Abstract
The Nkx6 gene family of homeodomain transcription factors consists of three members. For two, Nkx6.1 and Nkx6.2, important developmental roles in the central nervous system and pancreas have been demonstrated. Here we introduce the third member of the Nkx6 gene family, Nkx6.3, and identify similar and distinct patterns of expression for all three Nkx6 genes in the hindbrain and gut of the developing mouse embryo.
Nkx6.1 and Nkx6.2 are both expressed in the developing mouse spinal cord and hindbrain (Sander et al. 2000a; Vallstedt et al. 2001; Muller et al. 2003; Pattyn et al. 2003). Previous work has shown that Nkx6.1 is required for the formation of somatic spinal motor neurons as well as for the migration of branchial motor neurons in the hindbrain. Both functions of Nkx6.1 are partially compensated for by Nkx6.2 (Sander et al. 2000a; Vallstedt et al. 2001; Muller et al. 2003; Pattyn et al. 2003). In addition to neural tube, Nkx6.1 is also expressed in the embryonic pancreas. Inactivation of the Nkx6.1 gene in mice results in an almost complete loss of the insulin-producing beta cells in the pancreas (Sander et al. 2000b).
A homology search with the Nkx6.1 homeodomain revealed the presence of a yet uncharacterized third Nkx6 transcript in mice that we named Nkx6.3. This Nkx6.3 transcript was isolated from a RIKEN full-length enriched library from mouse male cecum (Gen-Bank Accession AK018683). All three NKX6 proteins have almost identical homeodomains and share an engrailed homology domain (eh1-domain) that mediates interaction with the corepressor protein groucho (Muhr et al. 2001). Outside the homeo- and eh1-domains, NKX6.2 and NKX6.3 are more similar in structure to each other than to NKX6.1, which contains poly-serine-and poly-alanine-rich regions in the N-terminal half of the protein. Further homology searches using the Ensembl database (
To generate a riboprobe, a portion of the 3’ untranslated region of Nkx6.3 (base pairs 1105–1927) was subcloned into pBKS (Stratagene; La Jolla, CA). Nkx6.2 transcripts were detected with a fragment containing base pairs 253–1244 from the mouse cDNA (GenBank Accession L08074). The mouse cDNA in situ riboprobe for Nkx6.1 (GenBank Accession AF357883) was generated as previously described (Qiu et al. 1998). Embryonic day (e) 12.5 and e16.5 embryos were fixed overnight in 4% paraformaldehyde, washed with PBS, and incubated overnight in 30% sucrose in PBS. The embryos were embedded in Optimal Cutting Temperature (OCT) (Tissue Tek; Sakura, Torrance, CA) and stored at −80C. Ten-μm sections were used to conduct ISH. ISH with [α-35S]UTP-labeled anti-sense riboprobes was performed on coronal sections from e12.5 and e16.5 embryonic brains and sagittal sections from e12.5 and e16.5 mouse embryonic gut regions (Susens et al. 1997). After ISH, Cresylviolet staining was conducted before mounting the slides with Entellan. Darkfield and brightfield images were captured with a Zeiss Axiovision camera and software on a Zeiss Axioplan microscope (Zeiss; Thornwood, NY). The images were processed in Adobe Photoshop (Adobe Systems Inc.; San Jose, CA).
All three Nkx6 genes are expressed in the developing nervous system. Whereas Nkx6.1 and Nkx6.2 are expressed in the spinal cord, hindbrain, and midbrain (Qiu et al. 1998; Vallstedt et al. 2001; Muller et al. 2003; Pattyn et al. 2003), we found Nkx6.3 mRNA to be exclusively confined to the caudal hindbrain region. In the caudal hindbrain, Nkx6.1-positive nuclei comprise the somatic hypoglossal motor nucleus (nXII), the visceral dorsal motor nucleus of the vagal nerve (dmnX), and the branchial motor nucleus ambiguous (nA) (Figure 1B). At e16.5, Nkx6.1 expression is maintained only in the nA but in addition appears in the neural epithelium (NE) and in the raphe nuclei (nRP, Figure 1H). Similar to Nkx6.1, Nkx6.2 is expressed in the nA at e12.5 and e16.5 but is not detected in the hypoglossal nucleus or dorsal motor nucleus of the vagal nerve (Figures 1D and 1J). In contrast to Nkx6.1 and Nkx6.2, which are expressed early in the developing neural tube (Vallstedt et al. 2001; data not shown), Nkx6.3 is not detected prior to e12.5 and is not found in motor nuclei. At e12.5, Nkx6.2 and Nkx6.3 are both expressed in the ventral–medial aspect of the medullary reticular formation (MDR) (Figures 1D and 1F). At e16.5, Nkx6.3 expression is confined to the gigantocellular nucleus (Gi, Figure 1L), a nucleus that is part of the MDR. Similar to Nkx6.3, Nkx6.2 is also expressed in the Gi nucleus at e16.5 but in a more restricted region (Figures 1J and 1L). Additional Nkx6.2 expression is found in the inferior olive (IO) at e16.5 (Figure 1J). Expression at e18.5 of all three Nkx6 genes is similar to that observed at e16.5 with the exception of Nkx6.2, which is broadly expressed in the Gi nucleus at e18.5 (data not shown).
Studies of CNS development in Nkx6.1 and Nkx6.2 mutant mice have shown that NKX6.2 compensates for NKX6.1 in regions of coexpression of both proteins. In Nkx6.1 mutants, the nA, although present, exhibits a significant reduction in size (Muller et al. 2003), whereas in Nkx6.1/Nkx6.2 double mutants, the nA is completely absent (Pattyn et al. 2003). Given the overlapping expression of Nkx6.2 and Nkx6.3 in the MDR, it is possible that Nkx6.3 compensates for Nkx6.2. In agreement with this hypothesis, homozygous Nkx6.2 mutant mice develop normally and have no obvious defects of their respiratory or vasomotor function (Cai et al. 2001), two processes that are controlled by the neurons of the MDR. It is therefore possible that defects will only become apparent in Nkx6.2/Nkx6.3 double-null mutant mice.
In addition to the CNS, all three Nkx6 genes are also expressed in the developing gut. During gut development, the pancreas arises from two separate rudiments, which are first detected at e9.5 (Slack 1995). At e10.5, Nkx6.1 and Nkx6.2 are both strongly expressed in the pancreatic epithelium, whereas Nkx6.3 is only detected at very low levels (Sander et al. 2000b; data not shown). Whereas expression of Nkx6.1 persists in the pancreatic epithelium (Figures 2H and 2N), Nkx6.2 and Nkx6.3 are no longer detected by in situ hybridization at e12.5 or thereafter (Figures 2J and 2L; data not shown). However, both Nkx6.2 and Nkx6.3 are detected by RT-PCR in the pancreas at e15.5 (data not shown), indicating that low amounts of mRNA are still present at later developmental stages. Additionally, we have detected low levels of NKX6.2 in the pancreas by immunohistochemistry throughout development. At e12.5, the expression domains of Nkx6.2 and Nkx6.3 in the developing gut have diverged from Nkx6.1. At this age, Nkx6.1 is detected in the anterior stomach mesenchyme (Figures 2B and 2H), but unlike Nkx6.2 and Nkx6.3 is not found in the epithelium of the posterior stomach or the duodenum (Figures 2D, 2F, 2J, 2L). Restricted expression of the Nkx genes is also observed at e16.5 in the developing gut. Whereas Nkx6.1 is expressed in the pancreas (Figure 2N), Nkx6.2 and Nkx6.3 are detected in different regions of the duodenum (Figures 2P and 2R). In the duodenum, Nkx6.2 expression is restricted to the mesenchyme and Nkx6.3 to the epithelium (Figures 2P and 2R).
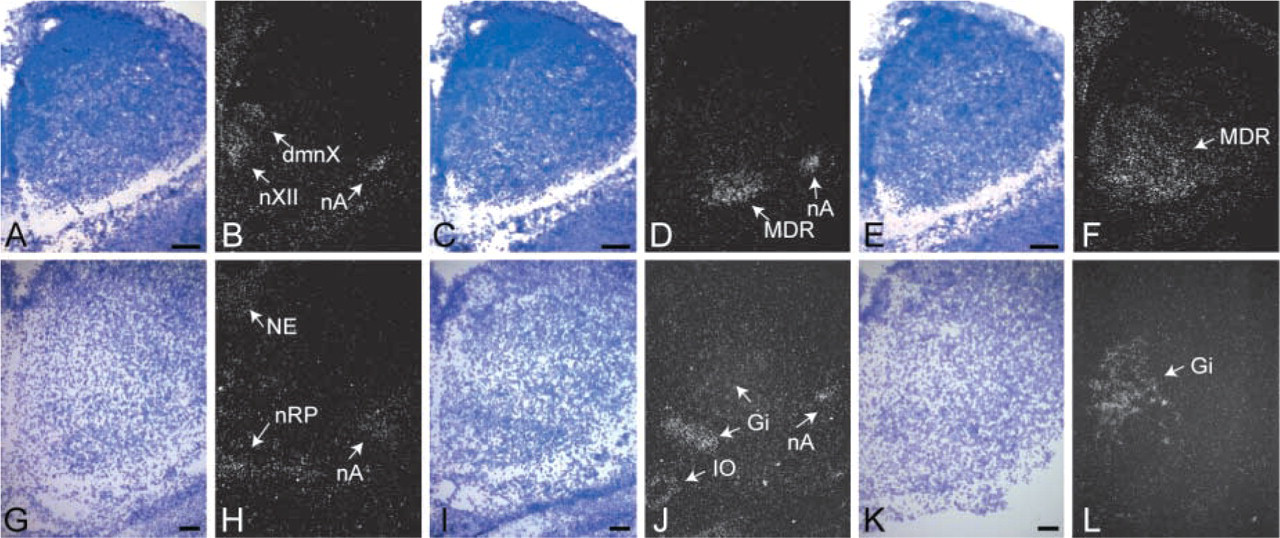
Nkx6 expression in the developing mouse hindbrain. Darkfield images of in situ hybridizations with riboprobes against Nkx6.1
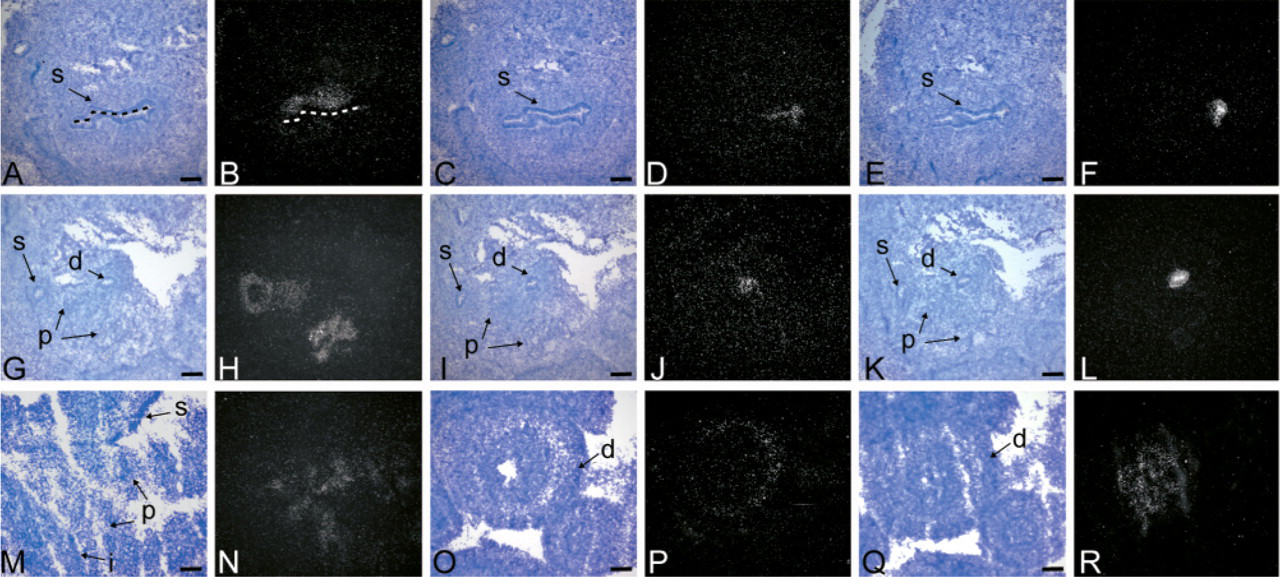
Nkx6 expression in the developing mouse gut. Darkfield images of in situ hybridizations with riboprobes against Nkx6.1
In summary, we have characterized the expression pattern of a novel Nkx6 transcription factor, Nkx6.3, and compared its expression to the two other Nkx6 family members, Nkx6.1 and Nkx6.2. We show that expression of Nkx6.3 in the developing embryo is confined to the gut and caudal hindbrain, two regions that also express Nkx6.1 and Nkx6.2. The data demonstrate that Nkx6.2 and Nkx6.3 share similar domains of expression. Once antibodies against NKX6.3 have been generated, it can be directly tested whether these two NKX6 factors are coexpressed in the same cells. This knowledge will be crucial to interpret the phenotypic alterations found in Nkx6 mutant mice.
Footnotes
Acknowledgements
This work was started at the Center for Molecular Neurobiology, Hamburg, Germany, and completed at the University of California, Irvine, with support from set-up funds from the University of California, Irvine, the National Institutes of Health Grants 1U19-DK61244 and RO1-DK068471 to MS. SBN was supported by a fellowship from the Humboldt Foundation.
We thank Dr. Bernd Fritzsch for helpful discussions.
