Abstract
During the course of diagnostic surgical pathology, pathologists have established a large collection of formalin-fixed, paraffin-embedded tissues that form invaluable resources for translational studies of cancer and a variety of other diseases. Accessibility of macromolecules in the fixed tissue specimens is a critical issue as exemplified by heat-induced antigen retrieval (AR) immunohistochemical (IHC) staining. On the basis of observations that heating may also enhance in situ hybridization (ISH) and the similarity of formalin-induced chemical modifications that occur in protein and in DNA, we designed a study to examine the efficiency of DNA extraction from archival formalin-fixed, paraffin-embedded tissues using an adaptation of the basic principles of the AR technique, i.e., heating the tissue under the influence of different pH values. Archival paraffin blocks of lymph nodes, tonsil, and colon were randomly selected. Each paraffin block was prepared in 34 microtubes. For each paraffin block, one tube was used as a control sample, using a non-heating DNA extraction protocol. The other 33 tubes were tested using a heating protocol under 11 variable pH values (pH 2 to 12) under three different heating conditions (80, 100, and 120C). Evaluation of the results of DNA extraction was carried out by measuring yields by photometry and PCR amplification, as well as kinetic thermocycling (KTC)-PCR methods. In general, lower pH (acid) solutions gave inferior results to solutions at higher pH (alkaline). Heating tissues at a higher temperature and at pH 6–9 gave higher yields of DNA. There appeared to be a peak in terms of highest efficiency of extracted DNA at around pH 9. The average ratios 260:280 of extracted DNA also showed better values for samples heated at 120C. PCR products of three primers showed satisfactory results for DNA extracted from archival paraffin-embedded tissues by heating protocols at pH 6–12, with results that were comparable to the control sample subjected to the standard non-heating, enzymatic DNA extraction method. This study is the first to document the use of heating at an alkaline pH for DNA extraction from archival formalin-fixed, paraffin-embedded tissues, a recommendation based on the principles of AR for protein IHC. These findings may lead to a more effective protocol for DNA extraction from archival paraffin-embedded tissues and may also provide enhanced understanding of changes that occur during formalin-induced modification of nucleic acids.
F
We developed the hypothesis that there may be analogies between “retrieval” of proteins for IHC and extraction of nucleic acids from archival paraffin-embedded tissue sections. Extraction of DNA from archival formalin-fixed, paraffin-embedded tissue was accomplished as early as 1985 using proteinase K and SDS as the major reagents (Goelz et al. 1985; Dubeau et al. 1986). High-temperature heating of the paraffin-embedded tissue was not applied as the primary means for extracting DNA, although heating the dewaxed paraffin sections at 100C was used as a primary step of PCR after regular DNA extraction (Shibata 1994). Subsequently, the heating AR technique was used for enhancement of in situ hybridization (ISH) methods applied to archival paraffin-embedded tissue sections (Sibony et al. 1995). Several different heating retrieval protocols have been documented to produce an improvement of signal by ISH or by fluorescence ISH (FISH) (Oliver et al. 1997; Bull and Harnden 1999; Kitayama et al. 1999; Gu et al. 2000). Successful application of the heat-induced retrieval method for ISH and FISH suggested that the high-temperature heating AR approach may perform in a similar manner for “retrieval” of nucleotides as for “retrieval” of antigens in the enhancement of IHC. In a related but separate approach, high-temperature heating methods have been used to improve extraction of nucleic acids from paraffin-embedded tissues (Banerjee et al. 1995; Faulkner and Leigh 1998). In this approach, in contrast to the AR technique, heat was not employed as the primary retrieval procedure for DNA extraction from archival paraffin-embedded tissue but was only part of a complex digestion protocol. Awareness that strong heating of tissue during DNA extraction could be used instead of enzyme digestion did not develop until careful comparative studies were carried out in recent years (Sepp et al. 1994; Frank et al. 1996). For example, Frank et al. (1996) pointed out that simple boiling gave results comparable with those of proteinase K digestion plus detergents followed by phenol-chloroform extraction of DNA from paraffin-embedded tissue. In addition, Coombs et al. (1999) reported a study that sought to optimize DNA/RNA extraction by comparing 10 protocols. They concluded that heating treatments at 90–99C by a thermal cycler, microwave, and simple boiling in solution [0.5% Tween-20, Tris-EDTA and Chelex-100 from Bio-Rad Laboratories (Hercules, CA)] significantly increased the efficiency of extraction of nucleic acids for use in molecular analysis.
The present study was designed to test the efficiency of DNA extraction from archival formalin-fixed, paraffin-embedded tissues based on application of the basic principles of the AR technique for proteins: heating the tissue under the influence of different pH value as previously documented (Shi et al. 1995). The goals included (a) development of a simpler and more effective protocol for DNA extraction and (b) enhancing understanding of the potential effect of the two major factors in relation to formalin/DNA interaction.
Materials and Methods
Tissues
Archival formalin-fixed, paraffin-embedded human tonsil, lymph nodes, and normal colon tissues were obtained from the Norris Cancer Hospital and Research Institute at the University of Southern California Keck School of Medicine (USC) from 1991 to 2000. Tissues had been processed as follows. Immediately after removal from the patients, all samples were routinely fixed in 10% neutral buffered formalin [average period of fixation was 24 hr at room temperature (RT)]. Fixed tissues were processed routinely through dehydration in graded ethanol, clearing in xylene, and embedding in paraffin blocks using automatic processing and embedding equipment (Tissue Tek VIP; Sakura Finetek, Torrance, CA). Selection of paraffin blocks was based on examination of one hematoxylin and eosin-stained slide to estimate the area of tissue and the total number of cells for comparability among different tissue sections. The study of human-derived tissue specimens has been exempted under 45 CFR 46.101 (b) and was approved by the Institutional Review Board (IRB #009071) at USC.
Each paraffin block was cut at 10 μm and collected in an autoclaved plastic microtube (1.5 ml). For each microtube, two sections of total 20-μm thickness were carefully collected to ensure that an equivalent amount of tissue was placed in each microtube.
Heating Procedure
A heat block (VWR Scientific Products; West Chester, PA) was used to attain a boiling heating condition (100C–105C). An autoclave for sterilization (Market Forge; New York, NY) was used to achieve a super high temperature heating at 120C. A water bath (Precision Scientific Inc; Chicago, IL) was used to maintain a temperature of 80C.
Universal Buffer Solution
A Britton and Robinson type of buffer was prepared for 11 buffer solutions ranging from pH 2.0 to 12.0. Briefly, the principal solution was made of 28.6 mM of each chemical as follows: citrate acid, KH2PO4, H3BO3, and diethylbarbituric acid. A solution of 0.2 N sodium hydroxide was added according to the original formula to reach the pH value demonstrated by pH meter (Oakton pHTestr 3; Singapore).
Design of Test
Samples from each of six paraffin blocks were prepared in 34 microtubes as described above, i.e., every tube contained an equal amount of two tissue sections of total 20 μm. A total of 204 tubes of tissue was included in this study. For each paraffin block, one tube was used as control sample by using a non-heating DNA extraction protocol. The other 33 tubes were tested using the heating protocol under 11 variable pH values (pH 2 to 12), and three different heating conditions (80, 100, and 120C). Evaluation of the results of DNA extraction was carried out by measuring yields by photometry (BioPhotometer; Eppendorf Scientific, Hamburg, Germany) and PCR amplification as well as the kinetic thermocycling (KTC)-PCR methods.
Non-heating DNA Extraction Protocol
Deparaffinization was carried out by adding 1 ml xylene to the microtube containing tissue sections for 30 min for two changes, followed by 100% and 75% ethanol for 30 min with two changes. After a washing step with PBS for 15 min in two changes 500 μl of lysis buffer (proteinase K 20 mg/ml, 50 μl, 1 M Tris-HCl solution 10 μl, 0.5 M EDTA 2 μl, 10% SDS 100 μl, and distilled water 838 ml) was added and incubated at 52C overnight until all tissue fragments were dissolved completely. Further extraction and purification procedures were performed by the following steps: addition of 500 μl phenol:chloroform:isopropanol alcohol (Invitrogen Life Technologies; Carlsbad, CA) at 25:24:1 to the dewaxed tissue, followed by mixing by vortex (Daigger Vortex; Scientific Industries, Bohemia, NY), and centrifugation (Eppendorf Scientific; Hamburg, Germany) at RT, 12,000 × g for 10 min. The supernatant fluid was removed to an autoclaved microtube using a 100-μl pipette, and one volume of chloroform was added, mixed by vortexing, and centrifuged at 12,000 × g for 5 min. The upper aqueous supernatant was carefully removed to another fresh microtube, adding 0.1 volume of 3 M sodium acetate and again mixed by vortexing, followed by addition of 1 volume of isopropanol, and incubation at −20C overnight. The precipitated DNA was centrifuged at 12,000 × g at 4C. The supernatant fluid was discarded and the precipitate washed once with 75% ethanol. The extracted DNA was collected after further centrifugation. The final yield of DNA was dissolved in 50 μl distilled water after drying completely in a hood.
Heating Protocol for DNA Extraction
A total of 500 μl universal buffer solution at various pH values ranging from pH 2 to 12 was added to each microtube containing a 20-μm tissue section and was heated at 80C, 100C, or 120C, respectively, using a water bath, heat block, or autoclave as described above. The heating time was 20 min for all temperatures. A cooling time of 5 min was allowed after heating. Further steps of extraction and purification were exactly as used for non-heating DNA extraction described above, except that enzyme digestion was omitted to compare the efficiency of DNA extraction more accurately.
Evaluation of DNA Yields
Using a photometer, the amount of DNA yield was measured according to the standard protocol recommended by the manufacturer.
The quality of DNA yields was evaluated by two laboratories [Drs. Datar and Shi, USC; Dr. Wu, Roche Molecular Systems (Alameda, CA)] independently using the PCR amplification with three primers, and the kinetic thermocycling (KTC)-PCR method.
PCR Analysis
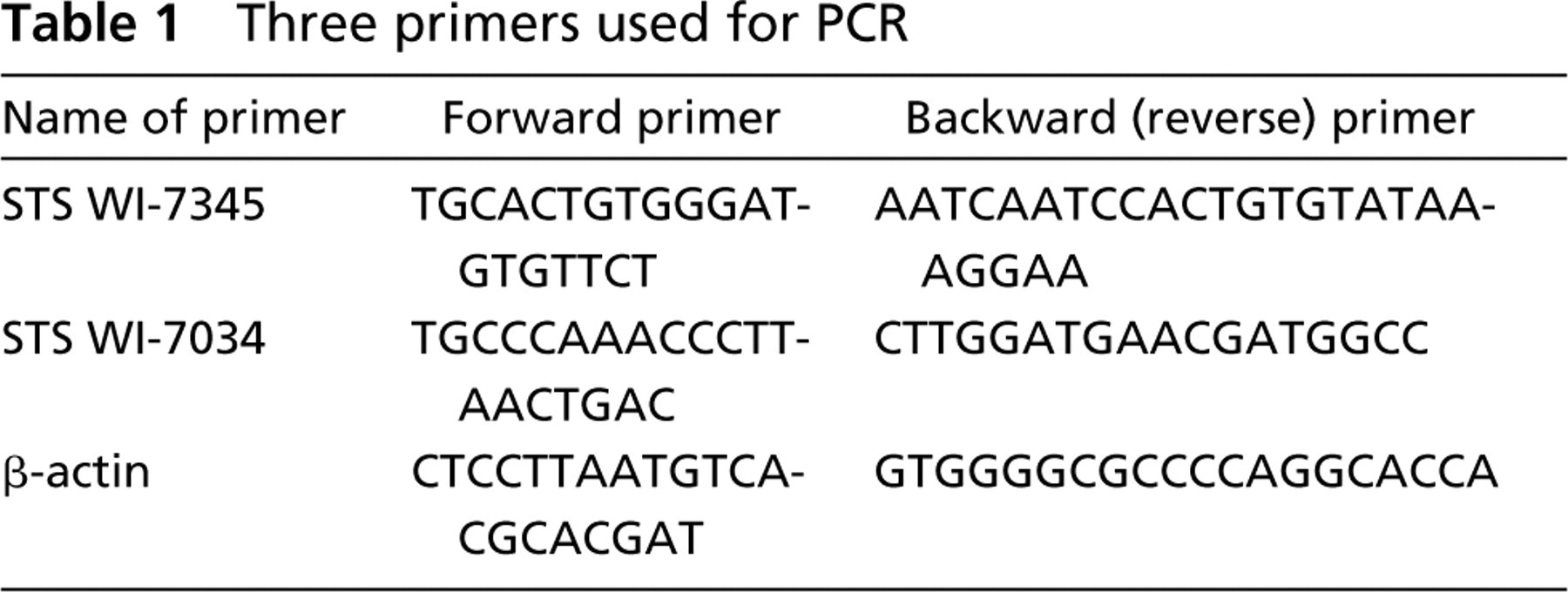
Each DNA extract was used as a template for PCR amplification, using three primer pairs as listed in Table 1. PCR tests were carried out based on groups of DNA samples extracted from six paraffin blocks under side-by-side identical conditions, i.e., each group consisted of 11 DNA samples that were extracted by heating at defined temperatures in buffer solutions of differing pH values, together with one sample extracted by the non-heating method as a control. Thus, for each paraffin block, three groups of heating at 80C, 100C, and 120C were tested, for a total of 36 PCR test results that were evaluated by gel electrophoresis.
PCR was performed by standard methods. The DNA sample diluted in 2 μl of distilled water containing 100 ng as template was added to 43 μl of 1.1 × ReddyMix PCR Master Mix [1.25 U Taq DNA polymerase, 75 mM Tris-HCl, pH 8.8, at 25C, 20 mM (NH4)2SO4, 1.5 mM MgCl2, 0.01% (v/v) Tween-20, 0.2 mM each of dATP, dCTP, dGTP, and dTTP]. The PCR protocol was then carried out with an initial 5-min denaturation step at 94C coupled to a repeating cycle of 1 min at 94C (denaturation), 1 min at 56C (annealing), and 1 min at 72C (extension) for 40 cycles, followed by a 10-min completion step.
Kinetic Thermocycling (KTC)-PCR
“Real-time” KTC-PCR assays (Higuchi et al. 1993) were carried out with the Applied Biosystems PE5700 instrument. Two μl of the genomic DNA was used for one PCR reaction. The KTC-PCR reaction was carried out in a solution of 10 mM Tris-HCl, pH 8.3, 50 mM KCl, 2.5 mM MgCl2, 200 μM dATP, 200 μM dCTP, 200 μM dGTP, 200 μM dTTP, 10 U of AmpliTaq-Gold, 0.2 × SYBR-Green (Molecular Probes; Eugene, OR), and 200 nM of each forward and reverse primer. Three sets of primers were used for the KTC-PCR assay, which amplify a 90-bp exon 3, a 168-bp exon 2, or a 368-bp exon 4 amplicon of the human p53 gene, respectively. The sequences of forward and reverse primers of exon 2 are 5′-TCATGCTGGATCCCCACTTTTCCTCTTG and 5′-TGGCCTGCCCTTCCAATGGATCCACTCA, respectively. The sequence of the forward and reverse primers of exon 3 are 5′-AATTCATGGGACTGACTTTCTGCTCTTGTC and 5′-TCCAGGTCCCAGCCCAACCCTTGCTCC, respectively. The sequence of the forward and reverse primers of exon 4 are 5′-GTCCTCTGACTGCTCTTTTCACCCATCTAC and 5′-GGGATACGGCCAGGCATTGAAGTCTC, respectively. The thermal cycling protocol was programmed as follows: an initial incubation at 95C for 10 min, then 40 cycles of 95C for 30 sec, 60C for 30 sec, and 72C for 45 sec, followed by a final extension at 72C for 10 min. Purified human genomic DNA purchased from Roche Molecular Biochemicals was used in the concentration range of 0.1–100 ng for the standard curves with primers of exon 3, 2, or 4.
Three primers used for PCR
The copy number of the amplifiable genomic DNA extracted from archival tissue sections was measured by KTC-PCR to examine the effect of pH of universal buffer solution and heating temperature on the yield. Three pairs of primers were used for KTC-PCR test. The first pair amplifies a 90-bp fragment from the region covering exon 3 of the p53 gene; the second pair amplifies a 168-bp fragment from region of exon 2. The third pair amplifies a 368-bp fragment from exon 4. A purified genomic DNA of a known copy number from Roche Molecular Biochemicals was used as a standard control. The experimental procedure was as described above. The results are shown in Figure 3.
Statistical Analysis
Student's t-test for paired samples was performed to compare the effects of different temperatures on DNA yield at pH 6–9. The results were considered significant when two-sided p values were less than 0.05.
Results
Measurement of DNA Yields
Total yields of DNA extracted from paraffin blocks correlated with the pH value of the buffer solution and the heating temperature, as summarized in Table 2 and Figure 1. Above pH 5, the amount of DNA yield was significantly increased. A peak of highest efficiency of extracted DNA was present at pH 9. In general, low pH (acid) was poorer than high pH (alkaline). Heating tissues at higher temperature produced greater yields of DNA, as shown in Figure 1 and Table 2. Figure 1 showed significant differences between 120C and 80C from pH 6 to pH 9 (T value = 7.899; p = 0.004) and between 120C and 100C from pH 6 to pH 9 (T value = 5.957; p = 0.0095). The average ratios 260:280 of extracted DNA also showed better values for samples heated at 120C. For example, the average ratios 260:280 for 80C, 100C, and 120C in buffer at pH 9 were 1.41, 1.44, and 1.60, respectively.
The control group, using the “routine” enzyme digestion protocol for DNA extraction from archival paraffin-embedded tissue sections, gave total yields for six samples of 44,275 ng compared to the “best” result of heating extraction of approximately 30,000 ng (Table 1). However, the quality of DNA extracted from paraffin-embedded tissue sections was equivalent based on comparable ratios 260:280 (1.58 for control sample vs 1.60 for heating at 120C in buffer solution of pH 9), as well as PCR products described below.
PCR Products
PCR products of three primers showed satisfactory results for DNA extracted from archival paraffin-embedded tissues by heating protocols at pH 6–12 with higher temperatures. The findings were comparable to those of the control sample obtained by a non-heating enzymatic DNA extraction method, as shown in Figure 2. In general, the better PCR products were seen at higher pH and higher temperatures. Some bands of PCR products were absent for samples obtained at 80C heating conditions, particularly when larger primers of 347 bp (STS WI-7034) and 541 bp (actin) were used.
KTC-PCR Products
The yields of amplifiable genomic DNA measured by Exon 3 primers to amplify a 90-bp fragment (Figure 3A) were affected by pH as well as heating temperature. If the pH was lower than 5, no amplifiable DNA was recovered regardless of the temperature at which the samples were heated. A heating temperature of 80C gave lower yields than either 100C or 120C in a pH range from 6 to 12. The best condition for yields of amplifiable genomic DNA by exon 3 primers was heating at 120C in the universal buffer with a pH of 10–12. Sections of 20 μm gave about 10,000 copies/μl measured by the exon 3 primer.
Average DNA yields (ng) extracted from archival paraffin-embedded tissue sections by heating protocols using buffer solutions of pH 2–12 a
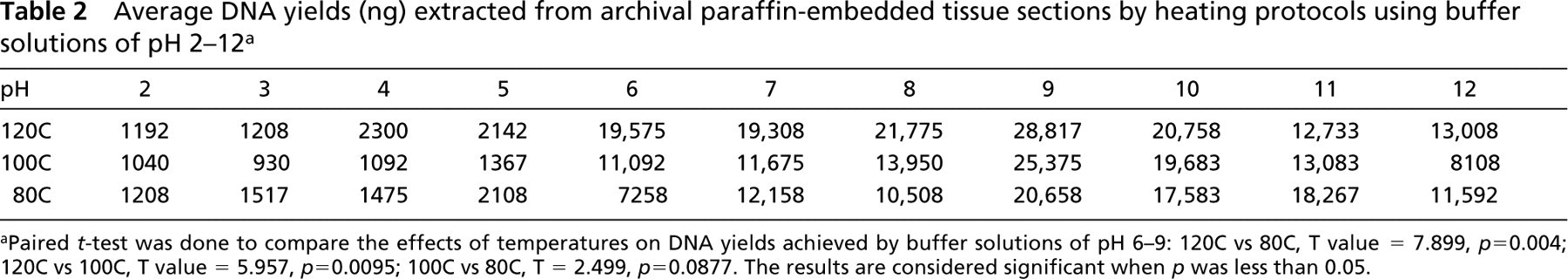
aPaired t-test was done to compare the effects of temperatures on DNA yields achieved by buffer solutions of pH 6–9: 120C vs 80C, T value = 7.899, p=0.004; 120C vs 100C, T value = 5.957, p=0.0095; 100C vs 80C, T = 2.499, p=0.0877. The results are considered significant when p was less than 0.05.
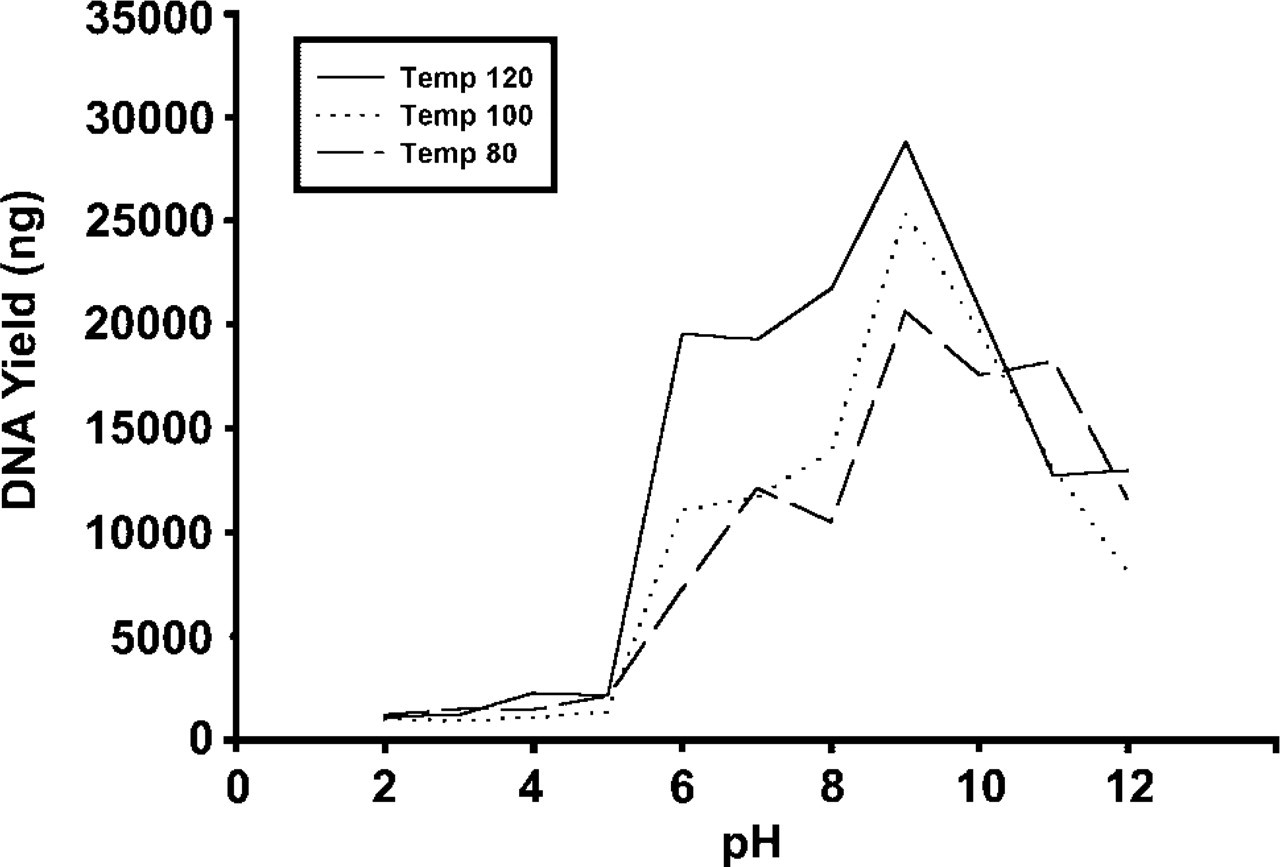
Comparison of DNA yields obtained by three heating temperatures of 80C, 100C, and 120C under the influence of various pHs ranging from 2 to 12. In general, low pH (acid) was poorer than high pH (alkaline). A peak for highest efficiency of extracted DNA was present at pH 9. Heating tissues at higher temperature appears to produce greater yields of DNA with a buffer solution of pH 6–9.
When the exon 2 primer was used to amplify a 168-bp fragment (Figure 3B), the results were consistent. The best condition appeared to be 120C with a pH of 10–12. However, the yield of amplifiable genomic DNA was lower, about 5500 copies/μl, indicating that the amplification efficiency of exon 2 is lower than that of exon 3 from the same genomic DNA isolated from paraffin-embedded tissue.
When the exon 4 primer was used to amplify a 368-bp fragment (Figure 3C), the results were consistent with those of both exon 3 and exon 2. The best result was achieved by heating tissues at 120C in a universal buffer solution at pH 11. However, the yield of amplifiable genomic DNA was even lower, about 1200 copies/μl, indicating that the amplification efficiency of exon 4 is lower than that of both exon 3 and exon 2 from the same genomic DNA isolated from paraffin-embedded tissue.
Discussion
Recent publications have demonstrated that high-temperature heating may improve DNA extraction from archival formalin-fixed, paraffin-embedded tissues (Frank et al. 1996; Faulkner and Leigh 1998; Coombs et al. 1999). However, to date these observations have not been equated to the principles derived from studies of heat-induced AR, which also is based on high-temperature heating of formalin-fixed, paraffin-embedded tissues (Shi et al. 1995). The present study has demonstrated that the same factors that affect the results of AR, i.e., heat and pH, may also influence the efficiency of DNA extraction from archival paraffin-embedded tissue. In our study, higher-temperature heating under an alkaline condition provided the most satisfactory results, as indicated by both quantity and quality of DNA yield (Table 2; Figures 1–3). One conclusion is that a simple heating method can be used to achieve the goal of analyzing gene alterations after PCR amplification. A recent article by Yamashita and co-workers (2001) documented an interesting method that could provide the basis for a simple heat-induced retrieval method for both DNA extraction and immunostaining simultaneously in a single archival paraffin-embedded tissue section. Although there is some evidence that the simple heating method for DNA extraction can be used alone to yield satisfactory results for detection of genomic alterations in DNA recovered from archival paraffin-embedded tissue sections, this approach has not been widely applied. One possible reason may relate to the finding that the quantity of DNA extracted from archival paraffin-embedded tissue by heating alone is lower than that extracted by an enzyme-based protocol. However, in our study the quantity was adequate and the quality, evaluated by both three-primer PCR and KTC-PCR methods, proved to be comparable, even superior for the heating as opposed to non-heat extraction protocols. In particular, KTC-PCR products demonstrated that higher-temperature heating at higher (alkaline) pH values may produce higher yields of high quality genomic DNA for the exons studied (Table 2; Figures 1–3). Further studies are required to validate wider applications for various kinds of exons.
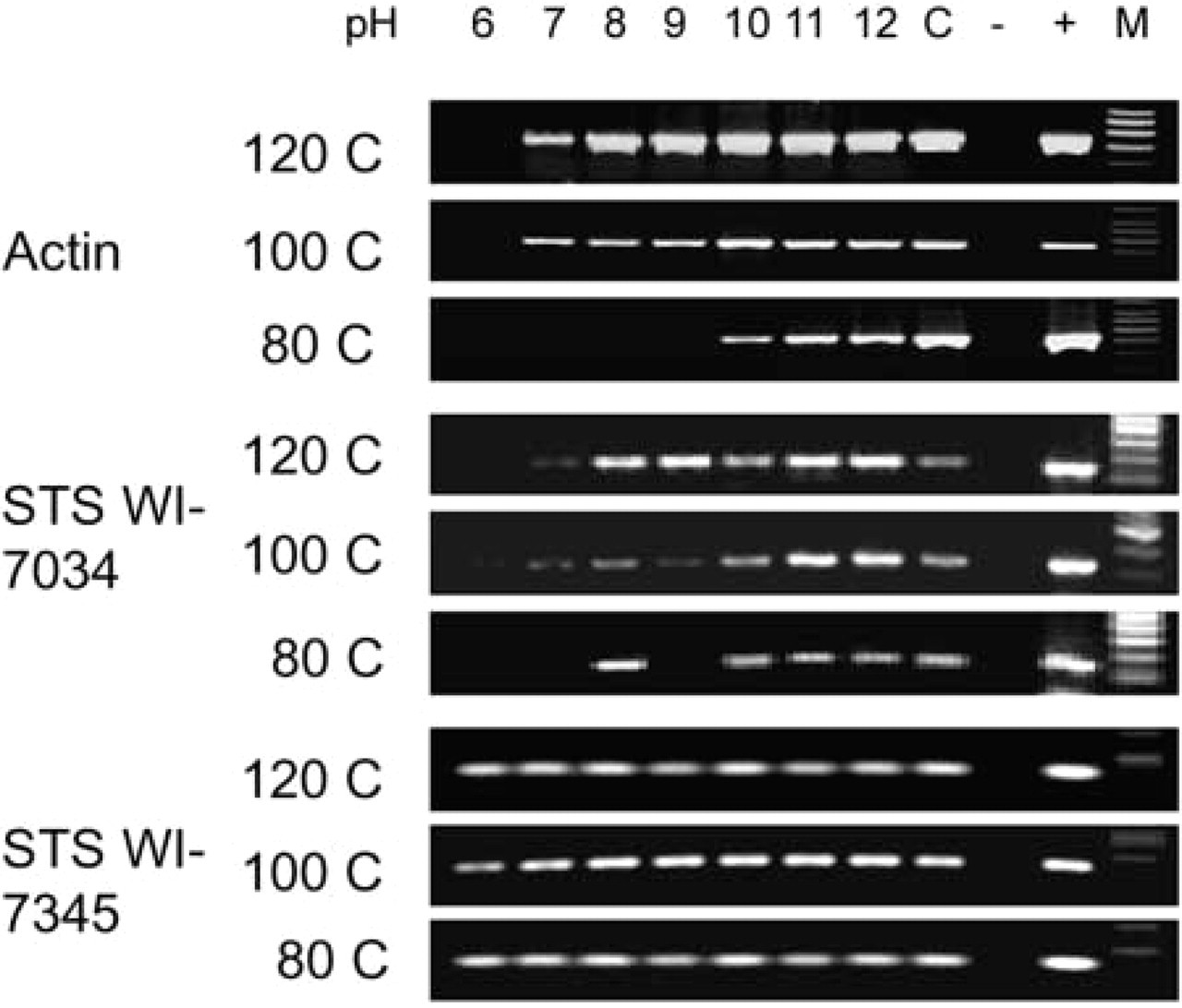
Gel electrophoresis of PCR products with three primers of STS WI-7345 (152 bp), STS WI-7034 (347 bp), and actin (541 bp) under the influence of two factors: heating at 80C, 100C, or 120C with various pH values ranging from 2 to 12. Because there are no bands from samples in buffer solutions of pH 2–5 under three heating conditions, Figure 2 shows PCR products from samples in buffer solutions ranging from pH 6 to 12 only. The pH values of buffer solutions are adjusted to within ± 0.3. C, control sample (DNA extracted from archival paraffin-embedded tissue sections without heating protocol); –, negative control (no template for PCR); +, positive control sample; M, GeneChoice DNA ladder.
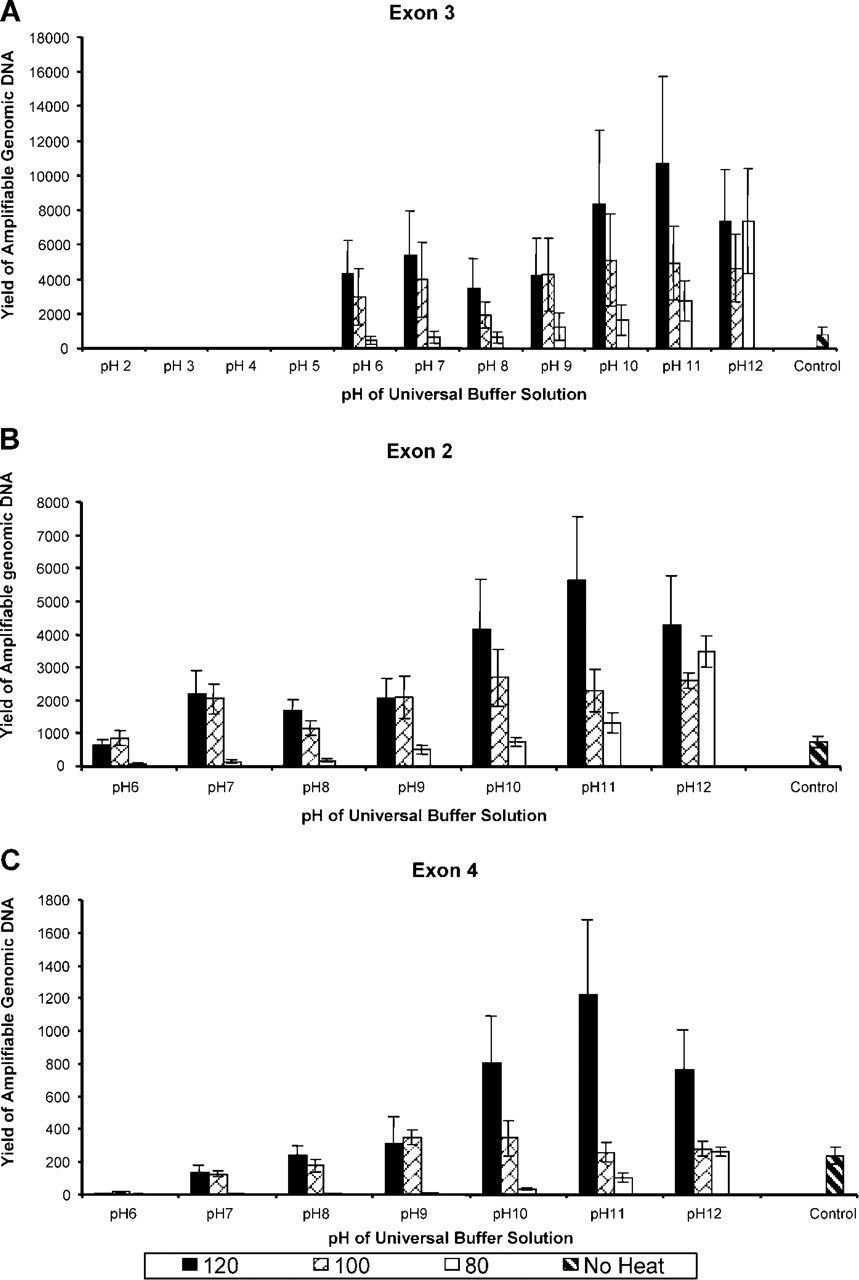
Amplifiable genomic DNA measured by KTC-PCR using exon 3 (
Alkaline lysis methods (at RT or 95C) have been used previously for DNA extraction from bacteria (Birnboim and Doly 1979), plant tissue (Wang et al. 1993), whole blood (Klintschar and Neuhuber 2000), or blood culture fluids and other fresh/frozen tissues (Kulski and Pryce 1996; Rudbeck 1998; Valsecchi 1998). However, the combination of high-temperature heating and alkaline pH has not been documented for DNA extraction from archival formalin-fixed, paraffin-embedded tissues. Furthermore, the ranges of temperature and pH values that affect the efficiency of the extraction process have not been previously explored.
The mechanism of action of heating in alkaline pH solution for DNA extraction may be similar to a hypothesis proposed for the mechanism of AR under different conditions of temperature and pH (Shi et al. 2000b). A strongly alkaline solution may denature and hydrolyze proteins, resulting in rupture of cell and nuclear membranes as well as breakage of crosslinkages caused by formalin fixation. Hydrolysis of proteins in tissue may also contribute to improved DNA extraction. Given the similarity of the chemical reactions of formaldehyde with nucleic acid and protein (antigen), it is perhaps not surprising that extraction or “retrieval” of the two macromolecules should be influenced by similar factors (Auerbach et al. 1977; McGhee and von Hippel 1977a, b; Masuda et al. 1999; Shi et al. 2001a).
Footnotes
Acknowledgements
Supported by an NIH grant 1 R21 CA91249–01.
We greatly appreciate Dr Andy E. Sherrod's kind help in providing some paraffin blocks for study. We also thank Ms Lillian Young for management of our laboratory.
