Abstract
A satisfactory protocol of protein extraction has been established based on the heat-induced antigen retrieval (AR) technique widely applied in immunohistochemistry for archival formalin-fixed, paraffin-embedded (FFPE) tissue sections. Based on AR, an initial serial experiment to identify an optimal protocol of heat-induced protein extraction was carried out using FFPE mouse tissues. The optimal protocol for extraction of proteins was then performed on an archival FFPE tissue of human renal carcinoma. FFPE sections were boiled in a retrieval solution of Tris-HCl containing 2% SDS, followed by incubation. Fresh tissue taken from the same case of renal carcinoma was processed for extraction of proteins by a conventional method using radioimmunoprecipitation assay solution, to compare the efficiency of protein extraction from FFPE tissue sections with extraction from fresh tissue. As a control, further sections of the same FFPE sample were processed by the same procedure without heating treatment. Evaluation of the quality of protein extracted from FFPE tissue was done using gel electrophoresis and mass spectrometry, showing most identified proteins extracted from FFPE tissue sections were overlapped with those extracted from fresh tissue.
T
Based on the principle of heat-induced AR technique, Ikeda et al. (1998) documented a protocol of protein extraction from FFPE tissue by heating sections in radioimmunoprecipitation assay (RIPA) buffer containing 2% SDS. Although they could not achieve identical SDS-PAGE pattern between fresh and FFPE tissues, eight common proteins were readily demonstrated by Western blotting technique indicating that the protein molecules ranging from 10 to 120 kDa remained well preserved. Subsequently, Yamashita and Okada (2005) reported a protocol of protein extraction from FFPE tissue sections by autoclaving 4-μm sections mounted on slides for 15 min, and followed by incubating tissue sections with 2% SDS containing urea and 2-mercaptoethanol. They demonstrated satisfactory results by Western blotting for two proteins that could not be identified in tissue sections without heating treatment. Prieto et al. (2005) reported the use of a commercial Liquid Tissue kit (Expression Pathology, Inc.; Gaithersburg, MD) used for protein extraction from FFPE tissue by heating tissue sections at 95C for 90 min. They demonstrated a large number of observable peptides by using mass spectrometry. Crockett et al. (2005) identified 324 proteins by mass spectrometry for proteins extracted from an archival FFPE cell block using enzyme digestion method. Also, Chu et al. (2005) developed a method of nondestructive molecule extraction using the heat-induced AR, and achieved extraction of proteins from a single FFPE tissue section. Recently, we also have established a satisfactory protocol of protein extraction from archival FFPE tissue sections, adapting the heat-induced AR principle, that was used in our laboratory for improving the detection of protein antigens in tissue sections. By this method, the SDS-PAGE pattern of protein extracted from FFPE tissue sections is closely comparable with that of protein extracted from fresh tissue sections taken from the same specimen.
However, two-dimensional PAGE has significant limitations for detailed protein analysis, including deficiencies in proteome coverage, dynamic range, sensitivity, and throughput. As a result, great efforts have been devoted to the development of non-gel-based proteome technologies, particular those based on combination of chromatography and electrokinetic separation techniques. An integrated proteome concentration and separation approach, involving inline combination of capillary isoelectric focusing (CIEF) with capillary reversed-phase liquid chromatography has been developed for advanced mass spectrometry (Chen et al. 2003). The utility of this method when married with AR-derived protein extraction methods has been evaluated in the present study using archival FFPE tissue.
All FFPE tissues used in the following studies were fixed in 10% neutral buffered formalin at room temperature for 24 hr. Archival human tissue blocks were stored within 5 years. Early studies in our laboratory evaluated the protocol of Ikeda et al. (1998) to extract protein from FFPE tissue sections. After deparaffinization by Octane (Sigma-Aldrich Inc.; St. Louis, MO), a total of 15 FFPE human breast tissue sections (10 μm each) were incubated at 100C for 20 min in RIPA buffer solution containing exactly the same chemical components as documented, followed by incubation at 60C for 2 hr (Ikeda et al. 1998). The supernatant was transferred to a clean fresh bottle after centrifugation, and the total amount of extracted protein was measured as 1650.3 μg/ml by a Biophotometer (Eppendorf Scientific; Westbury, NY). SDS-PAGE showed faint but visible bands ranging from 7 to 55 kDa, and Western blotting using monoclonal antibody to β-actin demonstrated a clear band with molecular weight 42 kDa (data not shown). To improve further this protocol, a comparative study between fresh and FFPE mouse liver tissues was carried out and showed much weaker bands by SDS-PAGE for FFPE tissue than for corresponding fresh tissue, although the whole range of protein bands was still superficially comparable between both samples (data not shown).
The next step, therefore, was to apply our experience of ‘retrieval’ of protein antigens in tissue sections, as evidenced by IHC methods, in an attempt to achieve better results of protein extraction from FFPE tissue, based on the principle of the AR technique, including the heating condition, pH, and various chemical solutions (Shi et al. 1995). Several solutions with various pH values ranging from pH 2 to 9, with or without 2% SDS, were tested to compare the quantity and quality of proteins extracted from FFPE tissues of both human and murine origin. The major conclusions may be summarized as follows.
Boiling FFPE tissue sections at 100C in Tris-HCl buffer solutions of pH 2 to 9 containing 2% SDS showed similar SDS-PAGE patterns and actin-positive bands as demonstrated by Western blotting, with a weaker band seen at low pH (pH 2.6). It was noted that heating the FFPE tissues at 120C yielded a particularly poor result for low pH solution.
2% SDS is a critical chemical component in the retrieval solution used to immerse FFPE tissue sections for heating treatment. All retrieval solutions tested without 2% SDS showed poor results whatever the heating conditions (data not shown).
Chemical solutions of 0.5% citraconic anhydride or 0.1 M sodium hydroxide did not yield satisfactory results.
An optimal protocol of protein extraction was thus devised, based on these experiments, and was applied in the following study. Five FFPE tissue sections (10 μm each) were deparaffinized by adding 1 ml Octane (Sigma), vortexing for 10 sec, followed by adding 0.075 ml methanol and vortexing again. After centrifugation, the upper layer of Octane and methanol was removed, and the residuum dried in a hood for 2-3 min. Fifty μl of 20 mM Tris-HCl buffer (pH 7 or 9) containing 2% SDS was then added to the dewaxed FFPE tissues sections, followed by heating at 100C on a heat block (VWR Scientific Products; West Chester, PA) for 20 min, then incubation at 60C in a incubator (Robbins Scientific; Sunnyvale, CA) for 2 hr. Two μl of the sample was used to measure yields of protein extracted from FFPE and from control fresh tissues subjected to an identical procedure. Identical measured amounts of proteins from each sample were loaded for analysis by SDS-PAGE and Western blotting to compare the results accurately. Although there were variations of protein yields among various protocols, the range of total yields was comparable (average 10,022.6 μg/ml in contrast to the fresh control of 11,524 μg/ml). A parallel study using human tissue from a resected renal carcinoma was performed to test and verify this optimal protocol. The SDS-PAGE patterns of protein samples of FFPE tissue sections obtained by boiling in pH 7 or pH 9 solution containing SDS, plus incubating in the same retrieval solution without heating, compared with samples extracted from fresh tissue taken from same case are shown in Figure 1. It is apparent that the SDS-PAGE patterns of proteins from samples extracted from FFPE after heat-induced AR treatment are comparable with patterns observed on samples derived from fresh tissues (Figure 1). Western blotting showed clear band of ki-67 using monoclonal antibody MIB-1 (DAKO; Carpinteria, CA) indicating a molecular weight range of 395/345 kDa (data not shown). Thus encouraged, we evaluated further protein samples by the more sophisticated and sensitive method of mass spectrometry in collaboration with Calibrant Biosystems, as described in the following section. This study of human tissue specimens has been exempted under 45 CFR 46.101(b) and was approved by the Institutional Review Board (IRB #009071) at the University of Southern California. University Animal Care Committee approved protocols were used in all experiments.
Analysis of four protein samples by mass spectrometry
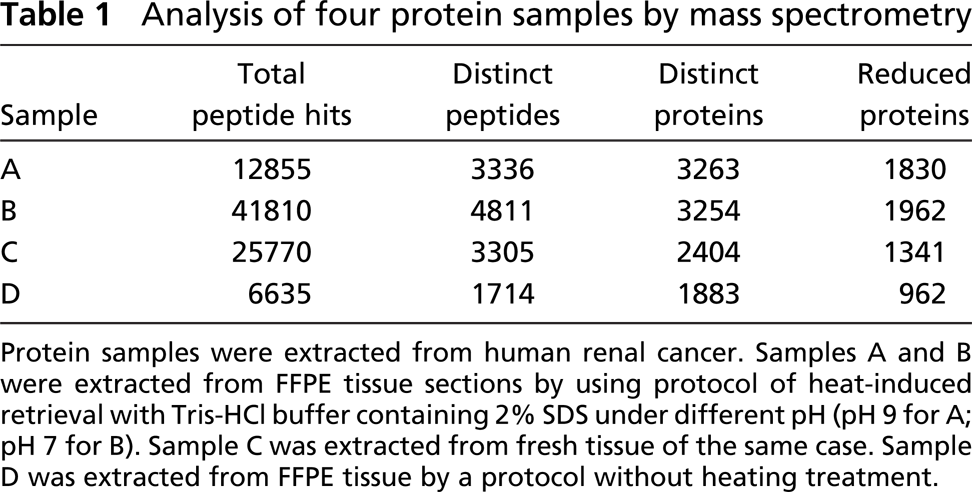
Protein samples were extracted from human renal cancer. Samples A and B were extracted from FFPE tissue sections by using protocol of heat-induced retrieval with Tris-HCl buffer containing 2% SDS under different pH (pH 9 for A; pH 7 for B). Sample C was extracted from fresh tissue of the same case. Sample D was extracted from FFPE tissue by a protocol without heating treatment.
Four samples of protein extracted from FFPE tissue of human renal cancer were analyzed by an online combination of CIEF with nano-reverse-phase liquid chromatography (RPLC) (Chen et al. 2003). Proteins were dialyzed overnight to remove residual SDS, denatured, digested, lyophilized and stored at−80C. Capillary isoelectric focusing was performed essentially as previously described, except an 84 cm long capillary was used and the peptide digest was loaded at a concentration of 1.67 mg/ml. Nano-RPLC of each CIEF fraction was performed using an Ultimate pump connected to a 15-cm pulled-tip fused silica capillary column packed with 3.5 μm Zorbax SB C18 particles (Ultimate-Dionex; San Jose, CA). Peptides were eluted using a 60-min linear gradient from 5% to 45% acetonitrile containing 0.02% heptafluorobutyric acid and 0.02% formic acid at a flow rate of 200 nL/min. The eluants were monitored using a LTQ (ThermoFinnigan; San Jose, CA) mass spectrometer, and mass spectra were acquired from 400 to 1400 m/z, followed by five data-dependent MS/MS scans with dynamic exclusion active for 3 min.
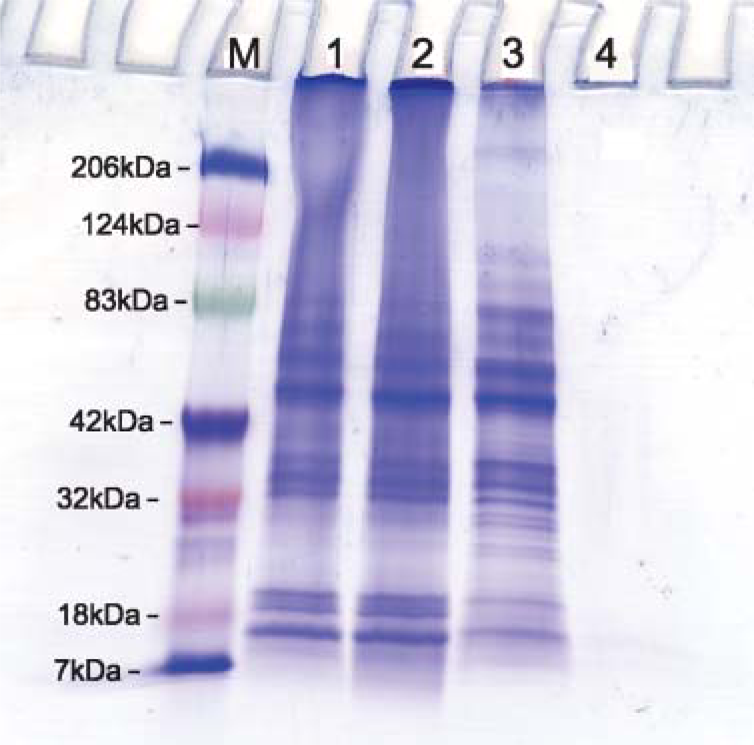
SDS-PAGE of total proteins extracted from FFPE tissue sections (Lanes 1, 2, and 4), and comparable frozen tissue sections (Lane 3) of renal carcinoma showing comparable bands among three lanes. (Lane 1) Protein extracted from FFPE tissue using heat-induced AR protocol by Tris-HCl + 2% SDS at pH 9. (Lane 2) Using the same AR solution at pH 7. (Lane 3) Protein extracted from fresh tissue. (Lane 4) Protein extracted from FFPE tissue without AR.
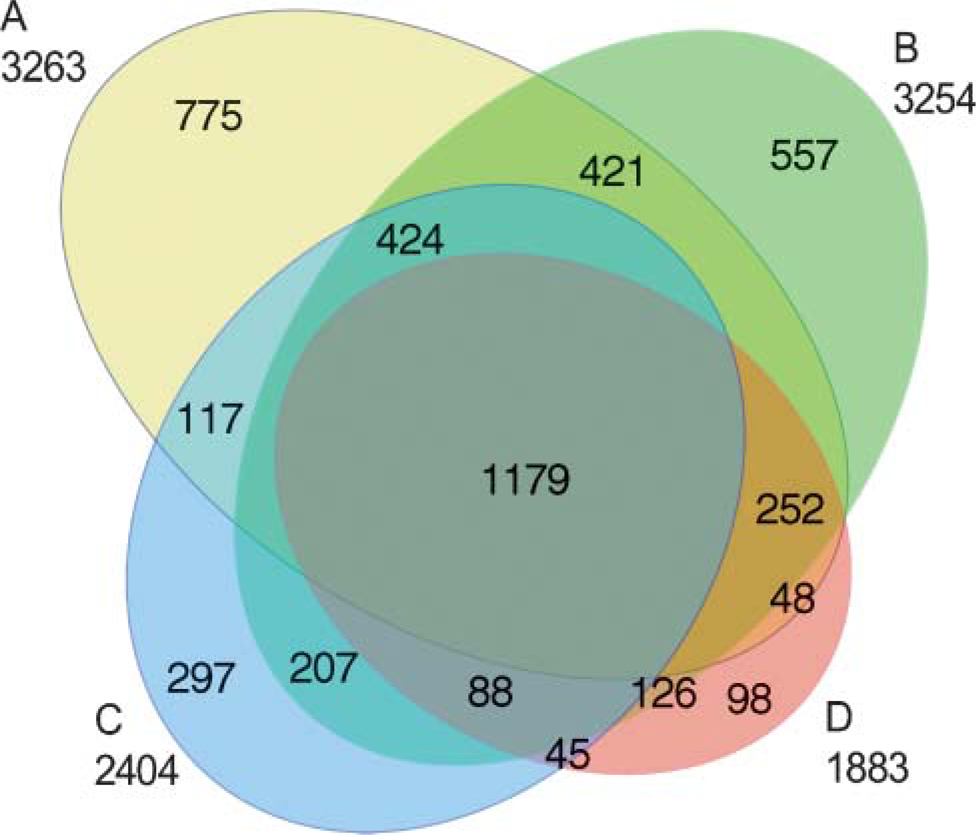
Diagram showing protein overlap of four samples as listed in the Table 1. Total number of distinct protein entries mapped by CIEF/LC/MS/MS. Total proteins are indicated for each sample below the label; subsets are indicated within the diagram. It indicates that most proteins extracted from FFPE tissue sections are overlapped with that extracted from the fresh tissue sections, although a small part of proteins that are not overlapped well among the four samples that need to be addressed in further studies.
The resulting MS data files were peak listed by ltq_dta.exe (ThermoFinnigan). Peptide and protein identifications were made using Open Mass Spectometry Search Algorithm (OMSSA) (National Center for Biotechnology Information, Bethesda, MD). Protein identifications are based on peptide identifications in the human International Protein Index (2.28) distributed and maintained by the European Bioinformatics Institute (http://www.ebi.ac.uk/IPI/IPIhuman.html). Peptide hits indicates the number of ions that met or exceeded the e-value cutoff of 0.05 set in OMSSA. Distinct peptides indicate the number of distinct peptide sequences identified. Distinct proteins indicate the number of distinct protein sequence entries in the IPI database which were mapped by one or more distinct peptide sequences. Reduced proteins indicate the number of distinct IPI protein sequence entries mapped by the number of peptide sequences unique to the database plus the number of identical protein clusters that contain identical distinct peptide mappings.
Total proteins of the four samples demonstrated by MS are listed in Table 1. It was surprising to see a large number of protein sequences clearly displayed after heat-induced AR, comparing favorably with the extract from fresh tissue. Also, most proteins of the four samples showed overlap (Figure 2), which indicates reproducible total proteins existed in the same tissue specimen. Potential reasons why fresh tissue yielded a lower number of total proteins may be the period of storage of this fresh tissue, diffusion and loss of protein, or denaturation during periods of warming (as on sectioning). A noticeable discrepancy of the quality of protein sample extracted from FFPE tissue by a protocol without heating treatment (Sample D) was found between SDS-PAGE (Figure 1) and MS (Table 1). Although SDS-PAGE showed a poor result of total proteins for this sample, a total of 962 proteins could be demonstrated by MS. In addition to the highly sensitive MS technique, protein digestion by trypsin and other reagents before MS analysis may contribute to improvement of protein quality as well.
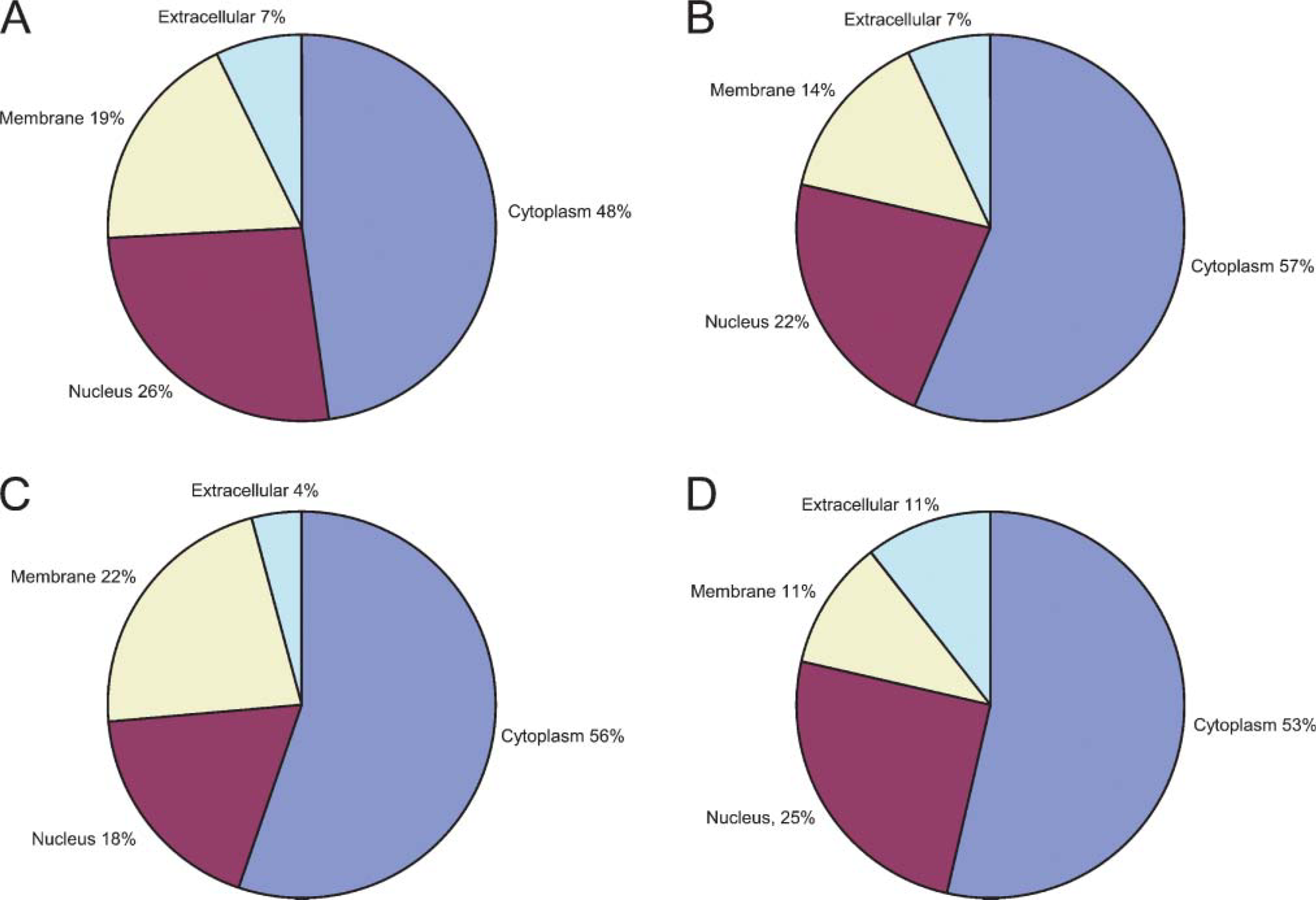
Diagram showing the gene ontology-predicted subcellular localization of proteins found in the four samples as listed in Table 1. Samples
The subcellular location of the identified proteins was assigned by the Gene Ontology database. More than half of the identified proteins have not been assigned subcellular locations by this database. The plots show the distributions of identified proteins with predicted locations over the total number of identified proteins with known locations. The plot distributions are in good agreement and show little variance (Figure 3).
Preliminary feasibility for extraction of nucleic acids and proteins from FFPE tissues based on the heat-induced AR principle has been documented, but numerous issues remain to be addressed to develop better techniques that enhance the quality and quantity of macromolecules extracted from FFPE tissues. Sophisticated molecular techniques, as exemplified herein by mass spectrometry, are able to be performed on protein extracted from FFPE tissue using the heating protocol described here, with the notation that variation among the four samples in terms of quality and yield require further study. However, this is characteristic of the random sampling nature of data-dependent experiments using MS. The apparent lower yield in frozen tissue may be addressed by use of a fresh prepared cell line sample compared with a FFPE aliquot. The presence of SDS was critical, and, with SDS, several AR protocols that had proved to be satisfactory for IHC on FFPE tissue sections also were satisfactory for protein extraction from FFPE tissue. Several strategies are available for future studies aimed at adapting AR-IHC protocols to AR-based protein extraction from FFPE tissues. It may be proposed that a two-phase approach be applied to protein extraction from the FFPE tissue: the first phase would be ‘antigen retrieval’ by high temperature heating to break down formalin-induced crosslinking (Rait et al. 2004; Sompuram et al. 2004; Yamashita and Okada 2005); the second phase would be the process of extraction of the heat-retrieved proteins from FFPE tissue, which may include solubilization or stabilization of extracted proteins (Chu et al. 2005). Further studies will be necessary to analyze the various factors that influence the efficiency of protein extraction from FFPE tissues. Such studies may have value not only in establishing better protocols for protein extraction, but also for understanding the mechanism of the heat-induced AR technique as applied to IHC.
Footnotes
Acknowledgements
This study was supported by National Institutes of Health Grant 1 R33 CA-103455-01. We greatly appreciate Puya Yazdi, a medical student at USC, and Hao Hao Huang, a student at UC Berkeley, for their efforts in doing some technical work for extraction and evaluation of protein from FFPE tissue sections.
