Abstract
Fluorescence in situ hybridization analyses were conducted to examine the presence or absence of the 180- and 350-bp knob-associated tandem repeats in maize strains previously defined as “one-knob” or “knobless.” Multiple loci were found to hybridize to these two repeats in all maize lines analyzed. Our results show that the number of 180- and 350-bp repeat loci do not correlate with the number of knobs in maize and that these tandem repeats are not independently sufficient to confer knob heterochromatin, even when present at megabase sizes.
Keywords
K
Two families of tandemly repeated DNA sequences have been identified within the maize knobs. Peacock et al. (1981) showed that a 180-bp tandem repeat is the main component in maize knobs. Ananiev et al. (1998a) more recently reported a 350-bp tandem repeat, TR-1, that is also associated with knobs. A correlation between the size of the knob and the copy number of the 180-bp repeat was previously demonstrated by radioactive DNA in situ hybridization (Dennis and Peacock 1984), such that highly repetitive loci associate with large knobs and less repetitive loci associate with smaller knobs. Fluorescence in situ hybridization (FISH) using the 180- and 350-bp repeats in two different maize lines has shown that these sequences may also be located in chromosomal regions without apparent knob formation (Ananiev et al. 1998a; Chen et al. 2000).
To further investigate the relationship between the locations of tandem repeats and knob formation, we conducted FISH analysis of the 180- and 350-bp repeats in six different maize accessions (Table 1). These accessions are inbred lines or varieties that were previously well characterized as either knobless or having one knob locus based on acetocarmine staining of meiotic pachytene chromosomes (Brown 1949; Crughtai and Steffensen 1987). Therefore, if the 180-and 350-bp repeats are only located in chromosomal regions associated with knobs, we would expect that these two repeats hybridize to either zero or one locus on the metaphase chromosomes of these maize accessions.
Maize accessions used in the FISH study
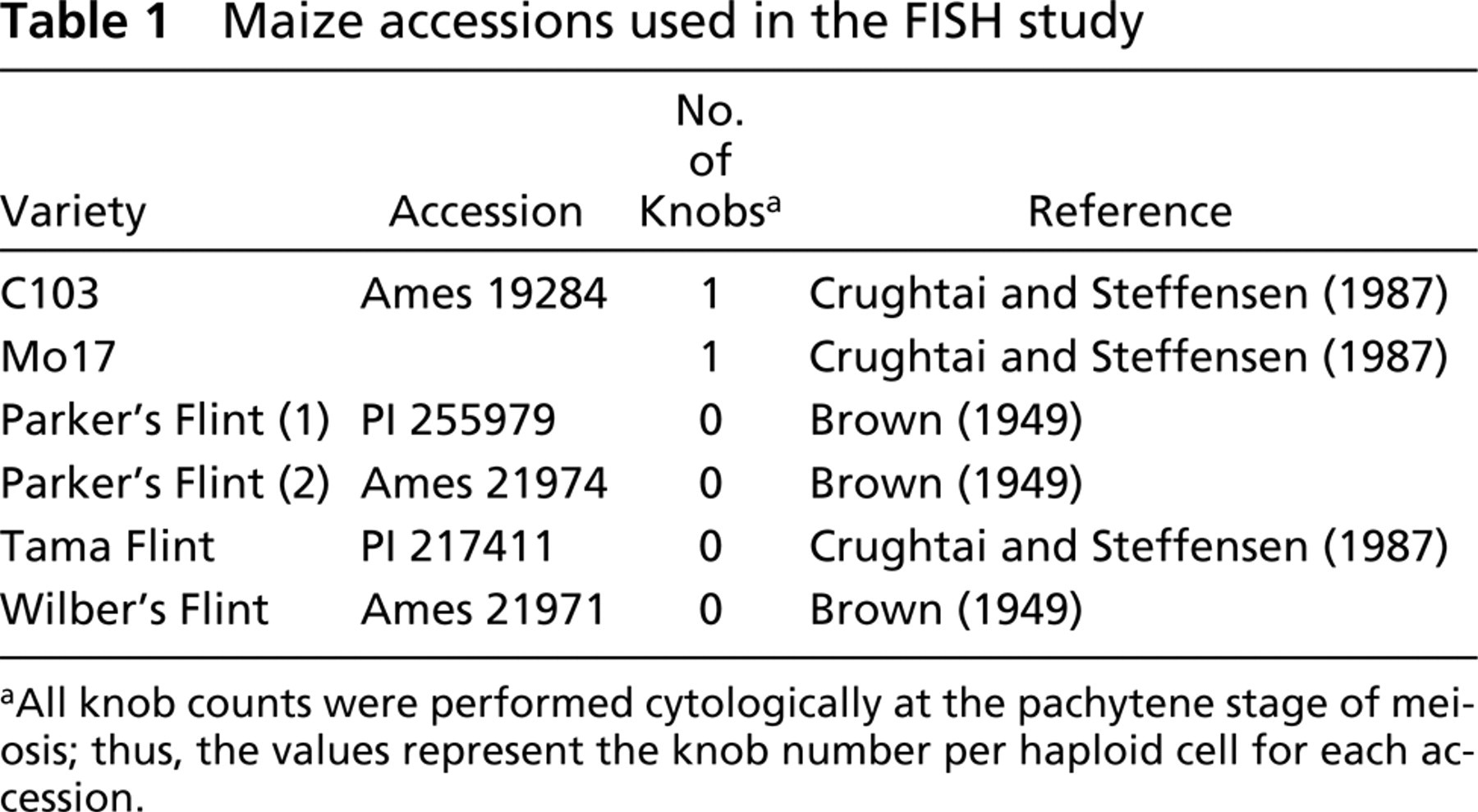
All knob counts were performed cytologically at the pachytene stage of meiosis; thus, the values represent the knob number per haploid cell for each accession.
C103 was classified as a one-knob variety (Crughtai and Steffensen, 1987). We observed one pair of chromosomes hybridized with the 350-bp repeat and two pairs of chromosomes hybridized with the 180-bp repeat (Figure 1A). Similarly, Mo17 was also classified as a one-knob line (Crughtai and Steffensen, 1987). We observed one pair of chromosomes hybridized with the 350-bp repeat (Figure 1B). Another pair of chromosomes showed a very large hybridization site to the 180-bp repeat. At least two other pairs of chromosomes showed weak hybridization to the 180-bp repeat, although the number of chromosomes with weak hybridization signals was not consistent in different cells.
Three knobless lines, Parker's Flint, Wilbur's Flint, and Tama Flint, were used in FISH analysis. We analyzed two different accessions of Parker's Flint. Both accessions contain one pair of chromosomes that hybridize to the 350-bp repeat. The 180-bp repeat probe generated several strong hybridization sites as well as a number of weak sites in the two Parker's Flint accessions (Figures 1C and 1D). Both Wilbur's Flint and Tama Flint showed one pair of chromosomes that hybridized to the 350-bp repeat and several chromosomes that hybridized to the180-bp repeat (Figures 1E and 1F).
The single location of the 350-bp repeat was ideal for fiber-FISH analysis. We analyzed the 350-bp locus in C103 and Mo17, and the microscopic sizes of the fiber-FISH signals were converted into kilobases using a 3.21 kb/μm conversion rate (Cheng et al. 2002). The 350-bp repeat array was estimated to be 842 ± 191 kb in C103 based on measurements of 15 fibers and 1282 ± 136 kb in Mo17 based on 7 fibers. Figure 1G shows two Mo17 signals located in the same microscopic field. The FISH signals derived from both C103 and Mo17 DNA fibers were largely contiguous, but small gaps with consistent locations were observed along the fibers. This indicates that the 350-bp repeat arrays are interrupted by other DNA sequences, most likely invasion of retrotransposons (Ananiev et al. 1998b).
In Wilbur's Flint, the 350-bp locus overlapped with weak FISH signals derived from the 180-bp repeat. Two-color fiber-FISH revealed short, interspersed 180-bp repeat signals within the 350-bp repeat arrays (Figure 1H). Interspersion of these two repeats was also reported in a 12-kb cosmid clone derived from maize chromosome 9 (Ananiev et al. 1998a). Short stretches of DNA sequences within the two repeats show some level of homology (Ananiev et al. 1998a; Hsu et al. 2003). It was suggested that the 350-bp repeat may have evolved from a 180-bp ancestral repeat (Ananiev et al. 1998a; Hsu et al. 2003). Interspersion of the fiber-FISH signals derived from these two repeats supports a common evolutionary origin for the two repeat families.
Previous studies showed that the 180- and 350-bp repeats are the main components of maize knobs (Peacock et al. 1981; Ananiev et al. 1998a,b). A correlation between knob size and the 180-bp repeat content has previously been documented using in situ hybridization (Dennis and Peacock 1984). Chen et al. (2000) observed that some maize 180-bp repeat loci do not associate with knob-like heterochromatin. We demonstrate that all four knobless maize accessions display multiple loci containing these two repeats, suggesting that these repeats do not necessarily confer the heterochromatic knob structure. In Mo17, the size of the FISH signals derived from the 350-bp repeat is significantly smaller than the major pair of FISH signals derived from the 180-bp repeat (Figure 1B). We assume that the single knob in this line is associated with the large 180-bp locus. Thus, the 350-bp array, which contains >1 Mb of DNA, is not associated with a distinct knob feature. These results suggest that the presence of high-copy 180- and 350-bp tandem repeats is not sufficient to induce a knob-like heterochromatic structure. The extreme condensation of knob heterochromatin may be conditioned by a combination of tandem repeats and other unknown factors.
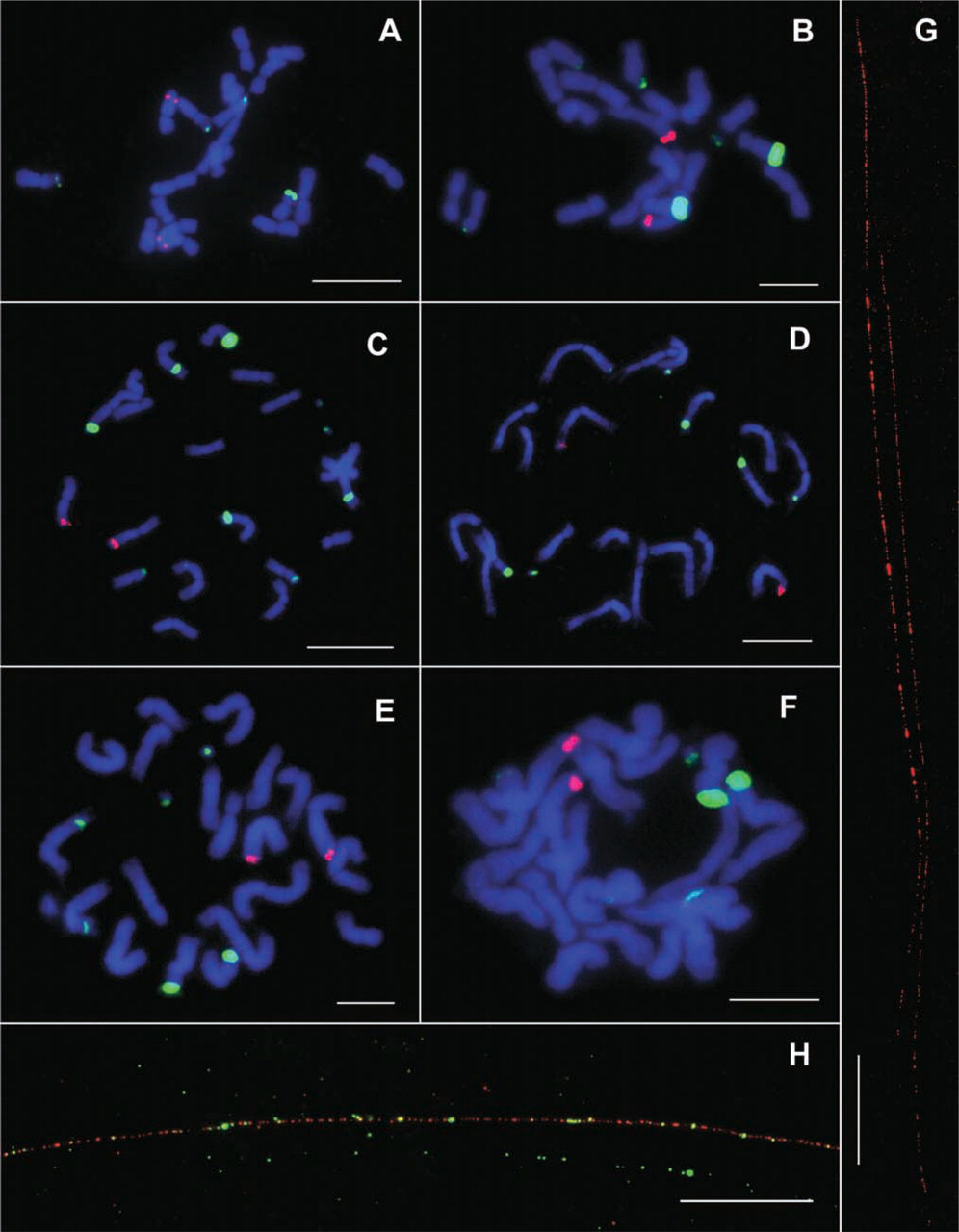
FISH and fiber-FISH analyses of the 180- and 350-bp repeats in one-knob and knobless maize lines. The 180-bp repeat is detected by a green color and the 350-bp repeat by a red color. (
