Abstract
The spindle checkpoint, the primary mechanism to ensure that two daughter cells receive the same amount of DNA, is compromised in many malignant tumors and has been implicated as a contributor to aneuploidy and carcinogenesis. The extent of expression and subcellular localization of the spindle proteins Aurora A, Mad2, and BUBR1 varies considerably in different immunohistochemical (IHC) reports from archival tumor tissues. Given the conflicting reports in the literature about the localization of these proteins, we examined the subcellular localization of Aurora kinase A, Mad2, and BUBR1 in normal and cancerous human tissues by IHC. In normal tissues, Aurora A was mainly localized to the nucleus when monoclonal or purified polyclonal antibodies were used, and Mad2 was localized to the nucleus, whereas BUBR1 was localized to the cytoplasm. In malignant tissues, Aurora A showed additional staining in the cytoplasm in the majority of tumors analyzed. Furthermore, BUBR1 was also localized to both the nucleus and cytoplasm in a significant fraction of tumors. Subcellular localization of Mad2 was similar in normal and malignant tissues. Thus, the validity of some earlier IHC studies of Aurora A, Mad2, and BUBR1 should be reconsidered, indicating that high-quality antibodies and a high-alkaline antigen-retrieval technique are required to achieve optimal results. We conclude that the subcellular localizations of these spindle proteins are different, although they have overlapping biological functions, and that Aurora A and BUBR1 undergo a shift in the subcellular localization during malignant transformation.
M
The role of these proteins during mitosis and their subcellular localization during cell cycle progression are not completely known. Results from immunohistochemical (IHC) studies concerning their subcellular localization in human archival tissues are contradictory. Most groups report cytoplasmic staining of Aurora A (Tanaka et al. 1999; Takahashi et al. 2000; Ehara et al. 2003; Hoque et al. 2003; Li et al. 2003a; Rojanala et al. 2004; Royce et al. 2004; Tong et al. 2004), whereas other studies show nuclear staining (Gritsko et al. 2003; Tanaka et al. 2005). Three different reports show that Mad2 is localized to the cytoplasm in adenocarcinoma of the colon and stomach (Tanaka et al. 2001; Li et al. 2003b; Li and Zhang 2004). However, Mad2 was localized to the nucleus in mammary tissue (Pati et al. 2004) with additional staining in the cytoplasm of breast cancer cells. To our knowledge, only one study exists that has evaluated BUBR1 expression by IHC (Shin et al. 2003). This study shows nuclear staining in normal colonic epithelium and predominantly cytoplasmic localization in colon cancer. Given the conflicting reports of the subcellular localizations of these proteins as measured by IHC, this study had two main goals. First, after optimization of IHC staining protocols, we wanted to investigate the subcellular localization of Aurora A, BUBR1, and Mad2 in various normal tissues. Second, we wanted to investigate putative changes in the subcellular localization of these proteins during neoplastic transformation.
Materials and Methods
Thirteen normal tissues, five pancreatic adenocarcinomas, five squamous cell carcinomas of the skin, and five diploid and five aneuploid colon cancers were included in the study, including five normal controls for each cancer. Tissue microarrays were made from formalin-fixed, paraffin-embedded tissue blocks using the Beecher Tissue Microarrayer (Beecher Instruments; Sun Prairie, WI) (Camp et al. 2000; Hoos and Cordon-Cardo 2001; Hoos et al. 2001; Packeisen et al. 2002, 2003). Three cores from each tissue block were placed into the recipient paraffin block with a core size of 0.6 mm or 1.0 mm.
Specificities of commercially available antibodies directed against the different spindle proteins were evaluated by IHC and Western analysis. After optimization of staining protocols, the best antibodies were selected for further investigation of the subcellular localization of Aurora A, BUBR1, and Mad2 in normal tissues and their neoplastic counterparts.
Aurora A, Mad2, and BUBR1 are expressed at high levels in cells with a rapid proliferation rate (Zhou et al. 1998; Tanaka et al. 2001; Grabsch et al. 2003). Normal tonsil is an easy accessible tissue containing highly proliferative cells in its germinal centers and was chosen as positive control. Earlier studies report high levels of these proteins in Hela cells (Honda et al. 2000; Kops et al. 2004); therefore, a Hela cell lysate was added as an unambiguously positive control for Western analysis. A piece of fresh tonsil tissue was fixed in formalin for IHC studies, and cell suspensions were prepared for Western analysis. This enabled us to directly compare IHC findings with the Western analyses. Four colon cancers were included in the Western analyses to investigate expression of Aurora A, Mad2, and BUBR1 in a cancerous tissue reported to overexpress these proteins (Takahashi et al. 2000; shichiri et al. 2002; Li et al. 2003b). Normal colon expresses these proteins at very low levels (Bischoff et al. 1998) and was used as negative control. Fresh colonic mucosa was fixed in formalin for IHC analysis, and cell suspensions were made for Western analysis.
IHC
Tissue sections were cut at 5-μm thickness, mounted on SuperFrost glass slides (Menzel-Glaser; Braunschweig, Germany) and incubated at 56C for 1 hr. Three different antigen-retrieval buffers were tested: (a) citrate buffer, pH 6.0; (b) EDTA buffer, pH 8.0; and (c) Tris-EDTA buffer, pH 9.0.
Xylene-dewaxed paraffin sections were exposed to 0.5% H2O2 solution for 10 min to block endogenous peroxidase, rinsed in tap water for 5 min, and subjected to antigen retrieval in Tris-EDTA buffer in a microwave oven. The oven was set to 800 W and allowed to reach boiling, at which point the oven setting was reduced to 300 W to maintain a temperature of >85C for an additional 20 min. When sections had cooled to room temperature, they were incubated for 1 hr with the relevant antibodies diluted with Antibody Diluent (Ventana Medical Systems; Tucson, AZ). Slides were rinsed for 10 min in IHC wash solution (Ventana Medical Systems) and incubated for 30 min with the HRP-labeled polymer conjugated secondary antibody (EnVision+; DakoCytomation, Carpintera, CA). After rinsing for 10 min in IHC wash solution, DAB1 (DakoCytomation) was applied as a substrate for 7 min, the slides were rinsed in distilled water for 10 min, counterstained with hematoxylin, and mounted. A Nikon (Tokyo, Japan) Eclipse E 600 light microscope was used to evaluate the staining results. One hundred randomly selected, positive cells in each tissue were evaluated to determine the subcellular localization of each protein. If >90% of the cells showed nuclear staining, localization of the protein in question was said to be primarily nuclear. If the number of cells with both nuclear and cytoplasmic staining exceeded 10%, the protein in question was localized to both the nucleus and cytoplasm.
Western Analysis
Samples for Western analysis were made from Hela cells and fresh frozen tissue that was cut into small pieces and boiled in 300 μl Laemmli buffer containing 5% β-mercaptoethanol and 0.5% PMSF. Protein concentrations were measured (RC DC Protein Assay; Bio-Rad Laboratories, Hercules, CA) and 40 μg of protein sample and 5 μg of a molecular mass standard (Precision Plus Protein Standards; Bio-Rad Laboratories) were loaded onto SDS-polyacrylamide gel. After electrophoresis, samples were transferred onto polyvinylidene difluoride membranes and incubated overnight at 4C with the primary antibodies of interest. Blots were then incubated with AP-labeled polymer-conjugated secondary antibody for 1 hr (Envision+; DakoCytomation) and developed using a NBT/BCI colorimetric procedure (Roche Diagnostics; Mannheim, Germany). Antibodies that did not produce specific bands in accordance with the published molecular mass of each protein, or that gave high levels of background, were excluded for further investigations.
Antibody Purification
Highly confluent HCT116 colon cancer cells expected to express Aurora A at very low levels were trypsinized, rinsed in PBS, and centrifuged at 1500 x g for 6 min and the supernatant discarded. The pellet was washed twice in PBS and resuspended in 10 ml PBS. One ml of this solution was treated with three cycles of 10-sec sonication on ice. The cell lysate was mixed with a polyclonal antibody against Aurora A (Santa Cruz Biotechnology; Santa Cruz, CA) to achieve relevant antibody concentration for IHC studies (1:300) and Western analysis (1:375). After incubation overnight at 4C, the sample was centrifuged at 2000 x g for 6 min, and the supernatant solution containing the absorbed polyclonal antibody was used for Western and IHC analyses.
The study protocol was in accordance with the Helsinki Declaration and approved by the regional ethical committee for scientific studies on human tissues.
Results
Antibody Specificity and Antibody Purification
For Aurora A and BUBR1, three different antibodies were tested and for Mad2 two different ones were tested (Table 1). The monoclonal antibody from Novocastra (Newcastle, UK) against Aurora A (46 kDa) showed a high degree of antigen specificity with low levels of background (Figure 1). Both antibodies (BD Transduction Laboratories; Franklin Lakes, NY and Immuquest; Cleveland, UK) against Mad2 (24 kDa) gave high-quality results on IHC and Western analyses (Figure 1). For the detection of BUBR1 (125 kDa), only the antibody from BD Transduction Laboratories gave optimal results (Figure 1). As seen from the Western blot analyses in Figure 1, the antibodies showed distinct bands at the molecular mass for each respective protein in lysates from Hela cells (He) and normal tonsils (T). In the normal colon, only a few weakly positive cells localized to the basal part of the crypts could be seen, and the levels were too low to be detected by Western analysis (Figure 1). The variability in band intensity in the colon cancers (#6, #13, #19, #20) of the different antibodies probably reflects different levels of protein in each cancer.
With the monoclonal antibody against Aurora A, we found primarily nuclear staining in contrast to the polyclonal antibodies tested (Table 1). These antibodies gave a high degree of cytoplasmic staining in addition to variable nuclear positivity. Corresponding Western blots showed multiple bands of different molecular masses other than 46 kDa (molecular mass of Aurora A). After antibody purification, the nuclear staining persisted with only faint staining of the cytoplasm with a reduced number of nonspecific bands on Western blot analysis (data not shown). Antibody purification requires the antibody to be exposed to cells that do not express the antigen of interest. Because Aurora A is expressed only in proliferating cells and has a limited half-life of ∼2 hr (Honda et al. 2000), exposing the antibody to confluent cell cultures fulfills this criterion. Purification of the antibodies against Mad2 and BUBR1 was not performed because the selected antibodies showed a high level of antigen specificity on Western analysis, and background staining on IHC was not evident.
The different antigen-retrieval buffers used in the heat-induced antigen-retrieval protocol gave different results with respect to nuclear staining and staining intensity. The use of an automated IHC slide-staining machine, NexES IHC (Ventana Medical Systems) gave weak or negative results for all proteins with each buffer tested. Applying a manual staining protocol with citrate buffer (pH 6.0), BUBR1 showed only weak staining of tonsils and proliferative phase endometrium. Aurora A and Mad2 were negative. EDTA buffer (pH 8.0) gave weak or moderately good staining quality in most tissues for Mad2 and Aurora A. For the detection of BUBR1, EDTA buffer gave equivalent results as with Tris-EDTA (pH 9.0) except for markedly weaker staining in colon cancer. To achieve optimal staining in all tissue types, Tris-EDTA buffer was necessary for all antibodies. One important observation concerning Aurora A was the lack of nuclear staining in tonsils, lymph nodes, and proliferative phase endometrium when EDTA buffer was used. Aurora A was localized to the nucleus in cells of these tissues only when Tris-EDTA buffer was used in the antigen retrieval.
Description of antibodies tested: clonality and results of IHC and WA analyses
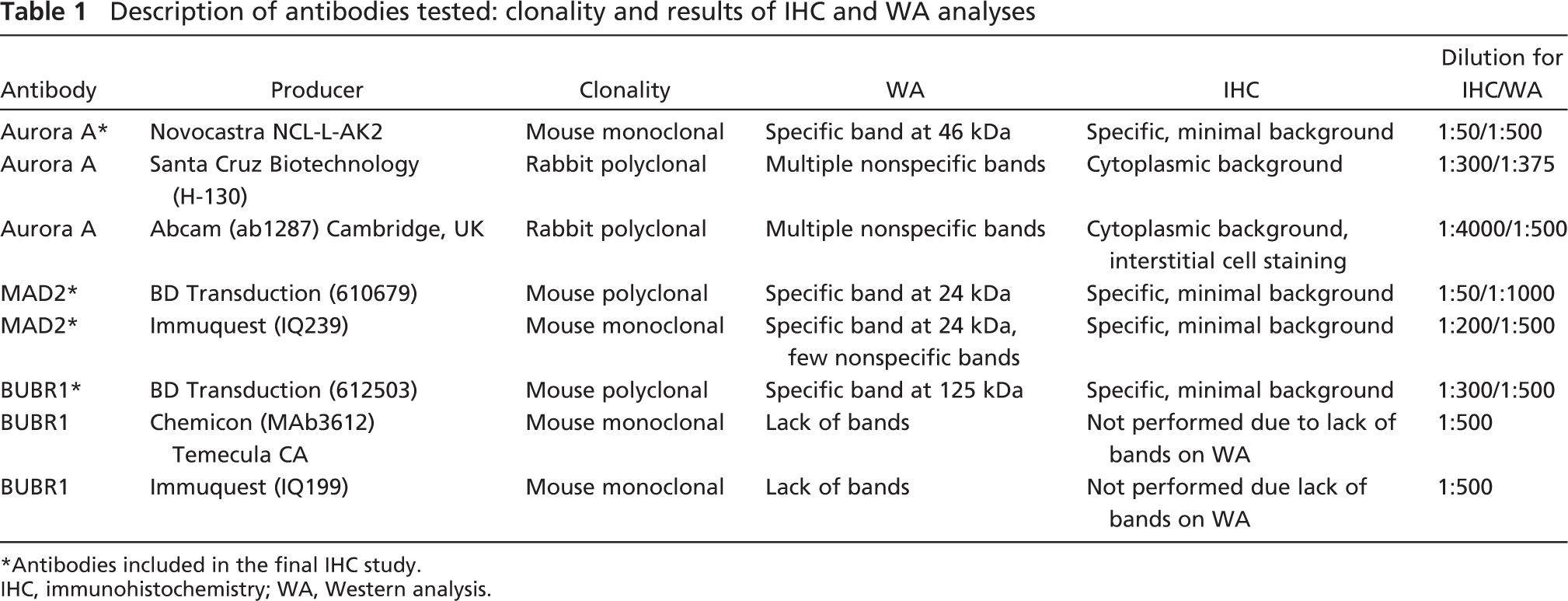

∗Antibodies included in the final IHC study.
IHC, immunohistochemistry; WA, Western analysis.
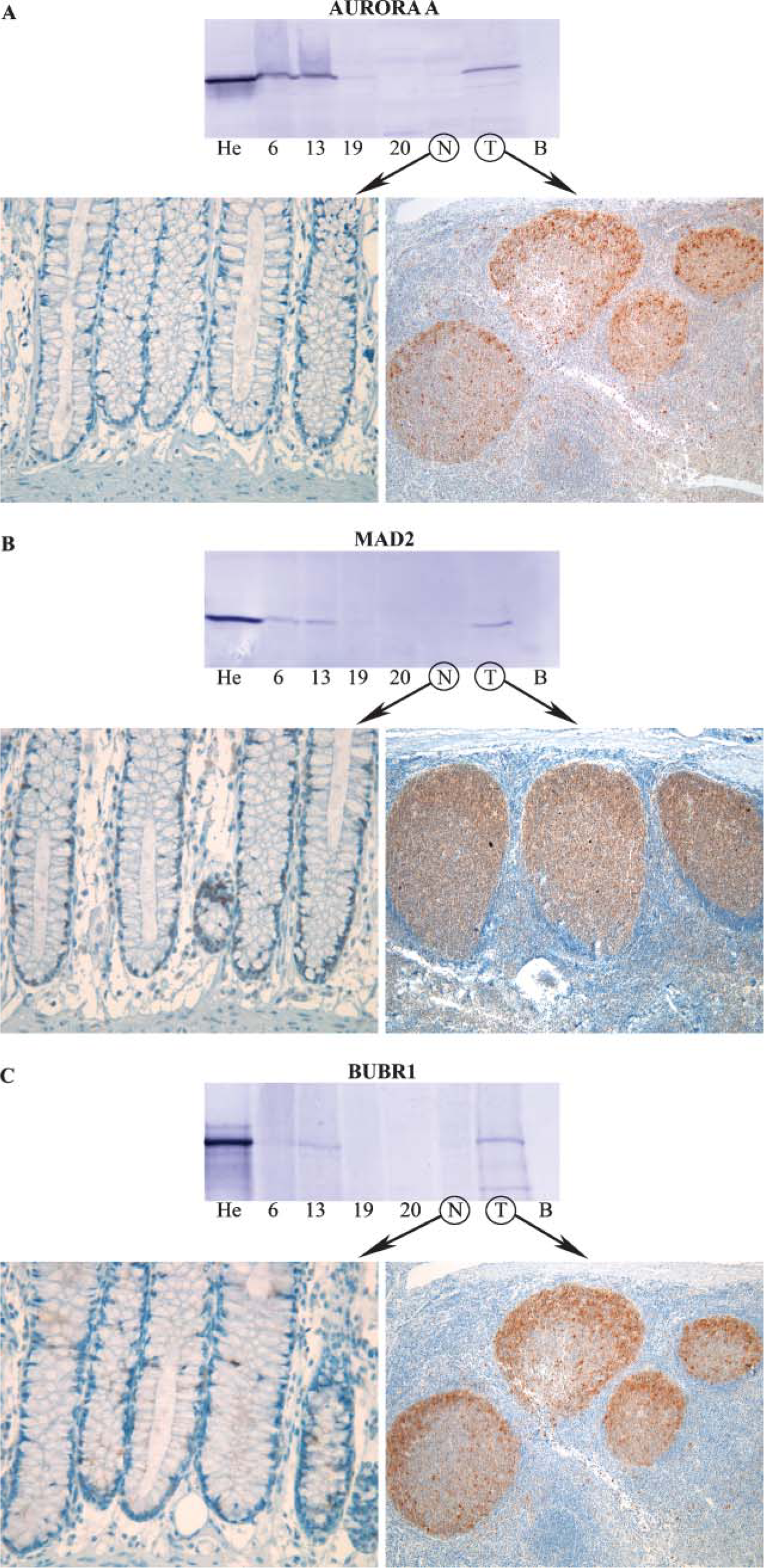
Western and immunohistochemical (IHC) analyses with antibodies against (
Subcellular Localization of Aurora A, Mad2, and BUBR1 in Normal Tissues
All proteins stained mitotic cells in all tissues as reported in the literature. Parenchymal cells of stable tissues such as liver, kidney, and pancreas did not express detectable levels of either protein. In pancreatic ducts, positive cells for Aurora A and Mad2 in the nucleus were rarely seen. The secretory phase endometrium is non-proliferating and consequently was negative for Aurora A and BUBR1. However, Mad2 showed weak perinuclear staining, reflecting that Mad2 is localized to nuclear pores during interphase (Campbell et al. 2001).
Aurora A was localized to the nucleus in all labile tissues examined (Table 2), and in highly proliferating cells such as spermatocytes and proliferative phase endometrium additional cytoplasmic staining was evident (Figures 2A and 2B). In gastric pits, Aurora A was mainly localized to the cytoplasm with 75% of the cells positive only in the cytoplasm. The remaining cells showed Aurora A both in the nucleus and cytoplasm. Both antibodies against Mad2 (Table 1) showed nuclear staining in all labile tissues examined. Additional cytoplasmic staining of Mad2 was seen in cells of highly proliferating tissues such as spermatocytes and proliferative phase endometrium (Figures 2C and 2D). On the other hand, BUBR1 stained the cytoplasm of all labile tissues examined (Figures 2E and 2F). Table 2 summarizes the subcellular localization of Aurora A, Mad2, and BUBR1. Studies of the spindle proteins on fresh frozen tissue (tonsils) gave equivalent results as described for formalin-fixed tissues. Negative controls where TBS was substituted for the primary antibody did not produce any visible background staining.
Subcellular localization of Aurora A, Mad2, and BUBR1 examined by IHC
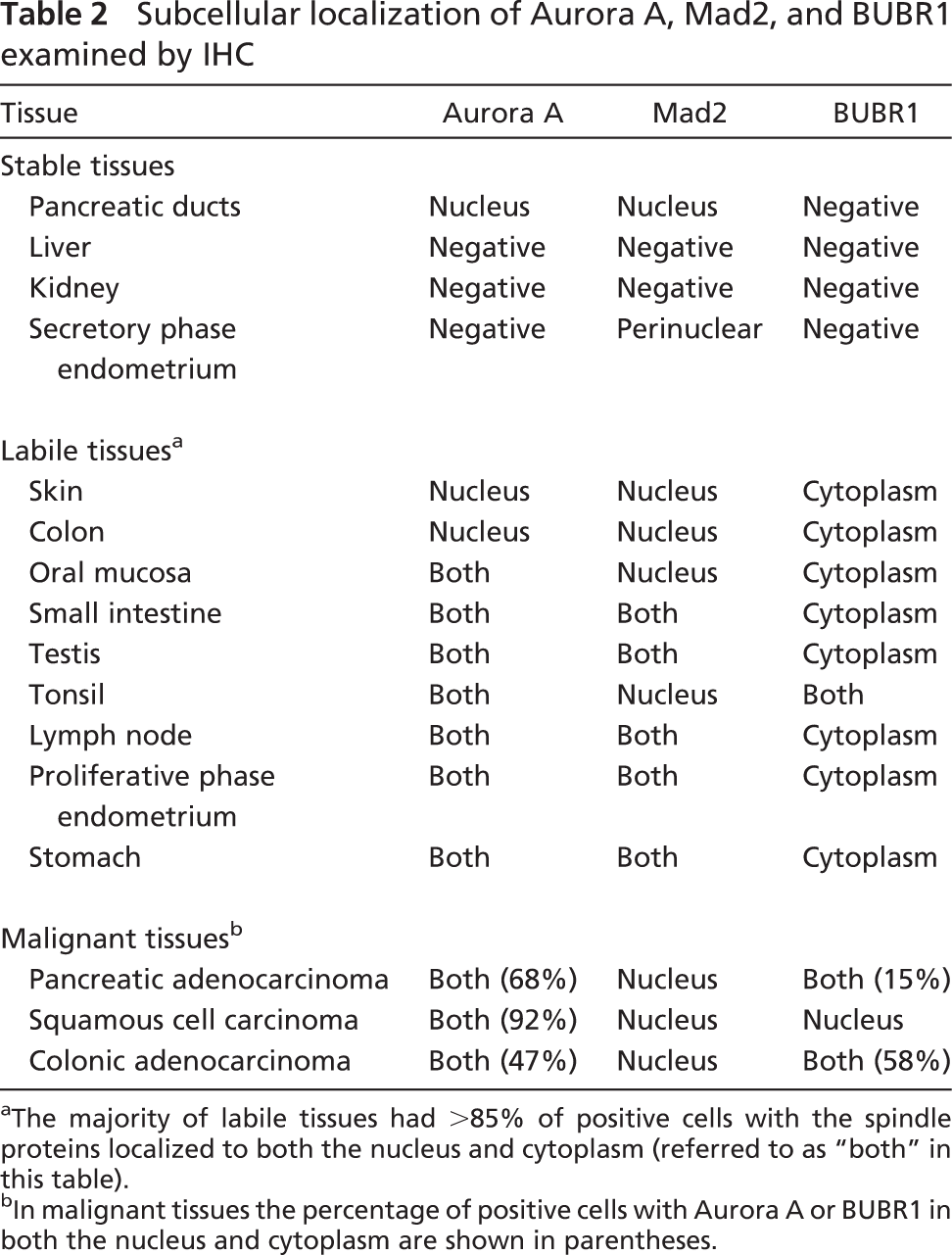
aThe majority of labile tissues had >85% of positive cells with the spindle proteins localized to both the nucleus and cytoplasm (referred to as “both” in bhis table).
bIn malignant tissues the percentage of positive cells with Aurora A or BUBR1 in both the nucleus and cytoplasm are shown in parentheses.
Subcellular Localization of Aurora A, Mad2, and BUBR1 in Malignant Tissues
Skin, colonic mucosa, and pancreas representing two labile tissues and one stable normal tissue, and carcinomas developing from these tissues, were selected for further IHC analysis. Normal tissues showed only nuclear staining for Aurora A and Mad2, whereas BUBR1 was localized to the cytoplasm (Table 2). Aurora A was localized to both the nucleus and cytoplasm in 68% and 47%, respectively, of positive cells of adenocarcinomas of pancreas and colon (Figures 3A and 3B). Furthermore, Aurora A was localized to both the nucleus and cytoplasm in >90% of positive cells in squamous cell carcinoma of the skin (Figures 3C and 3D). Remaining positive cells showed Aurora A localized only to the nucleus. Subcellular localization of Mad2 remained nuclear in malignant tissues; however, perinuclear staining was more evident. As for Aurora A, the subcellular localization of BUBR1 changed from cytoplasmic in normal tissues to also include the nucleus in malignant ones. In colon cancer, BUBR1 was localized to both the nucleus and cytoplasm in 43% of positive cells (Figures 3E and 3F), and 15% of the cells showed nuclear staining only. Remaining positive cells showed BUBR1 localized to the cytoplasm as in normal tissues. In pancreatic adenocarcinoma, BUBR1 was localized both to the cytoplasm and nucleus in 15% of positive cells. In squamous cell carcinoma this number was reduced to 7%, with remaining cells being positive only in the cytoplasm in both cancers.
The 10 colon cancers studied in this report were subjected to DNA ploidy analysis by flow cytometry according to Vindeløv et al. (1983) and are a part of an ongoing study in our laboratory. Even though there was an obvious change in the subcellular localization of Aurora A and BUBR1 from normal to malignant tissues, preliminary data revealed no association between the subcellular localization or expression levels of these proteins and ploidy status in colon cancers.
Discussion
Aurora A
The difference in molecular mass between the spindle proteins (Aurora A 46 kDa, Mad2 24 kDa, and BUBR1 125 kDa) is a likely explanation for the different sub-cellular localization in normal tissues shown in this report. Studies of nucleocytoplasmic transport show that proteins <50 kDa in size can pass through the nuclear pores without energy utilization (Gorlich and Mattaj 1996). This makes it possible for the small spindle proteins Aurora A (46 kDa) and Mad2 (24 kDa) to diffuse into the nucleus after their production in the cytoplasm. However, the large molecular mass of BUBR1 makes the association with nuclear structures possible only in mitosis under normal circumstances, due to loss of the nuclear membrane in mitosis.
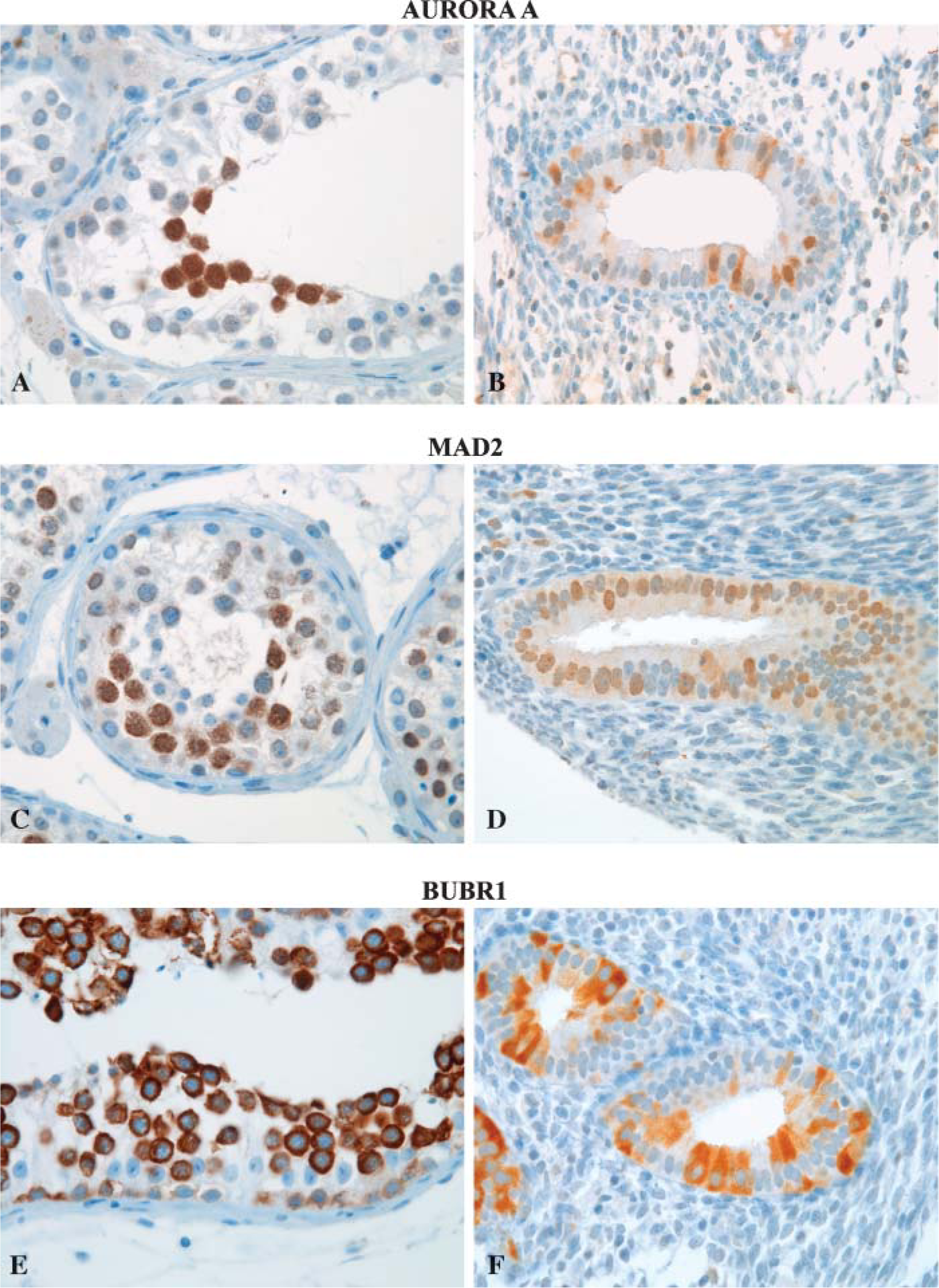
Examples of IHC of Aurora A (Novocastra) (
Several IHC studies report that Aurora A is localized only to the cytoplasm (Tanaka et al. 1999; Takahashi et al. 2000; Ehara et al. 2003; Hoque et al. 2003; Li et al. 2003a; Rojanala et al. 2004; Royce et al. 2004; Tong et al. 2004). Our study shows nuclear localization of Aurora A in a variety of different normal and malignant human tissues. The nucleus consists of a dense network of highly packed nucleic acids and proteins, and sufficient antigen retrieval is crucial to open the condensed DNA helices. The most important factors for retrieval of antigens masked by formalin fixation are pH and temperature (Shi et al. 1997). Our results indicate that an antigen-retrieval buffer with a high alkaline pH (Tris-EDTA, pH 9.0) is the most effective buffer to break cross-linkages and to increase membrane permeability to facilitate access for antibodies to nuclear antigens in formalin-fixed tissues. Studies on nuclear antigens and the effect of pH described by Morgan et al. (1997) are consistent with this finding. Some IHC studies of Aurora A did not apply antigen retrieval at all (Tanaka et al. 1999; Li et al. 2003a), and most other studies have used citrate buffer (standard citrate buffer has a pH of 6.0) in their antigen-retrieval procedure. If one compares two reports describing Aurora A in squamous cell carcinoma of the esophagus, Tanaka et al. (2005) show primarily nuclear localization of Aurora A using an antigen-retrieval buffer from DakoCytomation (JCode S1699) in an autoclave (121C for 5 min). Contrary to this, Tong et al. (2004) report cytoplasmic staining of Aurora A using sodium citrate buffer (heat induction not specified) in their antigen-retrieval technique. These methodological differences will have a major impact on the accessibility of nuclear antigens to the antibody. Lack of adequate antigen retrieval in the latter and other studies that have used citrate buffer in their procedure (Takahashi et al. 2000; Gritsko et al. 2003; Rojanala et al. 2004; Royce et al. 2004; Tong et al. 2004) may thus explain the lack of nuclear staining in these reports. However, a recent study proposed that the cause of decreased antigen availability caused by formalin fixation was due to a reduction of the electrostatic charges existing between antigens and antibodies, rather than the tenet of formalin-induced methylene bridges and protein denaturation (Boenisch 2006). Furthermore, they report that even without performing antigen retrieval and by prolonging the primary antibody incubation from 10 to 60 min, 35/40 antigens investigated were found to stain equally intense. We examined the staining intensity of the spindle proteins Aurora A, Mad2, and BUBR1 without doing antigen retrieval in our laboratory and extended the primary antibody incubation time from 1 to 18 hr (on sections with formalin-fixed tonsil). We could not detect positive cells after incubation with either of the antibodies. Hence, this shows that an antigen retrieval with a high alkaline pH is necessary for the detection of spindle proteins in formalin-fixed tissues, irrespective of the length of the primary antibody incubation time.
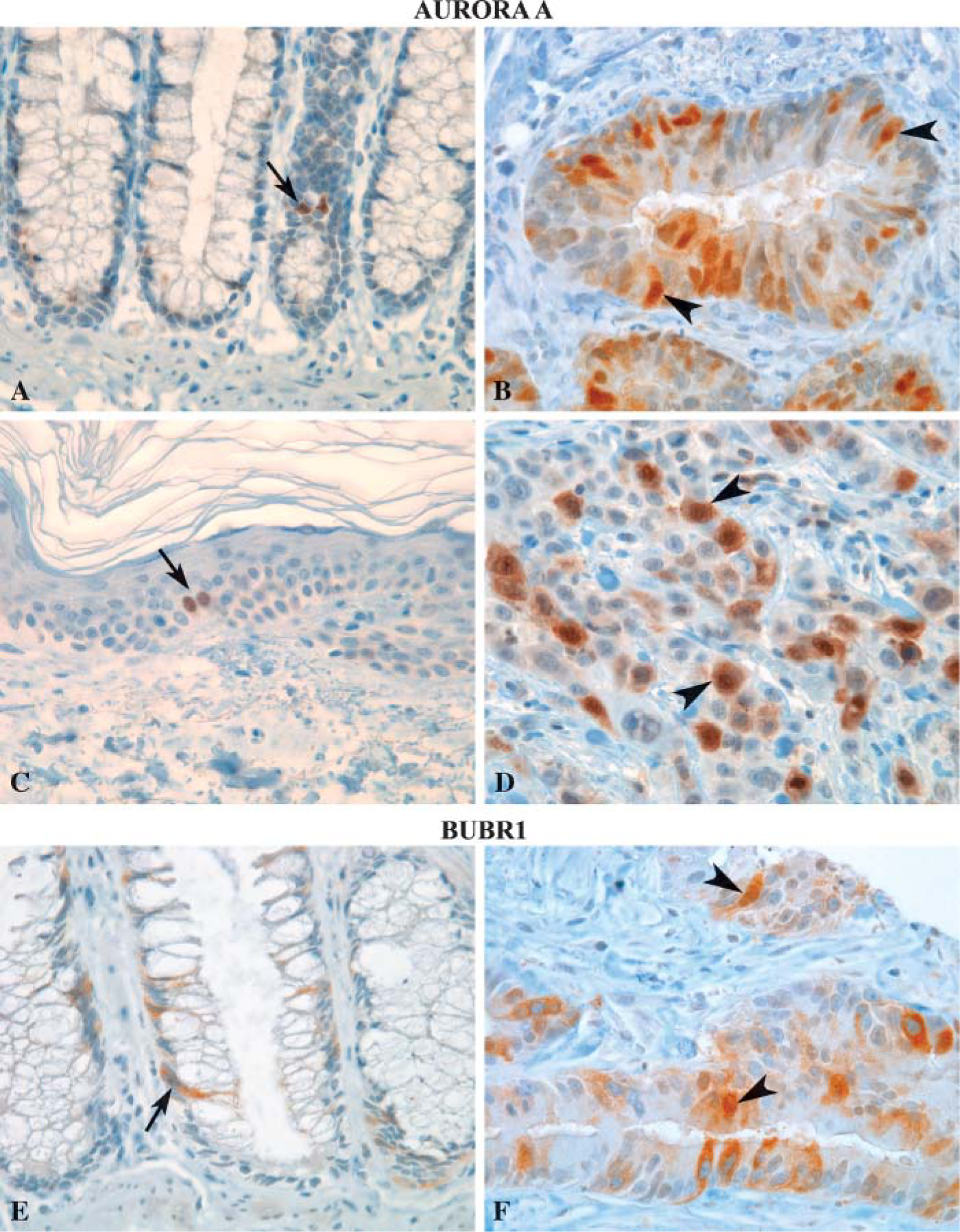
IHC analysis of Aurora A (
Although the antigen-retrieval protocol is optimized, the lack of antibody specificity may obscure the biological appearance of Aurora A within the cell. In previous IHC reports, the antibodies used to study Aurora A were all polyclonal, either produced by the investigators themselves (Tanaka et al. 1999; Takahashi et al. 2000; Hoque et al. 2003; Royce et al. 2004) or purchased commercially (Gritsko et al. 2003; Li et al. 2003a; Rojanala et al. 2004). Furthermore, all secondary antibodies were biotinylated, and heat-induced antigen retrieval was done prior to antibody incubation. We tested two different polyclonal antibodies that had affinities to antigens localized to the cytoplasm other than Aurora A. The unmasking of endogenous biotin is a known consequence of heat-induced antigen retrieval leading to increased level of background staining if followed by biotinylated reagents (Shi et al. 1997). Combining heat-induced antigen retrieval with an insufficient retrieval buffer, nonspecific polyclonal antibodies, and biotinylated reagents, one might get the false impression that Aurora A is localized only to the cytoplasm.
Rojanala et al. (2004) advocate that the diffuse cytoplasmic staining observed for Aurora A in tumors is indicative of the pathological states of cancer cells. This is not in agreement with studies in Hela cells showing that the limited amount of Aurora A in the cytoplasm binds to the centrosome and is not distributed diffusely in the cytoplasm (Honda et al. 2000; Meraldi et al. 2002; Krystyniak et al. 2006). Our IHC studies reveal that Aurora A is localized primarily to the nucleus, with additional staining in the cytoplasm in malignant and some labile normal tissues. The mechanism for this shift in subcellular localization is not known. One interpretation is that the expression of Aurora A may be more closely associated with proliferative activity than with malignant transformation.
Mad2 and BUBR1
Mad2 has also been reported to be distributed diffusely in the cytoplasm when studied by IHC (Tanaka et al. 2001; Li and Zhang 2004). With two antibodies we have shown that Mad2 is localized to the nucleus in agreement with IHC studies in mammary tissue (Pati et al. 2004) and cellular in vitro studies using immunofluorescence (Campbell et al. 2001; Shin et al. 2003). The increased perinuclear staining of Mad2 in cancer cells observed in our study may reflect an increased expression of Mad2 that shifted to the nuclear membrane where it colocalizes with Mad1 (Campbell et al. 2001). Our findings that BUBR1 is localized to the cytoplasm is in agreement with Shin et al. (2003) who detected BUBR1 in the cytoplasm of Hela cells by immunofluorescence. The reason for the localization of BUBR1 to the cytoplasm as compared with the nuclear localization of Aurora A and Mad2 is probably explained by the large molecular mass (125 kDa) of BUBR1. In malignant tissues the subcellular localization of BUBR1 also shifted to include the nucleus. Translocation of cytoplasmic proteins into the nucleus in transformed cells is known, e.g. the translocation of β-catenin from the cytoplasm into the nucleus is indicative of a disruption in the Wnt signal transduction pathway in colon cancers caused by mutations in the APC gene (Morin et al. 1997; Cheuk and Chan 2004). One might speculate that a similar mechanism may be responsible for the translocation of BUBR1 to the nucleus in cancers. However, BUBR1 and other checkpoint components are activated by the lack of microtubule occupancy (Yu 2002) and currently are not linked to signaling pathways activated by external stimuli. Aneuploid colon cancer cells have a robust spindle checkpoint (Tighe et al. 2001), and the transition into the nucleus of BUBR1 could reflect an activated spindle checkpoint in aneuploid cells. However, our preliminary data could not reveal any difference in the levels of BUBR1 or the other spindle proteins in aneuploid vs diploid cancers. More comprehensive studies are needed to investigate this matter further.
In conclusion, antigen retrieval, antibody specificity, and the type of staining protocol are critical aspects when performing IHC and will affect the final results considerably as demonstrated in this study. Knowledge of the subcellular localization is crucial when interpreting IHC expression and distribution of a protein of interest in normal and neoplastic tissues. We conclude that Aurora A and Mad2 have their prime subcellular localization in the nucleus, and that the cytoplasmic localization of Aurora A in some labile normal tissues may be linked to a higher protein synthesis in proliferating cells with a reduced cell cycle time. BUBR1 is a large spindle protein localized to the cytoplasm in normal tissues. In malignant tissues, Aurora A is localized to both the nucleus and cytoplasm in the majority of cancers examined, whereas Mad2 is localized to the nucleus in both normal and malignant tissues. The subcellular localization of BUBR1 shifts from the cytoplasm in normal tissues to also include the nucleus in malignant tissues. In some colon cancer cells, BUBR1 was localized only to the nucleus. Numerous in vitro studies have investigated these spindle proteins on the cellular level, but in order to understand their biological behavior in human cancer development, additional in vivo studies on human cancers are necessary.
Footnotes
Acknowledgements
The Norwegian Cancer Society provided sole financial support for this study. No conflicts of interest are declared.
The authors thank Professor Helge Scott for excellent technical advice.
