Abstract
In glioblastoma, a fraction of malignant cells consists of therapy-resistant glioblastoma stem cells (GSCs) residing in protective niches that recapitulate hematopoietic stem cell (HSC) niches in bone marrow. We have previously shown that HSC niche proteins stromal cell–derived factor-1α (SDF-1α), C-X-C chemokine receptor type 4 (CXCR4), osteopontin (OPN), and cathepsin K (CatK) are expressed in hypoxic GSC niches around arterioles in five human glioblastoma samples. In HSC niches, HSCs are retained by binding of SDF-1α and OPN to their receptors CXCR4 and CD44, respectively. Protease CatK cleaves SDF-1α to release HSCs out of niches. The aim of the present study was to reproduce the immunohistochemical localization of these GSC markers in 16 human glioblastoma samples with the addition of three novel markers. Furthermore, we assessed the type of blood vessels associated with GSC niches. In total, we found seven GSC niches containing CD133-positive and nestin-positive GSCs as a single-cell layer exclusively around the tunica adventitia of 2% of the CD31-positive and SMA-positive arterioles and not around capillaries and venules. Niches expressed SDF-1α, CXCR4, CatK, OPN, CD44, hypoxia-inducible factor-1α, and vascular endothelial growth factor. In conclusion, we show that GSC niches are present around arterioles and express bone marrow HSC niche proteins.
Introduction
Glioblastoma is the most aggressive and deadly primary brain tumor with a poor patient survival of only 12–15 months after diagnosis.1–6 In glioblastoma, a small fraction of the malignant cells consists of glioblastoma stem cells (GSCs) which are held responsible for therapy resistance, tumor maintenance,7–12 and recurrence.3,13–23 GSCs reside in a specific microenvironment, referred to as the GSC niche, which is considered to be a dynamic and complex milieu that protects GSCs against therapy and allows self-renewal and quiescence of GSCs3,7,14–16,21,22,24–32 and enables the GSCs to have a robust DNA damage response.33,34 Recently, evidence has been reported that the GSC niche has tumor-immunosuppressive capacities.35,36
The most widely used markers to detect GSCs are CD13312,37–48 and nestin.20,37,39,45–47,49–51 In a previous study, we have shown that CD133-positive and nestin-positive GSCs reside in hypoxic niches around a small fraction of arterioles with CD31-positive endothelium.46,111 CD133 expression has been reported to be upregulated in hypoxic conditions.52,53 In these GSC niches, we found expression of hematopoietic stem cell (HSC) niche proteins stromal cell–derived factor-1α (SDF-1α), osteopontin (OPN), and cathepsin K (CatK). 46 Hypoxia induces expression of hypoxia-inducible factor-1α (HIF-1α) and vascular endothelial growth factor (VEGF) in glioblastoma which are responsible for the upregulation of C-X-C chemokine receptor type 4 (CXCR4),54–57 SDF-1α,54–56 OPN,58–60 and CD44.13,58
In human bone marrow, SDF-1α is a chemoattractant which binds CXCR4-positive HSCs in hypoxic niches in bone marrow61–66 in close vicinity of arterioles and sinusoids.66–68 OPN and SDF-1α are produced and secreted by osteoblasts and endothelial cells in bone marrow and interact with their receptors CXCR4 and CD44 on HSCs, respectively, to retain HSCs in niches.61–66,69 CatK is a cysteine protease involved not only in bone degradation but also in SDF-1α cleavage and inactivation in bone marrow66,70–72 that release HSCs out of niches into the circulation.63,73,74 HIF-1α and VEGF are important factors for the production of HSC niche proteins and maintenance of HSCs in niches in bone marrow.61–66,69,75,76
CatK is one of the highest differentially expressed proteases in glioblastoma relative to normal brain.71,77 CatK can cleave and inactivate SDF-1α and thereby inhibit invasion of CXCR4-positive GSCs toward SDF-1α in vitro. 72 However, we have not yet been able to detect activity of CatK in glioblastoma despite its high differential expression.71,77 We assume that the activity of CatK is tightly regulated because of its strong hydrolytic activity explaining why we have found CatK protein expression but not CatK activity associated with GSC niches in glioblastoma. 71 OPN has been reported recently to maintain the stem cell phenotype in GSCs and stimulate double-strand DNA repair.78,79
Based on the proteins that are known to be crucial in HSC niches, we defined GSC niches to be positive for the GSC marker proteins, CD133 and nestin, that are important for the maintenance of HSCs80,81; niche markers SDF-1α, CXCR4, CatK, OPN, and CD44; and the hypoxia markers HIF-1α and VEGF. 111
The aim of the present study was to determine which markers in HSC niches are expressed in GSC niches in a larger number of human glioblastoma samples than in our first GSC study. 46 To specifically detect cancer cells in the glioblastoma samples, we localized immunohistochemically the isocitrate dehydrogenase 1 (IDH1)R132H mutation in IDH1R132H mutated and IDH1 wild-type glioblastoma samples. The IDH1 mutation is the most frequently occurring mutation (50–80%) in secondary glioblastoma.6,82–85 Furthermore, smooth muscle actin (SMA) and CD44 were included as novel markers. In addition, we aimed to determine around which type of blood vessels these markers are clustered.
Materials and Methods
Patients
Surgically obtained snap-frozen glioblastoma samples (all grade IV astrocytoma) from 18 patients (aged 38–74 years, anonymized to the researchers) were obtained from the Brain Tumor Bank maintained by the Department of Neuropathology at the Academic Medical Centre (AMC, Amsterdam, The Netherlands). Seventeen samples were IDH1 wild-type, and one sample was IDH1R132H mutated. Research was performed on “waste” material that is stored in a coded fashion. Consent for this project was reviewed and waivered, and the project was approved by the Medical Ethics Review Committee of the Academic Medical Center and University of Amsterdam (reference number W14_224 # 14.17.0286). Consent for removal of the tissue and its storage in the tumor bank for research purposes was obtained and documented in the patients’ medical charts.
Chromogenic Immunohistochemistry
Chromogenic immunohistochemistry (IHC) experiments were performed on serial cryostat glioblastoma sections of 18 glioblastoma samples. Data of 16 glioblastoma samples have been analyzed and are described in this article, because two glioblastoma samples had to be excluded due to tissue damage in too many sections of the series. We stained for blood vessel wall markers CD31 and SMA; GSC markers CD133 and nestin; niche markers SDF-1α, CXCR4, CatK, OPN, and CD44; and hypoxia markers HIF-1α and VEGF. We aimed to determine where these markers are localized in the 16 glioblastoma samples and whether a recognizable staining pattern could be visualized in the glioblastoma sections. In addition, we analyzed the type of blood vessels around which niches were found in the 16 glioblastoma samples.
Cryostat sections (7-μm thick) were cut at −25C on an HM560 cryostat (MICROM, Walldorf, Germany), picked up on glass slides, and stored at −80C until used. All staining procedures including controls were performed on serial sections of each glioblastoma sample.
Sections were air-dried at room temperature for 15 min before immunohistochemical staining. The sections were fixed in acetone (−20C) for 10 min and were air-dried afterward for 15 min. The sections were encircled with a PAP pen (Dako, Glostrup, Denmark), followed by three washing steps of 5 min with phosphate-buffered saline (PBS). The sections were treated with 100% methanol containing 0.3% H2O2 for 15 min to block endogenous peroxidase activity to prevent nonspecific background staining, followed by three washing steps of 5 min each using PBS. Then, sections were incubated with PBS containing 10% normal goat serum (Dako) and 0.1% bovine serum albumin (BSA) for 45 min to further reduce nonspecific background staining. Sections were subsequently incubated overnight at 4C with primary antibodies as indicated in Table 1.
Details of Primary Antibodies Used for Chromogenic Immunohistochemistry.
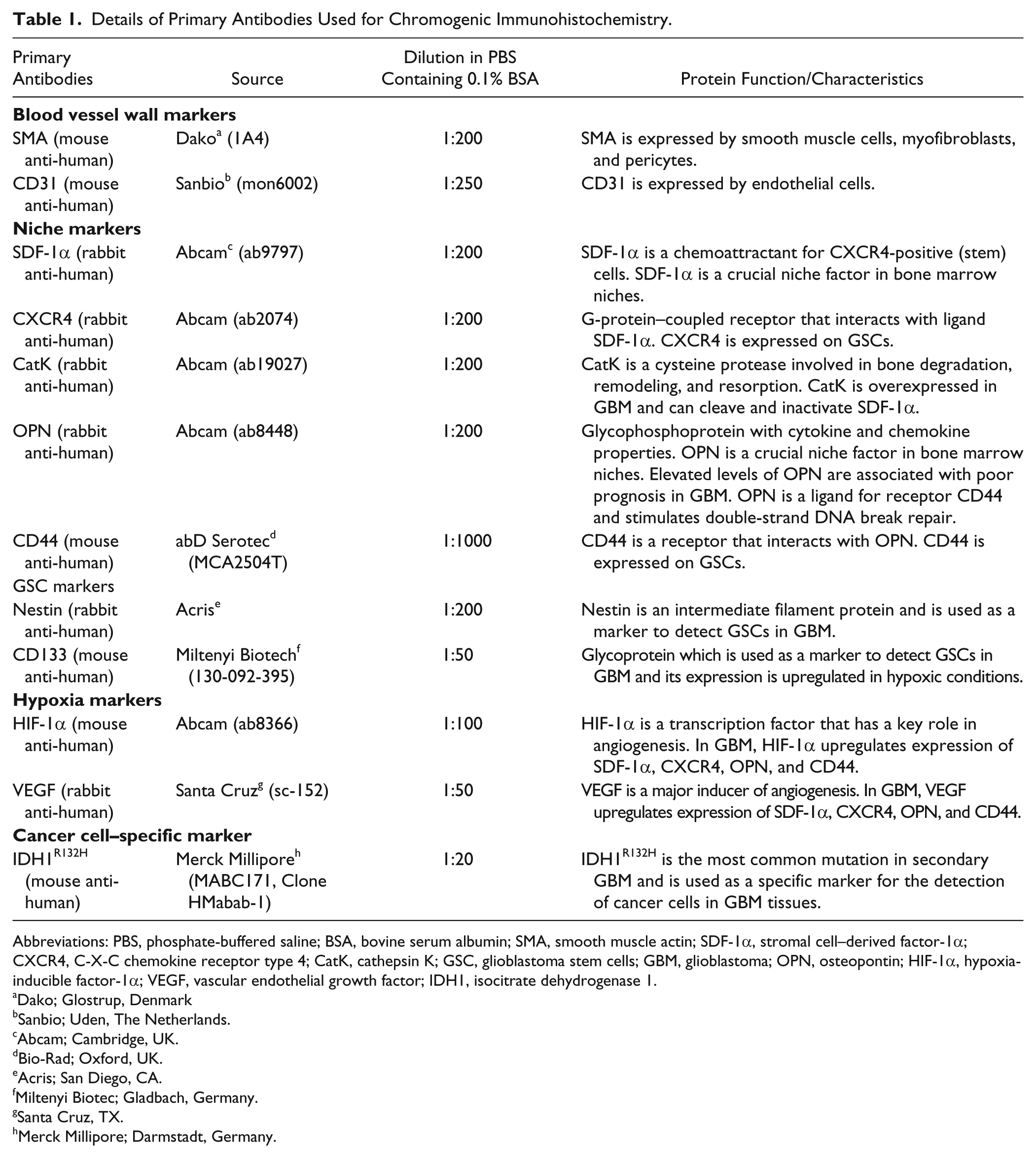
Abbreviations: PBS, phosphate-buffered saline; BSA, bovine serum albumin; SMA, smooth muscle actin; SDF-1α, stromal cell–derived factor-1α; CXCR4, C-X-C chemokine receptor type 4; CatK, cathepsin K; GSC, glioblastoma stem cells; GBM, glioblastoma; OPN, osteopontin; HIF-1α, hypoxia-inducible factor-1α; VEGF, vascular endothelial growth factor; IDH1, isocitrate dehydrogenase 1.
Dako; Glostrup, Denmark
Sanbio; Uden, The Netherlands.
Abcam; Cambridge, UK.
Bio-Rad; Oxford, UK.
Acris; San Diego, CA.
Miltenyi Biotec; Gladbach, Germany.
Santa Cruz, TX.
Merck Millipore; Darmstadt, Germany.
After incubation with secondary antibodies and three washing steps of 5 min each using PBS, the sections were incubated with AEC (peroxidase substrate kit; Vector Laboratories, Burlington, CA) for 20 min, followed by one washing step of 5 min using PBS. Sections were then placed in running tap water for 20 min and then in distilled water followed by incubation in hematoxylin (Fluka Biochemica, Sigma-Aldrich, St. Louis, MO) for nuclear counterstaining for 3 min. Sections were again placed in running tap water for 20 min and then in distilled water. All incubation steps were performed at room temperature, except for the overnight incubations with primary antibodies that were performed at 4C. Finally, the sections were covered with glycerin/gelatin mounting medium (Sigma-Aldrich).
Control incubations were performed either in the absence of the primary antibody or in the presence of rabbit serum or mouse serum in the same concentration as the primary antibody to determine the effect of the serum on nonspecific background staining.
Detection of IDH1R132H-mutated Glioblastoma Cells
An antibody against t expressed all over the sections, but the strongest he IDH1R132H mutation was used to specifically detect cancer cells in the glioblastoma tissue sections. 86 Sections were air-dried at room temperature for 15 min before immunohistochemical staining and were afterward fixed in 4% paraformaldehyde (Acros Organics, Geel, Belgium) containing 3.7% methanol (Merck Millipore, Darmstadt, Germany) diluted in PBS for 10 min at room temperature. The sections were encircled with a PAP pen (Dako), followed by three washing steps of 5 min with PBS. The sections were treated with 100% methanol containing 0.5% H2O2 for 15 min to block endogenous peroxidase activity to prevent nonspecific background staining, followed by three washing steps of 5 min each using PBS. Then, sections were incubated with PBS containing 10% normal rabbit serum (Dako) for 45 min to further reduce nonspecific background staining. Sections were subsequently incubated overnight at 4C with a primary antibody to detect the IDH1R132H mutation (rabbit anti-human IDH1R132H, dilution 1:20 [see Table 1] in PBS containing 0.1% BSA). After incubation with secondary antibody (goat anti-rabbit secondary antibody [Dako, 1:200 diluted in PBS containing 0.1% BSA]) and three washing steps of 5 min each using PBS containing 0.1% BSA, the sections were incubated with DAB substrate kit (Dako) for 10 min, followed by one washing step of 5 min using PBS. Sections were then placed in running tap water for 20 min and then in distilled water followed by incubation in hematoxylin (Sigma-Aldrich) for nuclear counterstaining for 15 min. Sections were again placed in running tap water for 20 min and then in distilled water. Finally, the sections were covered with glycerin/gelatin mounting medium (Sigma-Aldrich). Control incubations were performed in the absence of the primary antibody.
Fluorescence Immunohistochemistry
Sections were air-dried for 30 min at room temperature, encircled using a PAP pen (Daido Sangyo, Tokyo, Japan) and fixed for 10 min at room temperature using 4% paraformaldehyde (Acros Organics) containing 3.7% methanol (Merck) diluted in PBS. Sections were washed twice in PBS, followed by permeabilization using 0.1% Triton X (Sigma-Aldrich) in PBS and two washing steps in PBS. Blocking of nonspecific staining was performed by incubation in PBS containing 0.5% BSA (Sigma-Aldrich). Afterward, sections were incubated with primary antibodies at room temperature (rabbit-anti-human CatK [isotype IgG], ab19027, dilution 1:500, Abcam; mouse-anti-human SDF-1α [isotype IgG1], MAB310, dilution 1:600, R&D Systems, Minneapolis, MN; rabbit-anti-human SMA [isotype IgG2a], M0851 clone 1A4, dilution 1:500, Dako). Single, double, or triple staining procedures were used for the three primary antibodies. After 60 min of incubation at room temperature, sections were washed with PBS. Incubation with secondary antibodies was performed for 30 min at room temperature (goat-anti-rabbit CatK [IgG] conjugated with Alexa 488, dilution 1:500, Invitrogen, Carlsbad, CA; goat-anti-mouse SDF-1α [IgG1] conjugated with Alexa 647, dilution 1:500, Life Technologies, Carlsbad, CA; goat-anti-mouse SMA [IgG2a] conjugated with Alexa 568, dilution 1:500, Life Technologies). Removal of fluorescent lipofuscin from sections was performed by incubation for 30 min in 30-mM ammonium acetate (Merck Millipore) and 10-mM copper sulfate (Merck). Afterward, sections were washed for 30 sec using distilled water and four times for 5 min using PBS. Sections were mounted in ProLong Gold containing DAPI (Life Technologies, Bleiswijk, The Netherlands) and coverslipped. Preparations were hardened overnight at room temperature. Control incubations were performed in the absence of primary antibodies. Confocal imaging was performed using a confocal microscope (SP8-X SMD; Leica, Amsterdam, The Netherlands). Images of 1024 × 1024 pixels were taken as optical sections and were analyzed using LAS X software (Amsterdam, The Netherlands).
Hematoxylin–Eosin (HE) Staining
Glioblastoma cryostat sections were air-dried for 15 min. Then, the sections were fixed with Formol-Macrodex (4% formaldehyde, 7.2-mM CaCl2, 0.12-M Dextran-70, 0.12-M NaCl, and 7.96-mM CaCO3) for 10 min, followed by a washing step in distilled water for 5 min. Nuclei were stained with hematoxylin for 3 min; the sections were placed in running tap water for 20 min, after which the sections were placed in distilled water. Sections were then stained in eosin for 3 sec, dipped five times in distilled water, 15 times in 70% alcohol, 15 times in 96% alcohol, and 10 times in 100% alcohol. Afterward, sections were rinsed three times for 5 min in xylene. The sections were covered with the synthetic mountant Pertex (Histolab, Gothenburg, Sweden). All steps were performed at room temperature.
Microscopic Analysis of Staining Patterns
Sections were analyzed by light microscopy and images were taken using the QWin software and the Leica DMLB microscope (Leica; Mannheim, Germany). The staining patterns of IHC experiments were analyzed by five independent observers (V.V.V.H., J.R.W., H.K., W.T., and C.J.F.N.).
Results
Identification of GSC Niches in Glioblastoma Samples
In the 16 glioblastoma samples, we found seven GSC niches that expressed GSC markers, HSC niche markers, and hypoxia markers exclusively around arterioles, including the sample that was IDH1R132H mutated. Figure 1 shows the morphology of an arteriole that is the center of a GSC niche around the tunica adventitia of the arteriole, where we found GSCs in a single cell layer.
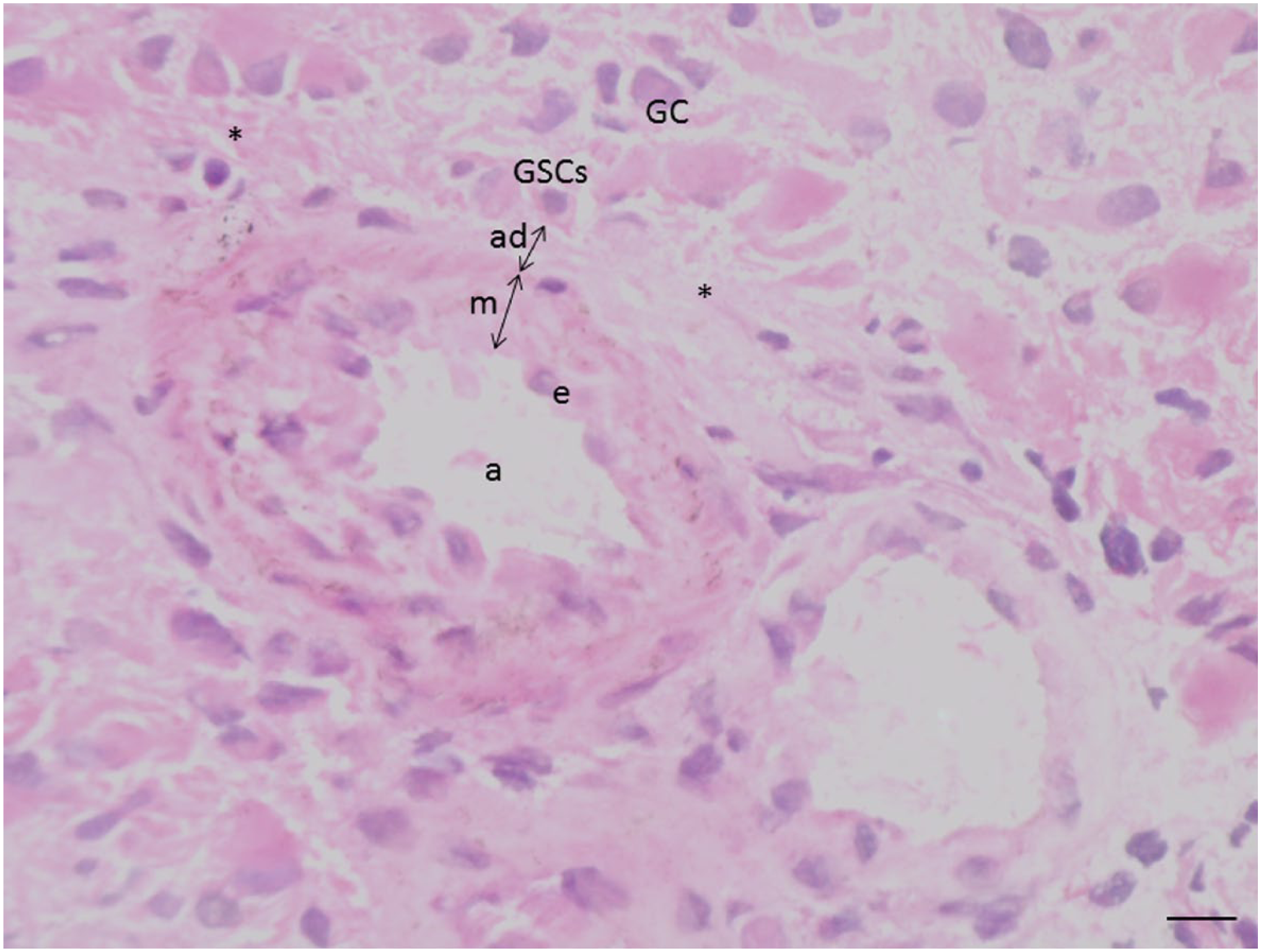
Image of an arteriole (a) in a glioblastoma section after hematoxylin–eosin staining surrounded by glioblastoma cells (GCs) and necrosis (*). The arteriolar wall consists of endothelium (e), tunica media (m), and tunica adventitia (ad). Glioblastoma stem cells (GSCs) are localized adjacent to the outer rim of the tunica adventitia. Bar = 20 µm.
Detection of Cancer Cells in Glioblastoma Samples
An antibody against the IDH1R132H mutation was used to specifically detect cancer cells in the glioblastoma tissue (Fig. 2). IDH1R132H mutated cancer cells were attached to the adventitia of arterioles (Fig. 2A) as well as widely distributed over the sections of the tumor sample (Fig. 2B). Staining of sections of IDH1 wild-type samples confirmed that the wild-type glioblastoma cancer cells are not IDH1R132H mutation positive (Fig. 2C).
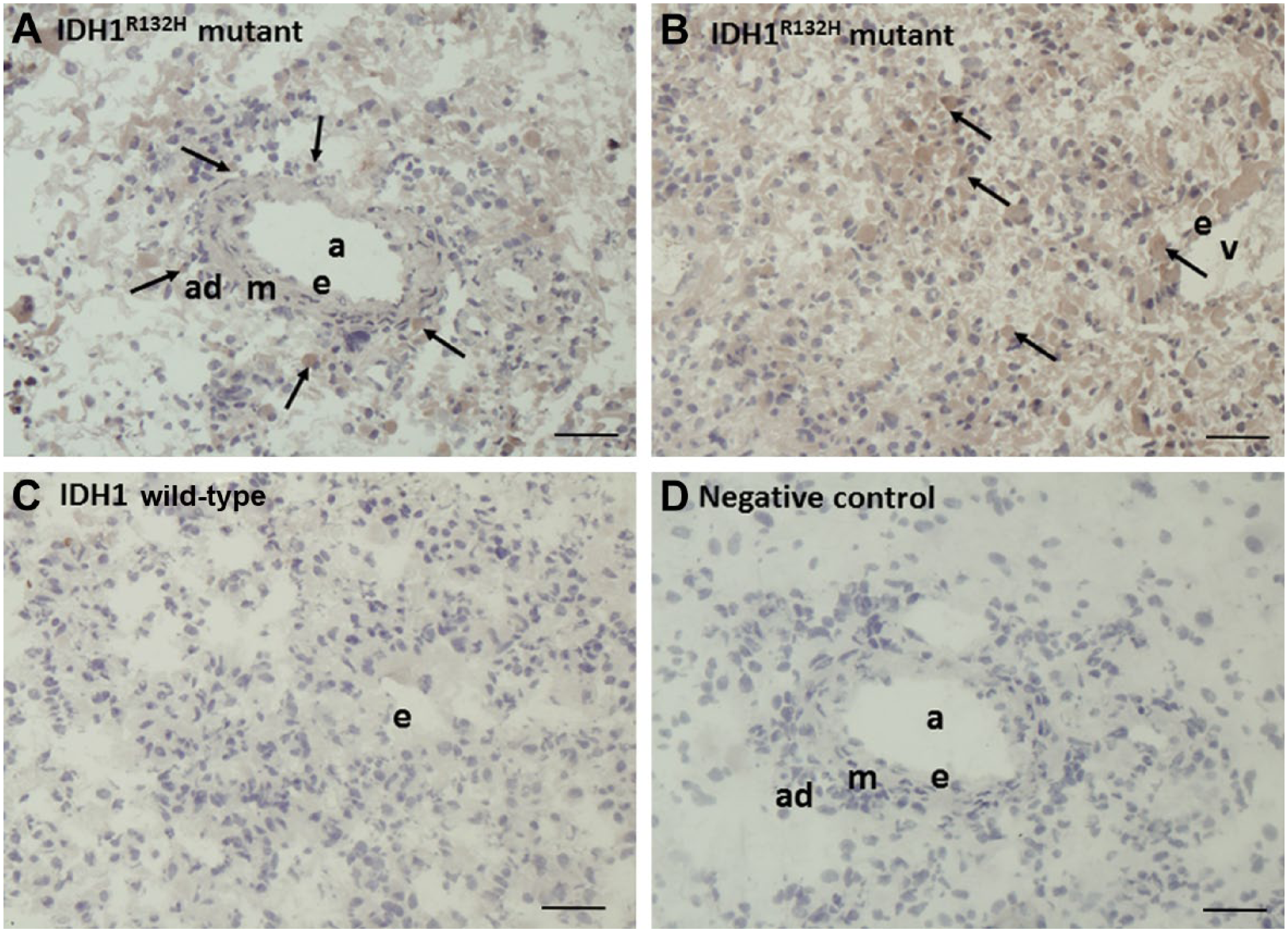
Immunohistochemical localization of the IDH1R132H mutation in serial cryostat sections of IDH1R132H-mutated (A, B) and IDH1 wild-type (C) human glioblastoma. (A, B) IDH1R132H is expressed in cancer cells adjacent to the arteriolar adventitia (A) and dispersed throughout the sections (B). (C) IDH1R132H is not expressed in IDH1 wild-type samples. (D) Negative control incubation performed in the absence of the primary antibody. Scale bar = 20 µm. Abbreviations: IDH1, isocitrate dehydrogenase 1; a, arteriole; ad, tunica adventitia; m, tunica media; e, endothelium; v, venule. Arrows indicate IDH1R132H-positive glioblastoma cells.
Expression of Blood Vessel Wall Proteins in Glioblastoma
Chromogenic IHC on glioblastoma serial sections showed that CD31 was expressed by endothelial cells of arterioles, venules, and capillaries (Fig. 3A and B). Expression of SMA was found in smooth muscle cells of arteriolar and venular walls (Fig. 3C and D).
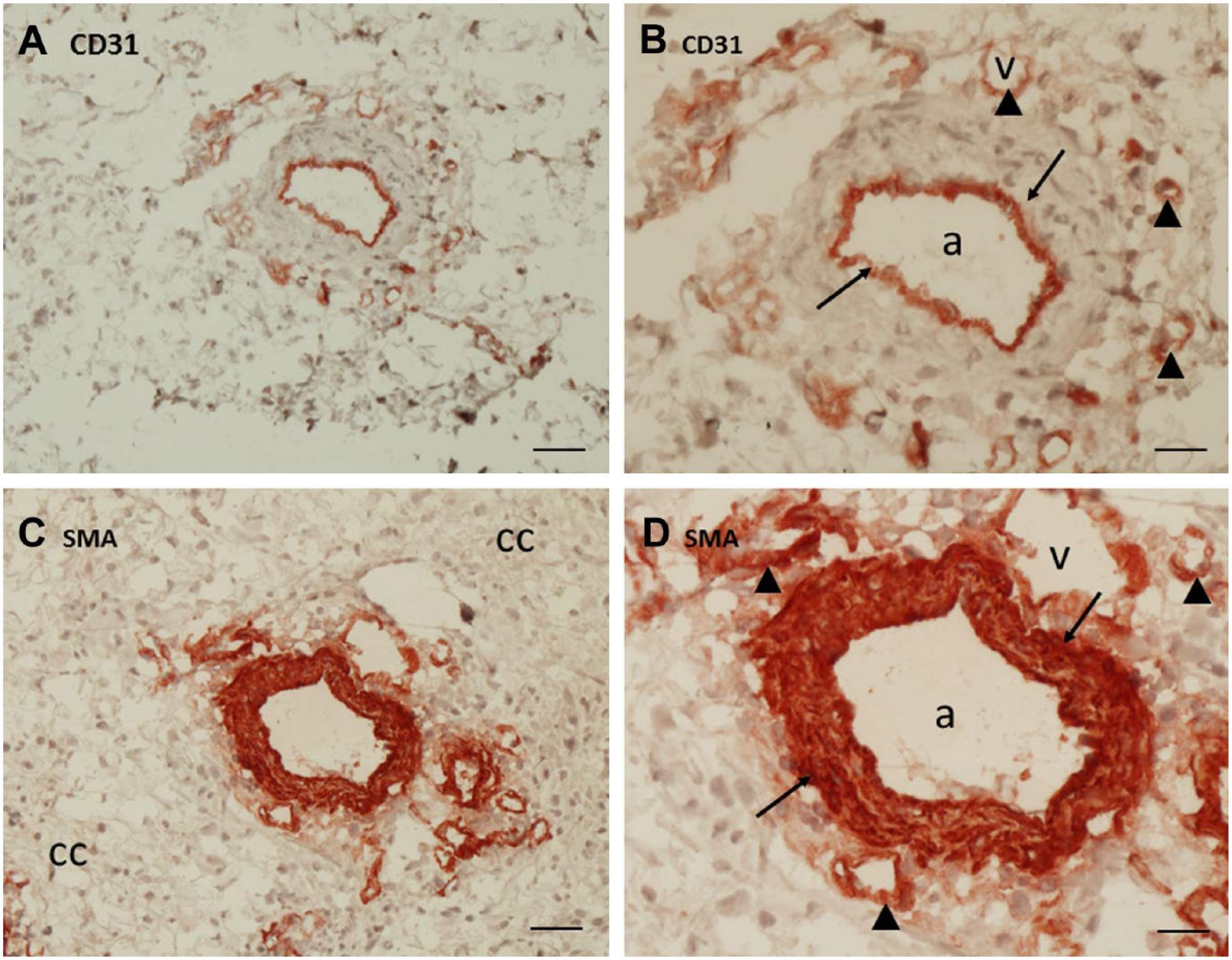
Immunohistochemical localization of CD31 (A, B) and SMA (C, D) in serial cryostat sections of human glioblastoma. (A, B). CD31 expression is present in endothelial cells of a large arteriole (arrows) and of small blood vessels around the large arteriole (arrow heads). (C, D). SMA expression is present in smooth muscle cells in the wall of the large arteriole (arrows) and in some of the small blood vessels around the large arteriole (arrow heads). A, C: bar = 40 µm; B, D: bar = 20 µm. Abbreviations: SMA, smooth muscle actin; a, arteriole; v, venule; cc, glioblastoma cells.
Expression of GSC Markers
We performed IHC using the most widely used GSC markers CD13312,37–47 and nestin.37,39,45–47,49–51,87 The staining revealed that strong CD133 and nestin expression was present in a cellular monolayer adjacent to the tunica adventitia of CD31-positive and SMA-positive large arterioles (Figs. 3 and 4). CD133 and nestin are not specific as GSC markers, because both markers were also expressed in endothelial cells, astrocyte-like cells, and cells in the arteriolar wall (Fig. 4A–D). Taken together, CD133 and nestin were selectively expressed adjacent to CD31-positive and SMA-positive arterioles.
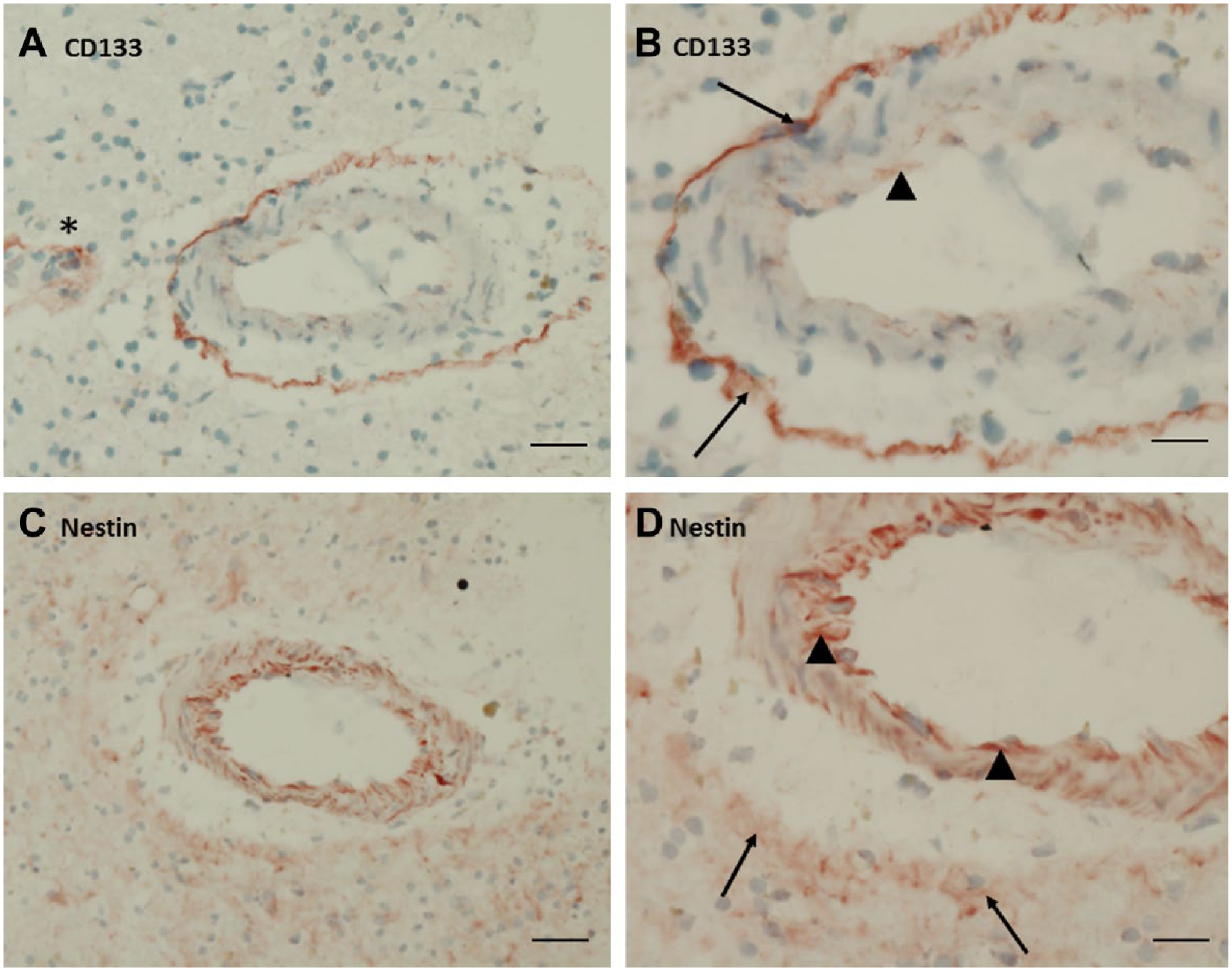
Immunohistochemical localization of CD133 (A, B) and nestin (C, D) in serial cryostat sections of human glioblastoma. (A, B) Strong CD133 expression was present in cells in a rim around a large arteriole (arrows). Low CD133 positivity was found in endothelial cells (arrow head). * indicates CD133 positivity in and around small blood vessels adjacent to the arteriole. (C, D) Low expression of nestin was found in a rim around a large arteriole (arrows) and strong expression was found in endothelial cells and in cells of the arteriolar wall (arrow heads). A, C: bar = 40 µm; B, D: bar = 20 µm.
Expression of Chemoattractive Signaling Proteins in Glioblastoma
Receptor CXCR4 and its ligand SDF-1α were strongly expressed around arterioles (Fig. 5A–D) in serial sections of glioblastoma. CXCR4 expression was found in a cellular monolayer adjacent to the tunica adventitia in a similar way as CD133 and nestin were expressed (Fig. 4). CXCR4 and SDF-1α were also found in endothelial cells (Fig. 5A–D). CatK, the protease that can inactivate SDF-1α, was also found around arterioles (Fig. 5E and F). SDF-1α and CatK were strongly expressed as a thin rim at the edge of the tunica adventitia (Fig. 5C–F). Low expression of CatK was found in endothelial cells (Fig. 5E and F). Receptor CD44 and its ligand OPN were found around arterioles (Fig. 5G–L). OPN was localized in a thin rim at the edge of the tunica adventitia whereas CD44 was found in a cellular monolayer adjacent to the tunica adventitia. OPN expression was also found in endothelial cells and cells in the arteriolar wall (Fig. 5G–J). CD44 was expressed all over the sections, but the strongest expression of CD44 was found in the cellular monolayer around arterioles (Fig. 5K and L). Taken together, the chemoattractive signaling proteins SDF-1α, CXCR4, CatK, OPN, and CD44 were selectively expressed around arterioles. Besides, confocal imaging data show clear co-localization of SDF-1α and CatK in the arteriolar adventitia of SMA-positive arterioles (Fig. 6).
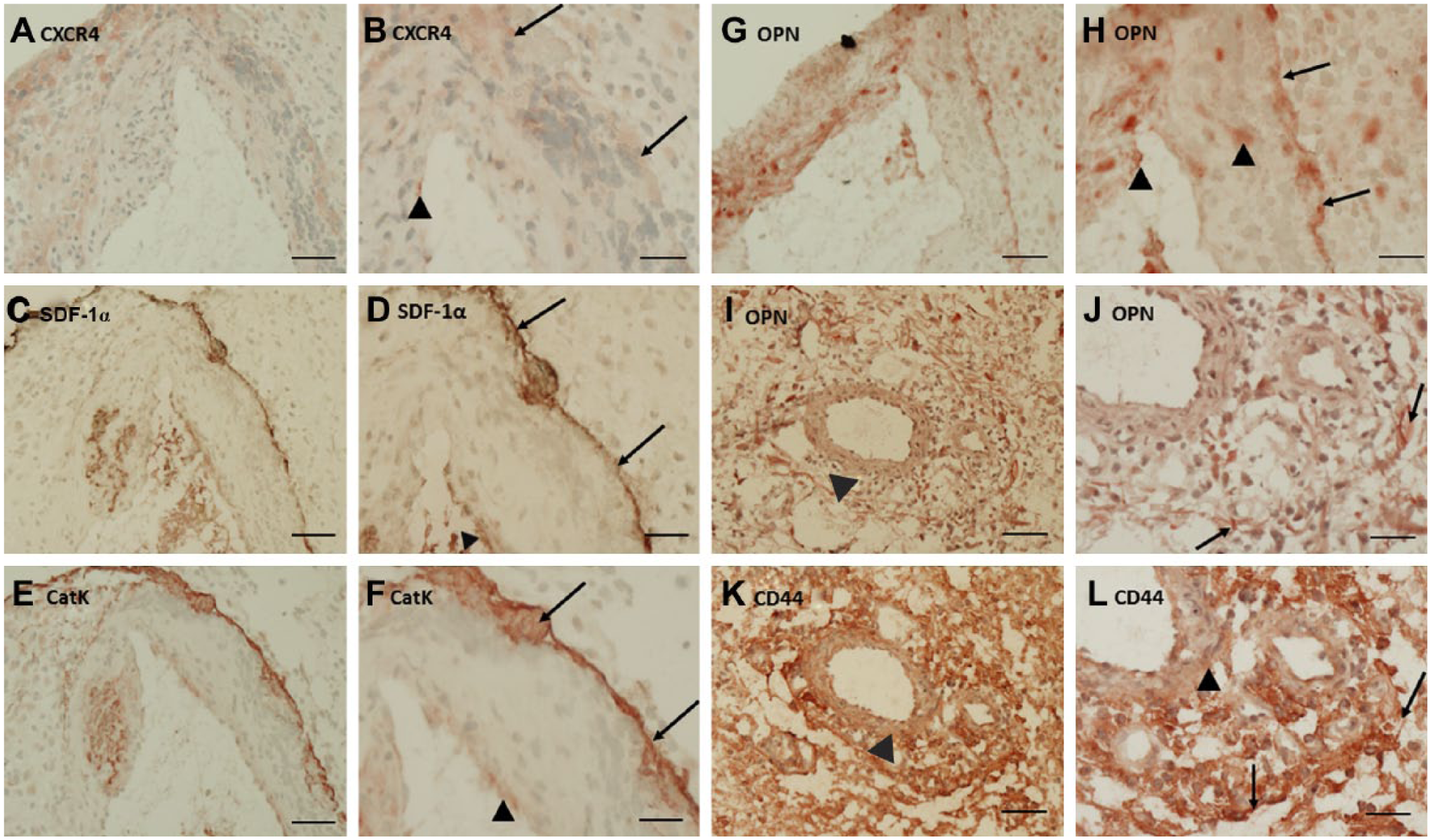
(A) Immunohistochemical localization of CXCR4 (A, B), SDF-1α (C, D), CatK (E, F), OPN (G–J), and CD44 (K, L) in serial cryostat sections of human glioblastoma. (A, B) CXCR4 expression was present in a cellular rim around a large arteriole (arrows) and in some endothelial cells (arrow head). (C, D). Strong SDF-1α expression was found in a thin rim around a large arteriole (arrows) and in endothelial cells (arrow head). (E, F). Strong CatK expression was found in a thin rim around a large arteriole (arrows) and low expression of CatK was found in endothelial cells (arrow head). (G–J). OPN expression was present in a thin rim around a large arteriole (arrows), in endothelial cells and cells in the arteriolar wall (arrow heads). (K, L) CD44 expression was found all over the section, but the positivity was strongest in a cellular rim around a large arteriole (arrows). A, C, E, G, I, K: bar = 40 µm; B, D, F, H, J, L: bar = 20 µm. Abbreviations: CXCR4, C-X-C chemokine receptor type 4; SDF-1α, stromal cell–derived factor-1α; CatK, cathepsin K; OPN, osteopontin.
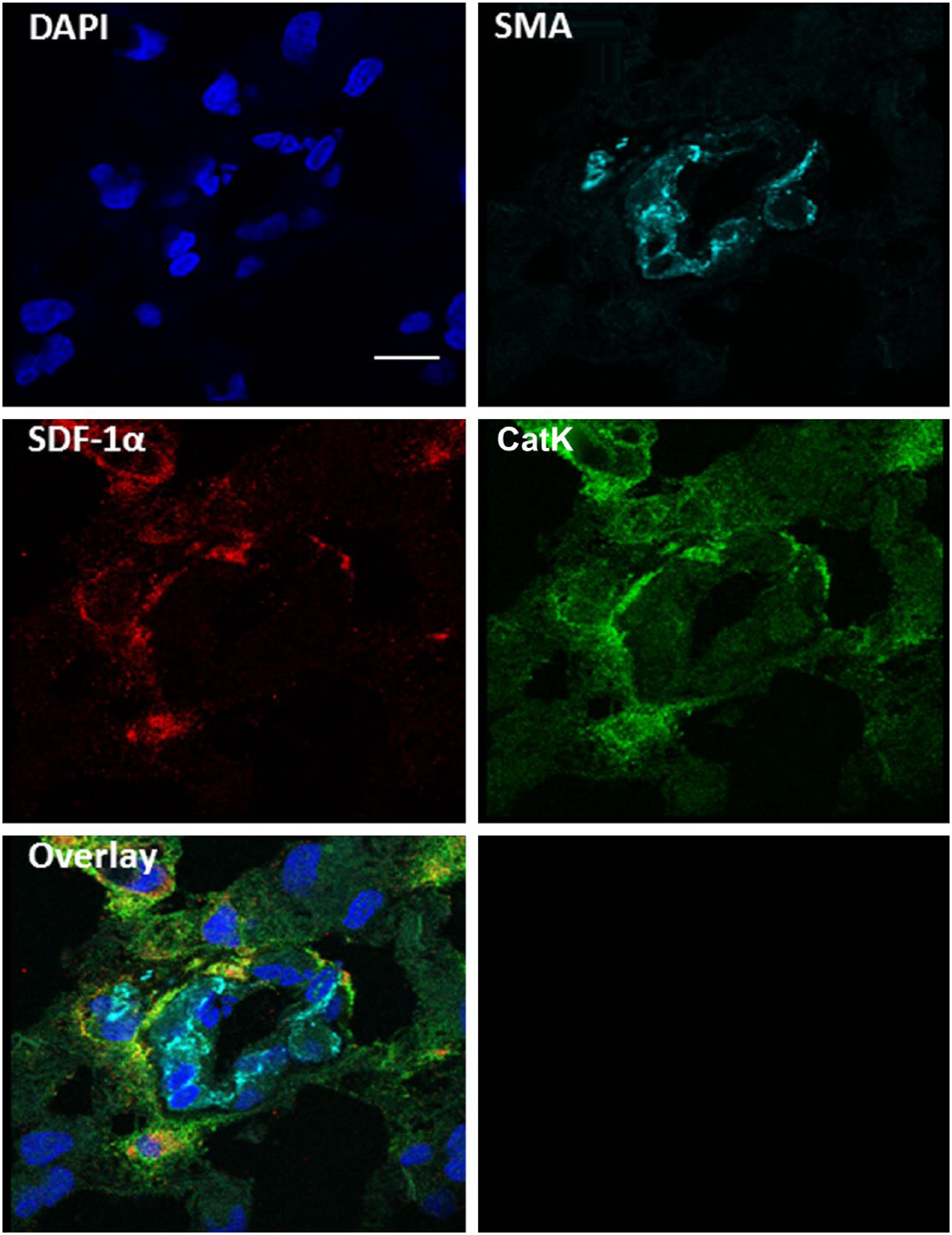
Co-localization of SDF-1α and CatK in niches around SMA-positive arterioles. Fluorescence immunohistochemical staining of SMA (light blue), SDF-1α (red), and CatK (green) in human serial cryostat sections of glioblastoma shows co-localization of SDF-1α and CatK around SMA-positive arterioles. DAPI was used as counterstaining to detect cell nuclei. Imaging was performed using confocal microscopy. Scale bar = 10 µm. Abbreviations: SDF-1α, stromal cell–derived factor-1α; CatK, cathepsin K; SMA, smooth muscle actin; DAPI, 4′,6-diamidino-2-phenylindole, dihydrochloride.
Expression of Hypoxia Markers in Glioblastoma
Glioblastoma sections were stained for hypoxia markers HIF-1α and VEGF (Fig. 7). HIF-1α was expressed in nuclei of cells around arterioles, in endothelial cells and in cells adjacent to necrotic areas (Fig. 7A and B). VEGF was highly expressed in individual cells around the tunica adventitia of arterioles, as well as in smooth muscle cells in the arteriolar wall and outside the niches in areas of differentiated glioblastoma cells (Fig. 7C and D).
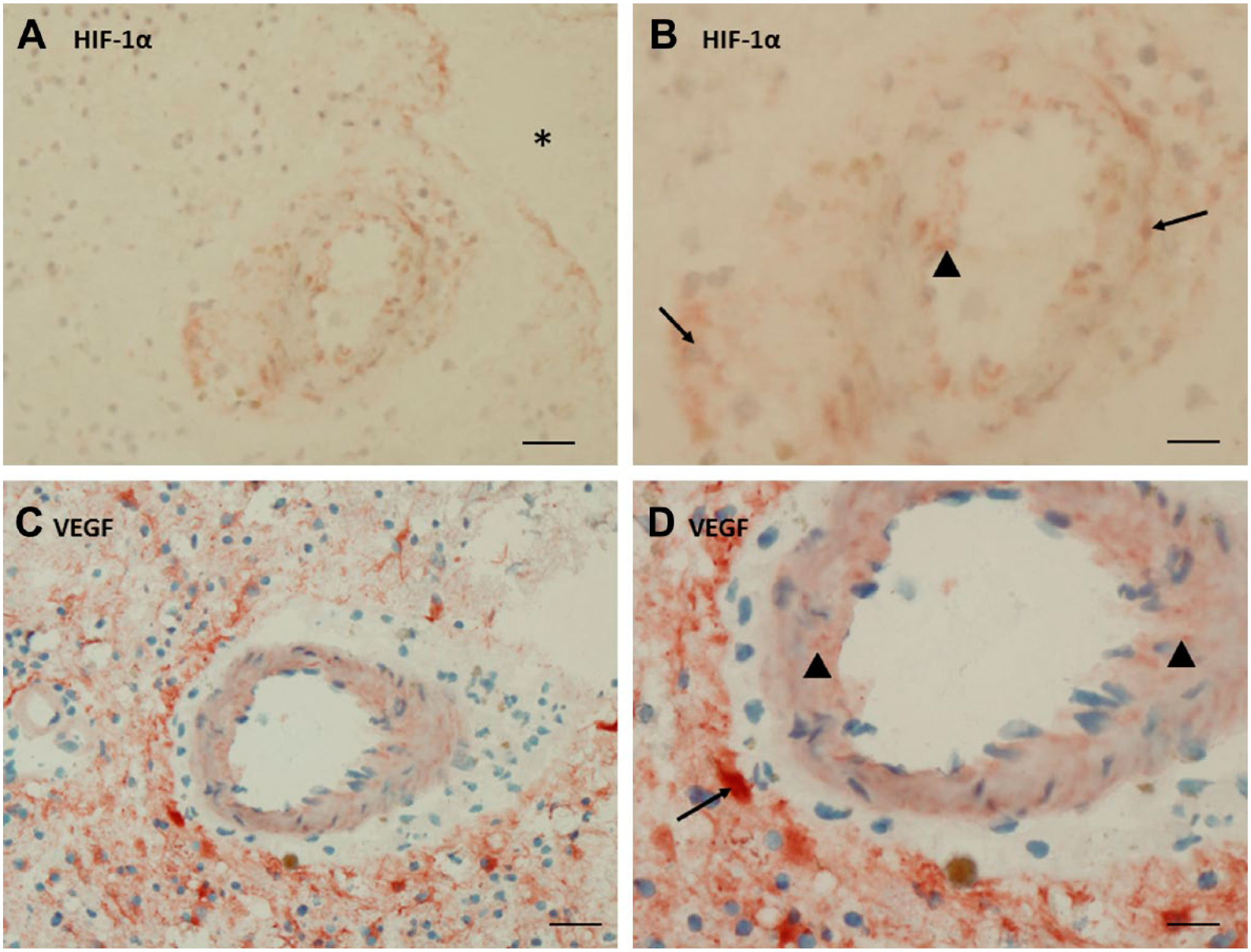
Immunohistochemical localization of HIF-1α (A, B) and VEGF (C, D) in serial cryostat sections of human glioblastoma. (A, B). HIF-1α expression was present in nuclei of cells in and around a large arteriole (arrows) and in endothelial cells (arrow head). * indicates necrotic area. (C, D). VEGF expression was found in astrocyte-like cells in a rim around a large arteriole (arrow) and in the arteriolar wall (arrow heads). A, C: bar = 40 µm; B, D: bar = 20 µm. Abbreviations: HIF-1α, hypoxia-inducible factor-1α; VEGF, vascular endothelial growth factor.
Histological Blood Vessel Analysis and Quantification
Our findings suggest that the markers for GSCs and their niches are only present in a rim around arterioles and not around any other type of blood vessel. To evaluate these findings quantitatively, we counted the number of CD31-positive and SMA-positive arterioles and venules and small CD31-positive blood vessels (capillaries and small venules) in sections of 16 samples of glioblastoma. We determined whether markers of GSCs and their niches were exclusively present around arterioles or also around venules or around small blood vessels. The results are summarized in Table 2 which shows that in total seven niches were found in the sections of the glioblastoma samples. Those seven GSC niches were all present around CD31-positive and SMA-positive arterioles. It means that 2% of the 335 CD31-positive and SMA-positive arterioles in all sections of the 16 glioblastoma samples were surrounded by GSC niches. GSC niches were not found around any of the 924 CD31-positive and SMA-positive venules or any of the 8085 CD31-positive small blood vessels in the sections (Table 2).
Quantitative Analysis of the Number of Blood Vessel Types in Cryostat Sections of the 16 Glioblastoma Samples.
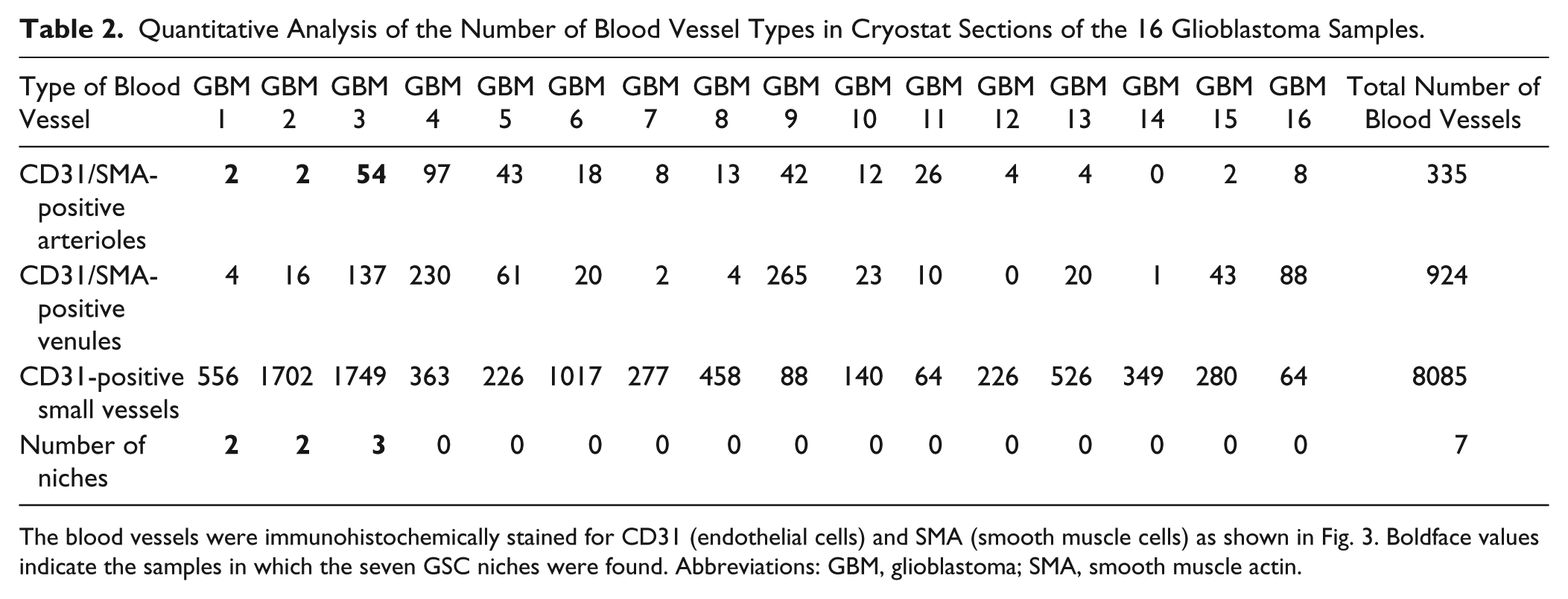
The blood vessels were immunohistochemically stained for CD31 (endothelial cells) and SMA (smooth muscle cells) as shown in Fig. 3. Boldface values indicate the samples in which the seven GSC niches were found. Abbreviations: GBM, glioblastoma; SMA, smooth muscle actin.
Discussion
In the present study, we aimed to determine immunohistochemically which bone marrow HSC niche proteins are expressed in GSC niches in glioblastoma. In addition, we determined the type of blood vessels that is associated with GSC niches. Our findings in the present study of 16 glioblastoma samples are in agreement with the findings of our previous study of five glioblastoma samples, 46 which indicates that our data of GSCs being localized exclusively around hypoxic periarteriolar niches are reproducible and correct.
In the 16 glioblastoma samples that were analyzed, we found seven GSC niches exclusively around arterioles with CD31-positive endothelium and a thick SMA-positive tunica media containing smooth muscle cells. No GSC niches were found around CD31- and SMA-positive venules or CD31-positive small blood vessels (Table 2). To determine whether the CD133-positive and nestin-positive cells are cancer cells, we stained for the IDH1R132H mutation that specifically detects cancer cells in the glioblastoma samples. As these IDH1R132H-mutated cancer cells are also positive for the stem cell markers CD133 and nestin, we have defined these cells as GSCs. In the present study, the number of periarteriolar niches was relatively low in comparison with our first study. 46 This discrepancy is likely due to the notorious heterogeneity in glioblastoma tumors. We do not have information about the exact region of the tumor where the samples are derived from. Talasila et al. recently reported that glioblastoma tumors have invasive areas and angiogenic areas. It is claimed that the angiogenic areas are mainly in the core of the tumors, whereas the invasive areas are more peripheral. 88 Because of the extensive necrotic and hypoxic areas around which GSC niches are found, we are convinced that the GSC niches are mainly present in the core of the tumors. Ongoing research is conducted to investigate this matter. Besides this type of heterogeneity, there is also heterogeneity of subclasses, the proneural, classical, and mesenchymal subtypes. 89
A schematic representation of our periarteriolar GSC niche concept is shown in Fig. 8A and B, based on the HSC niche concept (Fig. 8C and D), and it summarizes our present findings. The arteriolar wall consists of CD31-positive endothelial cells (tunica intima), SMA-positive smooth muscle cells (tunica media), and the stroma of the arteriolar adventitia (tunica adventitia). The niche factors SDF-1α and OPN are expressed in the thin rim directly adjacent to the tunica adventitia to retain CD133-positive and nestin-positive GSCs in the niche at the edge of the tunica adventitia by binding to the receptors CXCR4 and CD44 on GSCs, respectively. The endopeptidase CatK is expressed in the GSC niche in a similar way as SDF-1α and OPN. In the GSC niches, CatK is present in its inactive form. 77 When CatK is activated, it can cleave and inactivate SDF-1α, 72 liberating GSCs out of niches, where GSCs differentiate and become glioblastoma cells. In the HSC niche, HSCs are localized near arterioles close to bone and are retained in the periarteriolar niche compartment by SDF-1α–CXCR4 interactions and OPN–CD44 interactions. CatK can in turn release HSCs out of the niche, allowing HSCs to differentiate into hematopoietic progenitor cells in close vicinity of sinusoids (Fig. 8C and D). This emphasizes the similarities in the proteins that are expressed in both GSC niches and HSC niches, suggesting that the GSC niche functions similarly as the normal HSC niche in bone marrow.46,66 The images that we show are unfortunately not all of the same periarteriolar GSC niches. Due to frequently occurring necrosis in the glioblastoma samples, this is hard to achieve for all markers in one series of sections. Despite this problem, we show the most clear images of all markers in the niches. To confirm whether our observations were correct, we have included confocal imaging of SMA, SDF-1α, and CatK in Fig. 6 to show their co-localization in one periarteriolar GSC niche.
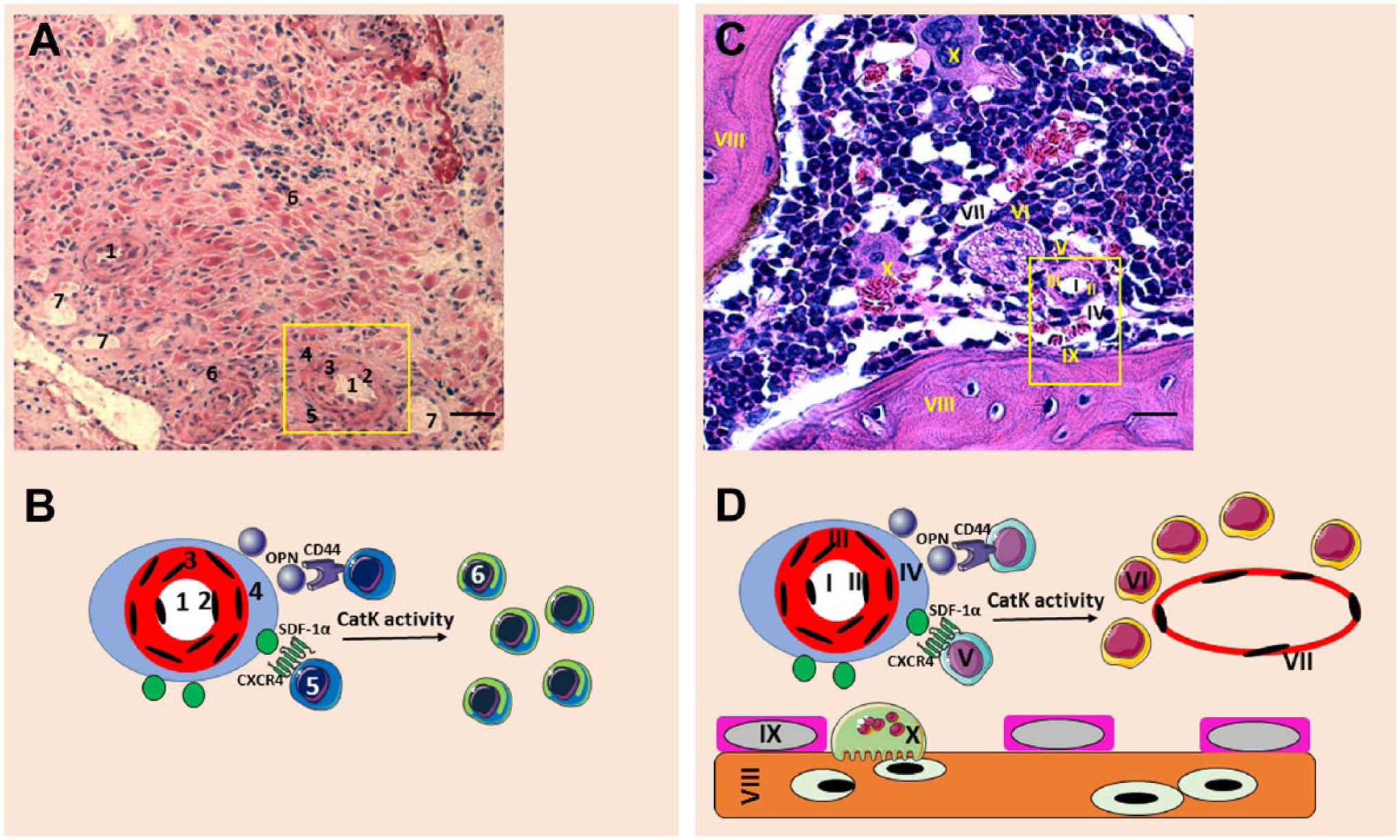
Overview of the similarities between GSC niches in glioblastoma and HSC niches in bone marrow. (A) Hematoxylin and eosin staining of human glioblastoma tissue sections showing a GSC niche around an arteriole (yellow rectangle). 1, lumen of arteriole; 2, endothelial cells; 3, smooth muscle cells; 4, arteriolar adventitia; 5, GSCs; 6, glioblastoma cells; 7, venule. Scale bar = 40 µm. (B) Schematic representation of the GSC niche showing that SDF-1α/CXCR4 interactions and OPN/CD44 interactions at the arteriolar adventitia retain GSCs in the GSC niche and that CatK activity releases GSCs out of the niche, so that GSCs differentiate into glioblastoma progenitor cells. The numbers in the illustration correspond with the numbers in 8A. (C). Giemsa staining of murine bone marrow tissue, showing an HSC niche around an arteriole (yellow rectangle). I, lumen of arteriole; II, endothelial cells; III, smooth muscle cells of the arteriolar tunica media; IV, arteriolar adventitia; V, HSCs; VI, hematopoietic progenitor cells; VII, sinusoid; VIII, bone; IX, osteoblast; X, osteoclast. Scale bar = 60 µm. (D). Schematic representation of the HSC niche showing that SDF-1α/CXCR4 interactions and OPN/CD44 interactions at the arteriolar adventitia retain HSCs in the HSC niche and that CatK activity releases HSCs out of the niche, so that HSCs differentiate into hematopoietic progenitor cells in vicinity of sinusoids and subsequently migrate out of the HSC niche into the circulation. The numbers in the illustration correspond with the numbers in 8C. Abbreviations: GSC, glioblastoma stem cell; HSC, hematopoietic stem cell; SDF-1α, stromal cell–derived factor-1α; CXCR4, C-X-C chemokine receptor type 4; OPN, osteopontin; CatK, cathepsin K.
We did not investigate the distribution pattern of white blood cells in glioblastoma samples in relation to the GSC niches. The localization of tumor-associated macrophages in glioblastoma (TAMs; M2 type of macrophages 36 ) and whether the stemness of GSCs is affected by TAMs are investigated at present.
The tunica adventitia of human arterioles and veins contain mesenchymal stem cell (MSC) niches.90–94 These niches are not in the same location as we found GSC niches. We found the GSC niches to be localized attached to the adventitia, whereas the MSC niches have been found at the other side of the adventitia adjacent to the tunica media.91,93,94 MSC niches in the arteriolar adventitia have been found to contain malignant stem cells, such as leukemic and glial stem cells. 92 Ergun et al. argued correctly that these niches are not perivascular but part of the vascular wall. 92 However, the GSC niches that we describe in the present study are perivascular, because the GSCs are found at the outer rim of the adventitia. To elucidate the exact localization of periarteriolar GSC niches and MSC niches in arteriolar walls and whether there is a functional relationship between periarteriolar GSC niches and arteriolar MSC niches needs to be further investigated. Nevertheless, it can be concluded that the tunica adventitia of arterioles (and possibly veins) is a microenvironment that is well suited for GSCs, most likely in combination with hypoxic conditions and expression of HIF-1α and VEGF.
Hypoxia markers HIF-1α and VEGF were expressed in the areas of periarteriolar GSC niches and in the vicinity of necrotic areas (Fig. 7). HIF-1α and VEGF are likely important to maintain the GSC niches, because they both upregulate CXCR4,54–57 SDF-1α,54–56 OPN,58–60 and CD4413,58 in glioblastoma. MicroRNA-21 (miR-21) is the most consistently overexpressed miRNA in glioma and is co-localized with HIF-1α and VEGF, but miR-21 expression was not associated with GSCs but rather with angiogenesis. 87 miR-21 expression has been associated with Notch-1 expression in cancer stem cells. 95
Hypoxia and necrosis around arterioles can be explained by the fact that oxygen is not released into tissues from erythrocytes in arterioles (and venules) which are transport vessels, but only in capillaries which are exchange vessels. 96 We bear in mind that GSC niches can also be present around SMA-positive venules, as these vessels are transport vessels where oxygen release does not occur. 96 However, we did not find GSC niches surrounding SMA-positive venules in our glioblastoma samples.
Calabrese et al. concluded that GSCs are found in close vicinity of capillaries on the basis of anti-CD34 IHC and defined this as perivascular niches. 97 However, the endothelial cell marker CD34 not only detects endothelium of capillaries 97 but also endothelium of large blood vessels. 98 Besides, only nestin was used as GSC marker in this study, which is also expressed by astrocytes in close vicinity of capillaries as was clearly shown in our previous study, 46 indicating that these astrocytes were wrongly taken as GSCs. In addition, the authors state correctly that GSCs in niches are in a quiescent state but show that endothelial cells of the capillaries induce proliferation of GSCs and are thus important for tumor propagation. 97 As quiescence of GSCs is maintained in the niches and not their proliferation, we conclude that Calabrese et al. have studied propagation of glioblastoma cells, which are possibly progenitor glioblastoma cancer cells in a similar way as hematopoietic progenitor cells around sinuses.66,111
The search for therapeutics that specifically target GSCs is increasingly gaining interest.20,99 For example, ibrutinib, a Bruton’s tyrosine kinase inhibitor, is promising to target GSCs. 100 Furthermore, proteins involved in invasive behavior and therapy resistance of glioblastoma, such as TROY and SGEF, may well be overexpressed especially in GSCs which warrants further research with respect to specific therapeutic targeting of GSCs.101,102 Alternatively, interactions between SDF-1α/CXCR4 and OPN/CD44 should be disrupted to enforce GSCs out of their protective niches to sensitize them to chemotherapy and irradiation. Richardson (2016) discussed along this line the application of a CXCR4 antagonist, and very recently, the effects of a brain-penetrating CXCR4 antagonist PRX177561 in preclinical models of human glioblastoma were reported.103,104,112 We suggest that this approach has therapeutic value for glioblastoma patients, as a similar approach has shown to be promising for acute myeloid leukemia (AML) patients.66,105 In AML, CXCR4-positive leukemic stem cells (LSCs) are protected by the normal HSC niches, which causes therapy resistance and a poor patient prognosis.106–108 In phase I/II clinical trials, AML patients are treated with the CXCR4 inhibitor plerixafor in combination with chemotherapeutic agents. Plerixafor treatment resulted in a twofold increased mobilization of AML cells in the peripheral blood, and the overall complete response rate was 46% which is higher than that of chemotherapeutic agents alone (21%). It is assumed that the underlying mechanism is the enforcement of LSCs out of the HSC niches.66,105
LSCs also express CD44 and CD44–OPN interactions can also facilitate anchorage of LSCs in HSC niches. 109 Neutralizing antibodies against OPN sensitized leukemic cells to Ara-C chemotherapy in acute lymphoblastic leukemia (ALL) xenografts.66,109,110 These studies indicate that inhibition of homing of LSCs in HSC niches sensitizes these cells to therapy.66,105,109,110 Therefore, we are convinced that inhibition of homing of GSCs in their niches may also be beneficial for glioblastoma patients.
In conclusion, bone marrow HSC niche proteins are also expressed in GSC niches in 16 glioblastoma samples. The hypoxic periarteriolar GSC niches are found around CD31-positive and SMA-positive arterioles and express GSC markers CD133 and nestin, niche markers SDF-1α, OPN, CatK, and CD44 and hypoxia markers HIF-1α and VEGF. The frequency of GSC niches around large arterioles is 2% in the glioblastoma samples that we analyzed.
Footnotes
Competing Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
VVVH and CJFN designed the study, performed experiments, analyzed and interpreted the data, gave intellectual input, revised the manuscript, approved the final version of the manuscript, and agreed to be accountable for all aspects of the work. JRW, HK, BB, MK, BS, RH, and WT performed experiments, analyzed and interpreted the data, revised the manuscript, approved the final version of the manuscript, and agreed to be accountable for all aspects of the work. ZT, MK, and RJM interpreted the data, revised the manuscript, approved the final version of the manuscript, and agreed to be accountable for all aspects of the work.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was financially supported by the Dutch Cancer Society (KWF; UVA 2014-6839; V.V.V.H., M.K., R.J.M., C.J.F.N.), René Vogels Foundation (V.V.V.H.), Jo Kolk Foundation (V.V.V.H.), and an AMC PhD scholarship (R.J.M.).
