasmic reticulum (arrows) of rat cardiomyocyte. Longitudinal section (inset) showed that SUR2A was also located in the cell membrane (arrowheads in inset). Bar = 1 μm; Inset = 500 nm.
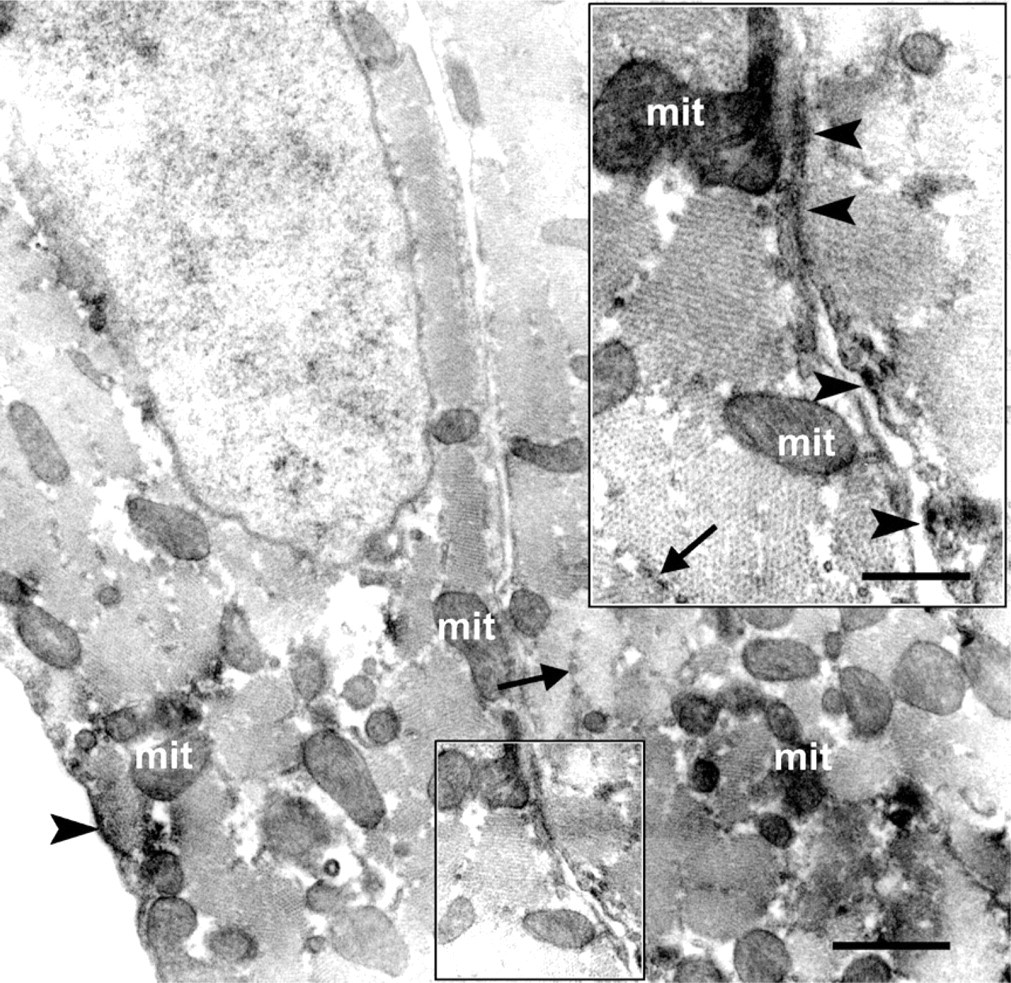
Immunoelectron micrograph showing that SUR2B was localized in the mitochondria (mit), endoplasmic reticulum (arrows), and cell membrane (arrowheads). Enclosed area in the rectangle was enlarged as an inset, showing that cell membrane of rat cardiomyocytes was positive to SUR2B antibody (arrowheads). Bar = 1 μm; Inset = 500 nm.
Discussion
Analysis of cells transfected with individual SUR subunit cDNA revealed that our newly raised anti-SUR2A and anti-SUR2B antibodies recognize only their respective proteins without cross-reaction to the others, although SUR2B antibody showed faint reaction to SUR1. In subcellular fraction experiments, the purity of every cellular fraction analyzed was confirmed by reaction with its own marker antibody. No more than background staining was observed in Western blotting and immunohistochemistry when either antibody was preabsorbed with its immunizing peptide. Experiments were also performed using the SUR2B antibody preincubated with SUR1, SUR2A, or SUR2B peptide in immunoblot analysis and immunohistochemistry. Results showed that the immunoreaction was blocked only by SUR2B peptide and not by SUR2A peptide or by SUR1 peptide, even though SUR1 shares 64% identity with SUR2B in the selective amino acid residues (14-mer) used for raising against SUR2B antibody. Thus, expressions and localizations of SUR2A and SUR2B in Western blot analysis and immunohistochemistry using these antibodies were convincible. The faint cross-reaction to SUR1 with anti-SUR2B antibody could not influence the localization of SUR2B, which was confirmed by the fact that adding SUR1 peptides into SUR2B antibody did not show any remarkable difference to SUR2B antibody-stained section (Figures 3B and 3E).
SURs are important regulators in KATP channels. SUR2A is prominent in cardiomyocytes (Chutkow et al. 1996; Inagaki et al. 1996), whereas RT-PCR of SUR2B has revealed its ubiquitous distribution in various cells and tissues including cardiomyocytes (Isomoto et al. 1996) and rat vascular smooth muscle (Wang et al. 2003). Until now, there have been no unequivocal data establishing the precise cellular localization of SUR2A and SUR2B subunits in cardiac myocytes. A previous study (Singh et al. 2003) showed that SUR2A colocalized with mitochondria in rat isolated ventricular myocytes in immunostaining but failed to demonstrate full-length SUR2A in immunoblot analysis of mitochondrial fractions. Our newly prepared anti-SUR2A antibody established, by Western blotting and immunoelectron microscopy, that full-length SUR2A is present not only in the cell membrane but also in the mitochondria of rat cardiomyocytes. A previous study revealed that SUR1 was strongly expressed at the sarcolemmal surface of ventricular myocytes (Morrissey et al. 2005). SUR2B was expressed in the microsome fraction in immunoblot analysis (Singh et al. 2003) and in t-tubules of rat cardiomyocytes in immunofluorescent staining in isolated rat cardiomyocytes (Singh et al. 2003; Morrissey et al. 2005) but not expressed in mitochondria. Our anti-SUR2B antibody established that SUR2B is widely expressed not only in the cell membrane, t-tubules, and microsome, but further revealed SUR2B is also in the mitochondria of rat cardiomyocytes both by immunoblotting and immunoelectron microscopy.
Kir6.1 and SUR2B are reported to form the smooth muscle type of KATP in blood vessels (Li et al. 2003; Seino and Miki 2003; Wang et al. 2003), although at least one study argues for a sulfonylurea-sensitive but ATP-insensitive K+ channel in vascular smooth muscle (Yamada et al. 1997); firm morphological evidence has not yet been provided. A recent report demonstrated that SUR2, not SUR1, is present in blood vessels, but the antibody used could not distinguish SUR2A from SUR2B (Morrissey et al. 2005). The antibodies described here have allowed discrimination between SUR2A and SUR2B, although the latter has faint affinity with SUR1, and our results provided evidence that both SUR2 subunits are expressed in the smooth muscles and endothelium of coronary blood vessels.
The uncertainty as to whether Kir6.1 is a subunit of the sarcolemmal (Seharaseyon et al. 2000; Kuniyasu et al. 2003) or mitochondrial KATP channel (Lacza et al. 2003; Singh et al. 2003; Zhou et al. 2005) applies to SURs as well. Originally, the molecular mass of SUR was determined to be 140–170 kDa by gel filtration chromatography and about ∼250 kDa by radiation inactivation analysis (Skeer et al. 1994), suggesting that the β-cell SUR is composed of more than one subunit in the native membrane. Later, high-affinity hamster SUR was cloned, and proteins of ∼140 kDa were detected by SDS-PAGE (Aguilar-Bryan et al. 1995). A 170-kDa and a 108-kDa SUR2 protein were detected in kidney mesangial cells (Szamosfalvi et al. 2002). Mitochondria also contain a 28-kDa SUR (mito-SUR), as revealed by binding studies performed in beef heart mitochondria with [3H]glibenclamide and SDS-PAGE (Szewczyk et al. 1997), and one of 63 kDa that was purified from rat brain mitochondrial fractions by DEAE–cellulose column chromatography (Bajgar et al. 2001). It is unknown why there were so many different sizes of mature SUR polypeptides detected in different tissues (Aguilar-Bryan et al. 1995; Szamosfalvi et al. 2002) or even in the same subcellular fractions (Szewczyk et al. 1997; Bajgar et al. 2001) in different tissues. It might be as a result of posttranslational modification or the susceptibility of SUR proteins to proteolysis. Our results revealed that SUR2B expressed in mitochondria fraction as a prominent polypeptide of ∼50 kDa and a weaker polypeptide of ∼120 kDa, suggesting that the short polypeptide may be a stable breakdown fragment from the 120-kDa polypeptide to proteolysis during the preparation of mitochondria. KATP channels are heteromultimers of SURs and Kir6.x with a (SURs-Kir6.x)4 stoichiometry with an estimated molecular mass of 950 kDa (Clement et al. 1997). Although core glycosylated SUR proteins located in the cell surface membrane have been detected at 100–250 kDa (Skeer et al. 1994; Aguilar-Bryan et al. 1995; Szamosfalvi et al. 2002), SUR in mitochondria (mito-SUR) was ∼28–63 kDa (Szewczyk et al. 1997; Bajgar et al. 2001). It is possible that mito-SUR protein may be truncated and active in a short form. Thus, the possibility could not be precluded that the short molecular SUR2B might be special for mito-SUR in mito-KATP. Further study is required to determine whether there are two different proteins, that is, sarcolemmal SURs and mitochondrial SURs, or whether they are the same proteins but expressed differentially in the cell membrane or in cellular organelles.
In conclusion, we established that SUR2A and SUR2B are widely expressed in rat heart. In cardiomyocytes, they are localized in the mitochondria, ER, and cell membrane.
Footnotes
Acknowledgements
This work was supported in part by research grants from Akita University (to HA) and in part by grants from Kitasato University School of Medicine (to KK).
We thank Prof. S. Seino (Kobe University Graduate School of Medicine) for his kind gift of plasmids pcDNA3-rSUR1, pCMV6c-rSUR2A, and pCMV6c-rSUR2B, and Prof. H. Kondo (Tohoku University Graduate School of Medicine) for his kind suggestions regarding this manuscript. We thank the staff of the Bioscience Research-Education Center of Akita University School of Medicine for their help, and Mr. Nibe for his assistance in this study.
