Abstract
Lipid peroxidation is a common feature of liver diseases, especially non-alcoholic fatty liver disease (NAFLD). There are limited validated tools to study intra-hepatic lipid peroxidation, especially for small specimen. We developed a semi-quantitative, fully automated immunohistochemistry assay for the detection of 4-hydroxynoneal (4-HNE) protein adducts, a marker of lipid peroxidation, for adaptation to clinical diagnostics and research. We used Hep G2 cells treated with 4-HNE to validate specificity, sensitivity, and dynamic range of the antibody. Staining and semi-quantitative automated readout were confirmed in human needle-biopsy liver samples from subjects with NAFLD and normal liver histology. The ability to detect changes in lipid peroxidation was tested in paired liver biopsies from NAFLD subjects, obtained before and after 4 weeks of treatment with the antioxidant vitamin E (ClinicalTrials.gov NCT01792115, n=21). The cellular calibrator was linear and NAFLD patients had significantly higher levels of 4-HNE adducts compared to controls (p=0.02). Vitamin E treatment significantly decreased 4-HNE (p=0.0002). Our findings demonstrate that 4-HNE quantification by immunohistochemistry and automated image analysis is feasible and able to detect changes in hepatic lipid peroxidation in clinical trials. This method can be applied to archival and fresh samples and should be considered for use in assessing NAFLD histology.
Introduction
Lipid peroxidation is the process in which oxidants, such as free radicals, react with double bonds in the carbon chain of fatty acids, especially polyunsaturated fatty acids. 1 In food production, lipid peroxidation products lead to distinctive flavors, for example, in dried cured meats such as Parma ham, 2 but within the human body these can lead to cell injury through modification of macromolecules. Lipid peroxidation products can diffuse from their site of origin and modify nucleic acids as well as proteins. 3 DNA modifications by lipid peroxidation can lead to mutations, 4 while protein modifications can lead to inflammation and apoptosis. 5 Lipid peroxidation products can be detected by a variety of methods including derivatization and high performance liquid chromatography, 6 analysis by liquid chromatography coupled to tandem mass spectrometry 7 or ELISA. 8 Most methods require a relatively large sample, specialized equipment and lead to the destruction of the sample. Quantification of lipid peroxidation by immunohistochemistry can be done with limited sample size in a non-destructive fashion while allowing for the analysis of histomorphology.
4-hydroxynonenal (4-HNE) is one of many alkenals formed during lipid peroxidation of polyunsaturated fatty acids, 9 and 4-HNE protein adducts have been described as a reliable and stable marker of lipid peroxidation in liver disease. 10 4-HNE protein modifications occur when the aldehydic group of 4-HNE is added to an amino group or cysteine, lysine, or histidine residues of the protein 11 and can be detected using immunohistochemistry. Visualization of 4-HNE adducts by immunohistochemistry in liver biopsies has been previously described12,13; however, rigorously validated assays have not been performed. In the current work, we aimed to develop a validated, semi-quantitative, fully automated immunohistochemistry assay that can be easily adapted to routine clinical or research applications. We then applied our newly developed assay to quantify 4-HNE adducts in liver biopsies of patients with non-alcoholic fatty liver disease (NAFLD) pre- and post-treatment with the lipid soluble antioxidant vitamin E (RRR-α-tocopherol) to address, for the first time, whether antioxidant treatment can decrease intrahepatic lipid peroxidation.
Materials and Methods
Cell Culture
The human hepatoma cell line, Hep G2 was obtained from ATCC (Manassas, VA) and cells were grown in DMEM (5.5 mmol/l Glucose, Corning, Corning, NY) supplemented with 10% FBS and 1% P/S (Sigma Aldrich, St. Louis, MO) at 37C and 5% CO2. 4-hydroxynoneal (4-HNE, 64 mmol/l; Cayman Chemicals, Ann Arbor, MI) was dissolved in molecular grade ethanol (Sigma Aldrich). In all, 80% confluent cells in a T75 flask were incubated with 0–200 µmol/l 4-HNE or ethanol control (0.3%) in completed DMEM for 6 hr. Cells were detached by scraping and pelleted (4000 rpm, 5 min, room temperature).
Cell Microarray
Scraped cells were fixed in 10% neutral buffered formalin (Sigma Aldrich) for 24 hr and then transferred to 70% ethanol. After fixation, cells were centrifuged and supernatant discarded, equal volume of warm 2%–3% low-melt agarose in 1X PBS was added and quickly vortexed for even dispersal and cooled in 4C for until the agar plugs were solidified. This gel matrix was then transferred to a histology cassette and processed in a standard tissue procedure and subsequently paraffin embedded to form a donor block. Cell microarray construction then followed using the same procedures previously described 14 using a semi-automated tissue microarrayer design (Pathology Devices, Westminster, MD). A core was removed from the donor block with 1.0 mm needle and placed in a hole prepared in a paraffin recipient block. The presence of cells was confirmed using a standard H&E staining.
Clinical Samples
Samples were obtained from patients participating in a clinical trial investigating the mechanism of vitamin E in patients with NAFLD (ClinicalTrials.gov, NCT01792115). In all, 21 patients with biopsy-proven NAFLD were randomized at a 1:1:1 ratio to receive one of three doses of natural vitamin E (RRR-alpha-tocopherol, Pharmavite, West Hills, CA), 200, 400, or 800 IU/d, for 24 weeks. Randomization was performed by the National Institutes of Health (NIH) Clinical Center Pharmaceutical Development Section using a random numbers table and block randomization with block size 6:3:3:3, stratified by non-alcoholic steatohepatitis (NASH) vs. NAFL and by diabetes. Compliance to the use of vitamin E was assessed with drug diaries and pill counts. Baseline needle biopsies were performed prior to initiating vitamin E treatment and a second on-treatment biopsy was performed after 4 weeks. The complete clinical trial protocol is available upon request from the corresponding author. Control samples with normal liver histology were from biopsies from patients with past history of chronic Hepatitis C infection, obtained 24 weeks after completing successful antiviral treatment with sustained clearance of virus (ClinicalTrials.gov, NCT00718172, n=5). Both trials were approved by the National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK) institutional review board and all subjects provided written informed consent.
Immunohistochemistry and Digital Image Analysis
Slides sectioned at 5 µm thickness were deparaffinized by xylene and rehydrated through a graded alcohol series. Heat-mediated antigen retrieval was performed using pH 6 citrate buffer (Dako, Carpinteria, CA) within a pressure chamber (Pascal; Dako). We used two-step temperature cycles (30 sec at 123.5C and 10 sec at 90C). The slides were cooled down for 20 min at room temperature after antigen retrieval, and then 3% H2O2 for 10 min was applied to block endogenous peroxidase. The sections were incubated with mouse monoclonal [HNEJ-2] anti-4HNE antibody (Abcam, MA; cat# ab48506) at 1:100 (1 µg/ml) or 1:200 (2 µg/ml) dilution for 1 hr at room temperature. HNEJ-2 has a high affinity for 4-HNE modified proteins, primarily HNE-histidine without any cross reactivity toward other aldehyde modifications. 15 The antigen–antibody reactions were detected with secondary EnVision+ Mouse-HRP System (Dako) and visualized with DAB+ (3,3-diaminobenzadine; Dako) for 10 min. Negative (IgG and omission of primary antibody) and positive controls were concurrently performed. Sections were lightly counterstained with hematoxylin, dehydrated with a series of ethanols, cleared in xylene and coverslipped before examination by light microscopy.
Stained slides were scanned with the NanoZoomer-XR (Hammatsu Photonics, Hamamatsu City, Japan) at 20× objective magnification. The images were analyzed using Visiopharm software v2017.7.1.3885 (Visiopharm, Hørsholm, Denmark). In the present study, we used two different algorithms for assessing 4-HNE adducts, one for cell line samples and another for tissues. The cell line experiments were conducted for Ab characterization in order to establish specificity, sensitivity, and dynamic range. In terms of cell line experiments, we calculated histoscores by multiplying the DAB staining intensity and the percentage of positive cells for a total possible range of 0–300, as described previously. 16 The DAB intensity was categorized as follows: 0 = negative, 1 = weak, 2 = moderate, and 3 = strong. The threshold value of RGB-B for negative, weak, intermediate, and strong were 40, 70, 100, and 140, respectively.
For human liver biopsy samples, the immunohistochemical score was expressed as the percentage of positive cells (possible range, 0–100). The threshold for positive cells was initially calibrated manually. After training the system to identify cells by digitally “painting” nuclei, we defined the nucleus size as 9.4 µm, with a sensitivity of 60% in RGB-G mode. Cytoplasm was defined by outlining the previously defined nucleus with a 4.9 µm width band using a threshold of 82% in HDAB-DAB mode. The similarity in color between lipofuscin, a yellow-brownish pigment consisting of highly oxidized proteins and lipids which are resistant to degradation and are considered aging pigments 17 and DAB creates the potential for false-positive detection. With the settings mentioned above, especially for HDAB-DAB mode, we successfully excluded potential false-positive due to lipofuscin during the algorithm setup for 4-HNE staining.
Histological Assessment of Human Liver Specimens
Histological samples were scored by a single pathologist (DEK) according to the NASH-CRN scoring system. 18
Detection of Circulating Markers of Lipid Peroxidation
Plasma samples were collected, aliquoted, and immediately stored in -80C. 9-hydroxyoctadecadienoic acid (HODE) and 13-HODE were measured as previously described. 19 In brief, 9- and 13-HODE were measured in 20 μl of protein-free plasma by online SPE-HPLC-MS/MS on a SCIEX 5500 QTRAP (SCIEX, Darmstadt, Germany) instrument. Detection of metabolites was carried out in multiple reaction monitoring (MRM) mode using negative electrospray ionization (ESI). Quantification was carried out using a seven-point standard curve of 9- and 13-HODE standards.
Western Blot
Cells were treated as described above, scraped into media at the end of the experiment and pelleted (5000 rpm, 5 min, 4C). Cells were homogenized in radioimmunoprecipitation assay (RIPA) buffer (Thermo Fisher Scientific) with added protease inhibitor (Roche, Switzerland). Protein was quantified using the bicinchonic acid (BCA) Kit (Thermo Fisher Scientific). Forty micrograms of protein per lane was separated by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) followed by transfer to a polyvinylidene difluoride membrane. Membrane was blocked for 1 hr in 5% milk in Tris-buffered saline Tween-20 (TBST) and HNEJ-2 antibody was incubated overnight (1:1000) at 4C with gentle agitation. Incubation with secondary antibody was carried out with gentle shaking for 1 hr at room temperature. Bands were visualized using enhanced chemiluminescence (ECL; Thermo Fisher Scientific), and band intensity was recorded with an Azure 300 (Azure biosystems). Band volume intensities were quantified with Fiji 20 and relative concentrations of 4-HNE protein adducts were calculated as the ratio of adducts to the amount of β-actin (Cell Signaling Technology).
Statistical Analysis
Data were analyzed using Graph Pad Prism 8 for Mac (GraphPad Software, San Diego, CA). Normality was assessed using the D’Agostino-Pearson omnibus normality test and significance was tested by Mann-Whitney or Wilcoxon matched-paired signed rank test, as appropriate. Differences in gender frequency between control and NAFLD group was calculated using Fisher’s exact test. For differences between doses a Kruskal-Wallis test was used with Dunn’s correction for multiple comparison. Correlations used either Pearson’s or Spearman’s correlation coefficients as appropriate.
Results
Anti- HNEJ-2 Can Quantify 4-HNE Protein Adducts in a Linear Fashion
To determine the ability of the antibody to detect and quantify 4-HNE in a linear fashion, we treated Hep G2 cells with varying doses of 4-HNE. The antibody detected 4-HNE protein adducts for all concentrations > 25 µM in a dose dependent manner (Fig. 1A) at both antibody dilutions tested. There is significant correlation between 4-HNE input and immunohistochemical signal at both 1:100 (R 2 = 0.97) and 1:200 (R 2 = 0.98) dilutions (Fig. 1B). The 1:200 dilution (2 µg/ml) was employed for all further analyses. Western blot analysis of 4-HNE adducts confirmed the dose-dependent detection (Supplemental Fig. 1). In summary, the employed antibody shows good sensitivity, specificity, and a broad dynamic range.
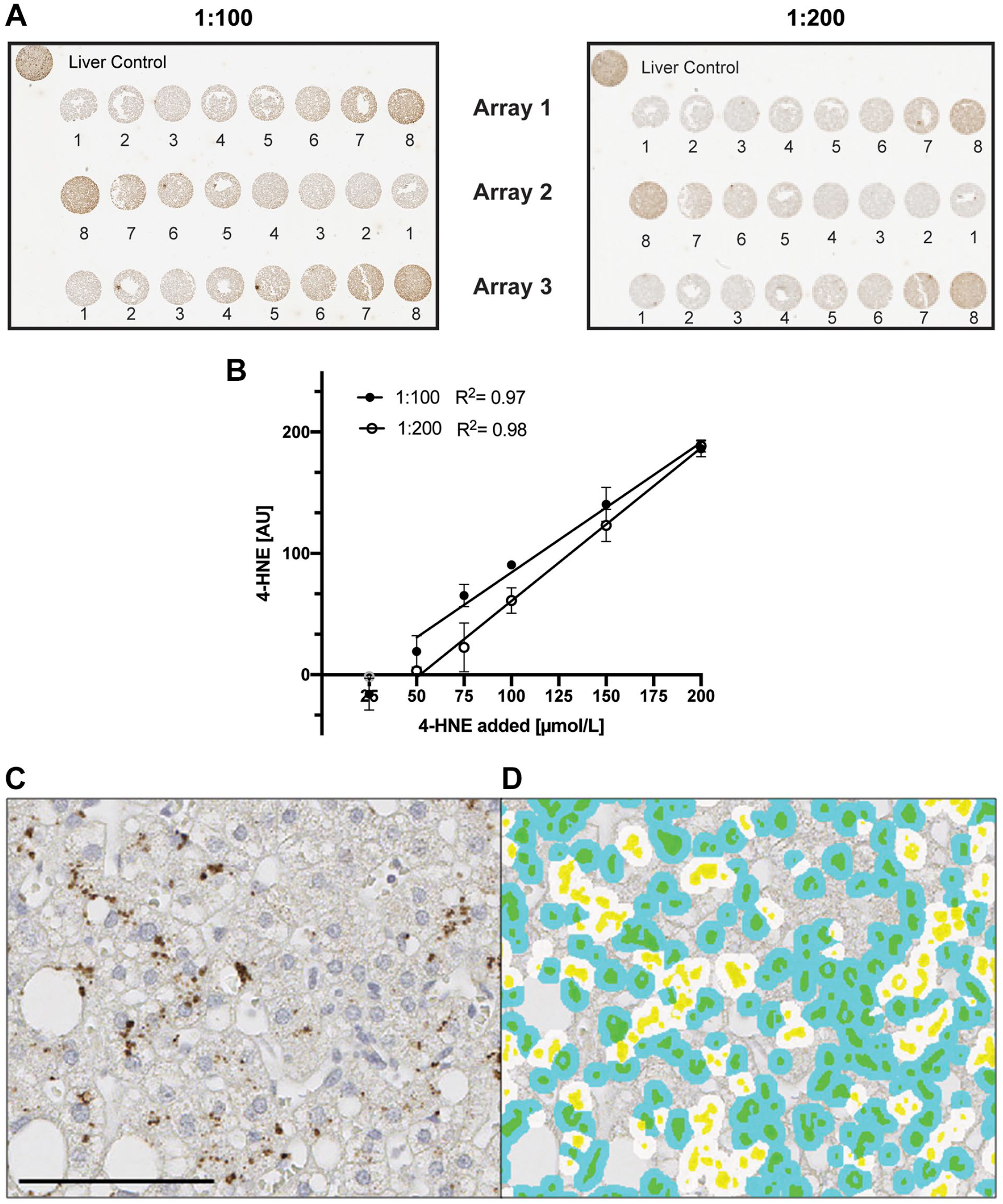
Calibration curves of cells treated with 4-HNE. Hep G2 cells treated with different doses of 4-HNE (1 = media, 2 = 0, 3 = 25, 4 = 50, 5 = 75, 6 = 100, 7 = 150, 8 = 200 µmol/l) were incubated with mouse monoclonal [HNEJ-2] at either 1:100 (filled circle, 1 µg/ml) or 1:200 (empty circle, 2 µg/ml) dilution for 1 hr at room temperature. (A) Shows three individual arrays for both dilutions. Background staining of the solvent control (0 µmol/l) was subtracted from all points. In the linear regression, 25 µmol/l was excluded for both dilutions as it did not differ from the background staining (B). Slopes were 1.071 (1:100) and 1.262 (1:200) while Y intercepts were -22.84 (1:100) and -65.44 (1:200), respectively. Quantification was performed using Visiopharm software. Data are shown as mean ± SD. Fig. 1C shows a representative image of 4-HNE staining in human liver with the corresponding image analysis (1D). Cytoplasmic 4-HNE was quantified and cells were classified as positive cells (white-colored spots) or negative cells (blue-colored spots). Yellow marks nuclei and green represents the combination of blue and yellow. Data are shown as mean ± SD. Scale bar, 100 µm. Abbreviations: 4-HNE, 4-hydroxynoneal; HNEJ-2, mouse monoclonal anti-4 hydroxynonenal antibody.
Anti- HNEJ-2 Can Detect and Quantify 4-HNE Protein Adducts in Liver Samples
Given the higher complexity of tissue compared to cells, we modified the algorithm of 4-HNE quantification to detect 4-HNE in clinical liver biopsy samples. While we employed an IHC score calculated using percent positive cells as well as DAB intensity for the cellular analysis, this approach was unsuitable for the clinical samples due to high background noise in tissue compared to cell samples. 4-HNE adducts could be detected in the cytoplasm in liver samples from NAFLD patients (Fig. 1C). The developed algorithm was able to automatically detect 4-HNE adduct cytoplasm-positive cells (Fig. 1D).
4-HNE Protein Adducts Are Increased in NAFLD Livers
4-HNE protein adducts were only minimally detectable in livers with normal histology (3.4 ± 5.9%) but were highly detectable in liver samples from NAFLD patients (24.2 ± 23.4%, p=0.02, Fig. 2), indicating increased levels of lipid peroxidation compared to controls. Interestingly, within subjects with NAFLD, the degree of 4-HNE positivity did not correlate with alanine aminotransferase (ALT), body mass index (BMI), or liver fat content (Fig. 3A–C). There was also no association between 4-HNE score and histological features of NAFLD (Supplemental Fig. 2A–F). Previous studies assessing hepatic lipid peroxidation in NAFLD used the circulating lipid peroxidation markers 9- and 13-HODE 21 ; however, we found that plasma 9- and 13-HODE do not correlate with hepatic 4-HNE staining (Supplemental Fig. 3G and H).
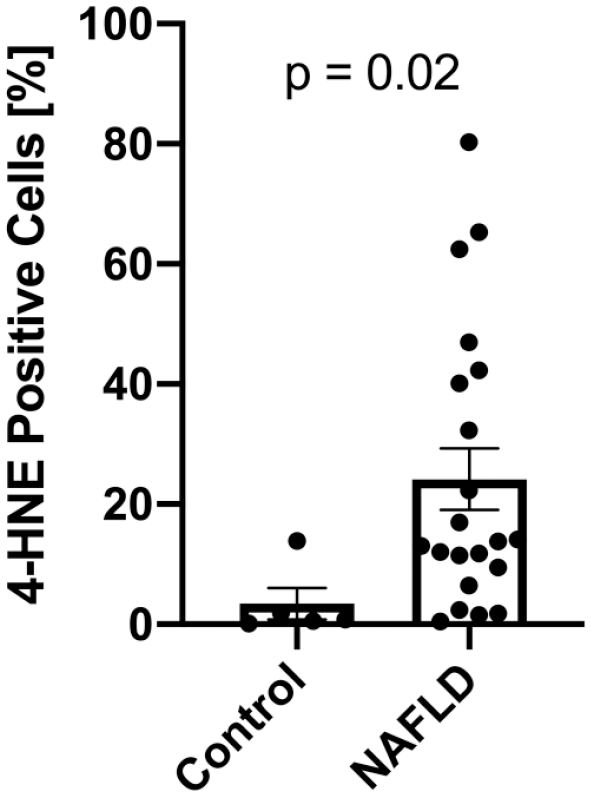
4-HNE staining is increased in NAFLD. Compared to controls with normal liver histology (n=5), patients with NAFLD (n=21) show a significant increase in 4-HNE positive cells. Data is shown as mean + SEM. Abbreviations: 4-HNE, 4-hydroxynoneal; NAFLD, non-alcoholic fatty liver disease.
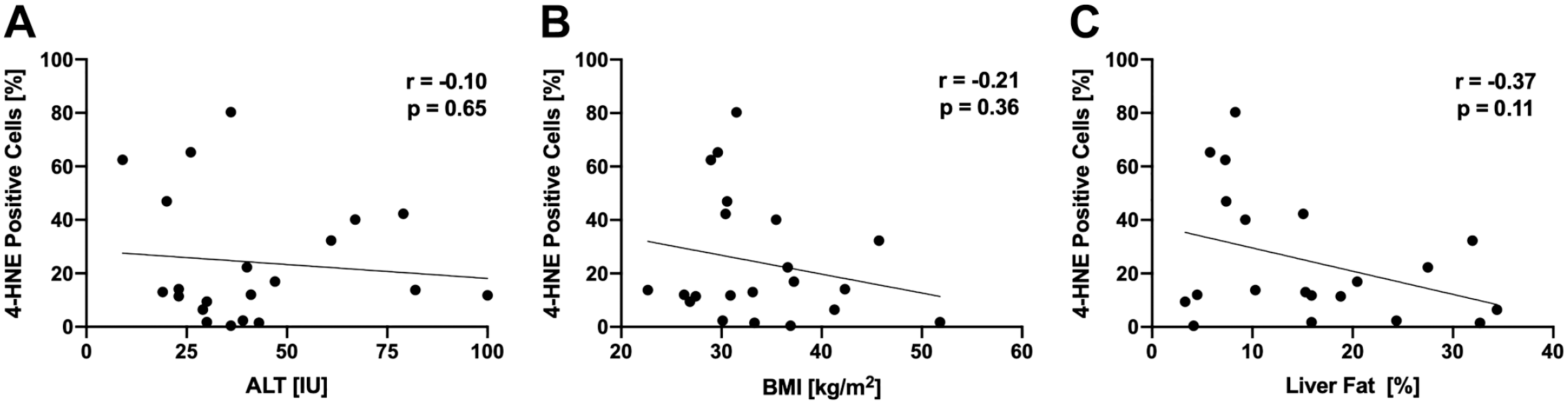
4-HNE staining is not correlated with (A) ALT, (B) BMI, or (C) liver fat determined by magnetic resonance spectroscopy. A Spearman’s rank correlation coefficient, B, C Pearson’s correlation coefficient. Abbreviations: ALT, alanine aminotransferase; BMI, body mass index; 4-HNE, 4-hydroxynoneal.
Four Weeks of Vitamin E Decreases Hepatic 4-HNE Protein Adducts in NAFLD
To determine the ability of the semi-quantitative algorithm to detect changes in endogenous 4-HNE, we analyzed paired biopsy samples from the patients with NAFLD before and on treatment with vitamin E, a known antioxidant. Treatment of NAFLD patients with vitamin E led to a significant (p=0.0002) decrease of 4-HNE adducts (24.16 ± 23.35 vs 7.28 ± 8.49 p=0.0002, Fig. 4A) irrespective of the given dose (Fig. 4B).
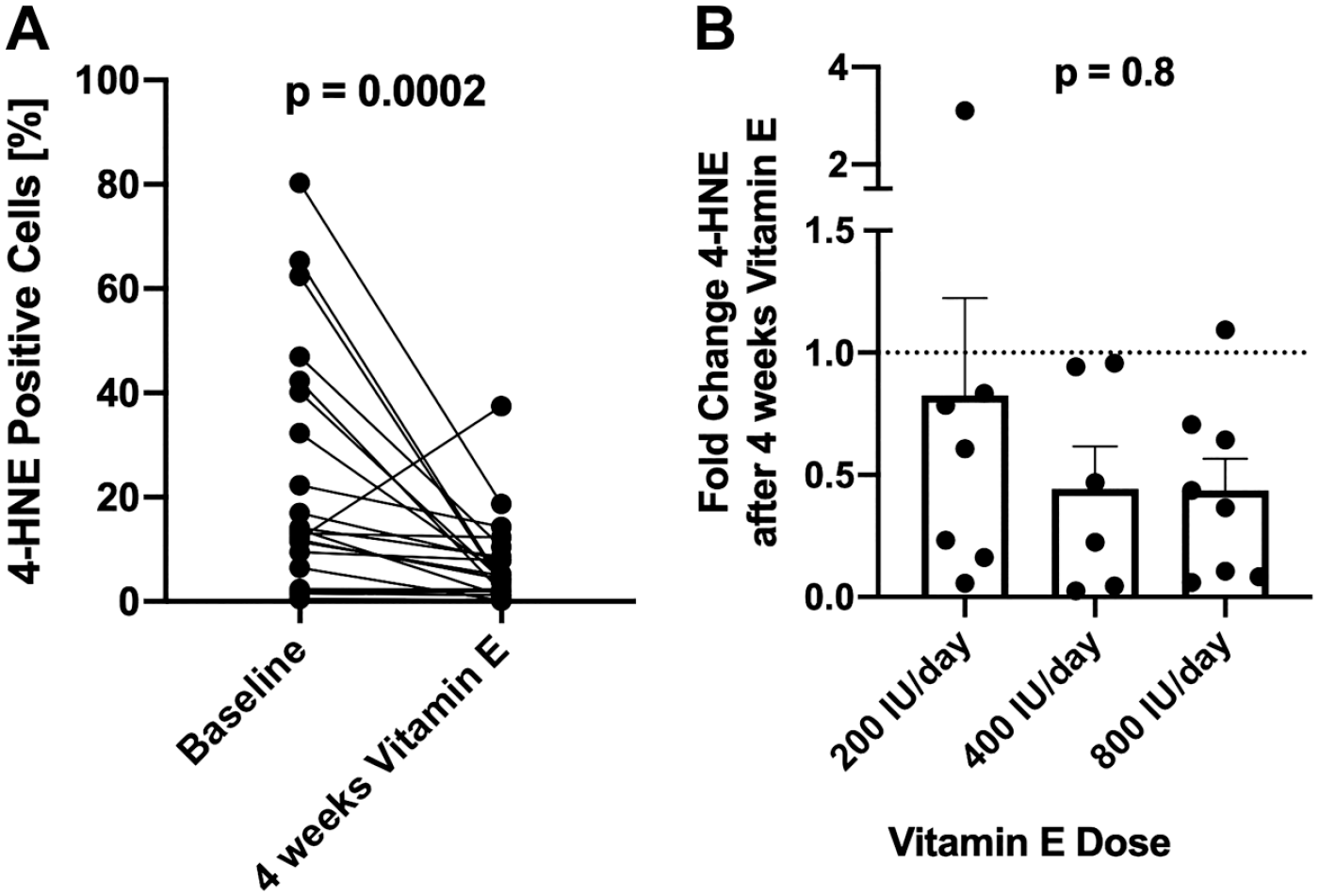
Four weeks of treatment with RRR-α-tocopherol (200–800 IU/day) decreased 4-HNE positive cells (A), with no differences between doses (B). Data are shown as mean + SEM. Abbreviations: 4-HNE, 4-hydroxynoneal; RRR-α-tocopherol, natural vitamin E.
Discussion
Lipid peroxidation is thought to play a pivotal role in the progression of many disease, especially those characterized by lipid accumulation like NAFLD. Routine assessment of lipid peroxidation in clinical or research settings is hindered by the absence of a validated, automated, semi-quantitative method that can be easily performed with a limited specimen size. In this work, we first validated the antibody and then developed an immunohistochemical (IHC) staining method for protein modifications by the lipid peroxidation product 4-HNE with automated image analysis suitable for clinical samples. While IHC data resulting from manual scoring methods has had good to excellent intra- and inter-observer reproducibility,22,23 manual estimation of negative and positive percentages of areas stained has been shown to have only poor reproducibility 24 making automated analysis the preferred solution. In contrast to other methods, our assay was validated against a standard curve generated from cells treated with exogenous 4-HNE, ensuring sensitivity and specificity of the detection of adducts by the antibody.
Patients with NAFLD have increased lipid peroxidation and antioxidant therapies like vitamin E25–27 or pentoxifylline 28 have been successfully tested in clinical trials. We demonstrate that 4-HNE quantification allows for detection of marked hepatic lipid peroxidation in patients with NAFLD compared to controls. Interestingly, the degree of lipid peroxidation was not associated with known clinical or histological parameters. This may be due to differences in the actual proteins modified by 4-HNE or their subcellular or zonal localization which we did not asses in this paper. Furthermore, it could possibly be a consequence of the relatively small sample size of this pilot experiment.
Lipid peroxidation in NAFLD has been primarily assessed in the circulation21,29–31 mainly by measuring the plasma lipid peroxidation markers 9- and 13-HODE. Lipid peroxidation was rarely assessed directly in the liver,12,13,32 and in fact, it is unclear if the circulatory markers of lipid peroxidation truly reflect intrahepatic levels. We did not observe a correlation between hepatic 4-HNE protein adducts and circulating 9- and 13-HODE, raising the question whether plasma 9- and 13-HODE are adequate markers of hepatic lipid peroxidation or whether they are more reflective of systemic lipid peroxidation.
To date, no assessment of intrahepatic changes in lipid peroxidation in response to antioxidants has been performed, although treatment with pentoxifylline in NAFLD has been shown to decrease circulating lipid peroxidation markers. 28 To this end, we investigated 4-HNE adducts in NAFLD patients before and after supplementation with vitamin E. To the best of our knowledge, we are the first to assess the effect of vitamin E supplementation or any other antioxidant on hepatic lipid peroxidation in humans. In accordance with the assumption that vitamin E acts as lipid peroxidation chain breaking antioxidant, we observed a significant decrease in 4-HNE protein adducts after 4 weeks of treatment. Vitamin E dose did not affect the magnitude of 4-HNE decrease, but this may be due to the use of supraphysiological doses which are 9–35 times the recommended daily allowance, 33 and is consistent with the lack of dose-dependent differences in clinical response parameters seen in that trial (under review).
One concern with automated immunohistochemistry algorithms is the presence of false positives due to lipofuscin. Due to its color, lipofuscin is hard to distinguish from DAB intensity. In our algorithm, we excluded lipofuscin as a confounder by defining the size of the nucleus and then outlining the cytoplasm with different color modes. Importantly, accumulated lipofuscin is not thought to be mobilized out of the hepatocytes rapidly, if at all; the fact that we could clearly show a significant decrease in the percent positive cells for 4-HNE with vitamin E treatment suggests we successfully avoided lipofuscin confounding.
A further limitation of our analysis is the absence of a true normal control. We chose to use samples from subjects who recovered from chronic hepatitis C. Normal histology human liver is not typically obtained due to obvious ethical concerns. “Healthy liver” controls are typically procured from non-tumor liver adjacent to a resected cancer or from deceased donors. Both of these sources are likely to be exposed to significant oxidative insults before, or just prior to tissue procurement, even when histological morphology appears normal. Our control samples were from subjects who were > 24 weeks past viral eradication, had normal ALT and normal histology. Thus, we believe they provide a better and more appropriate controls.
In summary, we have shown that 4-HNE adducts can be analyzed using fully automated image analysis and that this technique is useful in assessing lipid peroxidation in patients with NAFLD at baseline and in response to treatment. This approach may be used in studies to quantify changes in hepatic lipid peroxidation by medications or other interventions in clinical trials, even when specimen size is limited. Furthermore, our approach may be useful to study the pathophysiological consequences of lipid peroxidation protein adducts by combining staining for 4-HNE adducts with other techniques such as RNA scope. 34
Supplemental Material
2020-00106R1_Production_Supplemental_Material_online_supp – Supplemental material for 4-HNE Immunohistochemistry and Image Analysis for Detection of Lipid Peroxidation in Human Liver Samples Using Vitamin E Treatment in NAFLD as a Proof of Concept
Supplemental material, 2020-00106R1_Production_Supplemental_Material_online_supp for 4-HNE Immunohistochemistry and Image Analysis for Detection of Lipid Peroxidation in Human Liver Samples Using Vitamin E Treatment in NAFLD as a Proof of Concept by Maren C. Podszun, Joon-Yong Chung, Kris Ylaya, David E. Kleiner, Stephen M. Hewitt and Yaron Rotman in Journal of Histochemistry & Cytochemistry
Footnotes
Acknowledgements
The authors would like to thank Nevitt Morris, Wen-Chun A. Huang, and the Liver Diseases Branch clinical team for their excellent management of the clinical trial at the NIH Clinical Center.
Competing Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
All authors have contributed to this article as follows: planning (MCP, YR, SMH), execution (MCP, J-YC, KY), analysis (J-YC, KY, DEK), and writing (MCP, YR, SMH, J-YC). All authors have read and approved the final manuscript.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Intramural Research Programs of the National Institute of Diabetes and Digestive and Kidney Diseases and of the National Cancer Institute. Vitamin E for the clinical trial was provided by Pharmavite Inc under a clinical trial agreement. MCP received funding from the NIH Office of Dietary Supplements.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
