Abstract
Recently, we were able to show that satellite DNA amplification (satDNA-AMP) is present in advanced prostate cancer. A chromosome microarray study provided first evidence that satDNA-AMP appears to be largely independent of centromere-near/pericentric euchromatic copy number alterations. Therefore, it might be carefully suggested that satDNA-AMP could be a new and independent marker for advanced tumor progression:
Keywords
Introduction
Recent studies by ourselves 1 and others 2 have shown that satellite DNA is not only expressed as RNA, but is also amplified in advanced prostate cancer (PCa). As described in the study by Ariffen et al., 1 amplification of satellite DNA (satDNA-AMP) has already been observed in breast carcinomas, liposarcomas, retinoblastomas, melanomas, and gliomas. The satDNA-AMPs are mostly visible as homogeneously staining regions (HSRs) and less frequently as double minutes (DMs). In PCa, satDNA-AMPs have only been seen as HSRs, probably due to the small number of cases studied and reported for this phenomenon to date. 1
The expression of satellite DNA in the form of RNA has been reported not only in PCas 2 but also in the breast carcinoma cell line HeLa, 3 where the transcription of satellite DNA has also been shown to be essential for the maintenance of centromere structure and function. In addition, satellite RNA has been detected in human tissues under heat shock.4 –6 The latter is of interest in that satellite RNA has been shown to be involved in the recruitment of RNA processing (i.e. splicing factors) during stress response.5,7 It has been repeatedly shown that cellular stress is an inherent feature of carcinogenesis. 8
In addition, RNA expression of satellite DNA has been observed in prenatal development of insects9,10 and mammals.10,11 Interestingly, similarities between cancer and prenatal tissue, such as chromothripsis, chromosome instability, and/or expression of (proto)oncogenes in both cases, 12 have been observed in recent decades. Therefore, the expression of satellite DNA as RNA known from embryonic development,9 –11 which has now also been detected in tumors, 2 is not too surprising. However, the fact that heterochromatic satDNA-AMP in advanced tumors appears to be realized by HSR and/or DM formation has been overlooked until recently. 1
Materials and Methods
In our previous work on 31 primary PCas, 1 we found satDNA-AMPs of D1Z1, D2Z1, D3Z1, D4Z1, D15Z4, D20Z1, and DYZ1 in five cases (16%; see Table 1). In this study, we investigated whether these satDNA-AMPs are associated with nearby localized euchromatic DNA amplifications in the affected five advanced PCas. To this end, DNA was extracted from the formalin-fixed, paraffin-embedded (FFPE) blocks of these five samples and analyzed by 180k chromosome microarray (CMA) according to Aust et al. 13 As the studied PCa samples were 1–2 cm3 10 sections each, 10 µm were sufficient for DNA extraction. The oligo-array method applied here involves comparative hybridization of the tested tumor DNA and a reference genomic DNA (Human Female Genomic DNA; Promega, Mannheim, Germany) on a chip with 170,334 specific oligonucleotide sequences (Agilent Human Genome CGH Microarray 180K; Agilent, Waldbronn, Germany). As analysis software, ‘workbench7’ (CytoGenomics; Agilent, Waldbronn, Germany) was applied. The results were analyzed according to the oligonucleotide position provided by Agilent in the UCSC Genome Browser on Human Feb. 2009 (GRCh37/hg 19) version.
The Five PCas Studied Here, Their Gleason Scores, Their Satellite DNA Amplicons, and Regions With Copy Number Increases in Euchromatin.

Results
The results are shown in Table 1 and summarized in Fig. 1: No nearby euchromatic copy number alterations (CNAs) were detected in the majority of chromosomes affected by satDNA-AMP as HSR (see also Appendix Table 1). This is true for cases 1, 3, 11, and 13 for both affected chromosomes, although chromosome 1 shows gains and losses along its long arm (Fig. 1). In case 4, satDNA-AMPs were found originating from the chromosomes 3 and 4: Whereas chromosome 3 shows a centromere-near gain in copy number for the short arm, there was a pericentric deletion of euchromatin in chromosome 4. In none of the cases was oncogene amplification detected by CMA.
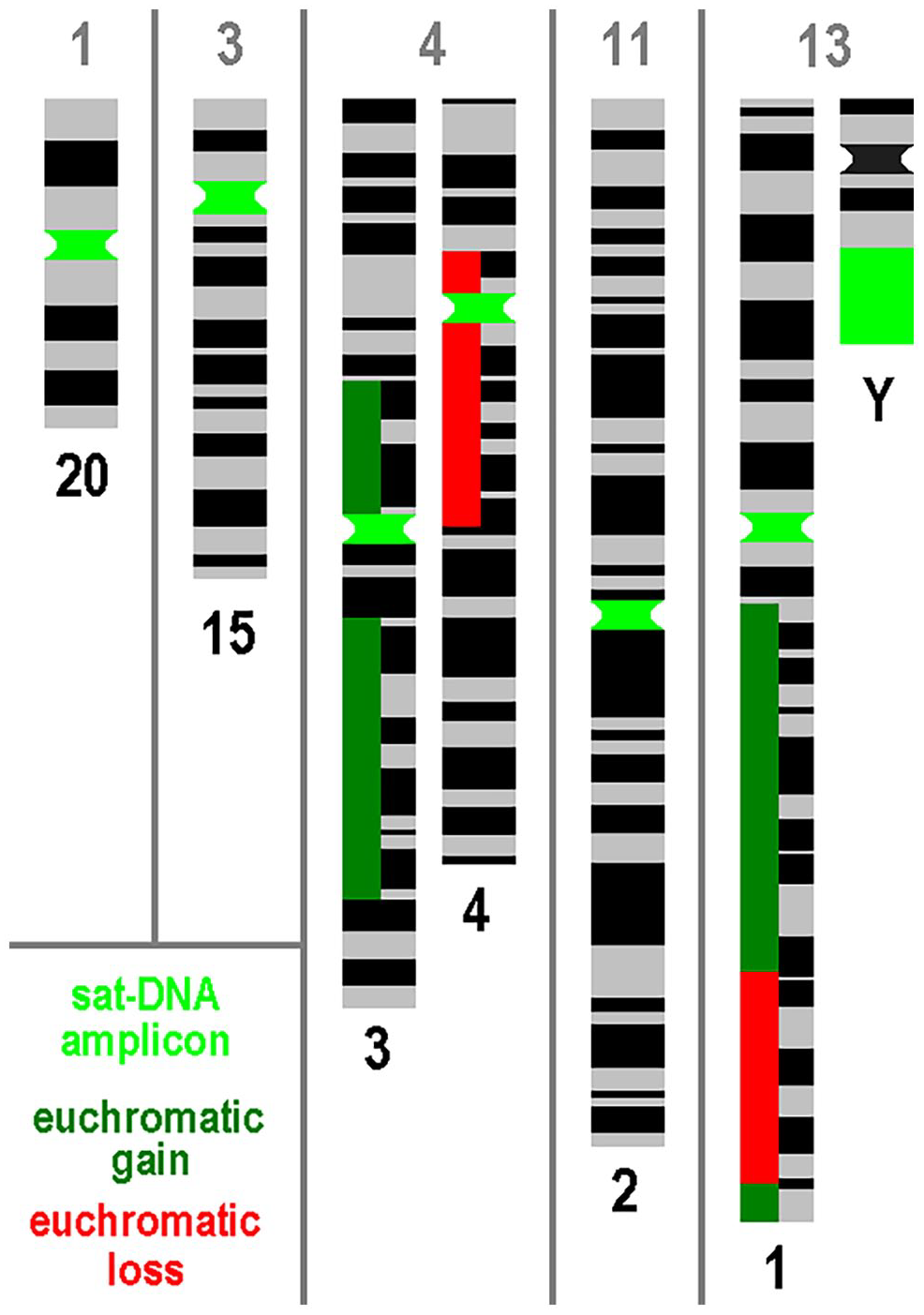
For the five PCas studied here (Nos. 1, 3, 4, 11, and 13), the affected chromosomes are shown with amplicons of satellite DNA (satDNA) together with the euchromatic gains and losses of these chromosomes according to Table 1 and Appendix Table 1. Abbreviations: DNA, deoxyribonucleic acid; PCa, prostate cancer.
In contrast, there were euchromatic centromere-near deletions in case 3 for chromosome 6p, case 4 for Yp and Yq, and euchromatic centromere-near gains in case 3 for chromosome 6p. All these euchromatic CNAs did not trigger satDNA-AMP in these chromosomes.
In total, the following euchromatic CNAs were found as gains two or more times in the five samples analyzed: 1q21q23.1~41, 6p22~21.31p21.1, and 7q11.23q11.23 and as losses: Xp22.32p22.31~11.3 and 8pterp22~11.2.
Discussion
In this study, satDNA-AMPs were not associated with nearby localized euchromatic DNA amplification in any of the five cases examined. Overall, the euchromatic CNAs detected were as expected for PCas.14,15 Only one affected chromosome 4, each showed a moderate gain or loss of centromeric euchromatin (see Fig. 1). Overall, this seems to indicate that satDNA-AMPs may occur largely independently of euchromatic CNAs in advanced tumors. If this is true, the rarely observed coamplification of centromere-related oncogenes, such as MDM2 and D12Z3 amplicons in lipomas, would be the exception rather than the rule for this type of amplicon formation. Should future studies identify satDNA-AMPs as DMs and HSR simultaneously in individual advanced tumors, satDNA-AMP, and amplification of oncogenes 16 would be closely related mechanisms.
It should be noted that satDNA-AMP is not yet routinely accessed or accessible. Satellite DNAs can only be quantified by molecular cytogenetics. 17 CMA, Sanger, and routine Next Generation Sequencing are blind to these DNA segments. This is, as these approaches do not cover heterochromatic regions and/or are not able to align or quantify highly repetitive sequences. 17 Similarly, Ljubić et al. 2 were only able to detect relatively small portions of alpha satellite DNA and not the centromeric alpha, beta, or satellite III DNAs, which are present as higher order repeat units, as studied by Ariffen et al. 1
Of course, the present study has several limitations: (1) It is a spadework study performed in only five cases. In addition, (2) it is known that DNA extracted from FFPE material is partially degraded and results may be less accurate than with cryofixed or fresh postoperative tumor material. 18 Still, as fluorescence in situ hybridization (FISH) and CMA studies on the here studied five FFPE-samples gave informative and easily evaluable results, DNA degradation did not seem to be an issue here. Furthermore, (3) tumor-specific euchromatic CNAs may not be complexly accessible as the tumor may be “contaminated” with normal cells. 19 Although (1) and (2) can only be circumvented in future studies, for (3) it can be said that comingling with normal cells was excluded by evaluation of the used sections by two experienced pathologists (E.J. and B.T.); the tumor percentage was between 75% and 100%. In addition, the Phosphatase and TENsin homolog (PTEN) gene deletion previously found in case 13 by FISH was also detected in the CMA analyses, and no such deletions were detected in cases 1, 3, and 4 by either FISH or CMA. Only in case 11 there was a discrepancy, as the PTEN deletion was found in ~25% of the cells by FISH, which was not visible in CMA (see Appendix Table 1); this can be explained by an insufficient proportion of cells with CNAs. 19
The findings that satDNA-AMP is not only present in advanced PCa samples 1 but satDNA is also expressed in such samples suggest that both events are correlated with each other. Additional evidence that satDNA-AMP is a feature of advanced tumors is provided by its presence in human 17 and murine-advanced tumor cell lines. 20 Thus, it can be cautiously concluded that satDNA-AMP could be a new and independent marker for advanced tumor progression.
In case satDNA-AMP is suited as a marker for advanced tumors, it is possible that not only the presence of satDNA-AMP, but also multiple centromeres due to (partially) polyploid tumors could trigger tumor malignancy via the expression of satDNA. In addition, chromosomal heteromorphisms that cause an excessive expansion of satellite DNA should be investigated to determine whether they predispose to a more severe cancer progression. Further studies in other tumor types should be done to check if satDNA-AMP and satDNA-RNA-overexpression are present there as well in advanced samples.
Footnotes
Appendix
Complete Data of CMA Analyses of the Five Presented Cases.

| Case 1 | ||
| Cytoband | Region Affected [GRCh37] | Kind of CNA |
| 1q21.2q23.1 | chr1:150257376-156860275 | Gain |
| 1q32.1q32.1 | chr1:201732114-203428584 | Gain |
| 5q34q35.1 | chr5:162956347-170262735 | Loss |
| 6p21.31p21.2 | chr6:35381089-39207142 | Gain |
| 7q11.23q11.23 | chr7:72365903-77469625 | Gain |
| 8pter-p22 | chr8:193250-15186829 | Loss |
| 9q33q34.11 | chr9:130052727-133273220 | Gain |
| 11q13.1q13.3 | chr11:64543890-68785854 | Gain |
| 12q24.31qter | chr12:125740414-132321220 | Loss |
| 16pterp13.1 | chr16:333753-5359467 | Gain |
| Case 3 | ||
| Cytoband | Region Affected [GRCh37] | Kind of CNA |
| 1q21.2q21.3 | chr1:150257376-151669638 | Gain |
| 6p22.2p22.2 | chr6:25947549-26285413 | Gain |
| 6p11.2p11.2 | chr6:57246930-57489926 | Loss |
| Xp22.32p22.31 | chrX:4886357-8422893 | Loss |
| Case 4 | ||
| Cytoband | Region Affected [GRCh37] | Kind of CNA |
| 1p36.21p36.13 | chr1:14336693-19510620 | Loss |
| 1q21.2q32.1 | chr1:149103194-203445250 | Gain |
| 2q31.1q31.3 | chr2:177202281-182606581 | Loss |
| 3p14.1p11.1 | chr3:69436836-88183074 | Gain |
| 3q13.32q26.2 | chr3:118575163-167925874 | Gain |
| 4p13p12 | chr4:42495208-46580752 | Loss |
| 4q11q22.3 | chr4:52689101-95362945 | Loss |
| 6p11.2p11.2 | chr6:57270000-57503312 | Gain |
| 6q14q14 | chr6:78979172-79023328 | Loss |
| 8pterp12 | chr8:161472-33254022 | Loss |
| 9pterp22.2 | chr9:204193-17697922 | Loss |
| 9q34.3qter | chr9:139722914-140240476 | Gain |
| 10q24.32qter | chr10:104490230-105279183 | Loss |
| 14q22.3q23.1 | chr14:56058978-58667420 | Loss |
| 14q23.3q24.1 | chr14:65795231-70110204 | Loss |
| 14q24.2q24.3 | chr14:73566341-74384090 | Gain |
| 14q24.3q31.1 | chr14:75412267-79535599 | Loss |
| 14q32.12q32.13 | chr14:92421603-96078236 | Loss |
| 17p13.2p13.1 | chr17:3903561-7744103 | Loss |
| 20p12.1p12.1 | chr20:13370091-14315907 | Gain |
| 21q22.3qter | chr21:43097982-45659989 | Loss |
| Ypterp11.2 | chrY:2537972-10012121 | Loss |
| Yq11q11.222 | chrY:13213476-21098877 | Loss |
| Case 11 | ||
| Cytoband | Region Affected [GRCh37] | Kind of CNA |
| 1p36.32p36.23 | chr1:2575457-8811612 | Loss |
| 16q22.1q22.1 | chr16:66955572-70442715 | Gain |
| 16q23.3q23.3 | chr16:82006620-83778055 | Loss |
| 21q22.2qter | chr21:40158830-42911329 | Loss |
| Xpterp22.33 | chrX:174905-2635590 | Gain |
| Xp22.32p11.3 | chrX:4886357-8422893 | Loss |
| Ypterp11.31 | chrY:124905-2585590 | Gain |
| Case 13 | ||
| Cytoband | Region Affected [GRCh37] | Kind of CNA |
| 1q21.2q41 | chr1:149378207-216090561 | Gain |
| 1q41q43 | chr1:216179405-241432469 | Loss |
| 1q43qter | chr1:242023918-249172869 | Gain |
| 6q14.1q16.2 | chr6:80113727-99769419 | Loss |
| 7p14.1p12.1 | chr7:40295934-53991940 | Loss |
| 7q11.23q11.23 | chr7:73100444-76734076 | Gain |
| 8p23.3p12 | chr8:193250-32336307 | Loss |
| 8q13.3q24.21 | chr8:71086949-119886848 | Gain |
| 8q24.23q24.3 | chr8:137364984-143388158 | Loss |
| 10q23.1q24.31 | chr10:82027464-101901252 | Loss |
| 11q14.1qter | chr11:78541181-134828734 | Loss |
| 12q12q12 | chr12:39213726-42672315 | Loss |
| 12q15q24.31 | chr12:68980497-125740473 | Gain |
| 13q13q31 | hr13:35831261-86543280 | Loss |
| 17q25.3qter | chr17:78787120-80993001 | Gain |
| Xp22.33p22.31 | chrX:4291483-8448318 | Loss |
Abbreviation: CNA, copy number alteration.
Competing Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
Made substantial contributions to conception and design of the study: Thomas Liehr.
Performed data analysis and interpretation: Thomas Liehr, Anja Weise, Jenny Holler, Elena Jamali, Melanie Wolter, Constanze Pentzold.
Performed data acquisition, as well as provided administrative, technical, and material support: Antonio Augusto Ornellas, Gilda Alves, Constanze Pentzold, Melanie Wolter, Thomas Liehr.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
