Abstract
Laser microdissection has opened a window to new technologies. The scientific fields of genomics, transcriptomics, and proteomics need pure samples for rendering reliable results. Homogeneous sample preparation is a prerequisite for modern molecular analyses, both qualitative and quantitative. Laser microdissection and pressure catapulting (LMPC) is a tool for isolating specific cells from complex tissues in a non-contact and contamination-free manner. Because LMPC technology is an optimal method for obtaining fast and reliable access to single cells, the possibility of automatic isolation of single fetal cells has the promise of being a big step forward in developing protocols for non-invasive prenatal diagnosis.
Keywords
A
Fetal cell recovery from maternal blood promises to be an attractive addition to non-invasive prenatal diagnosis, if reliable and consistent protocols can be developed. LMPC, in combination with the application of an automated scanning software (Metafer P; MetaSystems, Altlussheim, Germany or Cellenger; Definiens AG, Munich, Germany), allows automatic detection, isolation, and collection of single labeled cells and may, therefore, be a very helpful tool for developing such new protocols.
Principles of LMPC Technology
A pulsed UV-A laser is coupled to a regular research microscope and focused via the objective lenses to a micron-sized spot diameter. Within the narrow laser focal spot, forces are generated that allow ablation of material (laser microdissection) while the surrounding tissue remains fully intact. At the focal point, unwanted material is photofragmented into molecules and atoms, a phenomenon called “cold ablation” (Vogel and Venugopalan 2003). Inasmuch as this cutting is a fast, photochemical process without heat transfer, adjacent biological matter or biomolecules such as DNA, RNA, or proteins are not affected. Therefore, these molecules can routinely be isolated from the specimen for downstream analyses and applications.
Using the same laser, the separated cell(s) or selected tissue area can be lifted up and captured in a collection device. This is a totally non-contact process, inasmuch as only focused light is used for the transportation process. This so-called LPC technology (laser pressure catapulting) marks a breakthrough in modern laser capture methods and enables non-contact preparation of pure and homogeneous samples in a fast and efficient manner (Schütze et al. 2003).
Various targets, from parts of chromosomes to an entire living organism, such as the nematode Caenorhabditis elegans, are successfully transported without impairing the biological information or the viability of the specimen (Thalhammer et al. 2003). The same principle is applicable to the collection of live cells from a cell culture for subsequent re-cultivation (Stich et al. 2003).
The catapulted material subsequently will be spun down, analyzed directly, or used for further processing or various experiments.
Fetal Cells in Maternal Blood and LMPC
Current methods for the diagnosis of aneuploidy and monogenic disorders require invasive testing by chorionic villus biopsy, amniocentesis, or fetal blood sampling. These tests carry a procedure-related risk of miscarriage that is unacceptable to many couples. Isolation of fetal cells from maternal blood should allow accurate non-invasive prenatal diagnosis without the risks associated with invasive testing. The primary goal of identification is specificity; absolute certainty of fetal origin is required. Therefore, only single cells can be investigated. Inasmuch as fetal cells in maternal blood are rare events, automatic detection and collection systems are very helpful in isolating these single cells quickly. Application of automated scanning software (Metafer P or Cellenger) combined with an automated system for collecting single cells (PALM's LMPC technology) allows for fast access to the desired cells (see below). As soon as it is possible to clearly distinguish fetal cells from maternal cells, isolation of these cells from maternal blood will be no challenge for LMPC. Then a reliable non-invasive prenatal diagnosis can be offered, important especially for low-risk patients.
To overcome the limitations of non-invasive diagnosis caused by the rare appearance of fetal cells, cultivation may be an option. Cultivation of fetal progenitor cells from maternal blood offers the opportunity to produce significant numbers of fetal cells for prenatal molecular and cytogenetic analysis. A drawback of currently tested methods is the application of selective medium that never leads to clones of true single cells. LMPC allows the collection of single live cells directly out of a mixed culture without the necessity of applying techniques of selective stimulation (Figure 1).
Additionally, to ensure high-quality diagnosis, one single cell can be isolated early from a cultivated cell clone colony for a single-cell PCR, while the clone will grow further in the incubator. These results can later be compared with results obtained using the entire clone for diagnosis. Until now, no one has tried this approach, but it may be worth trying.
Further Applications of LMPC
Individual cells as well as small cell groups can be separated by LMPC and used directly for subsequent analyses. Paraffin and cryosections, even archival material, cell smears, cytospins, and chromosome preparations, are well-known applications of LMPC. New applications, for example, are the use of botanical or forensic samples as well as live cells from a cell culture or live material such as the nematode C. elegans.
One innovation is live-cell catapulting and reculturing, which demonstrates a new and easy method for clonal expansion of cells (Stich et al. 2003). Inasmuch as the viability of catapulted cells is not restricted, different cell types distinguishable by morphology, fluorescence, or transfection markers can be isolated quickly and reliably by LMPC. The preparation of stem cells and the selective elimination of specific cells from a culture, usually difficult procedures, are simple and fast with LMPC.
In the fields of pathology and oncology, the LMPC method is well-established. Unwanted material can be selectively eliminated by ablation. Reasonable expression analysis of tumor versus non-tumor cells is feasible only if no bulk material is used. Reliable results can be generated only by guaranteeing pure sample preparation by well-defined discrimination of tumor cells and non-tumor cells.
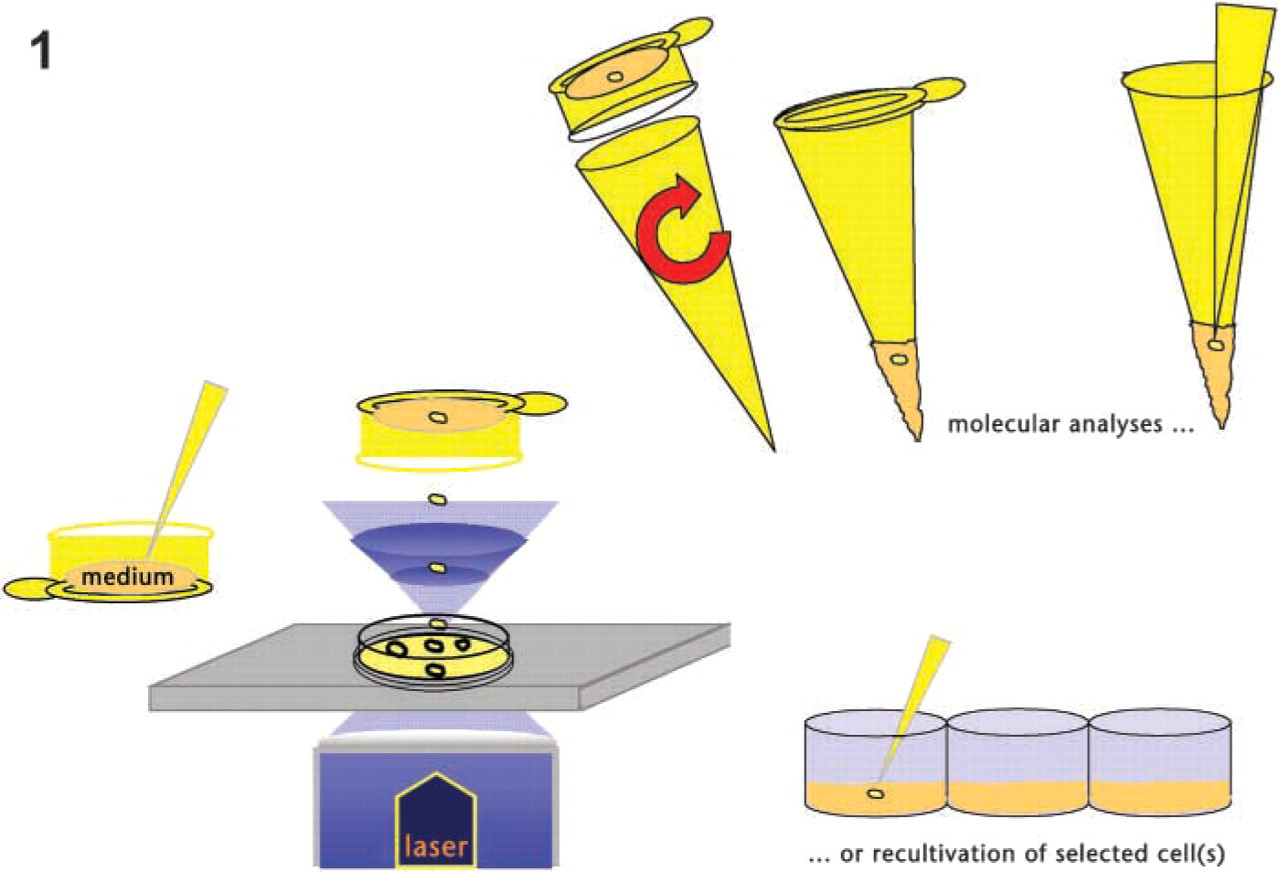
LMPC of live cells. The cell is catapulted into a medium-filled collection device. After simply spinning down into the tube, molecular analyses can be performed directly, or the cell(s) can be re-cultivated further.
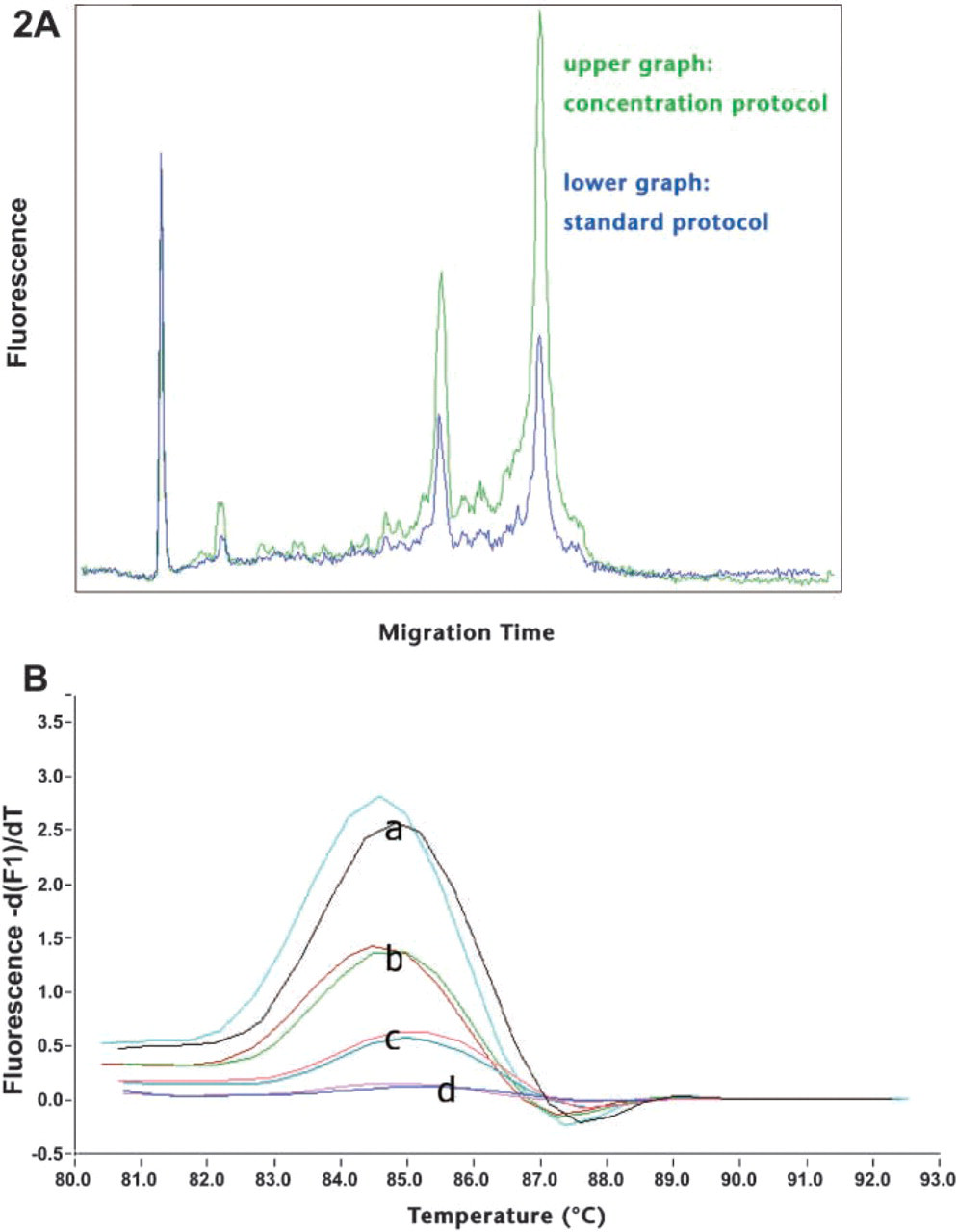
High-quality RNA of micro-dissected cells. (A) Frozen mouse liver section on a membrane slide stained with methylgreen; 2000 cells were catapulted, and RNA integrity was checked with Agilent's Bioanalyzer. Two different extraction protocols were used. Clearly visible 28S and 18S rRNA peaks show that with both protocols, high-quality RNA is extracted from microdissected cells. (
In forensics, freshly prepared or archived smears can be used, e.g., sperm or epithelial cell isolation. Even years after a crime, a genetic fingerprint made from specific cells can help to identify the culprit.
In cytogenetics, interesting single chromosomes, or chromosome arms or bands can be selected by LMPC to produce specific paint probes for reverse chromosome painting (Thalhammer et al. 2004). The selection of marker chromosomes for this purpose may be very helpful in the field of prenatal diagnosis.
High-quality RNA from Microdissected Cells
Many biological applications need intact high-quality RNA as starting material for basic and specialized experiments. Isolation of intact RNA, therefore, is the most critical step in performing many experiments and is essential for many techniques used in gene expression analysis. Homogeneous samples are prerequisites for reliable and reproducible results. Microdissected samples, therefore, lead to fast and precise results (Burgemeister et al. 2003).
RNA to be amplified for subsequent array hybridization must be of especially high quality (Figure 2A). Microarray chips are usually expensive, so only promising RNA should be applied to the chip.
Measuring expression profiles of individual cells is useful in a wide range of research and clinical applications. Using a new magnetic bead technology, we were able to reliably and reproducibly extract mRNA from even single cells (Figure 2B).
Automated Cell Recognition
The search for desired cells on a slide in, for example, the detection of rare fetal cells, can often be time consuming and exhausting. Image analysis software modules can be very helpful in detecting the desired cells automatically.
Metafer P and Cellenger are software modules capable of fast and automatic detection of labeled rare cells. Their reliable object recognition by intensity, color, and shape analyses allows the generation of a high-quality cell gallery for on-screen review and cell relocation.
An efficient detection algorithm has been designed by PALM for achieving integrated, interactive classifiers and rule sets for optimized recognition and accurate results.
Data of single cells assessed as fetal were transferred to the element list of the PALM RoboSoftware. Pure retrieval of those cells was then performed by non-contact LMPC. This type of investigation allows for automatic detection, selection, and collection of every cell type of interest in non-invasive prenatal diagnosis.
Footnotes
Acknowledgements
I would like to thank Sinuhe Hahn and Susanne Mergenthaler from University Women's Hospital in Basel for sending fetal cells for use in the development of the MetaferP software and for excellent collaboration. Thanks also to my colleagues in the PALM lab for brilliant work and to Renate Krause for expert help regarding the figures.
