Abstract
In this study, we describe the development of fluorescent oligonucleotide probes to variable regions in the small subunit of 16S rRNA in three distinct Giardia species. Sense and antisense probes (17–22 mer) to variable regions 1, 3, and 8 were labeled with digoxygenin or selected fluorochomes (FluorX, Cy3, or Cy5). Optimal results were obtained with fluorochome-labeled oligonucleotides for detection of rRNA in Giardia cysts. Specificity of fluorescent in situ hybridization (FISH) was shown using RNase digestion and high stringency to diminish the hybridization signal, and oligonucleotide probes for rRNA in Giardia lamblia, Giardia muris, and Giardia ardeae were shown to specifically stain rRNA only within cysts or trophozoites of those species. The fluorescent oligonucleotide specific for rRNA in human isolates of Giardia was positive for ten different strains. A method for simultaneous FISH detection of cysts using fluorescent antibody (genotype marker) and two oligonucleotide probes (species marker) permitted visualization of G. lamblia and G. muris cysts in the same preparation. Testing of an environmental water sample revealed the presence of FISH-positive G. lamblia cysts with a specific rDNA probe for rRNA, while negative cysts were presumed to be of animal or bird origin.
G
Classical approaches to parasitology, based on host specificity and trophozoite morphometrics, recognized over forty different species of Giardia (Kulda and Nohýnková 1978), but the development of molecular techniques and nucleotide sequencing of nucleic acids have firmly identified six species based on sequence of rRNA or unique morphological characteristics: Giardia lamblia, Giardia ardeae, Giardia muris, Giardia psittaci, Giardia agilis, and Giardia microti (van Keulen et al. 1993,1998; Adam 2000; Thompson et al. 2000). Initial morphological studies of Giardia led Filice (1952) to suggest that three morphological groups exist based on several criteria, including the shape of a microtubule-rich organelle called the median body. These three groups were: Giardia duodenalis, Giardia muris, and Giardia agilis. However, G. dudoenalis (syn. G. lamblia, Giardia intestinalis) should not be used as a species name because the described morphological criteria also fit the description for G. microti, G. ardeae, G. psittaci, characterized as distinct species on the basis of molecular sequencing of rRNA (van Keulen et al. 1993,1998). Therefore, in this manuscript, the term “Giardia lamblia“ will be used to describe isolates of Giardia that have been obtained from humans.
Phylogenetically, rRNA is conserved in function, organization, and sequence, and most cells contain thousands of copies within ribosomes, making this molecule an ideal target for development of genus- and species-specific probes. rRNA sequencing has been used in over 2000 microorganisms and cells, including five different species of Giardia, to establish phylogenetic relationships (van Keulen et al. 1993,1998). rRNA constitutes 90% to 95% of cellular RNA and contains three forms, namely a 16S small subunit (SSU), a 5S SSU, and a 23S large subunit (Waters and McCutchan 1990). Although rRNAs are highly conserved in structure, most of the diversity in nucleotide substitutions and deletions occurs in Giardia spp. within the 16S SSU, and these nucleotide alterations tend to be concentrated in nine variable regions.
Molecular approaches to characterizing or detecting Giardia spp. have included chromosomal band analysis (Campbell et al. 1990), PCR analysis, detection of heat shock protein genes (Abbaszadegan et al. 1992; Mahbubani et al. 1992), genetic analysis of glutamate dehydrogenase locus (Monis et al. 1996), sequence variation in rRNA and rDNA (Weiss et al. 1992, van Keulen et al. 1993,1998), sequence of giardin genes (Caccio et al. 2002), and the use of oligonucleotide microarrays (Wang et al. 2004). In the last two decades, non-radioactive hybridization methods using enzymatic or fluorescent markers have proved to be useful tools for multiple labeling of nucleic acid targets (Höfler, 1987; Nederlof et al. 1990; Speel et al. 1994). Detection of multiple mRNAs and antigens within the same cell or tissue has been described (Dirks et al. 1991; Trembleau et al. 1993), and our laboratory has also used fluorescent in situ hybridization (FISH) for the detection of 16S SSU rRNA in fixed Giardia trophozoites in archival pathology specimens of human small intestine (Macechko et al. 1998).
The aim of this study was to develop FISH methods for detecting rRNA in Giardia spp. cysts. Oligonucleotides specific for rRNA-variable regions in different species of Giardia were labeled with haptens or fluorochromes. A method was developed for identifying Giardia spp. cysts in environmental samples using immunochemical localization of cyst wall antigens, and speciation of these cysts was accomplished by simultaneous FISH with multiple fluorochrome-labeled probes emitting at different wavelengths.
Materials and Methods
Trophozoites and Cysts of Giardia
Trophozoites of G. lamblia (human isolates) and G. ardeae (from great blue heron) were grown in culture on a modified TYI-33 medium as previously described (Erlandsen and Bemrick 1987; Erlandsen and Rasch 1994; Macechko et al. 1998). Strains of G. lamblia included Nick, AB-Was, LT, Cam, MRr4, R, Egyp, C-1182, WB, and NO, and these strains were provided by Drs. Ted Nash (National Institutes of Health, Bethesda, MD), E.A. Meyer (Oregon Health Sciences University, Portland, OR), and Judy Issac-Renton (University of British Columbia, Vancouver, Canada) and tested for hybridization with rDNA oligonucleotide probes. Cysts of G. lamblia were obtained from Dr. Frank Schaeffer III, U.S. Environmental Protection Agency, Cincinnati, OH, and G. muris cysts were isolated from CF-1 mice inoculated with G. muris as previously described (Schupp and Erlandsen 1987). Environmental water samples containing Giardia spp. cysts were also obtained from treated drinking water during the investigation of a waterborne outbreak of giardiasis in Temagami, ON, Canada (Wallis et al. 1998).
DNA Probe Synthesis and Labeling
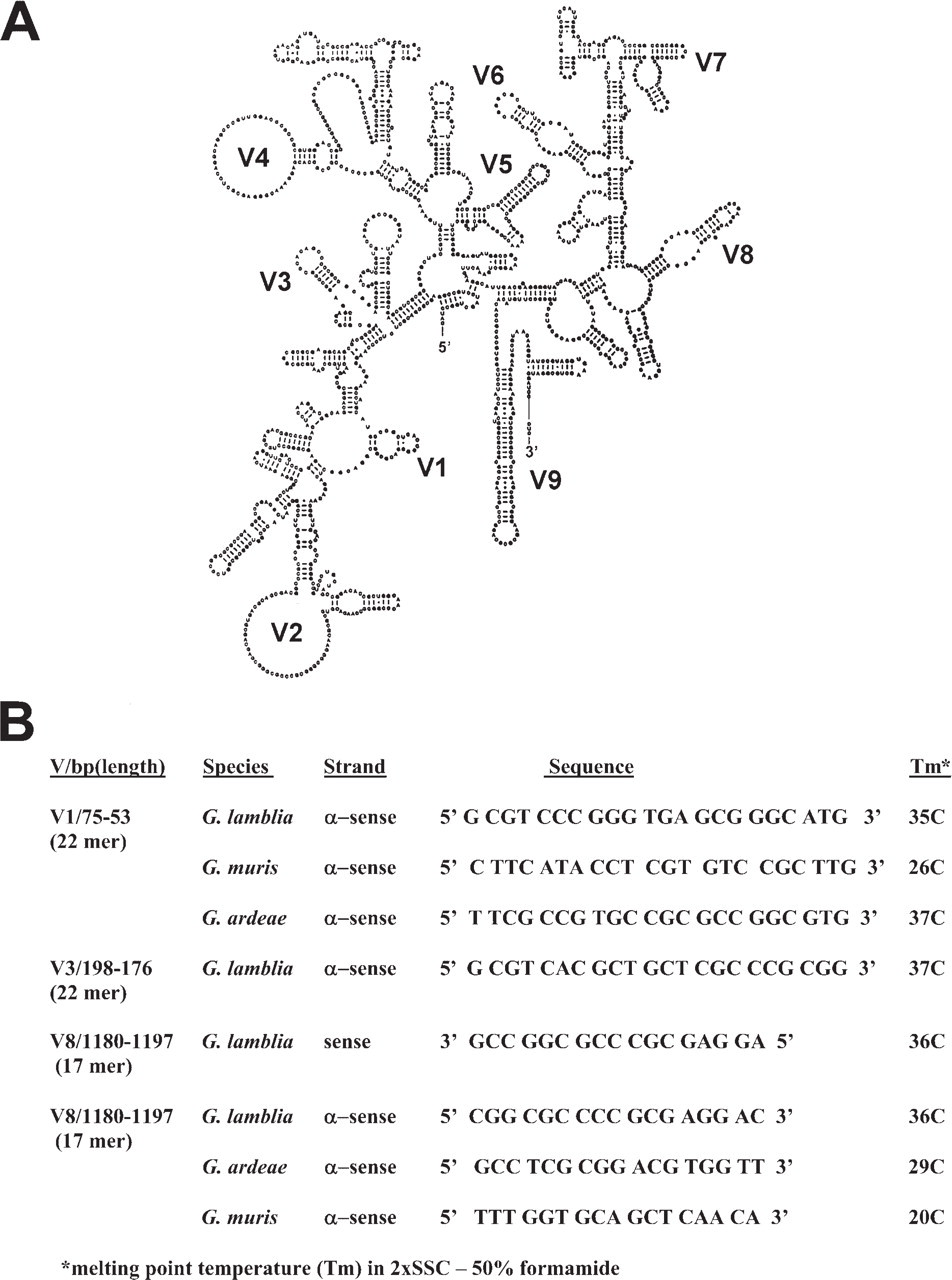
Sense and antisense oligonucleotide probes (17–22 mer) corresponding to rDNA in variable regions 1, 3, and 8 of the 16S SSU rDNA molecule (Figure 1) were synthesized on an Applied Biosystems 394 DNA synthesizer in the Microchemical Facility at the University of Minnesota. Probes were tailed at the 3’ end with digoxygenin-11-UTP and dATP with terminal transferase (Boehringer Mannheim, Indianapolis, IN) following the manufacturer's instructions to keep tail length below 100 base pairs, or oligonucleotides were labeled via a 5’ aminolink with one of the carboxymethylindocyanine dyes, including, FluorX-CTP (absorption 494 nm; emission 520 nm), Cy3-CTP (absorption 552 nm; emission 567 nm), or Cy5-CTP (absorption 648 nm; emission 665 nm) (Biological Detection Systems, Inc., Pittsburgh, PA; Amersham, Chicago, IL). All probes were purified by high-pressure liquid chromatography before use. The melting temperature (Tm) of each probe was calculated based on hybridization in 2X SSC (1X SSC = 0.15 M sodium chloride, 0.015 M sodium citrate) in 50% formamide.
Cell Fixation and Permeabilization
Trophozoites were attached to poly-
Prehybridization and Hybridization
Air-dried slides were rehydrated in 0.1 M PBS containing 7.5% sucrose, and charge was blocked by immersing for 10 min in 0.25% acetic anhydride in 0.1 M triethanolamine at pH 8.0. Slides were then rinsed for 2 min in 2X SSC, followed by dehydration in an ascending ethanol series (50%, 70%, 95%, 100%), and air dried. Cells were rehydrated in prehybridization mix [2X SSC, 50% formamide, 5X Denhardt's solution (0.02% polyvinylpyrrolidone, 0.02% Ficoll, and 0.02% bovine serum albumin), and 1 mg salmon sperm DNA] for 60 min at room temperature. Oligonucleotide probes (1 ng/μl) in 50% formamide in 4X SSC were denatured by boiling for 10 min, cooled immediately to 4C, and then added to slides that were heated for an additional 10 min at 85C, followed by hybridization overnight at 37C in moist chambers. Following hybridization, slides were rinsed sequentially in 4X SSC, 2X SSC, and 1X SSC.
(
Alkaline Phosphatase Incubation for Digoxygenin-labeled Probes
After the last stringency wash, slides were immersed for 2 hr in a blocking solution (0.1 M Tris, 1% normal sheep serum, 0.3% dry milk, 0.1% Triton X-100, and 0.05% sodium azide) on a slow rotary shaker. The slides were drained and then incubated overnight with a 1:100 dilution of sheep anti-digoxygenin Fab fragments conjugated with alkaline phosphatase (Boehringer Mannheim). Slides were then rinsed at room temperature for 30 min with TBS, pH 7.0, followed by 30 min in PBS, pH 8.0. Alkaline phosphatase activity was revealed by incubation in 5-bromo-4-chloro-3-indolyl phosphate and nitroblue tetrazolium. The enzyme staining reaction was stopped by incubating the slides in 10 mM Tris containing 0.1 mM EDTA at pH 8.0, and slides were mounted in an aqueous medium.
Immunocytochemistry and Multiple rDNA Hybridization
Multiple labeling of rRNA in cysts was accomplished by mixing 1 ng/μl each of oligonucleotide probes specific for G. lamblia (Cy5 conjugate) and G. muris (FluorX conjugate) and denaturing the probes by boiling for 10 min. These probes were cooled to 4C, and the mixture was applied to slides, incubated for 10 min at 85C, and hybridized overnight at 37C. Following stringency washes, single rDNA hybridization with Cy3-conjugated probes were directly immunostained for cyst wall antigens using a fluorescein-conjugated mouse monoclonal antibody (RD348; Meridian Diagnostics, Cincinnati, OH), whereas double rDNA hybridization with Cy5 and FluorX oligonucleotide conjugates were counterstained for cyst wall antigens using rabbit anti-Giardia lamblia cyst antibody (GLMB; provided by Judy Sauch, Environmental Protection Agency, Cincinnati, OH) and sheep-anti-rabbit IgG conjugated with Cy3.
Controls
Hybridization controls consisted of: (a) omitting the specific probe or use of unlabeled probe; (b) using sense and anti-sense probes for the same variable rDNA region; (c) RNase (100 μg/ml) digestions of cells in 2X SSC for 30 min at room temperature, oxidized RNase (no enzyme activity) at 100 μg/ml in 2X SSC for 30 min at room temperature; (d) use of 100:1 ratio of unlabeled probe to fluorochrome-labeled probe for hybridization; (e) exposure only to concentrations of free Cy3 dye equivalent to that in labeled probe to determine whether nonspecific staining occurred via absorption of free dye; (f) use of high-stringency (0.1X SSC at either 50C or 65C, depending on probe) exceeding probe Tm by 5C to 10C; and (g) comparing staining of specific oligonucleotide probes with other known Giardia spp.
Microscopy
A Zeiss photomicroscope (Carl Zeiss; Thornwood, NY) equipped with brightfield, phase contrast, and Nomarski optics was used with 20X, 40X oil, and 63X oil planapochromatic objective lenses for evaluating digoxygenin-labeled probes. Fluorescent imaging of conjugated probes was accomplished with a BioRad MRC-600 laser scanning confocal microscope (BioRad; Hercules, CA) equipped with a dual krypton-argon laser. Enhancement of images was achieved by processing in Confocal Assistant, a program developed by Dr. Todd Brelje at the University of Minnesota.
Results
Based on evolutionary comparisons in the SSU of rDNA in Giardia spp. (van Keulen et al. 1993), differences in oligonucleotide sequences in variable regions of the SSU were predicted as likely candidates for genus- and species-specific probes (Figure 1). Initial experiments were conducted using a 17-mer oligonucleotide sense probe in variable region 8. The probe was enzymatically tailed at the 3’ end with terminal transferase and digoxygenin-11-UTP plus unlabeled dATP to minimize added tail length. Hybridization of this sense probe to G. lamblia trophozoites and detection with anti-digoxygenin-AP Fab fragments and color development in nitroblue tetrazolium resulted in intense labeling of nuclei (Figures 2A and 2B). Hybridization signal with the same probe in G. lamblia cysts was reduced, highly variable, and seemed to be related to poor penetration of antibody markers through the semipermeable cyst wall (data not shown).
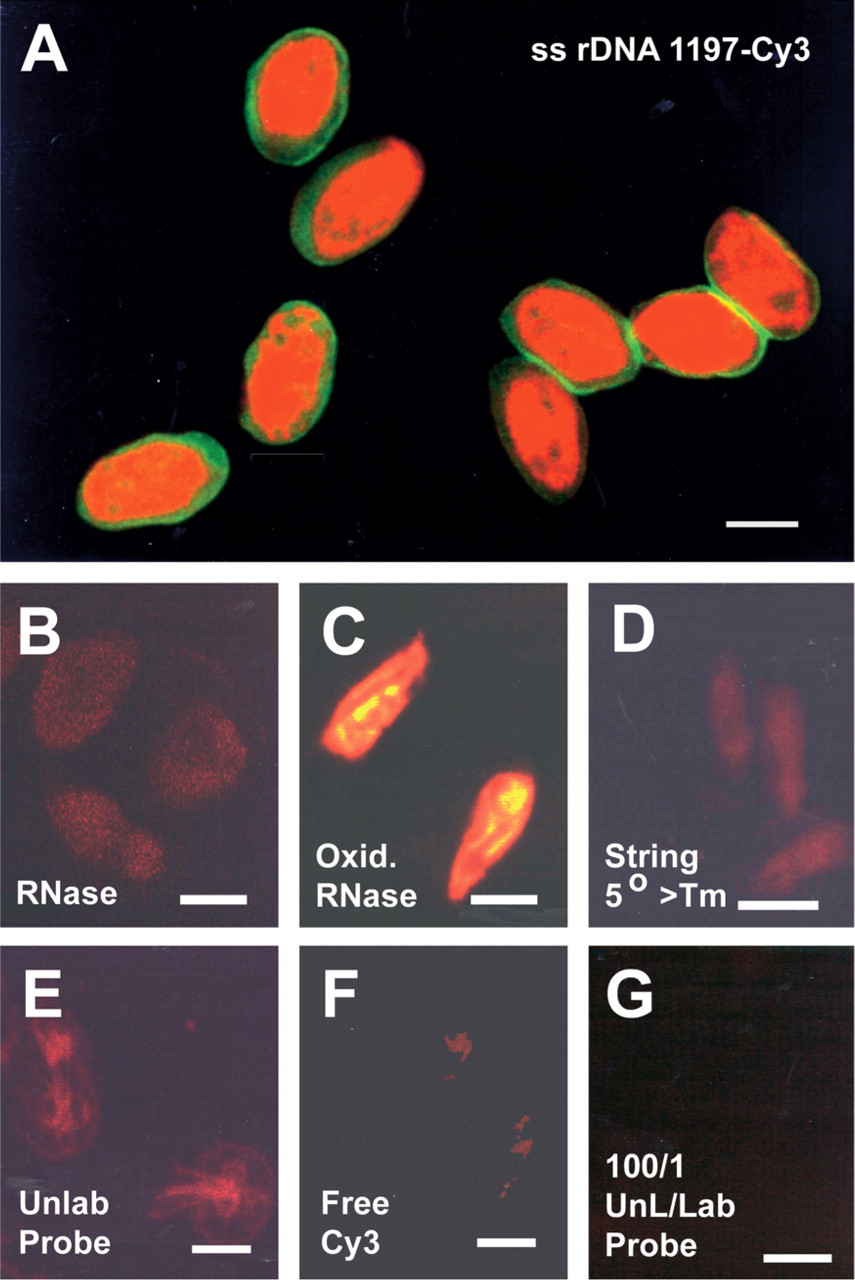
Hybridization of the antisense 17-mer oligonucleotide (1197–1180) probe in variable region 8, but labeled at the 5’ end with Cy3, produced intense and uniform labeling of cytoplasm in G. lamblia cysts and trophozoites (Figures 2C and 2E). In trophozoites, hybridization signal for rRNA was also detected within nuclei. Immunohistochemistry with fluoroscein-conjugated monoclonal antibody (RD348) specific for Giardia cyst wall antigen combined with FISH for rRNA (antisense probes 76–53, 198–176, and 1197–1180; Cy3-labeled at the 5’ end) successfully demonstrated the peripheral location of the cyst wall (green) surrounding the intense hybridization signal (red) in the cyst cytoplasm (Figure 3A). Controls employed for specificity of rDNA probes included the elimination of hybridization signal by digesting cells with RNase (Figure 3B), no diminished hybridization staining by using oxidized RNase, which lacks enzyme activity (Figure 3C), the use of high stringency in 0.1X SSC at temperatures exceeding the Tm by 5C (Figure 3D), the lack of hybridization signal when unlabeled probe was used (Figure 3E), the lack of binding of free Cy3 dye to cyst cytoplasm (Figure 3F), and the elimination of specific hybridization signal when a ratio of 100:1 (unlabeled to labeled) Cy3-labeled rDNA probe was employed (Figure 3G).
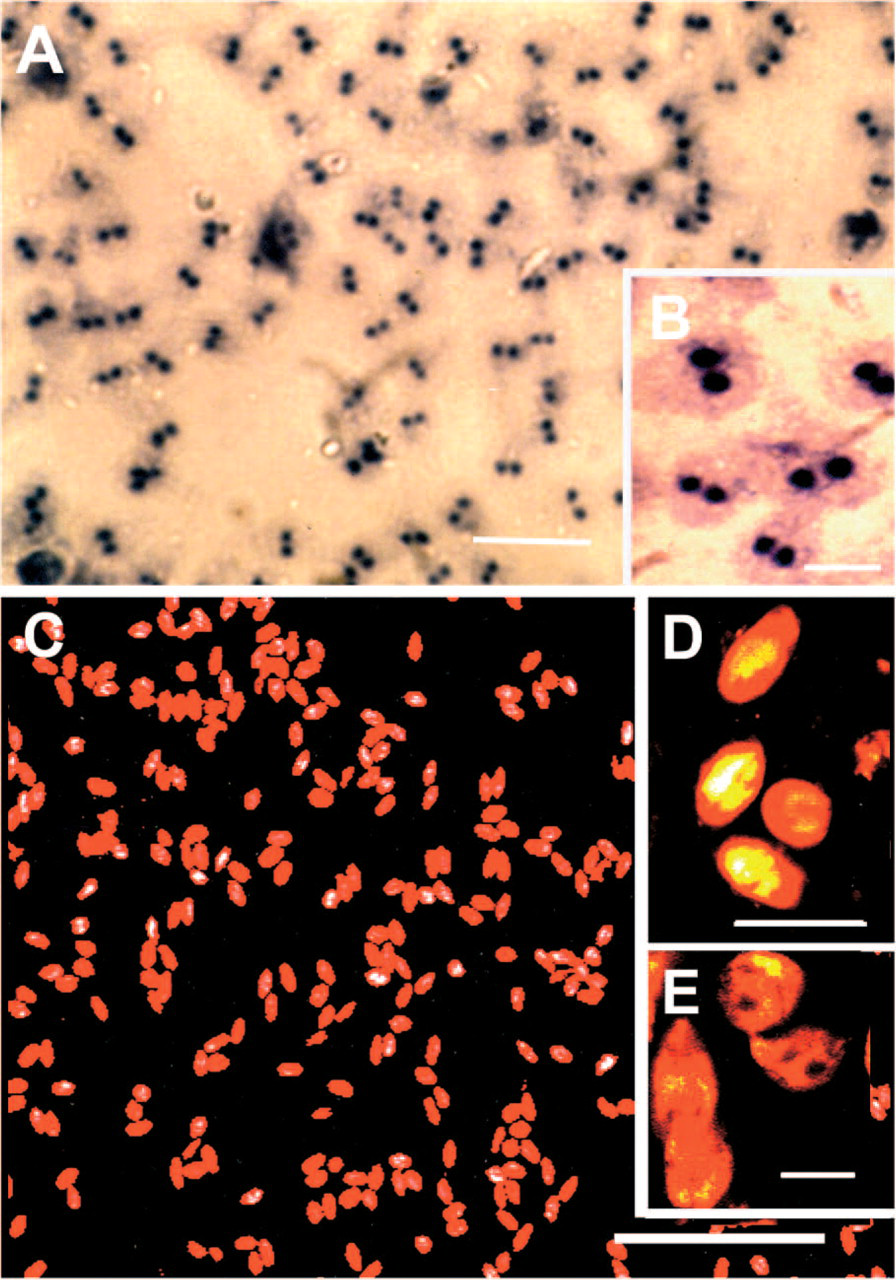
In situ hybridization of 17-mer SSU rDNA oligonucleotide probe (1197–1180) in G. lamblia cysts and trophozoites. (A) Fluorescent in situ hybridization (FISH) of G. lamblia trophozoites with sense rDNA probe tailed at 3’ end with digoxygenin-11-UTP and detected with sheep anti-digoxygenin antibody-alkaline phosphatase conjugate. Using sense probe, intense staining for alkaline phosphatase occurred in nuclei. (
Detection of rRNA in Giardia cysts by fluorochrome-labeled antisense oligonucleotide probes was achieved with cysts freshly isolated from fecal samples (human isolate, mouse isolate), and could also be accomplished with cysts stored in formalin for up to nine months (Figure 4A) with no decrease in hybridization signal. Experiments on specificity of oligonucleotide probes for individual species of Giardia clearly showed that the G. lamblia rDNA probes produced strong hybridization signals in trophozoites isolated from humans (ten strains tested), and lack of hybridization to G. muris cysts or G. ardeae trophozoites (Figure 4B); a lack of cross-reactivity was also observed with G. psittaci and G. microti cysts (data not shown). The G. muris oligonucleotide probes tested produced intense hybridization signal in cysts from mice (Figure 4C), but were entirely negative in production of signal in G. lamblia cysts (Figure 4D) or trophozoites of G. ardeae (Figure 4E).
Multiple labeling of populations of G. lamblia and G. muris cysts was investigated by combining the G. lamblia-specific probe labeled with Cy5 and the G. muris-specific probe labeled with FluorX, and performing hybridizations simultaneously (Figures 5A and 5B). Use of relatively pure or enriched populations of these two cysts resulted in minimal background staining. Intense specific hybridization signals for rRNA were detected in the cytoplasm of G. lamblia and G. muris. Giardia cysts could be identified by strong specific cytoplasmic hybridization signals, surrounded by an immunostained cyst wall (red) indirectly immunostained with rabbit anti-GLMB and goat anti-rabbit IgG labeled with Cy3.
To test the effectiveness of fluorchrome-labeled rDNA probes for cyst detection, two types of environmental samples were investigated. G. lamblia cysts of human origin were mixed with mouse (Figure 5C) or human fecal material (Figure 5D). High backgrounds of nonspecific staining of debris was observed with the Cy3-labeled SSU rDNA probe (75–53); however, recognition of Giardia cysts was readily accomplished at low magnification due to the combination of immunolabeling of the cyst wall for genus identification with the strong hybridization signal with species-specific oligonucleotide rDNA probes. In a second group of samples obtained from Canadian environmental water samples, combined immunostaining for Giardia cyst wall antigens and hybridization with a G. lamblia-specific rDNA probe (198–176) demonstrated the presence of G. lamblia cysts (Figure 5E) that were previously identified as genotype A by van Keulen et al. (2002), but also detected the presence of immunostained Giardia cysts that were negative for hybridization with the G. lamblia-specific rDNA probe and were interpreted as being of animal or bird origin (Figure 5F).
Discussion
The results presented here clearly demonstrate that variable regions in the 16S SSU of rRNA can be used to design species-specific rDNA probes for FISH detection of different Giardia spp. Contrary to the dogma that mRNA is unstable due to intrinsic RNase activity, cellular rRNA binds ribosomal proteins and is extremely stable and resistant to degradation. Data for oligonucleotide sequence-specific probes in three different variable regions (V1, V3, and V8) in rRNA from G. lamblia, G. muris, and G. ardeae were obtained from direct comparisons of sequence alignment of the entire 16S SSU for these three species (van Keulen et al. 1993). The availability of enzymatic labeling of either the 3’ or 5’ ends of oligonucleotides enabled us to make enzyme-labeled or fluorochromelabeled specific rDNA probes for individual Giardia probes. Detection of Giardia cysts with enzymatic methods was less desirable because of variations and irreproducibility of staining. Therefore, our efforts were concentrated on developing fluorochome-labeled oligonucleotide probes for species identification and on using immunochemical methods for recognition of cyst wall antigens. Combined use of immunochemical methods and FISH was valuable in examining environmental samples in which nonspecific staining similar to the wavelength of emission for the fluorochrome probe occurred. Strong nonspecific staining with fluorchrome-labeled rDNA probes for Giarda can be misinterpreted as a false positive. However, this misinterpretation was avoided by using genus-specific immunochemical identification of the cyst wall as the criterion for cyst recognition, and then evaluating the cyst cytoplasm for positive staining with Giardia species-specific SSU rDNA probes. The specificity of the rDNA probes to 16S SSU rRNA was demonstrated by use of both methodological and specificity controls. Treatment of cells with RNase and oxidized RNase indicated that rRNA was the target for the antisense SSU rDNA probes, and FISH with free fluorochrome dye and high ratios of unlabeled to labeled probe supported the need for fluorochrome-labeled oligonucleotide for strong FISH staining. Species specificity for rDNA probes to variable regions 1, 3, and 8 were established by testing each species probe against molecularly identified Giardia spp. isolated from other non-primate animals and birds, as demonstrated by the sequence of 16S SSU rRNA (van Keulen et al. 1993,1998).
FISH of G. lamblia cysts and controls with Cy3-labeled 22-mer SSU rDNA oligonucleotide probe (75–53, variable region 1). (
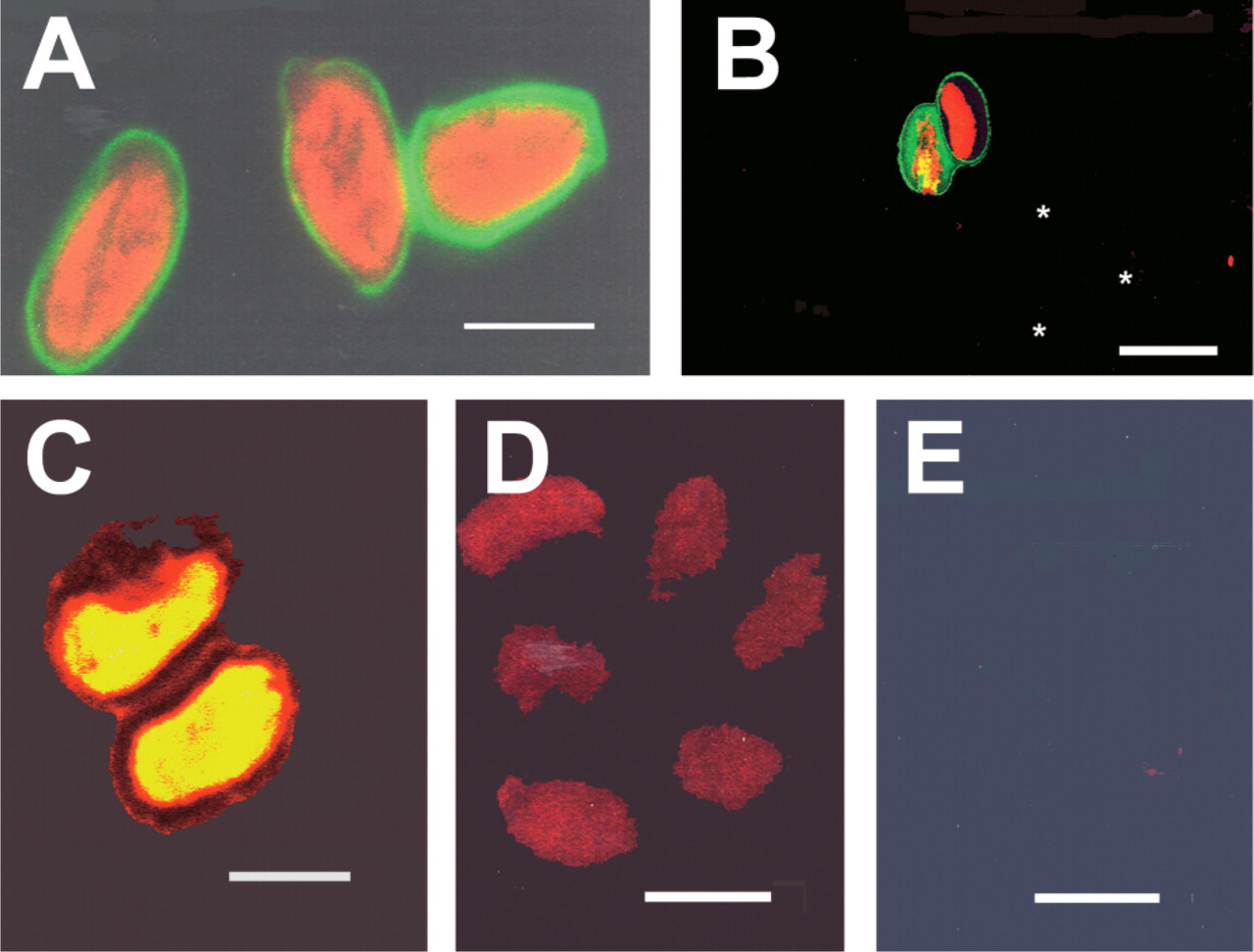
Specificity of SSU rDNA probes (antisense 75–53) for Giardia spp. (
In the last decade, genetic analysis has revealed considerable levels of diversity in G. lamblia (Thompson et al. 2000). Isolates of G. lamblia fall into two major assemblages (A, B) based on a unique trinucleotide in the 16S SSU rRNA molecule (Homan et al. 1992; van Keulen et al. 1995; Thompson et al. 2000); and more recently, primers specific for PCR amplification of the triose phosphate isomerase gene distinguished between assemblages A and B (Wang et al. 2004). Slight variations within each assemblage have been detected, and attempts to characterize which assemblage contains Giardia strains that are pathogenic in humans have led to contradictory results (Homan and Mank 2001; Read et al. 2002). Design of fluorescent oligonucleotide probes to distinguish multiple species of Giardia, and also to detect subtypes in assemblages within the G. lamblia species, would be valuable for epidemiologic evaluations of outbreaks of giardiasis and to address the question of zoonotic potential for this disease.
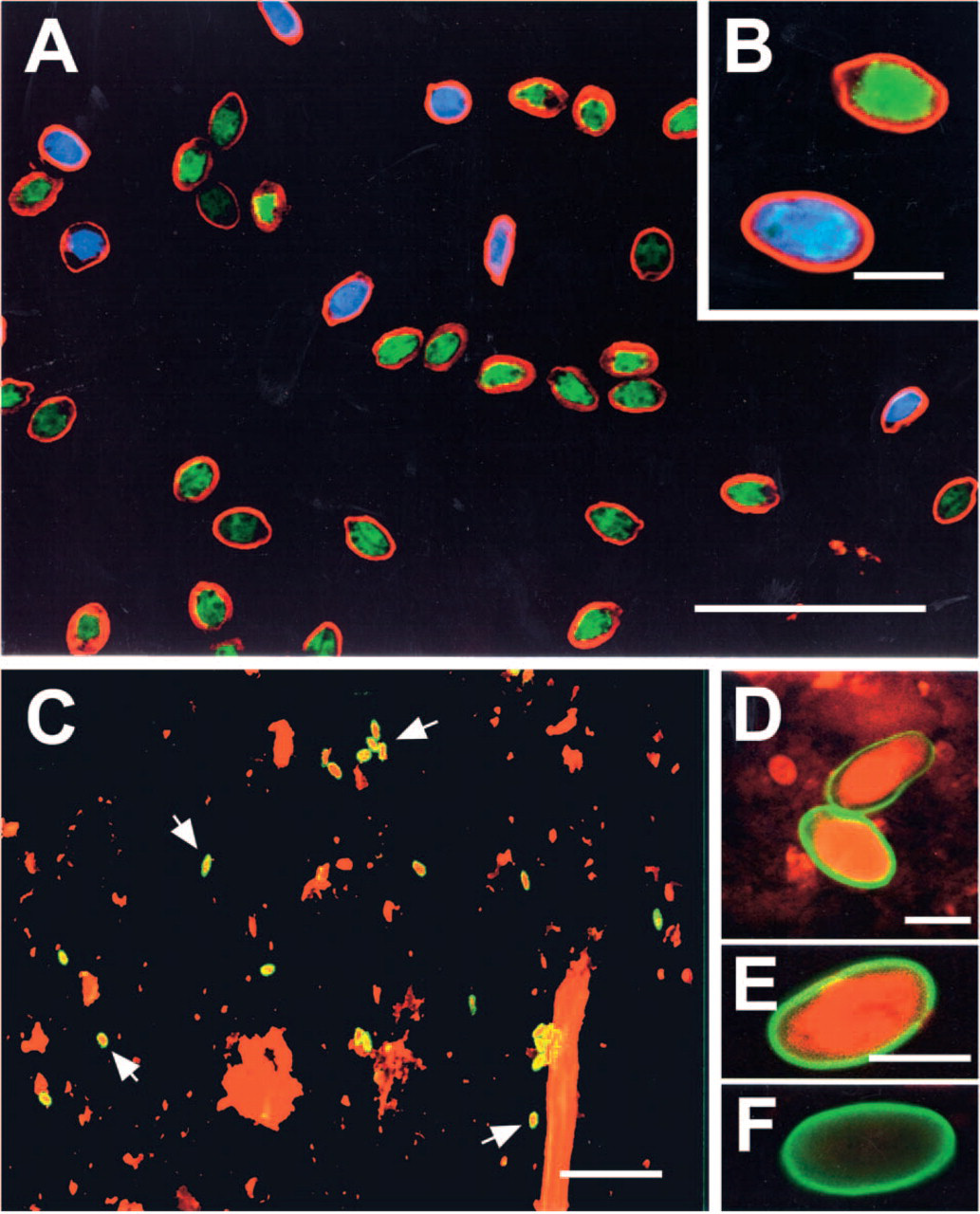
Multiple FISH of G. lamblia and G. muris cysts with specific SSU rDNA probes (75–53) conjugated with Cy5 (blue) and FluorX (green), respectively. (
Detection of Giardia in environmental water and fecal samples has been investigated using PCR amplification with gene probes for giardin genes (Mahbubani et al. 1992; Caccio et al. 2002), SSU rRNA (Abbaszadegan et al. 1991; Rochelle et al. 1997; Ghosh et al. 2000), 18S rRNA (Weiss et al. 1992), mRNA (Kauener and Stinear 1998) triose phosphate isomerase (McIntyre et al. 2000; Amar et al. 2003), variant-specific surface protein genes (Ey et al. 1997), and genes for heat shock protein (Rochelle et al. 1997) have been used to detect Giardia cysts with a sensitivity of less than one cyst per milliliter. However, detection in environmental samples was highly variable, occasionally giving negative results even when 105 cysts were present, or differentiation between known Giardia species was not achieved. Major drawbacks in using gene probes on environmental samples have been nonspecific absorption of probes on debris within the samples, generation of nonspecific DNA fragments, and inhibition of amplification due to contaminants within the sample.
In this study, we have shown that FISH with rDNA probes can positively identify G. lamblia in water samples under outbreak conditions (Figure 5E). The application of this technique to routine monitoring could be an important improvement in the analysis of water samples, especially using the enhanced recovery offered by US Environmental Protection Agency Method 1623, because results could be obtained more quickly than by PCR methods. The identification of cysts that are not G. lamblia and do not correspond to isolates known to infect humans would help refine risk evaluation and would prevent unnecessary boil water advisories. It would also require that “zero tolerance” regulatory policies such as that adopted in Canada during the late 1990s (since rescinded) need to be re-evaluated to allow for the occurrence of species that are not infective to humans.
In summary, we believe that further refinement of our microscopic FISH approach using immunochemical identification of genotypic markers (cyst wall antigens) in combination with species-specific fluorescent oligonucleotides may provide a convenient and specific method for the simultaneous detection of different G. lamblia strains or other species (animal or bird origin). Use of multiple fluorescent probes for genus- and species-specific genes, combined with either microarray technology (Wang et al. 2004) or the use of radio-labeled fluorescent DNA probes (Dauwerse et al. 1992) for FISH, together with spectral imaging, should permit simultaneous evaluation of multiple genes within the same cyst. This approach should prove to be valuable for epidemiologic investigation of environmental water samples or clinical human fecal samples.
Footnotes
Acknowledgements
This work was supported in part by the U.S. Environmental Protection Agency (Grant EPA 1R821404). The authors wish to thank Mr. Walt Jakubowski for discussions.
