Abstract
The triple-negative breast cancer (TNBC) subtype occurs in about 15% of breast cancer and is an aggressive subtype of breast cancer with poor outcome. Furthermore, treatment of patients with TNBC is more challenging due to the heterogeneity of the disease and the absence of well-defined molecular targets. Microribonucleic acid (RNA) represents a new class of biomarkers that are frequently dysregulated in cancer. It has been described that the microRNA miR-210 is highly expressed in TNBC, and its overexpression had been linked to poor prognosis. TNBC are often infiltrated by immune cells that play a key role in cancer progression. The techniques traditionally used to analyze miR-210 expression such as next generation sequencing or quantitative real-time polymerase chain reaction (PCR) do not allow the precise identification of the cellular subtype expressing the microRNA. In this study, we have analyzed miR-210 expression by in situ hybridization in TNBC. The miR-210 signal was detected in tumor cells, but also in the tumor microenvironment, in a region positive for the pan-leucocyte marker CD45-LCA. Taken together, our results demonstrate that miR-210 is expressed in tumor cells but also in the tumor microenvironment. Our results also highlight the utility of using complementary approaches to take into account the cellular context of microRNA expression.
Introduction
Triple-negative breast cancers (TNBC) are defined by the lack of estrogen receptor, progesterone receptor, and HER2 gene amplification and are among the most aggressive breast cancer subtypes, with high rates of tumor recurrence and poor survival. TNBC are still poorly characterized at the molecular level, and it is recognized that TNBC is a heterogeneous disease. Different subtypes have been defined at the molecular level: basal-like, immunomodulatory, mesenchymal, mesenchymal stemlike, and a luminal androgen receptor subtype.1,2 To add to the complexity, inflammatory cells such as tumor infiltrating macrophages (TAM) or lymphocytes (TIL) play an essential role in tumor development and treatment response in TNBC. A high level of TIL is associated with improved disease-free survival and overall survival3,4 while a high infiltration of macrophages is associated with poor outcome. 5
Because of the heterogeneity of the disease and the absence of well-defined molecular targets, the treatment of TNBC patients is still very challenging. The identification of new molecular markers could increase our understanding of the TNBC subtype and help to design new treatments.6,7
MicroRNAs (miRNAs) are 19–24 nucleotides non-coding RNAs that control gene expression by targeting messenger RNAs (mRNA) and inducing translational repression or RNA degradation. Since several years, miRNAs have emerged as a new class of molecular biomarkers, because they are frequently dysregulated in cancer, acting as tumor suppressors or oncogenes.8–10 In that context, several miRNA profiles of breast cancer have been established by microarray or next generation sequencing (NGS), in which the tumor miRNA signatures have been correlated to clinico-pathological and prognostic parameters such as tumor size, lymph-node invasion, receptors status, and resistance to chemotherapy. One of the miRNA identified in these profiles, the miRNA miR-210, is a hypoxia regulated miRNA and a target of hypoxia inducible factor 1 alpha (HIF1-alpha). As hypoxia is an important feature of solid tumors, the role of miR-210 in cancer progression has been extensively studied, and it was shown that miR-210 is overexpressed in numerous cancers including breast, lung, head and neck, pancreatic cancer, or glioblastoma.11,12 MiR-210 has been described mainly as an oncogene but it is also suggested that it can act as a tumor suppressor, for example, in esophageal squamous cell carcinoma. 13 MiR-210 has multiple functions in cancer cells but also in normal cells, and is involved in angiogenesis, control of stem cell differentiation, cell cycle regulation, DNA damage repair, mitochondrial metabolism, and immune response.11,14
MiR-210 was shown to be upregulated in TNBC compared with estrogen positive breast cancer, and the overexpression of miR-210 in breast cancer is correlated with a poor prognosis, being associated with aggressiveness and shorter time to distant metastasis.15–17 MiR-210 is also implicated in the transition from in situ to invasive breast carcinoma, and its expression was shown to be inversely correlated to breast cancer 1 (BRCA1) expression level in breast cancer samples. 18 BRCA1 is a tumor suppressor protein that preserves genome integrity, and about one third of the TNBC have a deficiency in DNA double-strand break repair mechanisms due to the inactivation of BRCA1 by mutation (in familial breast cancer) or epigenetic mechanisms. MiR-210 has also been identified as one of the miRNA upregulated in familial breast cancer compared with non-familial breast cancer. 19
An important consideration related to miRNA signatures obtained using NGS, microarray, or quantitative real-time PCR is that they are obtained using RNA extracted from frozen or formalin-fixed paraffin-embedded (FFPE) tumor samples with various percentages of tumor, stromal, or inflammatory cells. The main limitation of these techniques in cancer research is that they do not discriminate between miRNA expression in tumor cells or other cellular subtypes. However, there are more and more data reporting the importance of immune infiltrate in TNBC, and approximately 20% of TNBC are highly infiltrated with lymphocytes.3,4 It was also shown that about 35% of the TNBC are highly infiltrated with macrophages. 5 Without techniques allowing the isolation of pure tumor cells RNA, it is difficult to determine precisely the cellular origin of the miR-210 signal.
In situ hybridization (ISH) for miRNA has the advantage that it allows the precise histological localization of miRNAs in situ, at the cellular level. This is the technique of choice to identify precisely the cells expressing miR-210. In this study, we have performed ISH to visualize miR-210 localization in TNBC samples and showed that miR-210 is expressed not only in epithelial cancer cells but also in the tumor microenvironment (TME), particularly in inflammatory cells. This is an important consideration to understand the role of miR-210 in TNBC.
Our study also highlights the limitation in the interpretation of miRNA signatures obtained using RNA extracted from tumor samples and the utility of ISH to identify the cellular population at the origin of the expression, as the microenvironment plays an important role in cancer progression.
Material and Methods
Study Population
FFPE tissues were selected from the biobank of the Institute of Pathology and Genetics (IPG, Gosselies). We analyzed 13 normal mammary tissues (obtained from reduction mammoplasty) and 26 TNBC. The 26 analyzed TNBC lack estrogen receptor (ER) or progesterone receptor (PR) expression (Allred score of 0/8) and are negative for HER2 amplification (confirmed by fluorescent ISH). None of the patients received chemo- or radiotherapy before surgery.
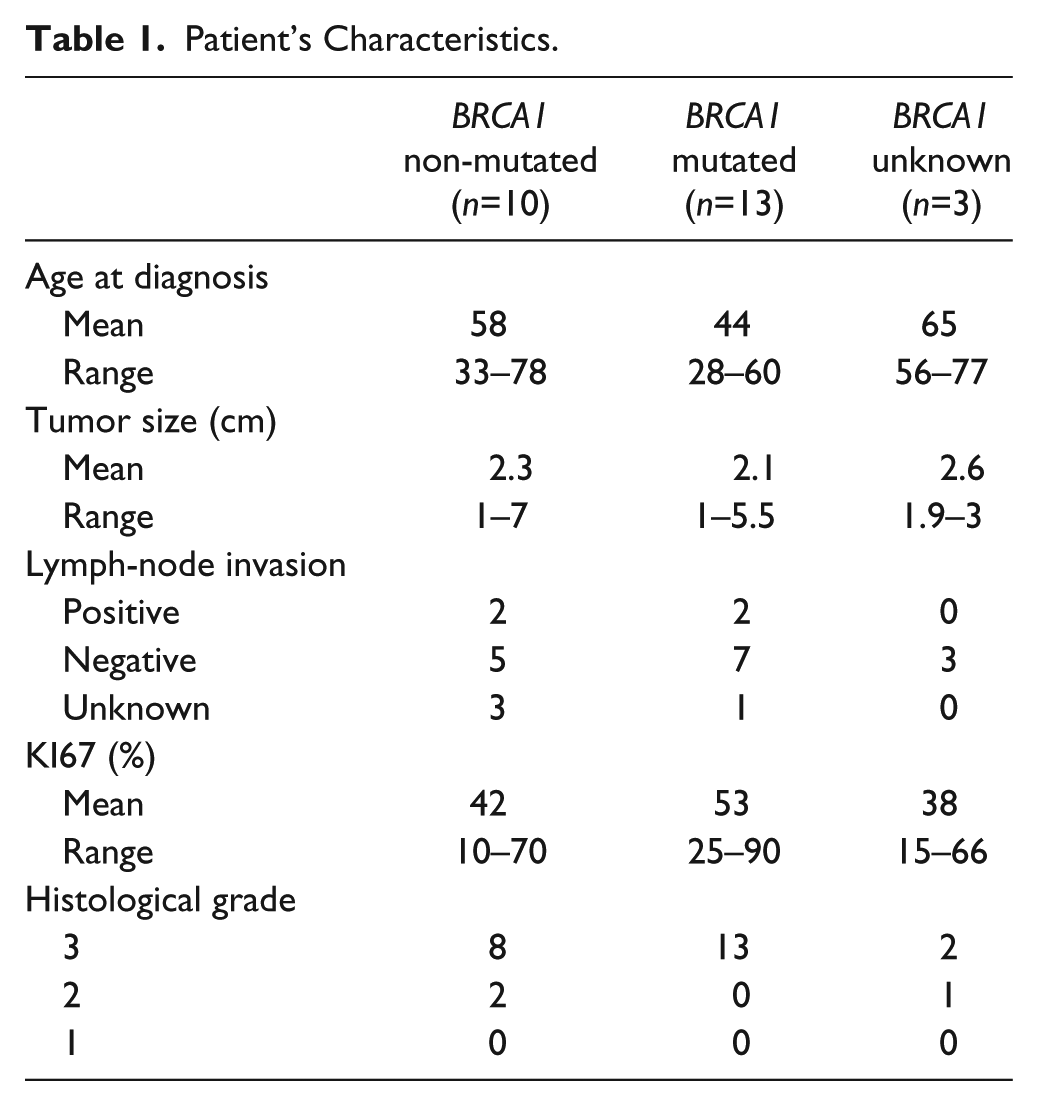
The germline BRCA1 status was known for 23 of the 26 TNBC analyzed (13 mutated cases and 10 non-mutated cases). The BRCA1 status was determined by geneticists with the patient agreement and according to the ethical rules in the context of a familial cancer investigation. All the experiments involving human tissues were conducted with the permission of the ethics committee (OM100) of the “Grand Hôpital de Charleroi” (GHdC). Patients’ characteristics are summarized in Table 1.
Patient’s Characteristics.
RNA Isolation
The tumor zone, selected by the pathologist on HE section, was macrodissected from unstained FFPE sections. Two to four 20 µm sections per tumor were used for RNA extraction. Isolation of total RNAs was performed using Ambion “Recover All Total Nucleic Acid Isolation kit for FFPE” (cat. no AM1975; Life Technologies Europe BV, Belgium) according to the manufacturer’s protocol.
Detection of miRNA by SYBR-Green Real-Time PCR
Total RNA (10 ng) was reverse transcribed in 10 µl reactions using the miRCURY LNA Universal RT miRNA PCR cDNA synthesis kit (cat. no 203301; Exiqon, Denmark), and the cDNA was diluted 80-fold. The miRNA specific primers for miR-210-3p (cat. no 204333) and RNU44 (cat. no 203902) were purchased from Exiqon. MiRNA levels were determined using the Exiqon ExiLENT SYBR Green master mix (cat. no 204403) according to the instructions of the manufacturer on a LightCycler 480. All RT-qPCR reactions were performed in triplicate. The real-time PCR results were analyzed using the 2−ΔCt method. The data were normalized to the expression of small nucleolar RNA 44 (RNU44), a widely used endogenous reference.
Knockdown of BRCA1 Using Small Interfering RNAs
The triple negative cell line SUM-159PT was purchased from Asterand Bioscience (Detroit, MI). The cells were maintained in Ham’s F12 medium (cat. no 11765; Life Technologies) supplemented with 5% fetal bovine serum (cat. no 10270106; Life Technologies), 1 µg/ml hydrocortisone (cat. no H4001; Sigma-Aldrich, Belgium), and 5 µg/ml insulin (cat. no. I9278; Sigma-Aldrich). The cell line was maintained at 37C in a humidified incubator containing 5% CO2. The cells (5 × 105) were seeded into each well of a 6-well plate and cultured in antibiotic-free culture media until they reached 70% confluency. To knock down endogenous BRCA1 protein levels in the SUM-159PT cell line, transient transfection was performed with RNAimax (cat. no 13778150; Life Technologies) using 10 nM “SMARTpool BRCA1” (cat. no M-003461; Dharmacon, GE Healthcare Europe) and a nonspecific control pool (cat. no. D-001206; Dharmacon). The RNAimax reverse transfection protocol was used according to the manufacturer’s instructions.
Protein Assays
Cell lysates were prepared and Western blotting was performed using standard protocol. Equal aliquots of total cell protein (15 µg per lane) were electrophoresed on sodium dodecyl sulfate (SDS)–polyacrylamide gradient gels (4–15% Mini-protean TGX gel, cat. no 456-1084; Bio-Rad Laboratories NV, Belgium), transferred to nitrocellulose membranes (cat. no 162-0214; Bio-Rad Laboratories). Detection was performed using the following primary antibodies: rabbit polyclonal BRCA1 (1:1000, cat. no 9010; Cell Signaling Technology Europe), and mouse monoclonal β-actin (1:50,000, cat. no A3854; Sigma-Aldrich). The proteins were visualized using the Lumi-Light PLUS Western Blotting Substrate (cat. no 12 015 196 001; Roche Applied Science, Mannheim, Germany).
Immunohistochemistry (IHC)
IHC was performed on 4-µm paraffin sections. Heat-induced antigen retrieval was performed in a PT-link pre-treatment module (DAKO, Agilent Technologies). Detection of CD45- leukocyte common antigen (CD45-LCA) was performed on a DAKO autostainer automate with ready-to-use antibody clone 2B11 + PD7/26 (cat. no. M0701; DAKO, Agilent Technologies). Detection of CD3, CD20, CD68, and CD56 was performed on an Omnis automate with ready-to-use antibody (Polyclonal Rabbit Anti-Human CD3 Ready-to-Use (IR503), Monoclonal Mouse Anti-Human CD20cy Clone L26 Ready-to-Use (IR604), Anti-Human CD68 Clone PG-M1 Ready-to-Use (IR613), and Monoclonal Mouse Anti-Human CD56, Clone 123C3 Ready-to-Use (IR628), DAKO, Agilent Technologies).
For the detection of HIF1-alpha and carbonic anhydrase IX (CAIX), sections were incubated overnight with mouse monoclonal anti-HIF1 alpha (dilution 1:25, BD transduction 610958) or mouse monoclonal anti-CAIX antibodies (dilution 1:1500, Santa Cruz sc-365900). The detection was performed with the EnVision+System-HRP Labeled Polymer anti-mouse (DAKO) following manufacturer’s instructions.
MiRNA Locked-Nucleic Acid (LNA) ISH
The detection probes hsa-miR-210-3p (cat. no 612239-360), U6 snRNA (cat. no 699002-350), and the scrambled miRNA (cat. no 699004-360), probes (all 3′ and 5′—end labeled with digoxigenin and LNA-modified) as well as the miRNA ISH buffer with proteinase K (cat. no 9000) were purchased from Exiqon (Vedbaek, Denmark). ISH was performed according to the “One-day miRNA ISH protocol” developed by Exiqon. Briefly, 6-µm FFPE sections were mounted on Superfrost Plus slides (Fisher Scientific, Merelbeke, Belgium). Sections were deparaffinized in fresh xylene and rehydrated. Then, 15 µg/ml of Proteinase K was applied onto the tissue and placed in the hybridization chamber at 37C for 10 min. Then, 50 pmol/µl of probes diluted with 1 × ISH buffer (Exiqon) were applied onto the tissue sections and incubated at 55C for 60 min. The slides were washed in decreasing concentration of saline-sodium citrate (SSC) buffer (5×, 1×, and 0.2×, cat. no S6639; Sigma-Aldrich) at 55C. The miRNA LNA-ISH probes were detected with anti-DIG-alkaline phosphatase antibody (1:800, cat. no 11 093 274 910; Roche Applied Science, Germany) and incubated at 30C for 120 min. The miRNA expression was visualized by 4-nitro-blue-tetrazolium and 5-bromo-4-chloro-3-indolynitrolphosphate substrate (cat. no 11 697 471 001; Roche Applied Science). The slides were counterstained with nuclear fast red (cat. no H-3403; Vector Laboratories Burlingame, CA).
Statistical Analysis
Differences in the expression levels between breast cancer and normal tissue were explored using the Student’s t-test. Data are presented as means ± standard error of the mean (SEM); p>0.05: nonsignificant (NS), p≤0.01: significant (**).
Results
MiR-210 Is Highly Expressed in TNBC Compared with Normal Tissue
To study the expression of miR-210 in TNBC, we first performed RT-qPCR analysis on RNA extracted from 13 normal breast tissues and 26 TNBC (13 TNBC with BRCA1 mutation, 10 non-mutated cases, and 3 cases with unknown BRCA1 status). The tumor zone was macrodissected to remove adjacent normal regions. Similarly to other previously published data,15,16 we found that miR-210 was significantly overexpressed in tumor samples compared with normal breast samples (Fig. 1A).
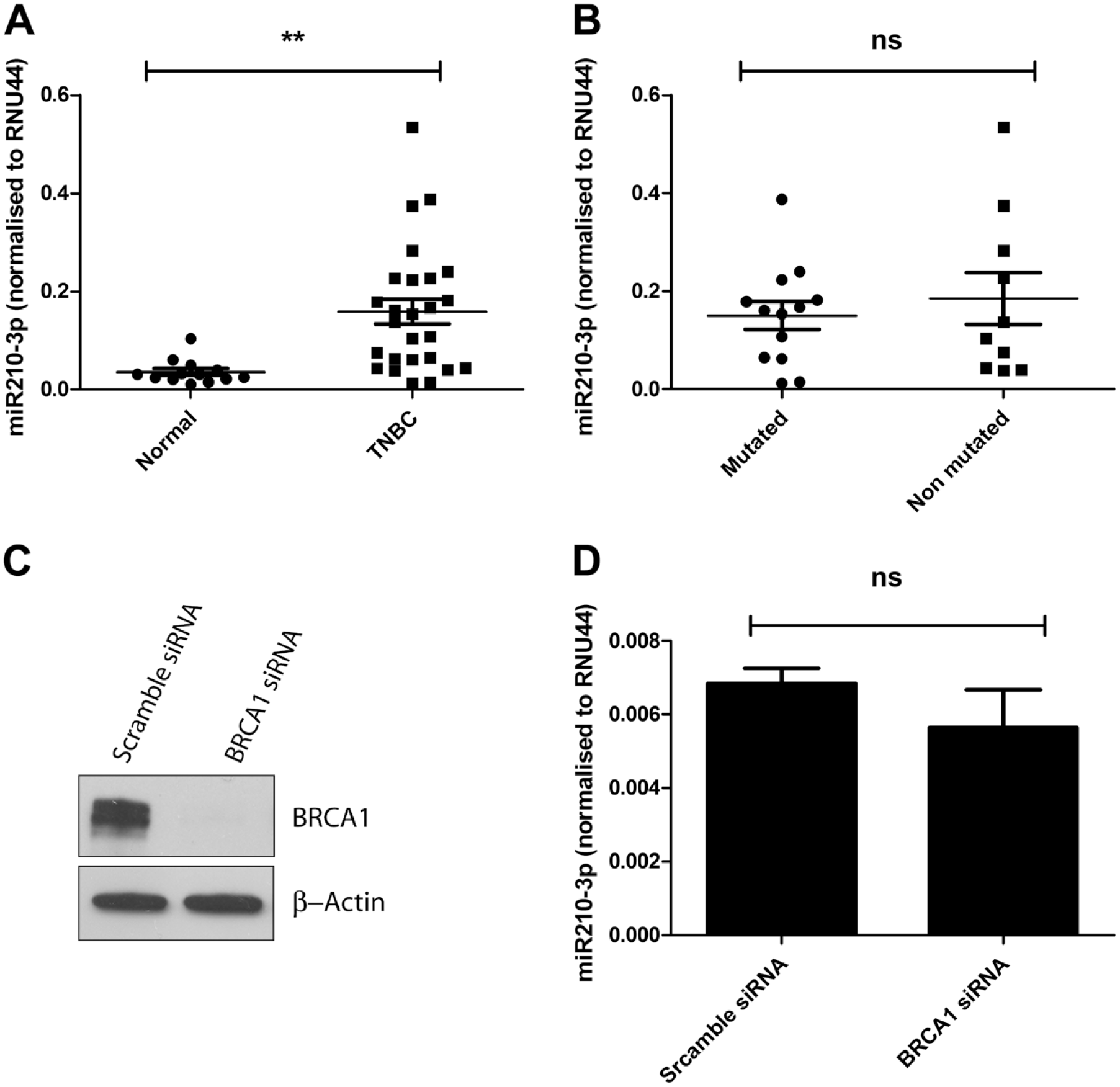
Analysis of miR-210 expression level by qRT-PCR in TNBC. A: distribution of the normalized expression level of miR-210 in 26 TNBC and 13 normal breast samples. MiR-210 is highly expressed in TNBC compared with normal samples (**, p=0.0016, bars represent means ± SEM). B: distribution of the normalized expression level of miR-210 in 13 TNBC with mutated BRCA1 and 10 TNBC with wild type BRCA1 gene: the miR-210 expression is similar between mutated and non-mutated samples (NS, p=0.5399, bars represent means ± SEM). C: Western blot analysis of BRCA1 protein expression in SUM-159PT cells transfected with scramble or BRCA1 siRNA for 72 hr. β-actin is used as loading control. D: analysis of miR-210 expression by RT-qPCR in SUM-159PT cells transfected with scramble or BRCA1 siRNA for 72 hr (normalized to RNU44): The depletion of BRCA1 by siRNA does not influence miR-210 expression (NS, p=0.3391, bars represent means ± SEM of three independent experiments). Abbreviations: qRT-PCR, quantitative real-time polymerase chain reaction; TNBC, triple-negative breast cancer; RNU44, small nucleolar RNA 44; SEM, standard error of the mean.
As triple negative cancers often present a deficiency in BRCA1 expression due to mutation or epigenetic mechanisms, we explored a possible link between miR-210 expression and BRCA1 mutational status (Fig. 1B). We analyzed miR-210 expression in non-mutated and mutated TNBC by RT-qPCR and found that miR-210 expression levels were highly variable in mutated and non-mutated cases, with low- and high-expressing samples. The miR-210 expression was comparable between the BRCA1 mutated and non-mutated breast cancer (Fig. 1B), suggesting that BRCA1 status does not affect miR-210 expression in TNBC.
To further validate these results, we analyzed the effect of BRCA1 silencing by siRNA on miR-210 expression in the triple negative cell line SUM-159PT (Fig. 1C and D). As illustrated in Fig. 1D, knockdown of BRCA1 expression in the TN cell line SUM-159PT does not affect miR-210 expression. These data are consistent with our RT-qPCR data on tumor samples.
Taken together, these results suggest that miR-210 is overexpressed in TNBC independently of BRCA1 status.
Cellular Localization of miR-210 Expression in TNBC
As TNBC are often infiltrated with immune cells, we hypothesized that the RT-qPCR signal could also originate from the non-tumor cells. We, thus, investigated the expression of miR-210 by ISH on FFPE sections of the TNBC samples previously analyzed by RT-qPCR.
The validation of the ISH protocol is illustrated in Fig. 2. As shown in Fig. 2A, the miR-210 probe signal was detected in the tumor cells, and no signal was detected in the TME. No signal was detected using the control scramble probe (Fig. 2B). Furthermore, no miR-210 signal was detected in normal glandular cells obtained from breast reduction (Fig. 2C), while a strong nuclear signal was detected using the U6 snRNA positive control probe (Fig. 2D).
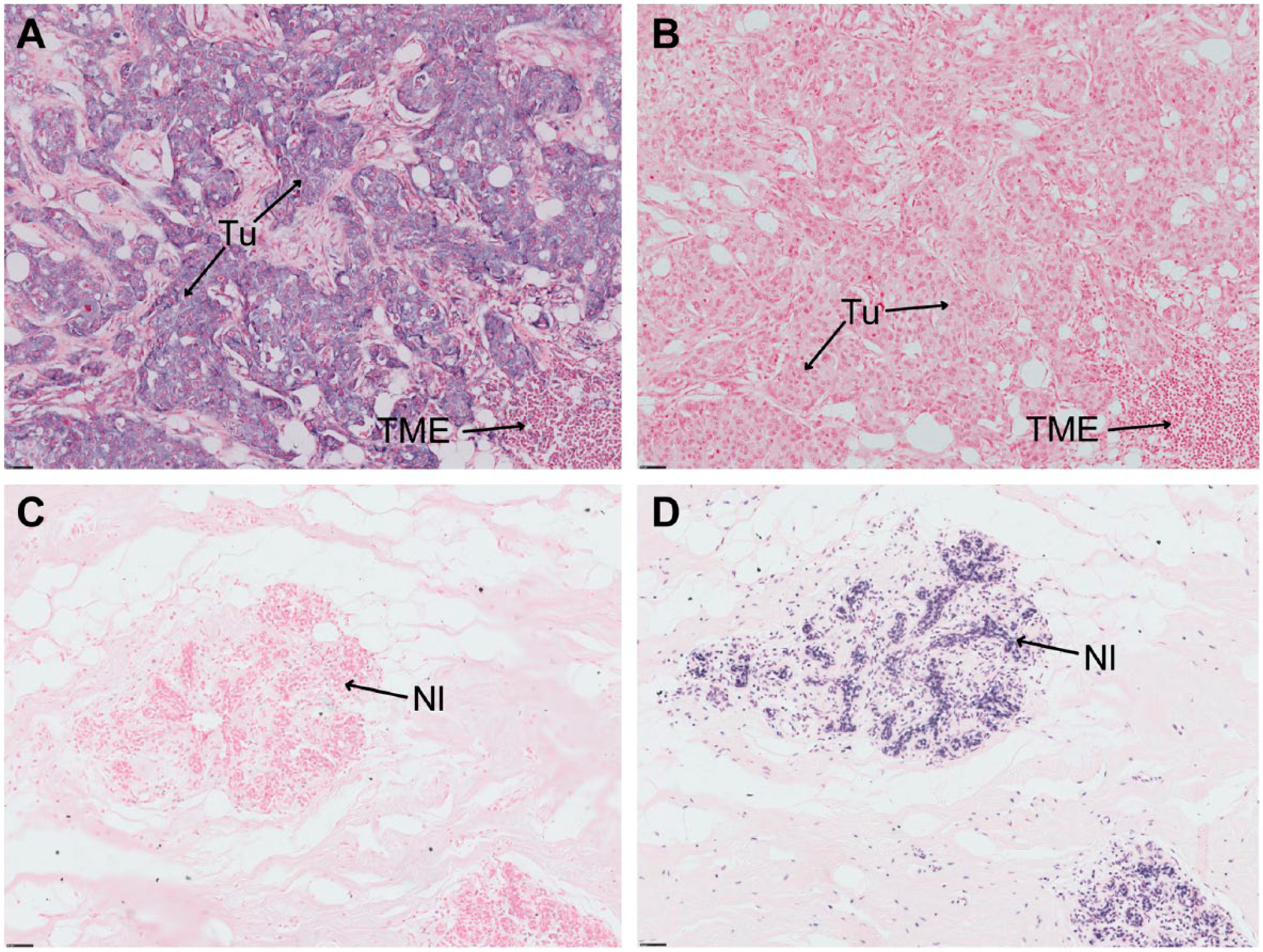
Validation of miR-210 in situ hybridization. A and B: The miR-210 signal is detected in epithelial cancer (A), and no signal is detected with the negative control probe (scramble probe) (B). C and D: Normal glandular cells do not express miR-210 (C) but express the positive control probe U6 in the nucleus (D). (Scale bar: 50 µm.) Abbreviations: TME, tumor microenvironment; Tu, epithelial tumor cells; Nl, normal glandular cells.
We then performed ISH analysis on 26 FFPE TNBC samples. The miR-210 signal was detected in tumor cells in 18/26 samples, and the signal intensity was heterogeneous between samples (Fig. 3). These results are consistent with the data obtained by RT-qPCR. However, a miR-210 signal was also detected in the TME in 13/26 samples (Fig. 3C–F). The proportion of miR-210 positive cells in the TME was very heterogeneous between samples, varying from a few positive cells to the majority of tumor infiltrating cells.
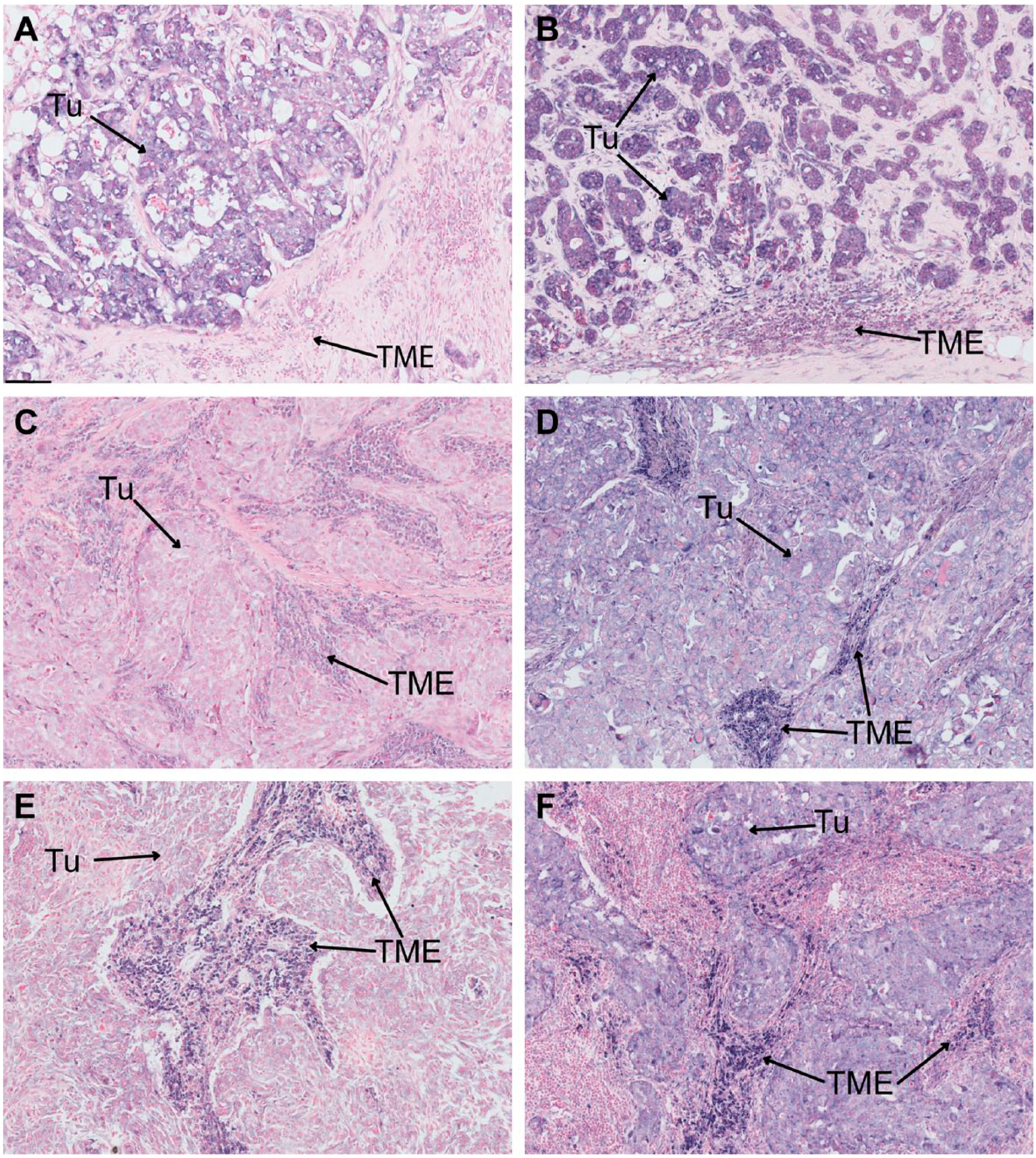
Detection of miRNA-210 expression in triple-negative breast cancer (TNBC) by in situ hybridization (ISH). A–D: miR-210 expression is detected in tumor cells. C–F. Strong miR-210 expression is detected in the tumor microenvironment. (Scale bar: 50 µm.) Abbreviations: Tu, tumor; TME, tumor microenvironment.
As the BRCA1 status was known for 23 of the 26 TNBC, we compared miR-210 expression between mutated and non-mutated samples. An miR-210 signal was detected in the tumor cells in 10 out of 13 BRCA1 mutated TNBC samples (77%). However, staining was also detected in the TME of 7/13 samples (54%). In non-mutated TNBC samples, miR-210 was detected in the tumor cells in 6/10 samples (60%). An expression in the TME was detected in 5/10 samples (50%).
These results indicate that in TNBC, miR-210 is expressed by the tumor cells but also by the TME, and that miR-210 distribution is relatively similar between BRCA1 mutated and non-mutated TNBC.
Identification of miR-210 Expressing Cells
As mentioned above, the expression of miR-210 was detected by ISH in non-tumor cells, in the TME. To identify the nature of these miR-210 expressing cells, we performed ISH analysis for miR-210 and IHC analysis for the pan-leukocyte marker CD45-LCA on adjacent sections for the 13 samples presenting an miR-210 signal in the TME. The leukocyte-common antigen CD45-LCA is expressed at the surface of several subtypes of immune cells, including the CD4+ and CD8+ T lymphocytes, the B lymphocytes, and the myeloid-derived cells, including tumor-associated macrophages. 20 Interestingly, we found that the miR-210 signal in non-tumor cells co-localized with regions positive for CD45-LCA (12/13 samples), suggesting that the miR-210 positive cells in these regions are immune cells (Fig. 4). We also performed ISH for miR-210 and IHC for CD3 (marker of T lymphocytes), CD20 (marker of B lymphocytes), CD68 (marker of macrophages), and CD56 (marker of natural killer cells) on adjacent sections to compare the localization of miR-210 signal with different markers of inflammatory cells. Our analysis on two samples might suggests that the miR-210 positive cells cannot be assigned to a single cellular subtype and may possibly be composed of a mixture of immune infiltrating cells (Supplemental Fig. 1A).
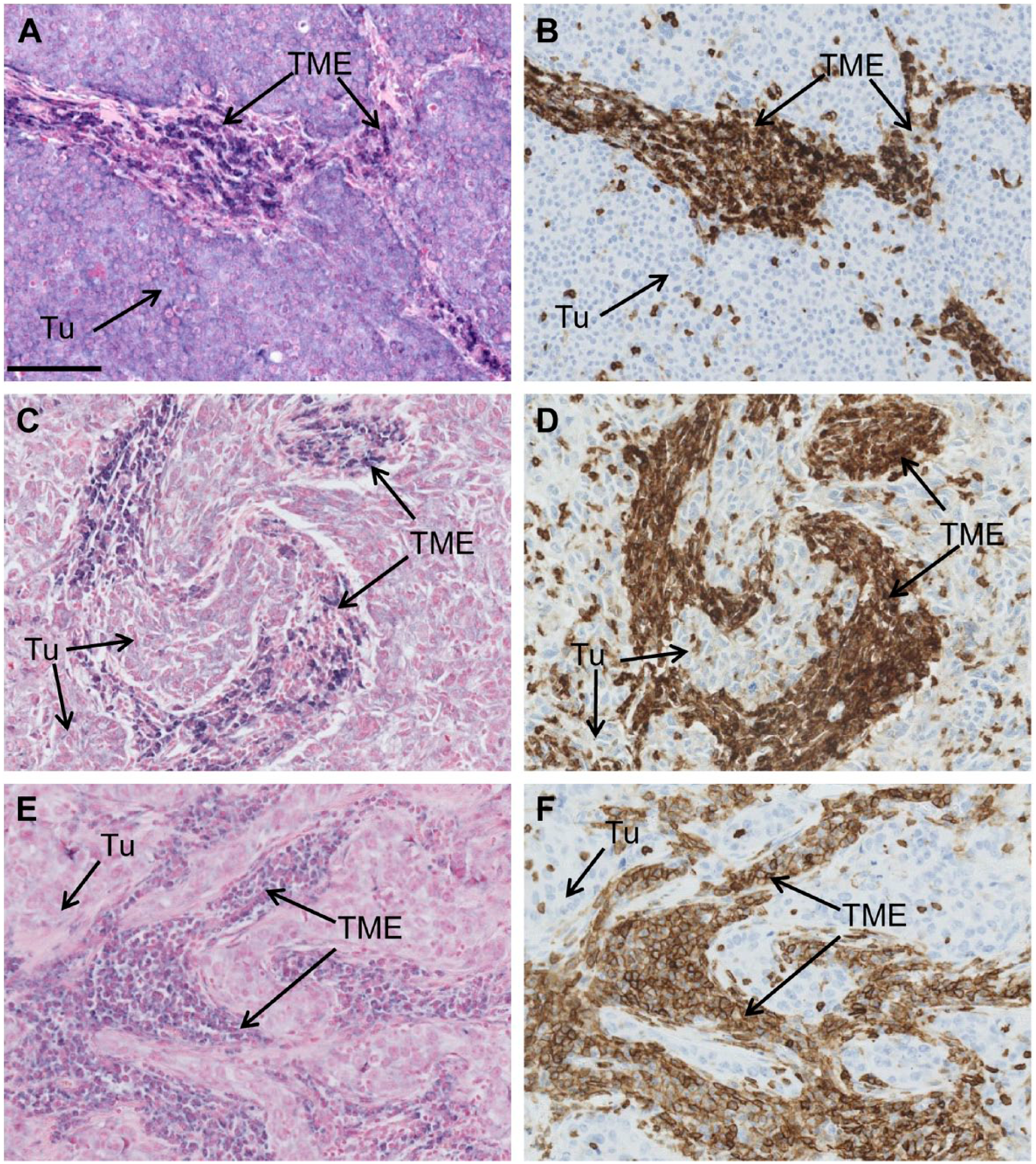
miR-210 positive cells in the tumor microenvironment co-localize with CD45-LCA expressing cells. A-C-E: detection of miR-210 by ISH; B-D-F: detection of CD45-LCA by IHC in adjacent sections of TNBC. (Scale bar: 100 µm.) Abbreviations: CD45-LCA, CD45-leukocyte common antigen; ISH, in situ hybridization; TNBC, triple-negative breast cancer; Tu, tumor; TME, tumor microenvironment.
Co-Localization of miR-210 and Hypoxic Markers
As miR-210 is a marker of hypoxia, we investigated whether the miR-210 expression in the TME could be correlated with HIF1-alpha accumulation or CAIX expression, two proteins expressed in hypoxic region of solid tumors. 21 IHC for CAIX and HIF1-alpha was performed on eight samples with miR-210 expression in the TME. In these samples, the CAIX expression or HIF1-alpha accumulation was only detected in the tumor cells or in the fibroblasts and not in the miR-210 positive immune cells (Fig. 5). These results suggest that miR-210 expression in these immune cells is HIF1-alpha independent.
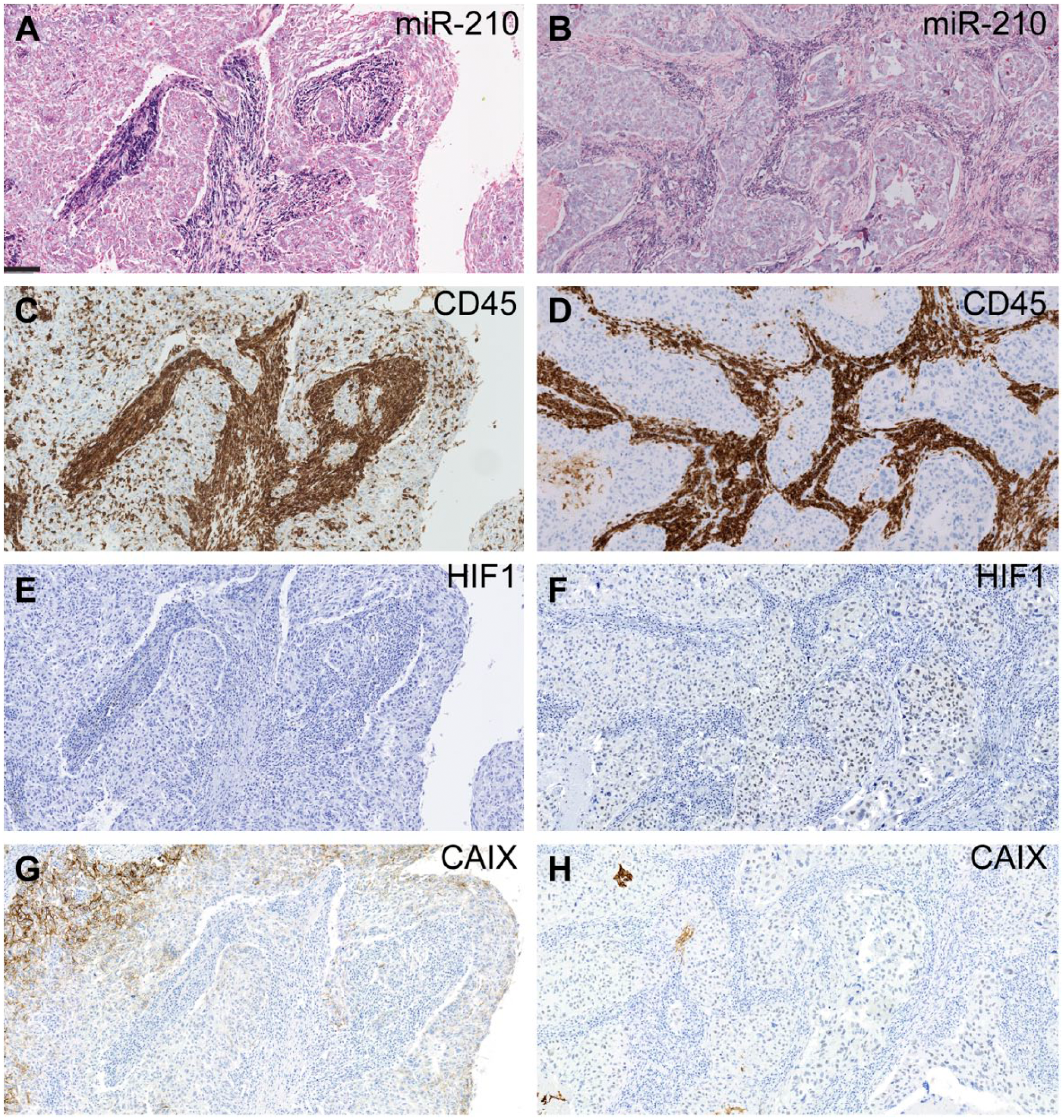
Detection of miR-210, CD45-LCA, HIF1-alpha and CAIX expression in adjacent sections of two representative TNBC samples. On the left (Fig. 5A-C-E-G), miR-210 is expressed in CD45-LCA positive cells, in the absence of HIF1-alpha expression. CAIX is expressed in the tumor cells. On the right (Fig. 5B-D-F-H), HIF1-alpha is detected in the tumor cells but not in the immune cells expressing miR-210. CAIX is not detected in the tumor cells. (Scale bar: 250 µm.) Abbreviation: CD45-LCA, CD45-leukocyte common antigen; HIF1-alpha, hypoxia inducible factor 1 alpha; TNBC, triple-negative breast cancer; CAIX, carbonic anhydrase IX.
Discussion
In this study, we confirm that miR-210 is overexpressed in TNBC compared with normal tissue, and we report for the first time the cellular localization of the miR-210 in TNBC by ISH.
As expected, the miR-210 signal was detected in tumor epithelial cells but was also present in non-epithelial cells in 50% of the 26 analyzed TNBC. The non-tumor cells expressing miR-210 co-localize with regions positive for the pan-leucocyte marker CD45-LCA, suggesting an immunological origin.
There are few data reporting miR-210 expression in the TME. The expression of miR-210 was detected by ISH in the stroma of non-small cell lung cancer, and it was shown that a high level of miR-210 expression in the stromal cells is associated with good prognosis. 22 In prostate cancer, miR-210 overexpression activates the fibroblasts and converts them into cancer-associated fibroblasts. 23
MiR-210 has numerous functions in tumor epithelial cells but also in the TME, which plays an important role in tumor progression. 11 The TME is composed of different cell types such as immune cells (macrophages, dendritic cells, natural killer cells, myeloid-derived suppressor cells, regulatory T cells, etc.), fibroblasts, or endothelial cells. 24 A role for miR-210 is described in several of these cell types, and evidences for a role of miR-210 in the control of immune response to tumor development are also accumulating. MiR-210 is involved in Th17 differentiation. 25 In B cells, miR-210 limit autoantibody production. 26 In regulatory T cells, miR-210 negatively regulates FOXP3. 27 All these reports suggest that miR-210 might modulate the immune response. It is, thus, clearly important to identify the cell type producing the miRNA to avoid misleading interpretations, even if the microenvironment itself might be part of the miRNA signature. An example of such wrong interpretation is illustrated by the miR-143/miR145 cluster that was described as a tumor suppressor, but recent studies rather suggest a critical role in the TME where its expression promotes tumorigenesis. 28
Multiple studies have consistently established that miR-210 is a robust target of hypoxia-inducible factors, and the induction of miR-210 is a consistent characteristic of the hypoxic response in both normal and transformed cells. 11 Interestingly, we found that miR-210 expression in CD45-LCA positive cells does not correlate with HIF1-alpha accumulation. This observation suggests that miR-210 expression is regulated by a mechanism independent of HIF1-alpha in these cells. While most studies reported that miR-210 is regulated in an HIF1-alpha dependent way, other studies reported miR-210 regulation in an HIF1-alpha independent manner. For instance, miR-210 can be regulated by HIF2 dependent 29 and HIF independent mechanisms. 30 Further investigations are needed to determine the mechanism of miR-210 regulation in the CD45-LCA positive immune cells of the TME.
An important consideration in relation to the prognostic signature established by NGS, microarray, or RT-qPCR is that they are derived from tissue samples with various percentages of tumor cells and stromal cells, which do not allow the precise identification of the cellular origin of the microRNA. This is also the case for the breast cancer prognostic signature, where the percentage of tumor cells in the analyzed samples is heterogeneous (when described), ranging from 30% to 80%.18,31–34 Considering that miR-210 has numerous functions in the TME, a high expression of miR-210 in the non-tumor cells might have a prognostic impact, and the significance of miR-210 expression in the infiltrating cells in TNBC clearly warrants further investigations.
Our results also highlight the necessity of using more standardized RNA extraction procedures and complementary approaches such as ISH to take into account the cellular context of miRNA expression in microarray and NGS analysis.
TNBC is a highly heterogeneous disease, and it is important to identify new molecular biomarkers that will help to understand their pathophysiology and to develop targeted therapies. MiRNA represents a new class of such biomarker. MiR-210 had been extensively studied in TNBC using diverse approaches such as NGS, microarray, or RT-qPCR. However, these techniques do not allow the precise identification of the origin of the miRNA signal, and data obtained using NGS or microarray are often contaminated with RNA extracted from non-tumor cells such as fibroblasts or leukocytes. This work highlights the importance of taking into account the stromal bias in gene expression studies. ISH allows the detection of altered miRNA expression at a single-cell resolution. Here, we show that in TNBC, miR-210 is highly expressed in the TME. The significance of this expression warrants further investigation and may allow a better stratification of TNBC, which will help in developing more targeted therapies.
Footnotes
Acknowledgements
We thank Pascale Hilbert and the Anatomical Pathologists of the Institute of Pathology and Genetics (IPG) for their help and discussion. We are also grateful to Sandy Haussy for experimental support.
Competing Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
FA-S, ABA, AM, IB, and PD conceived and designed the experiments. FA-S, ABA, AM, IB, and SS carried out the experiments. IB, AM, and PD analyzed the data. IB, AM, PD, and J-LC wrote the manuscript.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by a grant from the Institute of Pathology and Genetics—scientific research fundings (Gosselies, Belgium).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
