Abstract
Data on the in situ structural–functional organization of ribosomal genes in the mammalian cell nucleolus are reviewed here. Major findings on chromatin structure in situ come from investigations carried out using the Feulgen-like osmium ammine reaction as a highly specific electron-opaque DNA tracer. Intranucleolar chromatin shows three different levels of organization: compact clumps, fibers ranging from 11 to 30 nm, and loose agglomerates of extended DNA filaments. Both clumps and fibers of chromatin exhibit a nucleosomal organization that is lacking in the loose agglomerates of extended DNA filaments. In fact, these filaments constantly show a thickness of 2–3 nm, the same as a DNA doublehelix molecule. The loose agglomerates of DNA filaments are located in the fibrillar centers, the interphase counterpart of metaphase NORs, therefore being constituted by ribosomal DNA. The extended, non-nucleosomal configuration of this rDNA has been shown to be independent of transcriptional activity and characterizes ribosome genes that are either transcribed or transcriptionally silent. Data reviewed are consistent with a model of control for ribosome gene activity that is not mediated by changes in chromatin structure. The presence of rDNA in mammalian cells always structurally ready for transcription might facilitate a more rapid adjustment of the ribosome production in response to the metabolic needs of the cell.
Keywords
T
To study the relationship between the structure and function of chromatin, special attention has been paid to ribosomal genes. It has indeed been known that, in higher eukaryotes, the rRNA genes are present in multiple copies and are organized as tandem repeats. Each repeating unit comprises a transcribed region flanked by a stretch of DNA known as the intergenic spacer. The transcription unit contains, in addition to the genes coding for 18, 5.8, and 28S rRNAs, two external transcribed spacers at their ends and two internal transcribed spacers separating the coding sequences (Hadjiolov 1985). At the electron microscope level, the transcription unit of transcribing ribosomal repeats, visualized in fully spread chromatin preparations, appears to be constituted by a central DNA axis along with axis-associated RNA polymerase I particles connected with nascent growing RNA transcripts whose length progressively increases from the transcription initiation site to termination site, thus giving rise to the well-known “Christmas tree” structure (Miller and Beatty 1969). The transcription unit DNA of transcribing ribosomal genes is in a completely extended, non-nucleosomal configuration. It is clearly distinguishable from spacer chromatin that is often characterized by variable numbers of particles, irregular in shape and distribution, and generally somewhat smaller than those of nucleosomes (Franke et al. 1979; Scheer and Zentgraf 1982; Puvion-Dutilleul 1983). In mammalian cells (Puvion-Dutilleul et al. 1977; Puvion-Dutilleul and Bachellerie 1979), the chromatin of the very long intergenic spacer appeared to be identical to the bulk of transcriptionally inactive chromatin, whose regular, beads-on-a-string appearance indicates a nucleosome organization (Franke et al. 1979; Scheer and Zentgraf 1982; Puvion-Dutilleul 1983). Ribosomal gene expression appears therefore to be tightly related to the structure of chromatin harboring the rDNA. The absence of nucleosomes in transcriptionally active ribosomal genes has also been demonstrated by in vivo psoralen cross-linking assays, which showed that ribosomal genes associated with nascent RNA transcripts are free of regularly spaced nucleosomes, whereas inactive gene copies are characterized by regularly spaced nucleosomes (Conconi et al. 1989; Dammann et al. 1993, 1995).
The structural–functional organization of ribosomal genes has also been investigated in situ at the electron microscope level, mainly in mammalian cells but with much greater difficulties. The drawback for this type of investigation is the high compactness of the nuclear structures that does not allow the visualization of the Christmas trees and therefore the precise identification of the transcribing ribosomal genes. Nevertheless, the development of a series of electron microscope techniques, by allowing (a) the precise localization of the ribosomal genes within the nucleolar components, (b) the definition of the structural organization of chromatin, and (c) the identification of the sites of rDNA transcription provided important information on the structural–functional relationship of ribosome genes in situ. The purpose of the present work was to critically review the results obtained in this field of investigation and to discuss them in relation to data resulting from in vitro studies. Before tackling these issues, the ultra-structural organization of the mammalian cell nucleolus will be described briefly.
Organization of Nucleolar Structure
Although recent studies indicate that the nucleolus may be involved in processes such as longevity (Guarente 1997), mitotic entry (Olson et al. 2000), and tumor surveillance (Visintin and Amon 2000), rRNA transcription, rRNA processing, and ribosome assembly represent the major functions of the nucleolus. At the ultrastructural level, in routinely prepared samples, four distinct morphological components are constantly visualized within the mammalian nucleolus, independent of the cell type and nucleolar functional activity: the fibrillar center (FC), the dense fibrillar component (DFC), the granular component (GC), and the nucleolus-associated chromatin (Thiry and Goessens 1996) (Figure 1). The FCs are spherical structures of different sizes with a very low electron opacity and are composed of loosely interwoven fibrils. Their number is highly variable, ranging from one to several tens per nucleolus. The DFC is more electron opaque than the FCs and is composed of tightly packed fibrils. Usually, the DFC is located around the FCs. The GC is composed of granules of ∼15 nm in diameter, which surround the fibrillar components. The nucleolar chromatin can be subdivided into perinucleolar and intranucleolar chromatin. Perinucleolar chromatin may constitute a regular shell of variable thickness or may be arranged in small clumps around the nucleolus. Extensions of this perinucleolar chromatin penetrate deeply into the nucleolar body and come in contact with the FCs. At this place, the DFC surrounding the FCs is interrupted. In ultrathin sections, these invaginations appear as small compact clumps or more extended fibers of variable thickness.
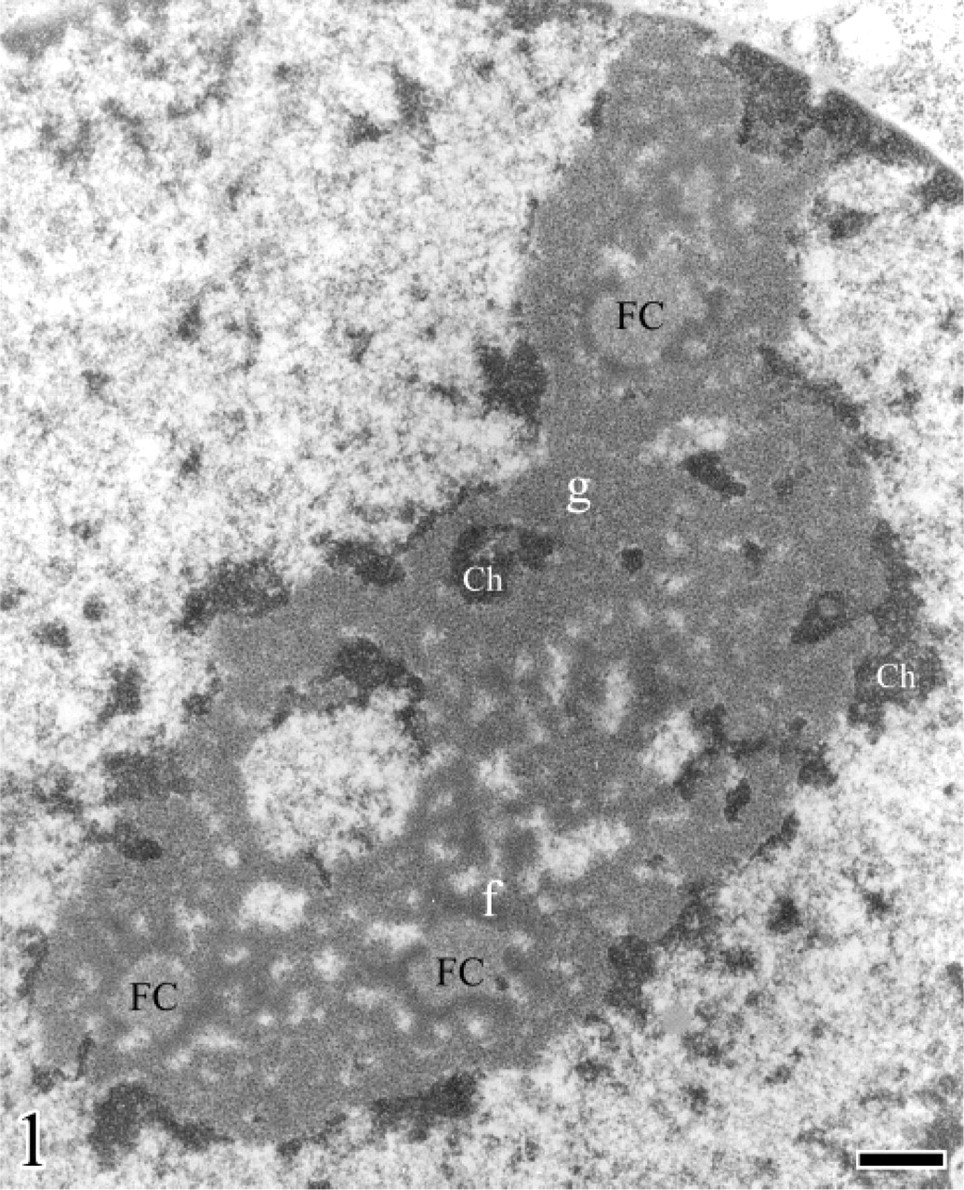
Electron microscopy of a nucleolus from human Hep-2 cells (larynx epidermoid carcinoma). The granular (g) and dense fibrillar (f) components as well as the fibrillar centers (FC) and the condensed chromatin (Ch) associated with the nucleolus are shown. Bar = 0.15 μm. From Figure 2–10, if not otherwise specified, samples were formaldehyde fixed and Epon embedded.
Localization of Ribosomal Genes within the Nucleolar Components
The precise location of the ribosomal genes in the nucleolar components has been defined by a series of cytochemical, immunocytochemical, and hybridization techniques carried out at the EM level. In actual fact, the major effort of these studies was to identify the structures within the nucleolus that represent the inter-phase counterpart of the nucleolar organizer regions (NORs) of metaphase chromosomes (Goessens 1984). In these peculiar chromosome regions, so called because nucleoli are reformed around them at the end of telophase, hybridization experiments have in fact exclusively detected ribosomal genes. A set of argyrophilic, non-histone proteins is associated selectively with the NORs (AgNOR proteins). By exploiting this cytochemical property of the NORs and adapting the silver staining procedure to electron microscopy, it was shown that the only structures stained by silver in interphase nuclei were the FCs and, to a lesser extent, the portion of the DFC around them (Bourgeois et al. 1979; Hernandez-Verdun et al. 1980; Howell 1982; Derenzini et al. 1990; Ploton et al. 1987). However, from a strict morphological point of view only the FCs should be identified as the interphase counterpart of the metaphase NORs because no structure similar to the DFC was ever seen around the NORs. The FCs' equivalence to the metaphase NORs was then accomplished by demonstrating the presence of ribosomal genes within them. The presence of DNA in the FCs was strongly suggested by data obtained by combining, at EM level, the silver staining procedure for the AgNOR proteins with the Feulgen-like osmium ammine staining specific for DNA (Derenzini et al. 1981a; Hernandez-Verdun et al. 1982): a constant overlapping of silver grains with DNA filaments was in fact observed in interphase cell nucleoli of human cell lines (Hernandez-Verdun et al. 1984; Derenzini et al. 1990). Subsequently, the presence of DNA within the FCs was unambiguously demonstrated in various cell types by different immunocytological approaches, in particular, using the terminal deoxynucleotidyl transferase–immunogold technique which allows the specific detection of DNA on ultrathin sections in very well-preserved cell structures, identified by routine uranium and lead staining (Thiry 1992a, b, 1993, 1995; Thiry et al. 1993). EM in situ hybridization studies (Thiry and Thiry-Blaise 1989, 1991; Puvion-Dutilleul et al. 1991, 1992) have further indicated that some rRNA genes are generally located in the intra- and perinucleolar chromatin but are also present in the FCs. Finally, the protein composition also indicated the correspondence between NORs and FCs. Immunocytochemical studies have in fact shown that proteins of the ribosomal gene transcription machinery, such as the RNA polymerase I, the DNA topoisomerase I, and the upstream binding factor are present both in the NORs and the FCs (Thiry and Goessens 1996). Taken together, these data support the notion that in mammalian cells the FCs of interphase nucleoli and the mitotic NORs are at least partly equivalent structures.
Structure of Ribosomal Genes In Situ
The study of the structural organization of chromatin in situ took advantage of the application at EM level of a Feulgen-like staining reaction which utilizes an osmium ammine complex as DNA electron-opaque tracer (Cogliati and Gautier 1973; Olins et al. 1989). By means of this technique, only DNA is stained in situ and the resolution of the stained structures is very high, not limited by the very thin end-reaction product. For a detailed description of the technique and its application to detect the in situ organization of the DNA-containing structures, see the following references (Derenzini et al. 1982a, 1990; Derenzini 1995). In thin sections of mammalian cells stained with the Feulgen-like osmium ammine procedure, only the structures known to contain DNA appear to be electron opaque, whereas all the other nuclear components are completely unstained (Figure 2). As far as the nucleolus is concerned, it can be easily identified by the presence of an almost continuous, electron-opaque ring (the perinucleolar chromatin) that delimits an electron-translucent space (the nucleolar area). In mammalian cells characterized by an intense ribosome biogenesis, many deeply stained structures are present within the nucleolus (the intra-nucleolar chromatin). This chromatin appears to be constituted of compact, irregularly shaped clumps and of extended fibers frequently exhibiting a thickness varying from 11 to 30 nm. A series of studies carried out at high magnification revealed that the chromatin clumps and fibers are composed of repeating roundish subunits with a diameter of ∼11 nm. These subunits are visualized as being composed of an ∼3-nm-thick DNA filament, which encircles an unstained inner core. The shape, size, and structural organization of these particles have indicated that they can be identified with nucleosomes (Derenzini et al. 1981b, 1982a, b, 1983a, b, 1984; Derenzini 1995). As shown in Figure 3 and at higher magnification in Figure 4, the organization of this subunit is not always obviously visible within the chromatin structures. In fact, the diameter of nucleosomes is ∼6–8 times smaller than the thickness of the sections used for electron microscopy. Overlapping of nucleosomes in the chromatin structures makes the visualization of a single subunit very frequently impossible. Furthermore, nucleosomes are differently oriented in the space, so that only when the electron beam is perpendicular to the bases of nucleosomes are the typical ring-shaped particles with a diameter of 11 nm clearly defined. These considerations can explain why the subunit organization can be more easily appreciated at the periphery of the small compact chromatin clumps and in the extended chromatin fibers. In addition to chromatin clumps and fibers, fuzzy, loose agglomerates of very thin contorted DNA filaments are visualized within the nucleolar area by the Feulgen-like osmium ammine reaction. The thickness of these DNA filaments is 2–3 nm, corresponding to that of the DNA double-helix molecule. The filaments never give rise to the nucleosome-like configuration (Figure 3 and Figure 4). This peculiar structural organization of DNA filaments was observed only within the nucleolus, being absent in the extranucleolar compartment. The size and number of these loose agglomerates of DNA filaments were highly variable. Usually they were small and numerous in highly transcriptionally active nucleoli, such as in proliferating cells, whereas only a solitary large agglomerate was present in transcriptionally inactive nucleoli such as in resting lymphocytes (Derenzini 1995; Derenzini et al. 1982b, 1983b,c,1984,1990). In the Feulgen-like osmium ammine-stained sections, the very high specificity of the reaction does not allow us to define the relationship among these chromatin structures and the other components of the nucleolus, which are in fact completely unstained. This drawback was, however, overwhelmed either by combining the selective DNA staining with the silver staining for the NOR proteins or by studying the distribution of the selectively stained DNA structures in nucleoli with a very simple organization such as that of ring-shaped nucleoli. In the first case a constant overlapping of the silver deposits was observed only on the loose agglomerates of extended DNA filaments, whereas no association was found with the clumps and the 11-to 30-nm-thick fibers of intranucleolar chromatin (Derenzini et al 1981a, 1990; Hernandez-Verdun et al. 1982, 1984). The colocalization of the silver deposits and DNA filaments indicated that the latter are mainly located in the FCs. The fact that the FCs were the site of major accumulation of the 2- to 3-nm-thick DNA filaments was also indicated by studies on the distribution of the intranucleolar chromatin structures in lymphocyte nucleoli (Derenzini et al. 1983c, 1990). In human resting lymphocytes the nucleolus is very simply organized. After routine uranium and lead staining, a solitary, large, roundish FC appears to be present in the middle of the nucleolar area, encircled by a rim of ribonucleoprotein components (Figure 5). In sections selectively stained for DNA, small, highly compact clumps of chromatin are visible within the nucleolar area. A large, solitary, roundish, lightly electron-opaque agglomerate of very loosened DNA structures is always present in the central portion of the nucleolus (Figure 5). By its size, shape, and localization, this agglomerate corresponds perfectly to the large FC as visualized in nucleoli either stained with uranium and lead salts or silver stained for AgNOR proteins (Figures 5A and 5B). At higher magnification, the loosened DNA structures appear to be constituted of short contorted DNA filaments, 2- to 3-nm thick, which are variously oriented within the space and frequently arranged as loops and open circles (arrows) (Figure 6). The presence of very loosened DNA structures in the middle portion of resting lymphocytes can also be detected by using the DNA staining procedure proposed by Testillano et al. (1991). Although this method does not allow us to clearly visualize the substructural organization of chromatin, which appears in fact to be homogeneously compacted, it nevertheless demonstrates the presence of extended DNA structures in the nucleolar area occupied by the FC (Figure 7). The structural organization of the DNA filaments and their relationship with the intra- and perinucleolar chromatin can be better investigated both using thicker sections (∼100 nm) than usual from Lowicryl-embedded samples or high magnification stereo-pair pictures taken from specimens at differently tilted angles (Derenzini et al. 1983a, 1987b; Derenzini 1995). In Figure 8, a Lowicryl section from a resting lymphocyte is shown in which the DNA filaments appear to be uniformly distributed within the agglomerate and to arise from the compact masses of peri- and intranucleolar chromatin. Three-dimensional observation of paper-printed stereopair pictures is not easy: the reader does not always own the necessary optical device, the distance between the two pictures is not always optimal for the stereological analysis and, of particular importance for high magnification pictures, the optical device visualizing the texture of the print paper makes it difficult to detect very thin structures. For these reasons, we reported in Figure 9 one picture resulting from the computer-assisted superimposition of two pictures taken at differently tilted angles. In this case the DNA filaments can be followed for very long tracts throughout the loose agglomerate. These filaments, which never gave rise to nucleosome-like structures, are in continuity with the compact clumps of chromatin bordering on the FCs. The transition between clumps and filaments was always sharp.
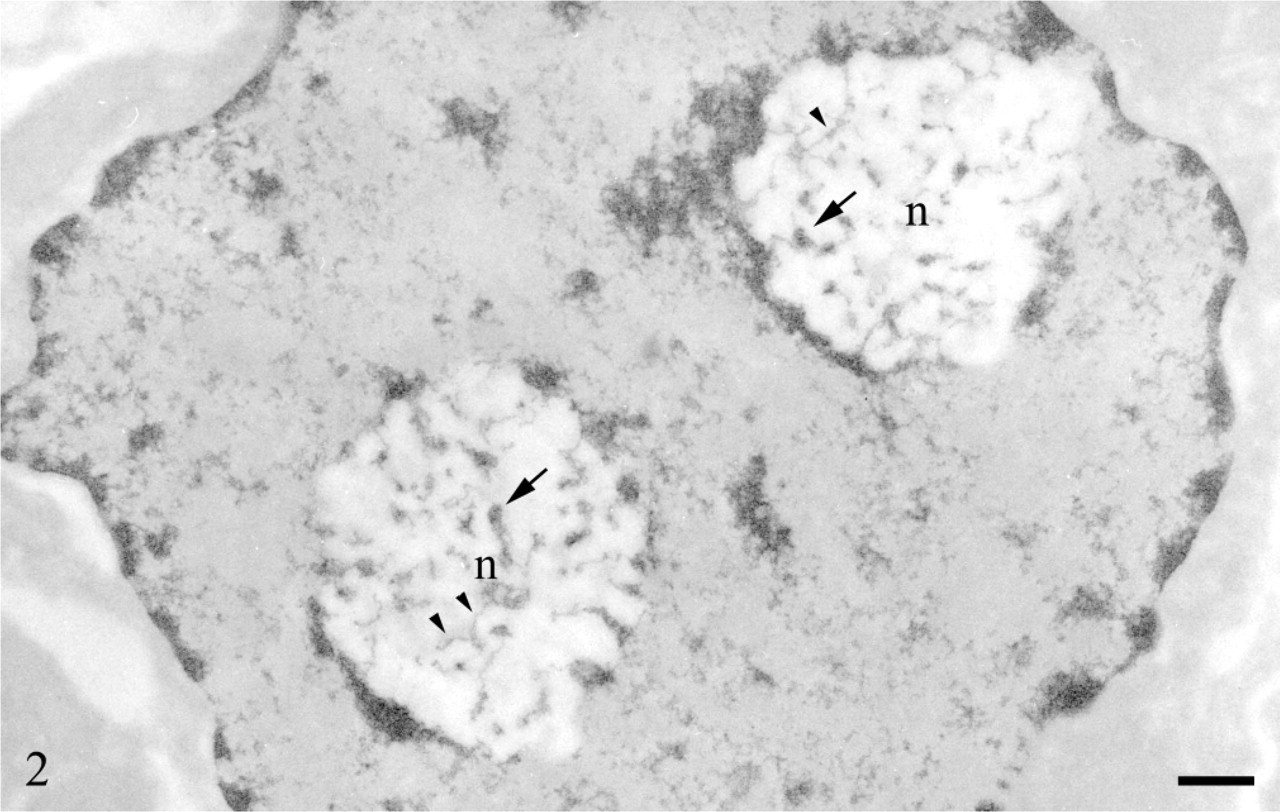
Regenerating rat hepatocytes, 24 hr after partial hepatectomy. Feulgen-like osmium ammine reaction. Only the DNA-containing structures are rendered electron opaque. In the two nucleolar bodies (n), which look more electron-translucent than the nucleoplasm, many DNA-containing structures are visible. They appear as small compact clumps (arrows) and elongated fibers (arrowheads) with variable thickness (intranucleolar chromatin). The nucleolar bodies are surrounded by highly compact DNA structures (perinucleolar chromatin). Bar = 0.5 μm. With permission, Biggiogera et al. Biol Cell 87:121–132, 1996.
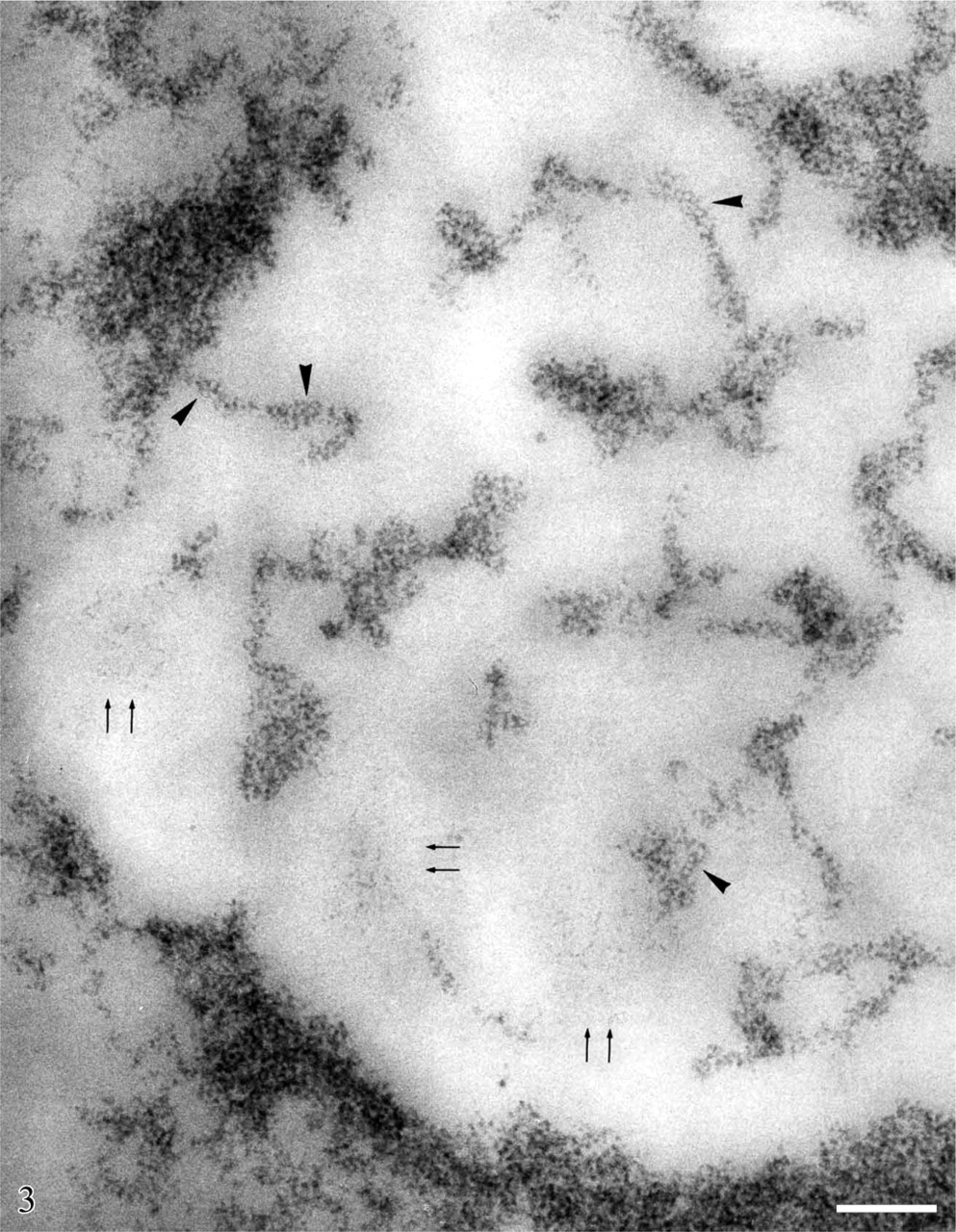
Regenerating rat hepatocytes, 24 hr after partial hepatectomy. Feulgen-like osmium ammine reaction. Portion of a nucleolar area showing the structural organization of the intranucleolar chromatin clumps and fibers. They appear to be constituted by roundish subunits with a diameter of ∼11 nm, made up of thin (2- to 3-nm thick) DNA filaments encircling an unstained inner core (arrowheads). These subunits represent the ultrastructural pattern of the nucleosomes in situ, when only DNA is stained. Among the chromatin clumps and fibers, small agglomerates of very thin (2- to 3-nm thick), extended DNA filaments are present (arrows) which never show the subunit organization of the compact chromatin clumps and fibers. Bar = 0.1 μm.
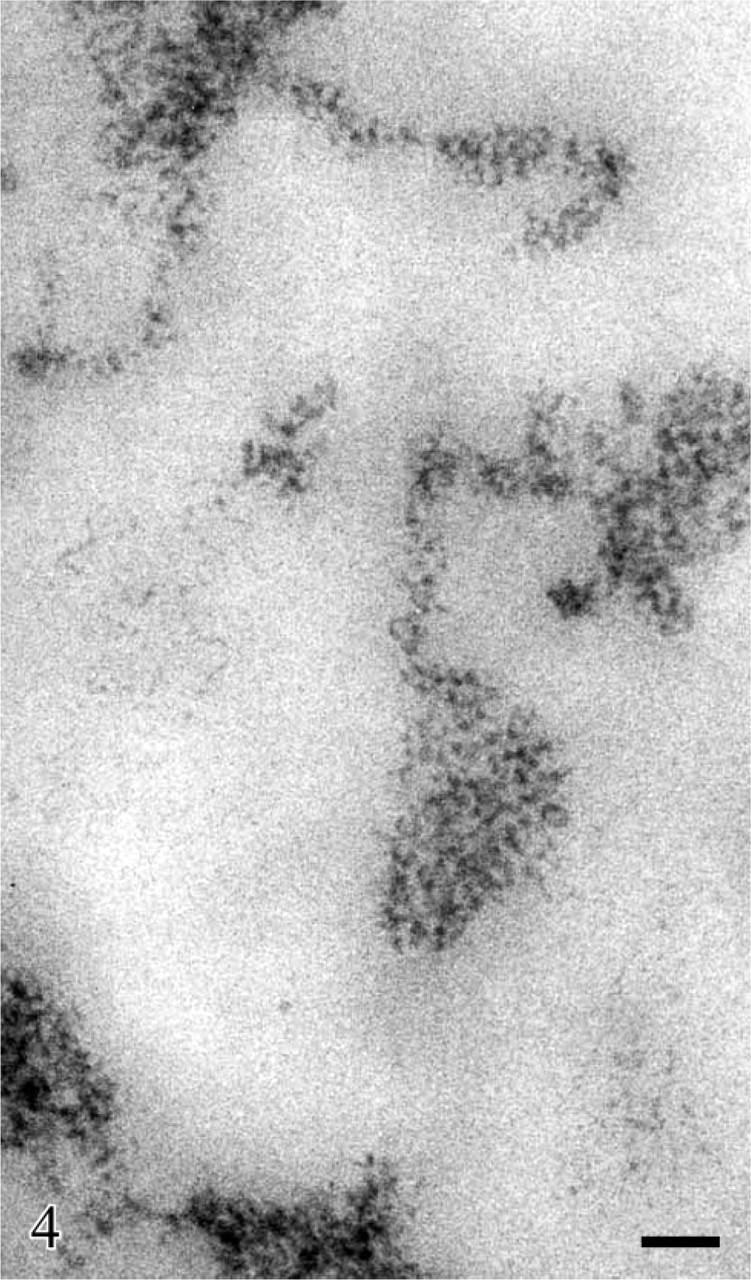
Detail of Figure 3 shown at higher magnification to appreciate the particulate organization of the chromatin clumps and fibers and the extended filaments in the loose DNA agglomerates. Bar = 0.05 μm.
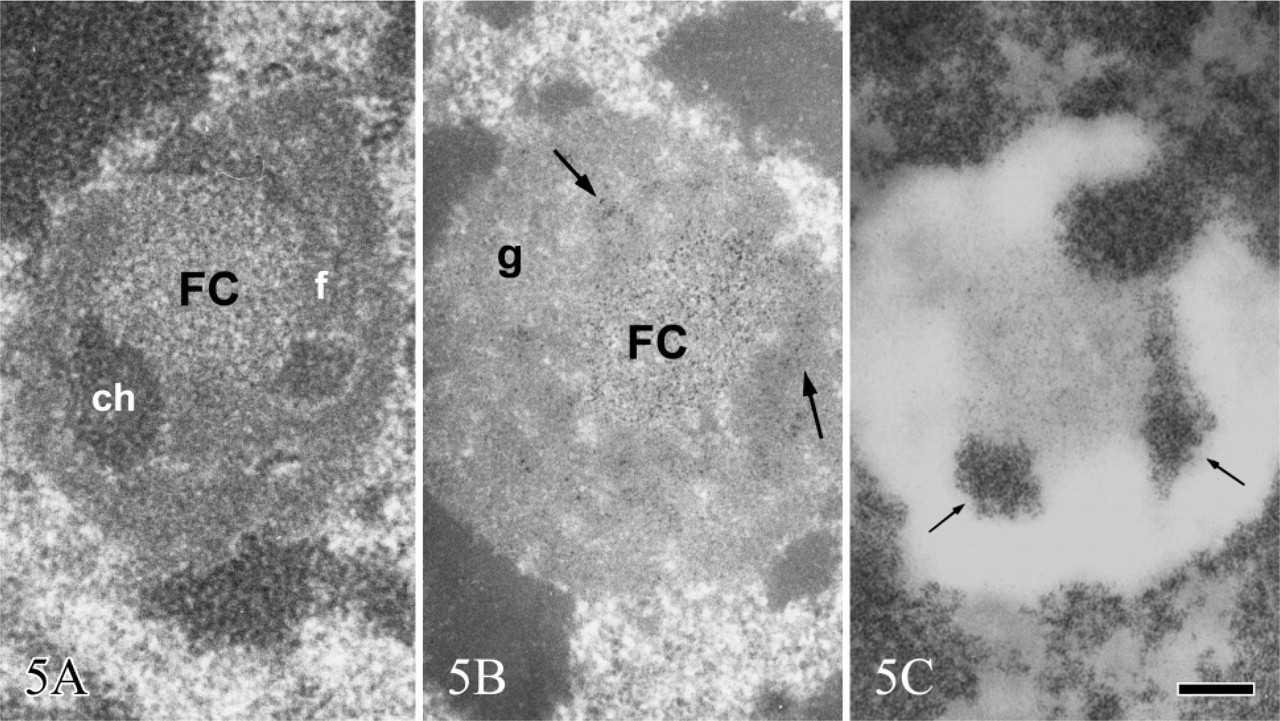
Human circulating lymphocytes. (
The presence of the extended DNA filaments in the FCs also fits with the fact that the DNA of the FCs is particularly vulnerable to enzymatic attack. On ultra-thin sections, initiation of transcription by Escherichia coli RNA polymerase appears to occur preferentially in the FCs (Thiry and Goessens 1991). DNase-I-sensitive sequences have also been predominantly detected in the FCs of Ehrlich tumor cell nucleoli (Thiry 1991).
In conclusion, these in situ data clearly indicated that the rDNA is in the form of extended and contorted DNA filaments, 2- to 3-nm thick, in the FCs, and presents a nucleosomal organization in peri- and intranucleolar chromatin.
Identification of Nucleolar Sites Where prerRNA Synthesis Occurs
Identification of the FCs as the nucleolar sites where ribosomal genes are located together with the enzymes necessary for ribosomal transcription (see above) led us to re-visit the established notion, mainly derived from early autoradiographic studies (reviewed by Fakan and Puvion 1980), that prerRNA synthesis occurred in the dense fibrillar component. Scheer and Rose (1984) first questioned this notion indicating the FCs as the site of rRNA transcription on the basis of the colocalization of ribosomal genes and RNA-polymerase I molecules in these nucleolar components. Since then, many studies have been carried out during the past 20 years with the aim to precisely localize active ribosomal genes within the nucleolar fibrillar components. At the present time, no univocal answer has yet been given to this problem, despite the use of the same techniques and, frequently, also of the same kind of cells by different research groups. Accordingly, actively transcribing ribosomal genes have been localized either in the FCs, or in the FCs together with the FC/DFC border zone, or in the DFC together with the DFC/FC border zone (Huang 2002). It is beyond the scope of the present review to critically discuss the available results on the precise localization of rRNA transcription in the nucleolar fibrillar components. Nevertheless, we think it convenient to focus on some data on which an agreement has been reached, as well as some new findings obtained by high-resolution tomographic analysis, which are useful to define the structural–functional organization of the ribosome genes in situ. Among the numerous technical approaches used to detect the sites of rRNA transcription at EM level in situ, the non-isotopic method for labeling the rRNA transcripts was considered by the different research groups to represent the most suitable procedure to localize actively transcribing ribosomal genes within the nucleolar component. The method consists of a short incubation of permeabilized cells with bromouridine-5′-triphosphate (BrUTP) followed by fixation, embedding, and revelation of the sites containing nascent Br-RNA by the immunogold procedure in thin sections (Hozak et al. 1994). By applying this technique some authors mapped the nascent rRNA transcripts on the DFC and, to a lesser extent, on the DFC/FC junctional zone. The DFC was therefore considered to represent the site where the steady-state Christmas trees were located, not excluding that ribosome genes with few prerRNA transcripts might also be present in the cortical zone of the FCs (reviewed in Raska 2003). Considering that the RNA polymerase I takes approximately only 140 sec to transcribe one rRNA gene with an elongation rate of 95 nucleotides per second (Dundr et al. 2002; Cheutin et al. 2004), whereas rRNA transcripts rapidly enter the processing sites, other authors used isolated nucleoli for a very short BrUTP labeling time (up to 1 min) to obtain precise information on the sites of rRNA transcription. They found that nascent transcripts were mainly located in the cortex of FCs and, to a lesser extent, in the DFC (Cheutin et al. 2002). In the same paper, the spatial arrangement of RNA polymerase I molecules within the whole volume of the FCs was investigated by electron tomography carried out on thick sections of cells labeled with anti-RNA polymerase I antibodies prior to embedding. Tomographic analysis showed that RNA polymerase I molecules were located in the cortex of FCs where they were grouped as several coils, 60 nm in diameter. Together these observations allowed the authors to propose a model in which actively transcribing ribosome genes are located within the cortex of the FCs (Cheutin et al. 2002; Ploton et al. 2004). Elongating rRNA transcripts with their extremity near the RNA polymerase I molecules are present in the cortex of the FCs, whereas with the other extremity, transcripts are present within the DFC (Figure 10). This model fits nicely with the finding that the DNA filaments observed in the FCs after specific osmium ammine staining were frequently organized as loops and open circles and that the 5′ external transcribed spacer rRNA—a selective marker of nascent transcripts—was located both at the periphery of the FCs and in the DFC (Puvion-Dutilleul et al. 1997).
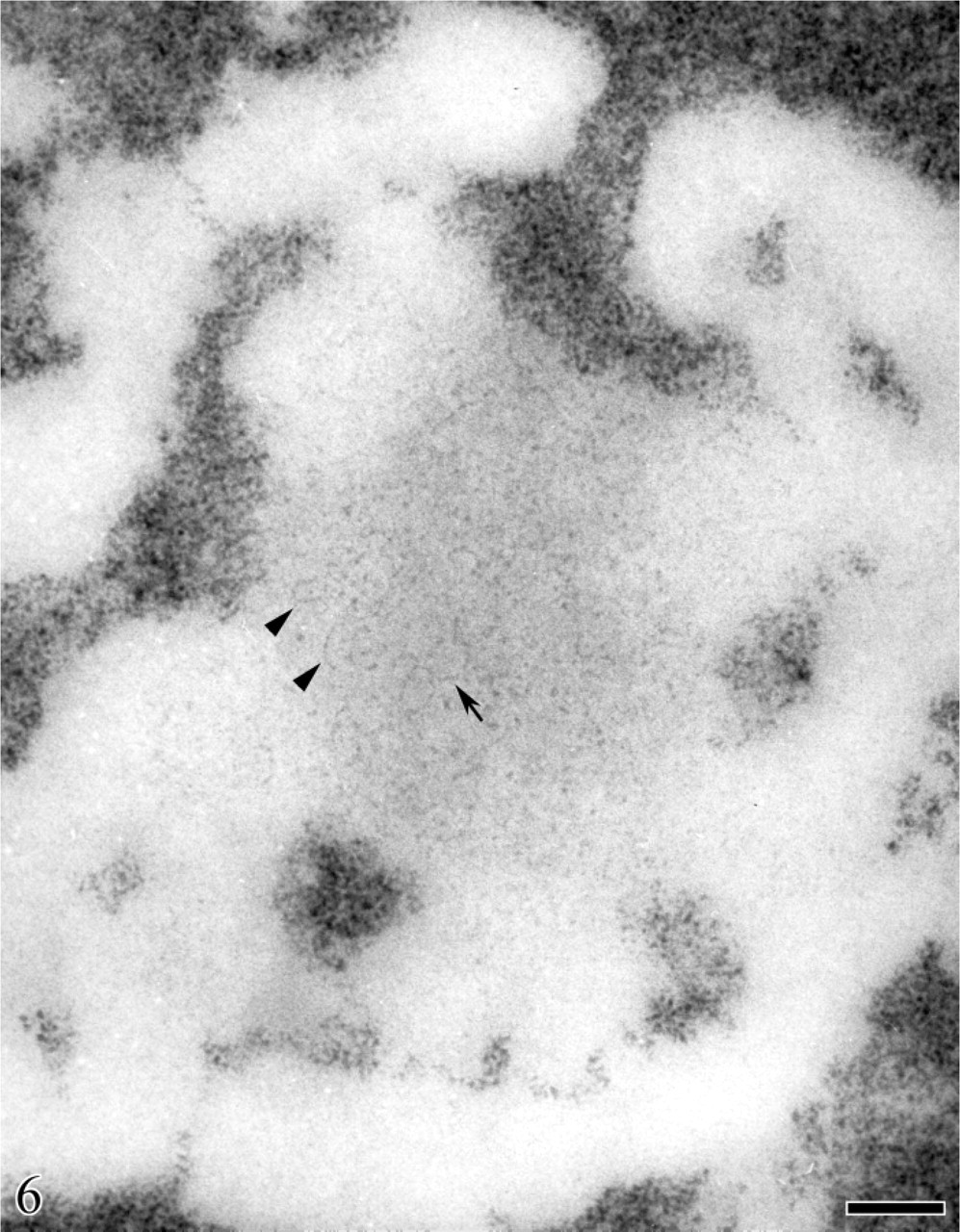
Human circulating lymphocyte. Feulgen-like osmium ammine reaction. High magnification of a nucleolar area showing a solitary, large agglomerate of extended DNA filaments. The DNA filaments are frequently arranged as loops (arrow) and open circles (arrowheads). Bar = 0.1 μm.
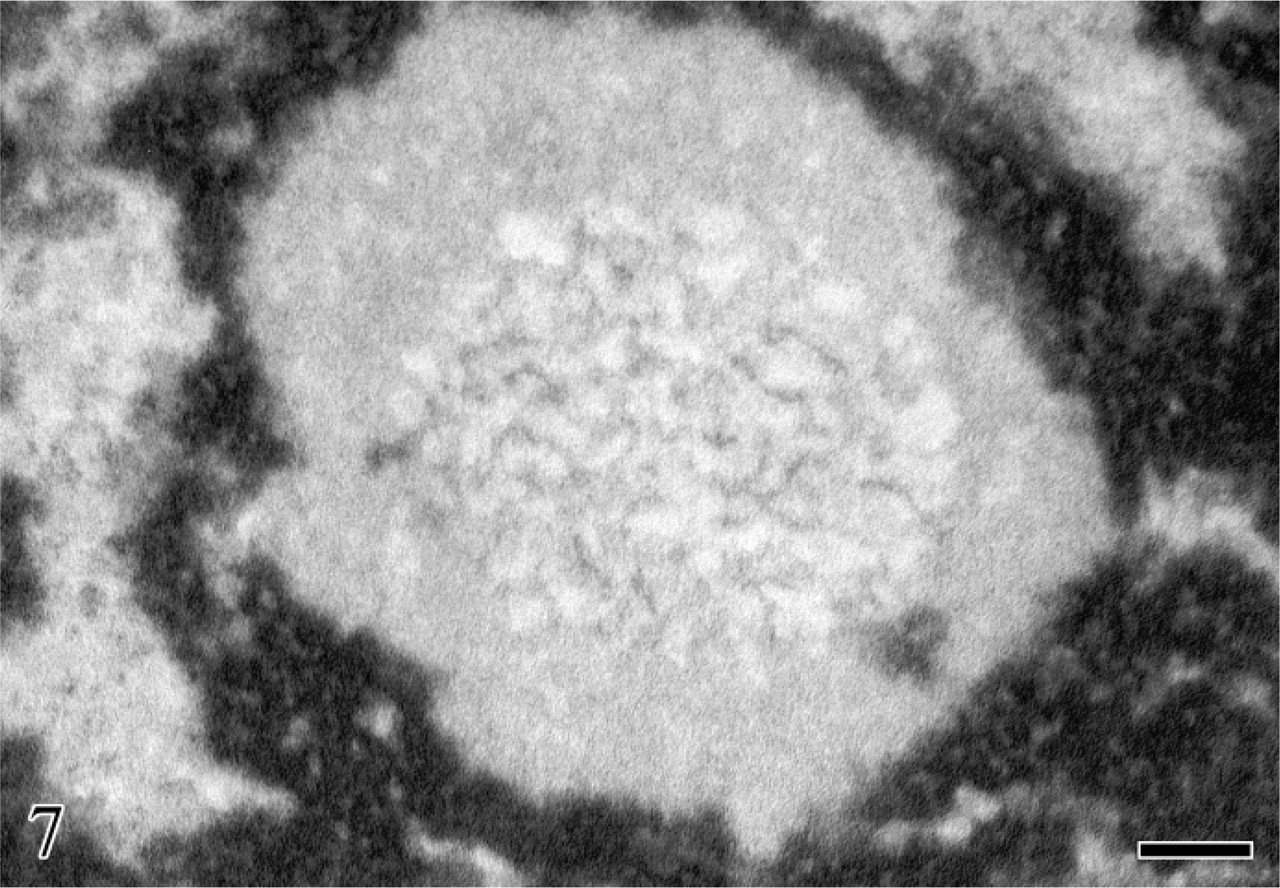
Human circulating lymphocyte. Glutaraldehyde fixation; en-bloc NAMA-Ur reaction for DNA staining. In the central zone of the nucleolar area numerous thin DNA-containing structures are visible. Bar = 0.1 μm.
In conclusion, even though a consensus for the precise location of transcribing ribosome genes in the fibrillar nucleolar components has not yet been reached, different research groups agree on the fact that the inner portion of the FC is not the site of rRNA synthesis. This is particularly evident in large FCs of transcriptionally silent cells, such as human circulating lymphocytes.
Structural–Functional Relationship of Ribosomal Genes In Situ
The observation that DNA in an extended non-nucleosomal configuration is present in the very small FCs of cells characterized by a very high rDNA transcription rate, such as regenerating rat hepatocytes, is consistent with the available evidence coming from both morphological analysis of spread chromatin preparations and DNase digestion experiments carried out on transcribing ribosome genes, which showed the absence of a nucleosomal organization of chromatin associated with rRNA transcripts (Scheer and Zentgraf 1982). In fact, in the case of very small FCs it is reasonable to assess that the few DNA filaments with a non-nucleosomal configuration present in this nucleolar component were all engaged in rRNA synthesis. On the other hand, this may not be the case for the extended DNA filaments present in the inner portion of the large FCs, especially if FCs of transcriptionally silent nucleoli were considered, where rRNA transcripts have never been found. One hypothesis is that they could correspond to intergenic spacers of transcribing rRNA genes. It is indeed well known that in mammalian cells the intergenic spacers are very long. In the mouse they represent about two thirds of 44 kb of rDNA repeats. However, by EM in situ hybridization, an intergenic spacer sequence upstream from the spacer promoter has been shown to be predominantly located outside the FCs in the chromatin clumps that interrupt the surrounding DFC (Thiry and Thiry-Blaise 1991). This DNA, unlike the DNA contained in the FCs, exhibits a nucleosomal configuration. This is in agreement with results observed in spread preparations of transcribing ribosomal repeats from mammalian cells showing that the intergenic spacer presents a nucleosomal organization (Puvion-Dutilleul et al. 1977; Puvion-Dutilleul and Bachellerie 1979). We can therefore exclude that the extended DNA present in the inner portion of the FCs may be constituted by the intergenic spacers of transcribing rRNA genes. A second hypothesis is that the extended DNA filaments would represent transcriptionally inactive rRNA genes that are, nevertheless, in a non-nucleosomal configuration. The presence of extended DNA filaments in the FCs independent of transcription is not surprising because a series of studies have shown that rDNA in an extended, non-nucleosomal structure is present in the FCs also when the rRNA gene transcription is blocked by drugs such as actinomycin D (Derenzini et al. 1983c; Hernandez-Verdun et al. 1984; Puvion-Dutilleul et al. 1992). Moreover, when RNA synthesis is stopped during mitosis, the DNA present in the metaphase NORs is always maintained in an extended non-nucleosomal configuration (Hernandez-Verdun and Derenzini 1983; Heliot et al. 1997).
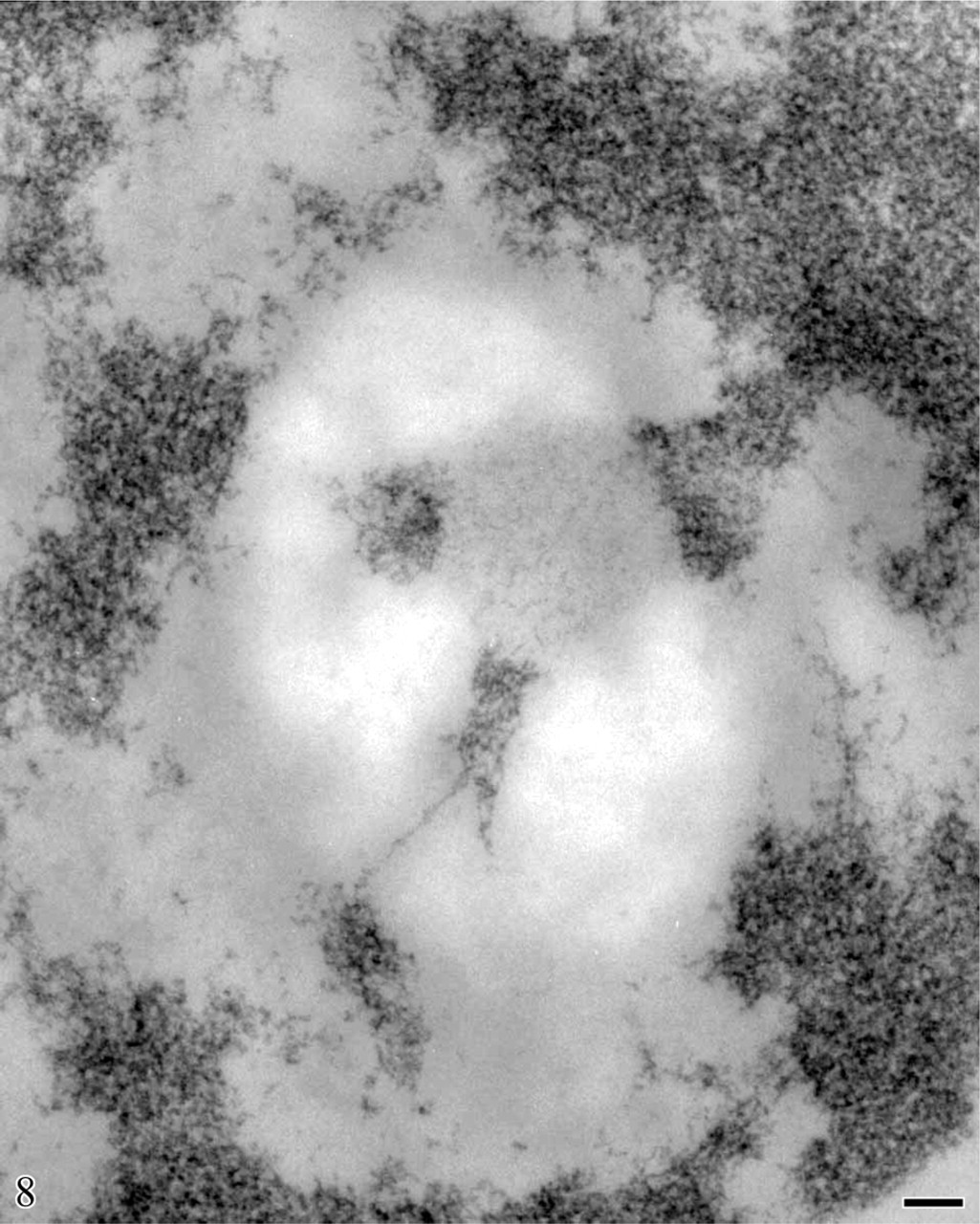
Human circulating lymphocyte. Feulgen-like osmium ammine reaction. Lowicryl embedding. Extended DNA filaments appear to be in continuity with the clumps of the compact intranucleolar and perinucleolar chromatin. Bar = 0.1 μm.
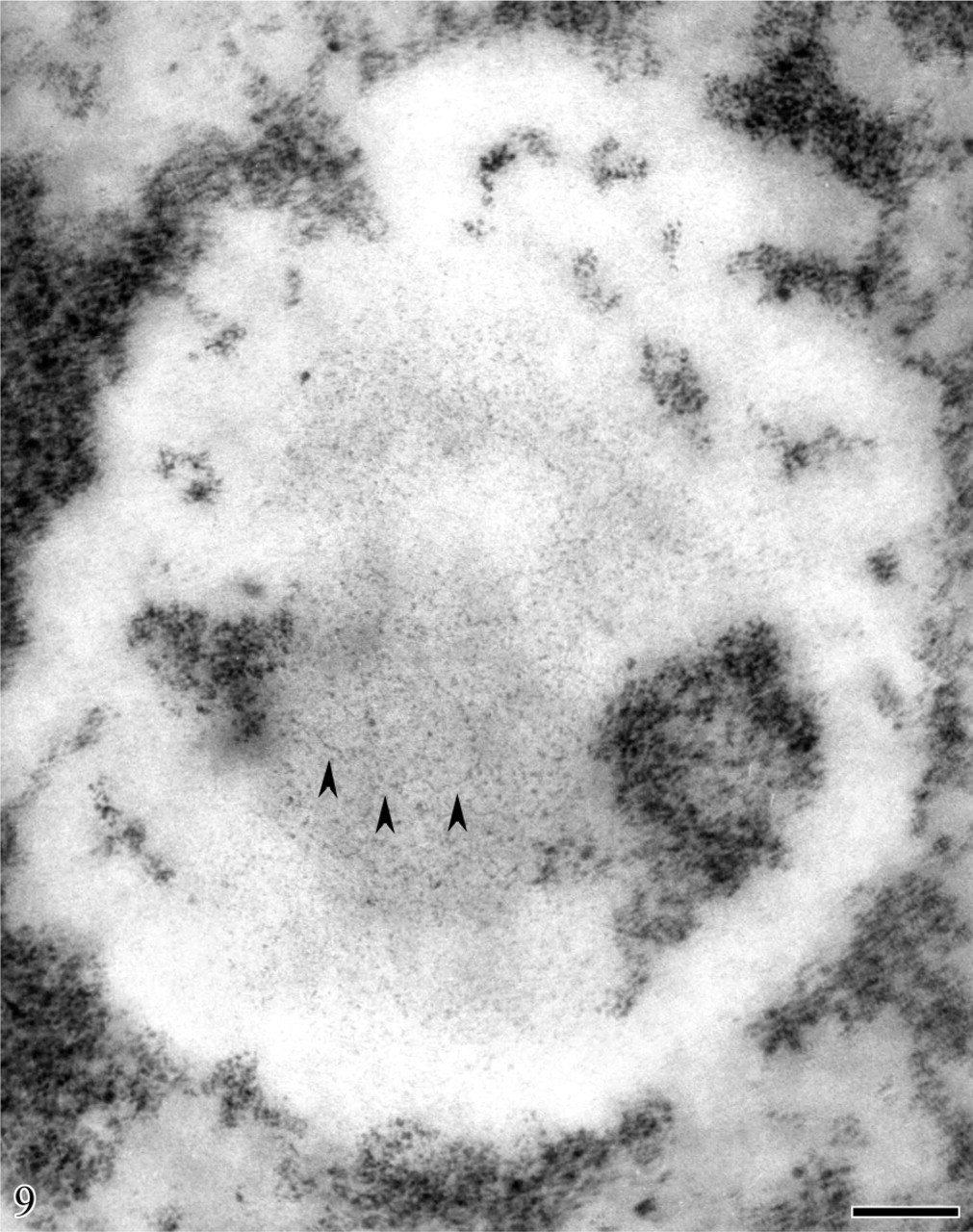
Human circulating lymphocyte. Feulgen-like osmium ammine reaction. Computer-assisted superimposition of stereo-pair micrographs taken with tilt angle ±5. Very long DNA filaments without nucleosome-like organization are visible (arrowheads). They appear to be in continuity with the intranucleolar chromatin clumps. Bar = 0.1 μm. Modified, with permission, Derenzini et al. Chromosoma 95:63–70, 1987.
The presence of transcriptionally inactive ribosome genes with a non-nucleosomal structure both in the FCs and metaphase NORs is consistent with results coming from the psoralen photo-crosslinking analysis of ribosome chromatin organization. By means of this assay, Conconi et al. (1989) showed that two distinct and coexisting types of ribosomal chromatin were present in Friend cells: one that contained nucleosomes and represented the inactive copies and one that lacked a repeating structure and corresponded to the transcribed genes. The relative amounts of the two types of ribosomal chromatin did not change either with variations of polymerase I transcriptional activity or during the cell cycle phases, thus indicating that the regulation of rRNA synthesis does not occur by changing the chromatin structure from a closed (nucleosomal) to an open (non-nucleosomal) configuration and vice versa, and that the portion of open chromatin can be both transcriptionally active and inactive. Regarding the extended, non-nucleosomal structure of the transcriptionally inactive rDNA present in the inner portion of the large FCs, this rDNA may constitute ribosome genes that are structurally ready for transcription. Stimulation of ribosome biogenesis could therefore occur by simple activation of RNA polymerase I transcription on ribosome genes that are already in an open, transcribable structure. Data on the changes of the extended rDNA distribution that occurred in resting human lymphocytes after phytohemagglutinin (PHA) stimulation indicated that this was in fact the case (Derenzini et al. 1983c, 1987a, 1993). A 24-hr treatment of lymphocytes with PHA induced an 8-fold increase of rRNA synthesis and a fragmentation of the solitary, large rDNA agglomerate of the resting lymphocytes into numerous rDNA agglomerates with a volume ∼15-fold smaller (Figure 11). These small agglomerates of extended DNA filaments corresponded to very small FCs surrounded by the DFC, thus indicating that the portion of the extended rDNA located in the inner part of the solitary FC of resting lymphocytes had become transcriptionally active. These structural changes that occur in large FCs of transcriptionally silent mammalian cells upon stimulation of rRNA synthesis are schematically illustrated in Figure 12.
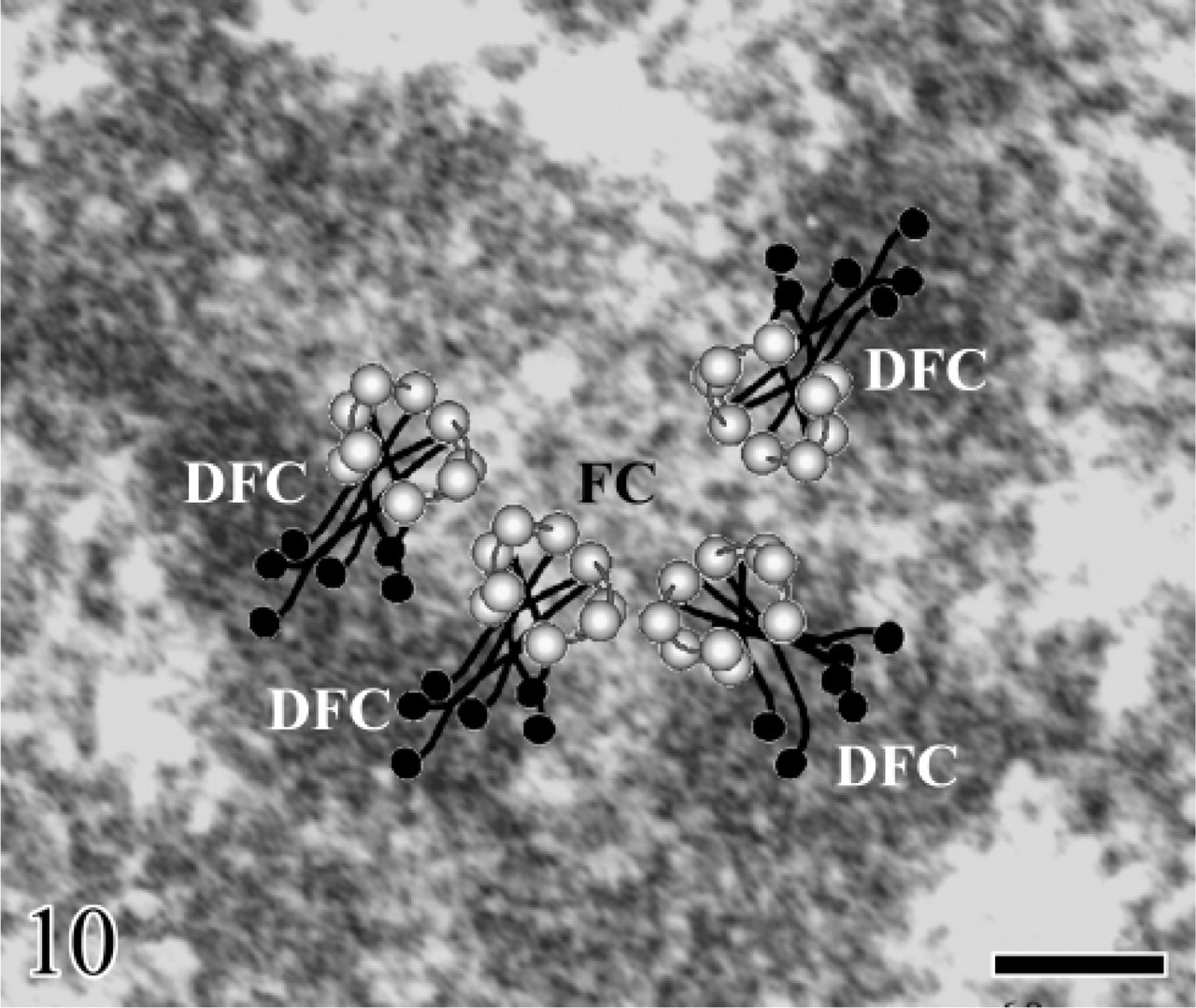

Image of one FC (300 nm in diameter), observed in a classically counterstained ultrathin section, was merged with a cross-section of the transcribed rDNA gene drawn to the same scale. Cross-sections of the rDNA gene, which is organized as four long coils, appear as four open rings localized within the cortex of FC. The open rings of rDNA are covered with several RNA polymerase I molecules (light circles) from which elongating rRNA molecules are emerging (black lines). The growing rRNA molecules, each ending in a terminal knob (black circles), enter the surrounding DFC. Bar = 60 nm. With permission, Cheutin et al. J Cell Sci 115:3297–3307, 2002.
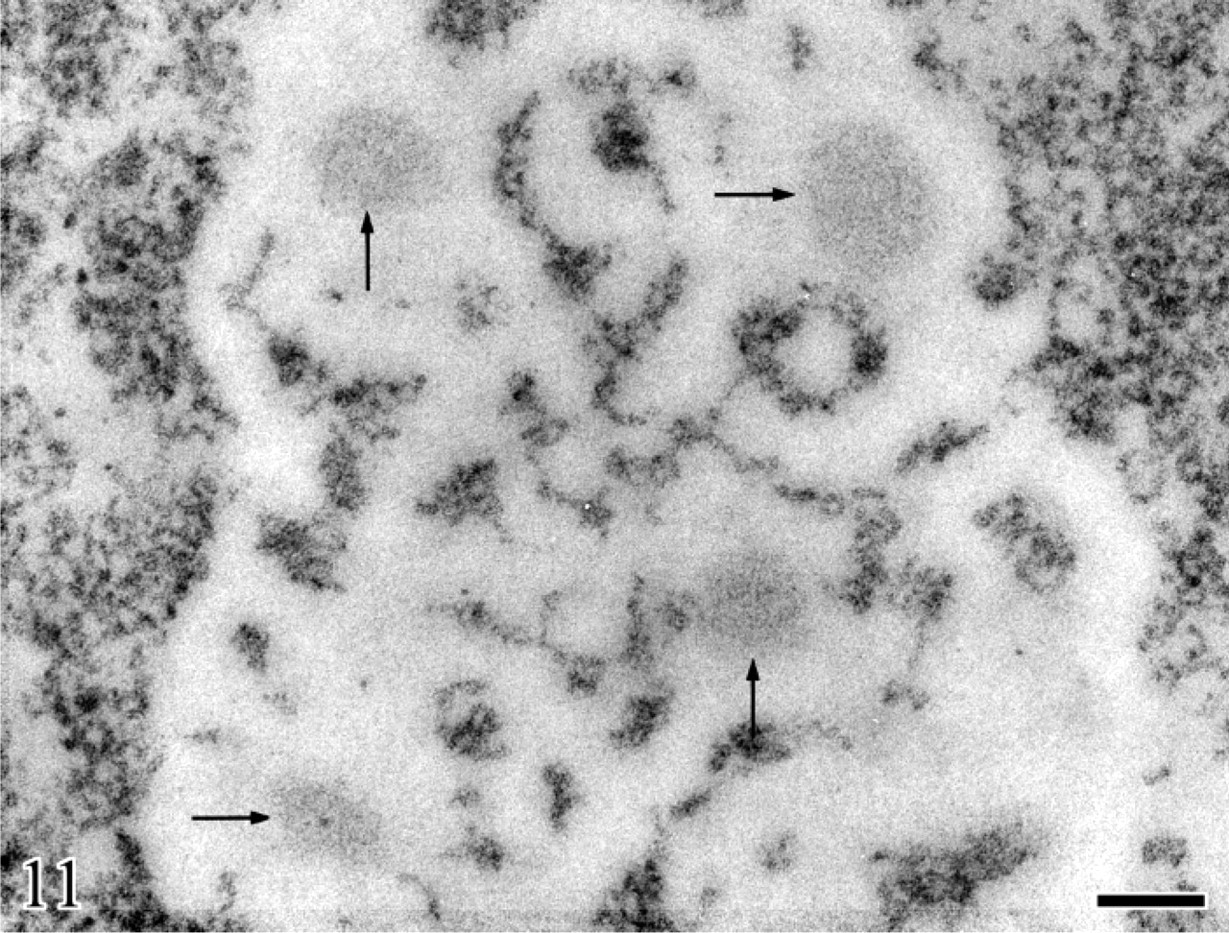
Human circulating lymphocyte, 48 hr after phytohemagglutinin stimulation. Feulgen-like osmium ammine reaction. In the enlarged nucleolar area, four well-separated loose agglomerates of DNA filaments are visible (arrows). They are smaller in size than the solitary agglomerate present in the nucleolus of resting lymphocytes. Bar = 0.2 μm. With permission, Derenzini et al. Exp Cell Res 145:127–143, 1983.
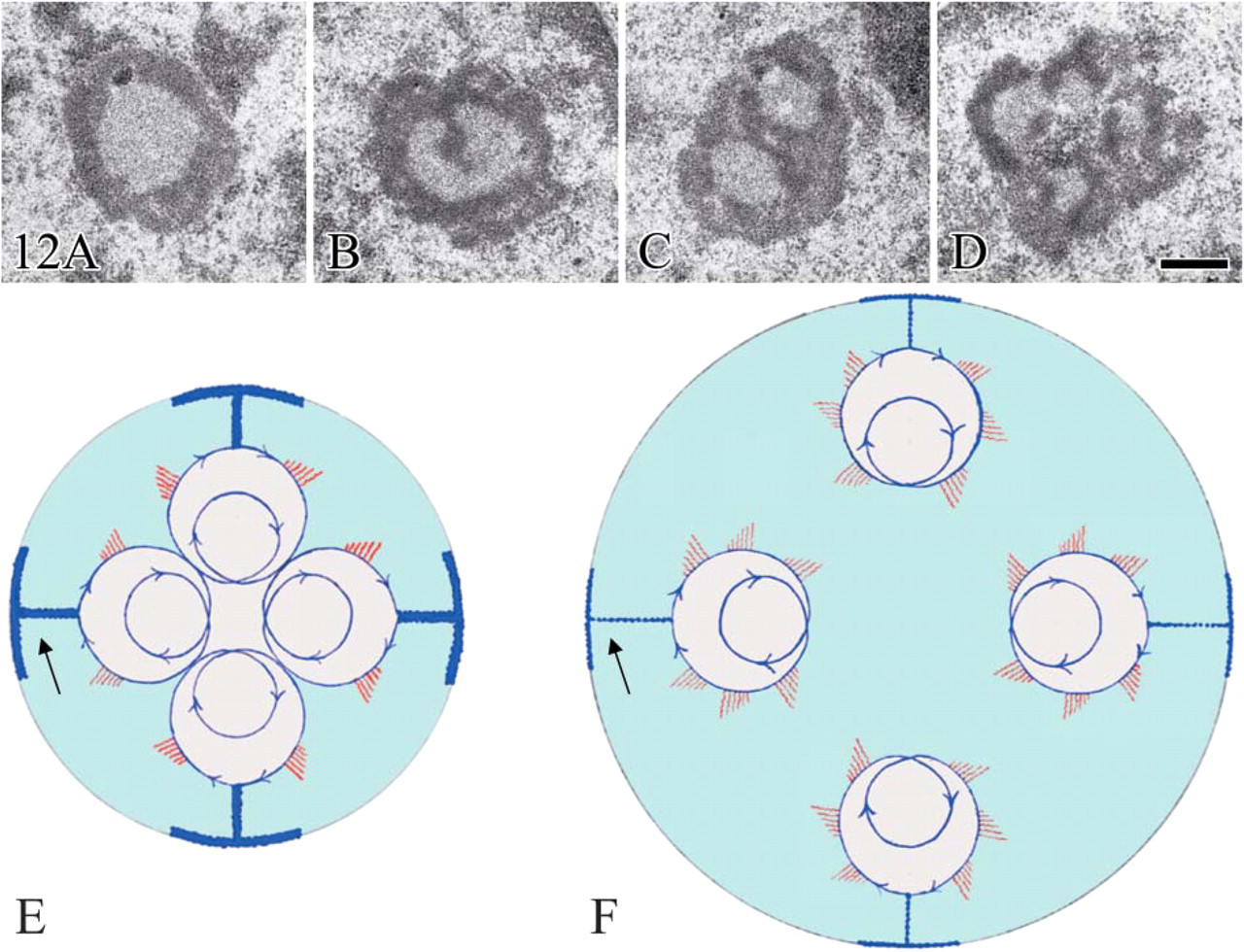
Structural changes occurring in human lymphocyte nucleoli after stimulation of rDNA transcription by 20 hr phytohemagglutinin treatment. Aldehyde osmium fixation, uranium and lead staining. (
Conclusions
A peculiar type of chromatin organization appears to emerge in the mammalian cell nucleolus. Results presented in the current review indicated that ribosomal genes located within the FCs are in an extended non-nucleosomal organization independent of their transcriptional activity. It is very interesting to note that FCs are only detected in the amniotes, i.e., they have been acquired recently during evolution (Thiry and Lafontaine 2005). The FCs contain all the molecules necessary for rRNA synthesis and, very likely, also for maintaining ribosomal chromatin in an extended configuration. Little is known about the mechanisms that establish the “open” chromatin state of rRNA genes, even if the action of an ATP-dependent chromatin remodeling complex has been postulated (Grummt and Pikaard 2003) as well as an ATPase activity actually located in the FCs of human fibroblasts (Fox et al. 1981). The FC could represent an evolution-linked structural–functional unit by which—by maintaining rRNA genes in a “ready to transcribe” state throughout the cell cycle—mammalian cells may develop a more rapid and appropriate adjustment of the ribosome production according to environmental conditions without being obliged to previously modify the chromatin structure. In this context, the importance of the FCs has been stressed by the observation that in cells lacking FCs, unlike in cells with FCs, natural or drug-induced rRNA synthesis inhibition induced changes of the extended rDNA filaments that are transformed into small clumps of condensed chromatin (Thiry et al. 1991; Mineur et al. 1998).
Footnotes
Acknowledgements
This work was supported by grants from Pallotti's Legacy for Cancer Research, Italian Ministry of Education (MIUR; grants for selected basic research topics) and University of Bologna (funds for selected research topics) (MD and GP); Ligue Régionale de la Marne, Ligue Régionale de l'Aube et Association pour la Recherche sur le Cancer, Grant #4497(M-FO'D and DP); and the National Funds for Scientific Research, Belgium (MT).
