Abstract
Sixty blood samples from pregnant women during gestational weeks 9–28 were investigated. Cell-free fetal DNA was extracted from maternal plasma or serum to be detected by nested PCR for determination of fetal gender. The SRY gene as a marker for fetal Y chromosome was detected in 34/36 women carrying a male fetus. In 3/24 women carrying female fetuses, the SRY sequence was also detected. Overall, fetal sex was correctly predicted in 91.7% of the cases. Therefore, the new, non-invasive method of prenatal diagnosis of fetal gender for women at risk of producing children with X-linked disorders is reliable, secure, and can substantially reduce invasive prenatal tests.
D
The aim of this investigation was the development of an efficient nested PCR-based method for the detection of Y-chromosome-specific sequence (SRY) gene in fetal DNA extracted from plasma or serum of pregnant women.
Patients
Sixty pregnant women at a gestational age ranging from 9 to 28 weeks and undergoing an invasive prenatal diagnostic procedure for reasons such as advanced maternal age, maternal serum, and ultrasound screening were recruited for this study.
Cytogenetics
Karyotypes of the fetuses during the first trimester were determined in chorionic villus samples using semi-direct methods. Karyotypes of the fetuses during the second or third trimester were determined by conventional banding cytogenetics method of cord blood samples obtained by cordocentesis. All fetal karyotypes were normal.
DNA Extraction from Plasma or Serum Samples
Blood samples from the 60 women were centrifuged at 1850 × g for 10 min and the supernatants were collected and stored at −20C. DNA was extracted from 500 μl plasma or serum using the Diatom DNA Kit (IsoGene; Moscow, Russia) according to the manufacturer's instructions. DNA was eluted in 50 μl of water and 5–10 μl was used as a template for the PCR reaction.
PCR Analysis
Two rounds of PCR were carried out to amplify the fragment of the SRY gene. The first PCR analysis was performed in a total volume of 50 μl. The PCR mixture contained 5–10 μl extracted DNA, 5 μl PCR buffer, 250 μM of dNTP mixture, 20 pM “external” SRY-specific oligonucleotide (Y1 5'-GTG TCC TCT CGT TTT GTG AC-3', Y2 5'-GTA ATC ATC GCT GTT GAA TAC-3') and 0.5 U Taq-polymerase (Sileks; Moscow, Russia). The PCR conditions were preheating at 95C for 3 min, then 30 cycles of 94C for 30 sec, 57C for 30 sec, 72C for 3 min, and then 72C for 3 min. The size of the PCR fragment was 369 bp. The second round was also performed in a total volume of 50 μl. The PCR mixture contained 5–10 μl of the product of the first PCR round, 5 μl PCR buffer, 250 μM of dNTP mixture, 20 pM “internal” SRY-specific oligonucleotide (Y3 5'-TGG CGA TTA AGT CAA ATT CGC-3', Y4 5'-CCT AGT ACC CTG ACA ATG TAT-3') and 0.5 U Taq-polymerase (Sileks). PCR conditions were preheating at 95C for 3 min, then 30 cycles of 94C for 30 sec, 57C for 30 sec, 72C for 1 min, and then 72C for 3 min. The size of the final PCR fragment was 130 bp.
The control PCR for a fragment of amylo-1,6-glucosidase gene (AGL) was carried out as standard assay in a total volume of 50 μl. The part of the AGL gene that is located on 1p21 was amplified. Oligonucleotide primers GL3.5 5'-CCG AGC TTA TTC TGT AGA AG-3' and GL3.6 5'-ACA TGC TCC TGA TGA CTT AC-3' were designed to amplify exons 33–34 of AGL. The size of the PCR product was 469 bp.
Nucleotide sequences for the SRY gene and AGL gene were obtained from the Gene Data Bank.
PCR products were analyzed in 2% agarose gel containing ethidium bromide.
PCR fragment of amylo-1,6-glucosidase gene was detected in all samples investigated, confirming the presence of DNA in the samples. Cytogenetic analysis revealed that 24 fetuses had normal female karyotype—46,XX, and 36 fetuses had normal male karyotype—46,XY.
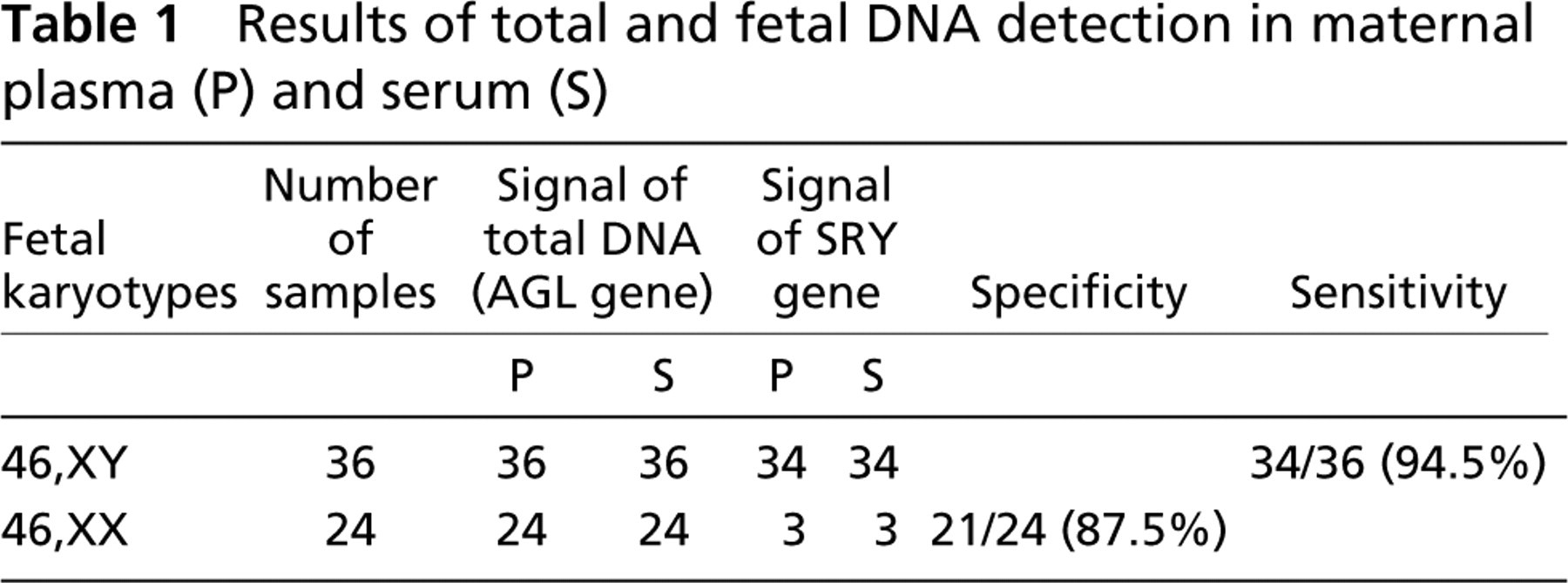
Specificity of PCR of SRY sequence was 87.5%: specific PCR fragment for Y chromosome was amplified in three DNA samples extracted from plasma or serum of 24 women carrying female fetuses. Thus, false-positive results were 5% (3/60 samples investigated). The sensitivity of the method was 94.5%: specific PCR fragment for Y chromosome was not amplified in two DNA samples extracted from plasma or serum of 36 women carrying male fetuses. Thus, false-negative results were 3.3% (2/60 samples investigated). Detection rate of this method (the coincidence of results of cytogenetic analysis with PCR results) was 91.7% (55/60 samples investigated) (Table 1).
Results of total and fetal DNA detection in maternal plasma (P) and serum (S)
It is well known that fetal cells circulate in maternal plasma of pregnant women. It has been shown that fetal cells crossing into maternal plasma are composed of different cell types such as erythroblasts, trophoblasts, and lymphocytes. Fetal white blood cells can even accumulate in maternal spleen or liver and remain there for a long time after labor (Bianchi et al. 1996). Thus, using fetal white blood cells for diagnosis may cause false-positive results. Only fetal nucleated red blood cells (erythroblasts) rapidly disappear after delivery and, consequently, can be correlated with the current pregnancy.
Some authors have reported successful applications of fetal cells for prenatal diagnosis of chromosome aneuploidy (Bianchi et al. 1997). However, low concentration of fetal cells in the maternal circulation (on average, one fetal cell per 106 maternal cells) (Simpson and Elias 1994) combined with a low success rate in sorting of these cells (e.g., by monoclonal antibodies) hampers detection and isolation of fetal cells. Thus, attempts have been tested to detect free fetal nucleic acids instead of fetal cells in maternal blood because fetal cells are exposed to apoptosis (Van Wijk et al. 2000). As maternal cells also are destroyed, the blood of pregnant women contains different DNA fragments (=cell-free DNA) of fetal and maternal origin. The presence of nucleic acids in human plasma has already been described (Mandel and Metais 1948). Prior to 1997, it had not been demonstrated that Y-chromosome-specific sequences could be detected by PCR in plasma or serum of pregnant women carrying male fetuses (Lo et al. 1997).
To detect small amounts of fetal DNA, real-time QF-PCR is the method of choice (Zhong et al. 2000). In cases of known X-linked disease, the possibility to screen for Y-chromosome gene presence in maternal blood allows women to avoid invasive procedures (Rijnders et al. 2004).
Two rounds of PCR were performed using two pairs of primers for Y-chromosome-specific sequence in the present approach (SRY gene region). The efficiency of nested PCR assay used was 91.7%, correlating well with published data (Lo 2000). PCR fragment for Y chromosome was amplified in three DNA samples extracted from plasma or serum of 24 women carrying female fetuses. Two of the three women were primagravida. In these cases it was probable that there was DNA contamination of samples. A third woman had earlier termination of her pregnancy, and in this case SRY-positive result might be due to a probable previous male pregnancy. It may be that the lymphocytes from this previous pregnancy are the source of cell-free DNA. The persistence of fetal DNA in maternal plasma long after delivery has been reported (Invernizzi et al. 2002). A long presence of entire fetal cells in maternal circulation is well known (Bianchi et al. 1996), and it is likely that fetal lymphocytes being degraded by plasma nucleases are followed by the appearance of cell-free DNA.
It is likely that the use of multiplex PCR assay with few Y-chromosome-specific gene markers will allow increased sensitivity and specificity of fetal DNA detection in maternal serum and/or plasma.
In summary, the presence of fetal cell-free DNA in the maternal circulation is a good, low-cost approach for the future development of novel strategies that are expected to provide non-invasive techniques for early prenatal diagnosis.
