Abstract
SPINT2 is a tumor suppressor gene that inhibits proteases implicated in cancer progression, like HGFA, hepsin and matriptase. Loss of SPINT2 expression in tumors has been associated with gene promoter hypermethylation; however, little is known about the mechanisms of SPINT2 deregulation in prostate cancer (PCa). We aimed to analyze SPINT2 expression levels and understand the possible regulation by SPINT2 promoter hypermethylation in PCa. In a cohort of 57 cases including non-neoplastic and PCa tissues, SPINT2 expression and promoter methylation was analyzed by immunohistochemistry and methylation-specific PCR, respectively. Methylation status of the SPINT2 promoter was also evaluated by bisulfite sequencing and 5-aza-2’-deoxycytidine treatment. Oncomine and TCGA databases were used to perform in silico PCa analysis of SPINT2 mRNA and methylation levels. A reduction in SPINT2 expression levels from non-neoplastic to PCa tissues was observed; however, none of the cases exhibited SPINT2 promoter methylation. Both bisulfite sequencing and 5-aza demonstrated that SPINT2 promoter is not methylated in PCa cells. Bioinformatics approaches did not show downregulation of SPINT2 at the mRNA level and, in corroboration with our results, SPINT2 promoter region is reported to be unmethylated. Our study suggests an involvement of SPINT2 in PCa tumorigenesis, probably in association with a post-translational regulation of SPINT2.
Introduction
Prostate carcinoma (PCa) is the most diagnosed cancer and the second leading cause of cancer-related death among men in the United States (Siegel et al. 2013). Besides the improvements in early detection and treatment options, cancer cell invasion and metastasis are the main cause of death in patients with PCa (Mundy et al. 2002). Therefore, it is important to understand and clarify the mechanisms involved in PCa to overcome this fatal end.
SPINT2 is a serine protease inhibitor Kunitz type 2, also known as hepatocyte growth factor (HGF) activator (HGFA) inhibitor type 2 (HAI-2). It displays inhibitory activity in membrane serine proteases, such as matriptase and hepsin, as well as secreted proteases such as HGFA and plasmin (Delaria et al. 1997; Marlor et al. 1997; Qin et al. 1998; Kirchhofer et al. 2005). Matriptase and hepsin have been implicated in the degradation of matrix components and transmembrane molecules, as well as in the modulation of cell-substratum adhesion (Del Rosso et al. 2002). Moreover, matriptase, hepsin and HGFA are able to cleave pro-HGF into its active form, and thus trigger the HGF/c-MET signaling pathway, which is implicated in tumor growth, angiogenesis, invasion and metastasis (Benvenuti et al. 2007; Owen et al. 2010).
The importance of SPINT2 in cancer malignancy has been highlighted in several cancer types. Low levels of SPINT2 have been described in ovarian carcinoma (Nakamura et al. 2009b), cervical cancer (Nakamura et al. 2009a), endometrial cancer (Nakamura et al. 2011), uterine leiomyosarcoma (Nakamura et al. 2010) and malignant gliomas (Hamasuna et al. 2001). In cervical cancer, SPINT2 downregulation is correlated with a higher tumor stage, the presence of lymph node involvement, and metastasis (Nakamura et al. 2009a). Recently, some reports indicate SPINT2 as a tumor suppressor gene, as hypermethylation of its promoter is a frequent cause of SPINT2 downregulation (Fukai et al. 2003; Schuster et al. 2003; Chiba et al. 2005; Morris et al. 2005; Kongkham et al. 2008; Morris et al. 2008; Tung et al. 2009; Dong et al. 2010). In PCa, a significant decrease of SPINT2 protein has been reported in poorly differentiated tumors as compared with lower Gleason score tumors (Bergum et al. 2010; Tsai et al. 2013). However, the process underlying SPINT2 down-regulation in PCa is still unknown. Herein, we prompted to clarify the profile of SPINT2 expression in a series of PCa cases and examine whether SPINT2 promoter methylation is the mechanism responsible for SPINT2 downregulation. Additionally, using bioinformatics databases such as Oncomine and TCGA, we further extend and support our findings.
Materials & Methods
Tissue Samples and Cell Lines
Fifty-seven formalin-fixed paraffin-embedded (FFPE) prostate samples were retrieved from the Department of Pathology of the Alto Ave-Guimarães Hospital, Guimarães, Portugal. All patients are of Portuguese origin and presented at a median age of 63 years (range, 49–74 years). The present study was previously approved by our local ethics committee.
Four PCa cell lines were used: 22RV1 (primary PCa), LNCaP (lymph node metastasis), PC3 (bone metastasis) and DU145 (brain metastasis). Human melanoma (MW9) and medulloblastoma (UW228) cell lines were used as controls. All cell lines were cultured in RPMI with 10% fetal bovine serum and 1% antimitotic/antibiotic at 37°C in an atmosphere of 5% CO2.
RNA Isolation and SPINT2 Expression Analysis
Total RNA from PCa cell lines was extracted using Trizol reagent (Invitrogen; Carlsbad, CA) and 1 µg of RNA was converted into cDNA using the Phusion RT-PCR kit (Finnzymes; Woburn, MA), according to the manufacturer’s instructions. cDNA was used to determine the levels of SPINT2 expression using primers described by Kongkham and collaborators (Kongkham et al. 2008). Beta-actin was used to normalize expression levels (Pinto et al. 2014). Quantitative real time RT-PCR (RT-qPCR) was performed in a CFX96 detection system (Bio-Rad; Hercules, CA), using SSOfast Evagreen supermix (Bio-Rad) as described by the manufacture. Optimized cycling conditions for qPCR were as followed: enzyme activation for 30 sec at 95°C; 45 cycles of denaturation at 95°C for 5 sec, and annealing/extension for 5 sec at 59°C. Melting curve was assessed at 65°C–95°C (in 0.5°C increments) at each 5 sec/step. The expression levels were normalized to beta-actin by the delta-CT method, and the data are the relative expression. Results are presented as the mean ± SD of three independent experiences in triplicate.
Western Blotting
For each cell line, 20 µg of protein was loaded onto polyacrylamide gels, separated by SDS-PAGE and electrophoretically transferred into nitrocellulose membranes. Blots were blocked with 5% milk for 2 hr at room temperature and protein was detected by overnight incubation (4°C) using rabbit anti-human SPINT2 antibody (1:1000, HPA011101; Sigma-Aldrich, St. Louis, MO). After washing, the membranes were incubated for 1 hr at room temperature with horseradish peroxidase–conjugated secondary antibody (1:5000; Santa Cruz Biotechnology, Dallas, TX). As a loading control, glyceraldehyde 3-phosphate dehydrogenase
Bisulfite Genomic Sequencing
DNA isolated from PCa (22RV1) and medulloblastoma (UW228) cell lines was treated with sodium bisulfite using the EZ DNA Methylation Golf Kit (Zymo Research Corporation; Irvine, CA) according to the manufacturer’s instructions. After treatment, the DNA promoter region of SPINT2 gene was amplified by touchdown PCR using primers already described (Dong et al. 2010) that cover 85 CpG sites of the SPINT2 promoter region. PCR products were gel-purified and Sanger sequenced (STABVIDA services) to determine SPINT2 methylation status.
Methylation-specific PCR (MSP) Analysis
DNA isolated from FFPE PCa tissue, as previously described (Viana-Pereira et al. 2011), or PCa cell lines, as previously described (Moniz et al. 2013), was treated with sodium bisulfite using the EZ DNA Methylation Golf Kit (Zymo Research Corporation) according to the manufacturer’s instructions. Methylation of the SPINT2 promoter was determined by methylation-specific PCR (MSP), performed as previously described (Morris et al. 2005; Kongkham et al. 2008). This region was amplified using two sets of primers that were designed and validated by bisulfite sequencing to amplify either methylated or unmethylated promoters of bisulfite-modified DNA. In a total reaction volume of 20 µl was 1 µl of DNA, 1× incomplete buffer (Invitrogen), 1.5 mM of MgCl2 (Invitrogen), 200 μM of dNTPs (Invitrogen), 0.2 μM of each primer and 1 U of Taq Platinum (Invitrogen). PCR cycling conditions were denaturation for 5 min at 95°C, followed by 38 cycles of 45 sec at 95°C (denaturation), 45 sec at 60°C (annealing) and 45 sec at 72°C (extension). The melanoma cell line (WM9) and blood were used as methylated and unmethylated SPINT2 promoter controls, respectively.
5-Aza-2’-Deoxycytidine Treatment
The four PCa cell lines (22RV1, LNCaP, PC3 and DU145) and the control melanoma cell line (WM9), which is known to bear a hypermethylated SPINT2 promotor, were treated with the demethylation agent 5-aza-2’-deoxycytidine (5-aza) (Sigma-Aldrich). Briefly, 3×105 cells/well were seeded onto 6-well plates and grown in RPMI medium supplemented with 10% FBS and 1% penicillin-streptomycin. Cells were treated with 5 µM of 5-aza for 72 hr, with medium and 5-aza replaced every 24 hr. As a control, cell lines were treated with DMSO. After treatment, SPINT2 methylation status and mRNA SPINT2 expression were evaluated by MSP and RT-qPCR, respectively.
SPINT2 Immunohistochemistry
Tissue microarray (TMA) blocks were constructed to contain three representative areas of each PCa (1-mm diameter core) and adjacent non-neoplastic tissues. Immunohistochemical staining was applied using the anti-human SPINT2 antibody (1:500, HPA011101, Sigma-Aldrich) and the streptavidin-biotin peroxidase complex system (UltraVision Large Volume Detection System Anti-Polyvalent, HRP; LabVision Corporation, Fremont, CA), according to the manufacturer’s instructions.
The level of SPINT2 staining was semi-quantitated by intensity scoring: 0, negative; 1, weak; 2, moderate; 3, strong (Bergum et al. 2010; Martinho et al. 2012; Martinho et al. 2013). Tumor tissues that presented scores between 0 and 1 were classified as negative for SPINT2 staining, those that were scored as 2 demonstrated moderate positive staining, and those scoring 3 as strongly positive. Placental and muscle tissues were used as positive and negative controls, respectively. Additionally, the primary antibody was omitted from the staining as another negative control. Images were taken in Nikon Eclipse 50i Y-THM microscope (Nikon; Tokyo, Japan).
In silico Analysis
SPINT2 mRNA levels were retrieved from Oncomine database (Rhodes et al. 2007; http://www.oncomine.org). Ten prostate samples datasets were assessed: Grasso (28 prostate gland, 59 PCa, 35 metastasis) (Grasso et al. 2012); Tomlins (28 prostate gland, 30 PCa, 19 metastasis) (Tomlins et al. 2005); Taylor (29 prostate gland, 131 PCa, 19 metastasis) (Taylor et al. 2010); Yu (23 prostate gland, 64 PCa, 24 metastasis) (Yu et al. 2004); Lapointe (41 prostate gland, 62 PCa, 8 metastasis) (Lapointe et al. 2004); Singh (50 prostate gland, 52 PCa) (Singh et al. 2009); Holzbeierlein (4 prostate gland, 40 PCa, 9 metastasis) (Holzbeierlein et al. 2004); LaTulippe (3 prostate gland, 23 PCa, 9 metastasis) (LaTulippe et al. 2002); Chandran (10 PCa, 21 metastasis) (Chandran et al. 2007); and Vanaja (8 prostate gland, 27 PCa, 5 metastasis) (Vanaja et al. 2003). SPINT2-negative and SPINT2-positive patients were categorized based on the median-centered intensity values of SPINT2 probes.
SPINT2 promoter methylation was evaluated using seven specific methylation probes (cg10154122, cg27176870, cg18354742, cg15375239, cg19337852, cg16655905 and cg23090870) collected from TCGA (https://www.tcga-data.nci.nih.gov/tcga) database, and comprised 201 patients. For all probes, intensity values greater than 0.5 were considered methylated and intensity values less than or equal to 0.5 were considered non-methylated.
Statistical Analysis
Pearson’s Chi-Square (χ2) test was carried out using SPSS 19.0 (SPSS Inc., Chicago, IL) to evaluate the correlation between SPINT2 expression and clinical characteristics. For in silico analysis, a one-way ANOVA was used to compare SPINT2 expression among normal, PCa, and metastasis samples using GraphPad Prism software version 6 (GraphPad Software, La Jolla, CA). A two-way ANOVA (Bonferroni’s post-hoc test) was used to compare mRNA expression levels before and after 5-aza treatment. The threshold used for statistical significance was p<0.05.
Results
Expression of SPINT2 Protein in Prostate Tissues
The expression of SPINT2 protein was analyzed by immunohistochemistry in a total of 57 patients; yet, due to technical issues, we could only obtain immunostaining results in 45 PCa and 52 adjacent non-neoplastic tissues (Fig. 1). Because only three samples—two non-neoplastic and one PCa—were classified as SPINT2 negative with a score of 0 or 1, we have grouped these samples with the moderate positive cases (score 2) and sub-divided the staining as low/intermediate (score 0–2) and high (score 3) expression. SPINT2 staining was exclusively detected in the prostatic acini, in the cytoplasm, and in the membrane of cells. In most samples, PCa presented weaker and less diffuse staining than normal parenchyma. We observed a significant reduction in the level of SPINT2 expression (high vs low/intermediate staining) from non-neoplastic to tumor tissues (43/52 (83%) low staining to 28/45 (62%) high staining, respectively; p=0.038; Fig. 1A and 1F). No significant association among the SPINT2 protein expression and clinicopathological parameters was observed (Table 1). Analysis of mRNA (Fig. 2A) and protein (Fig. 2B) expression in PCa cell lines showed that the majority of cell lines exhibited SPINT2 expression.
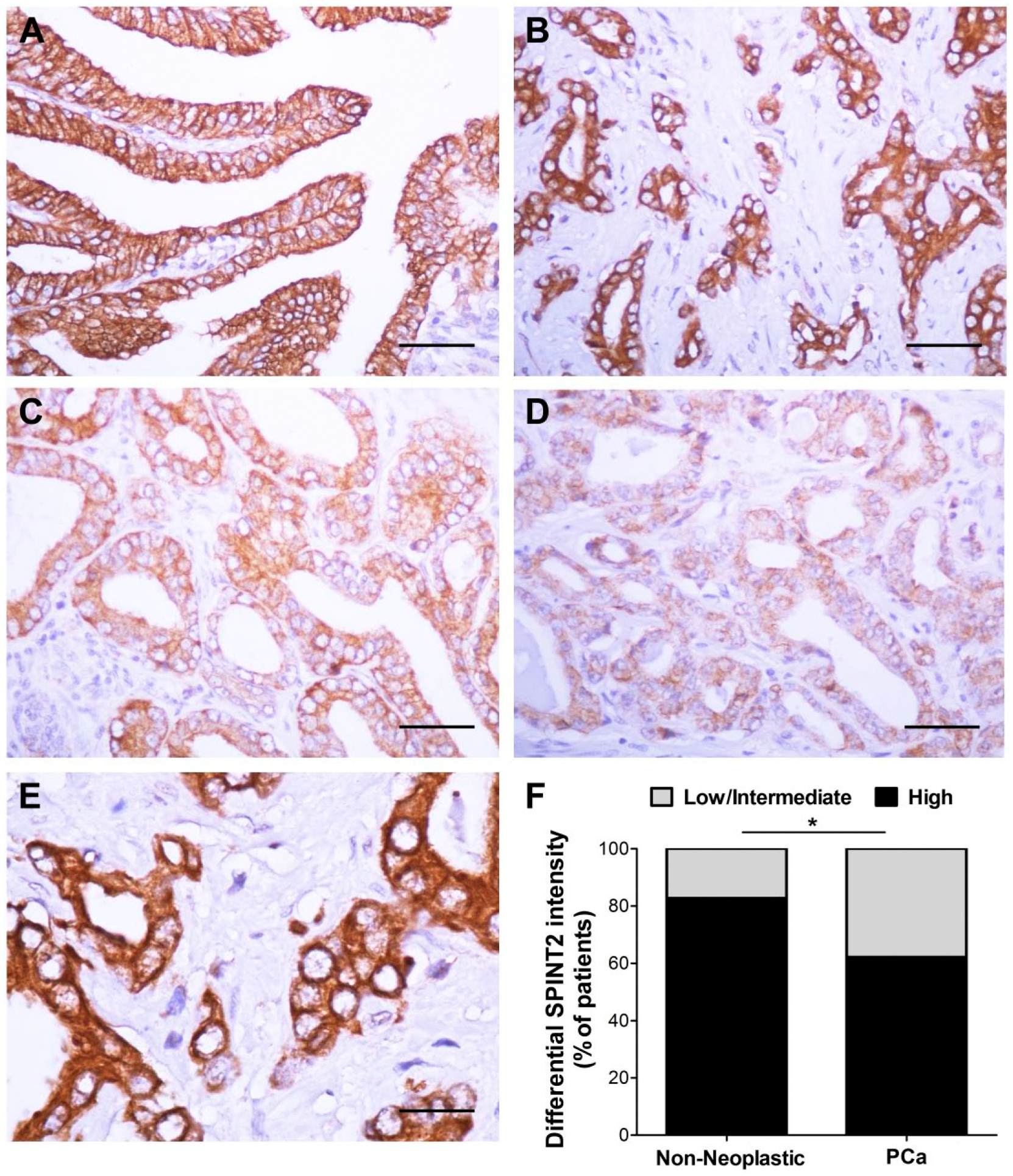
Staining for SPINT2 protein expression in non-neoplastic prostate and prostate carcinoma tissues. (A) Adjacent non-neoplastic prostate (intensity score 3); Prostate carcinoma exhibiting (B) strong staining (intensity score 3), (C) moderate staining (intensity score 2), and (D) weak staining (intensity score 1). (E) High magnification of SPINT2 staining shows SPINT2 localization in the membrane and cytoplasm. (F) Comparison between the percentage of non-neoplastic and neoplastic prostate tissues with high and low/intermediate SPINT2 expression. SPINT2 expression is reduced in prostate carcinoma. *p<0.05; **p<0.01; ***p<0.001. Scale (A–D) 50 µm; (E) 20 µm.
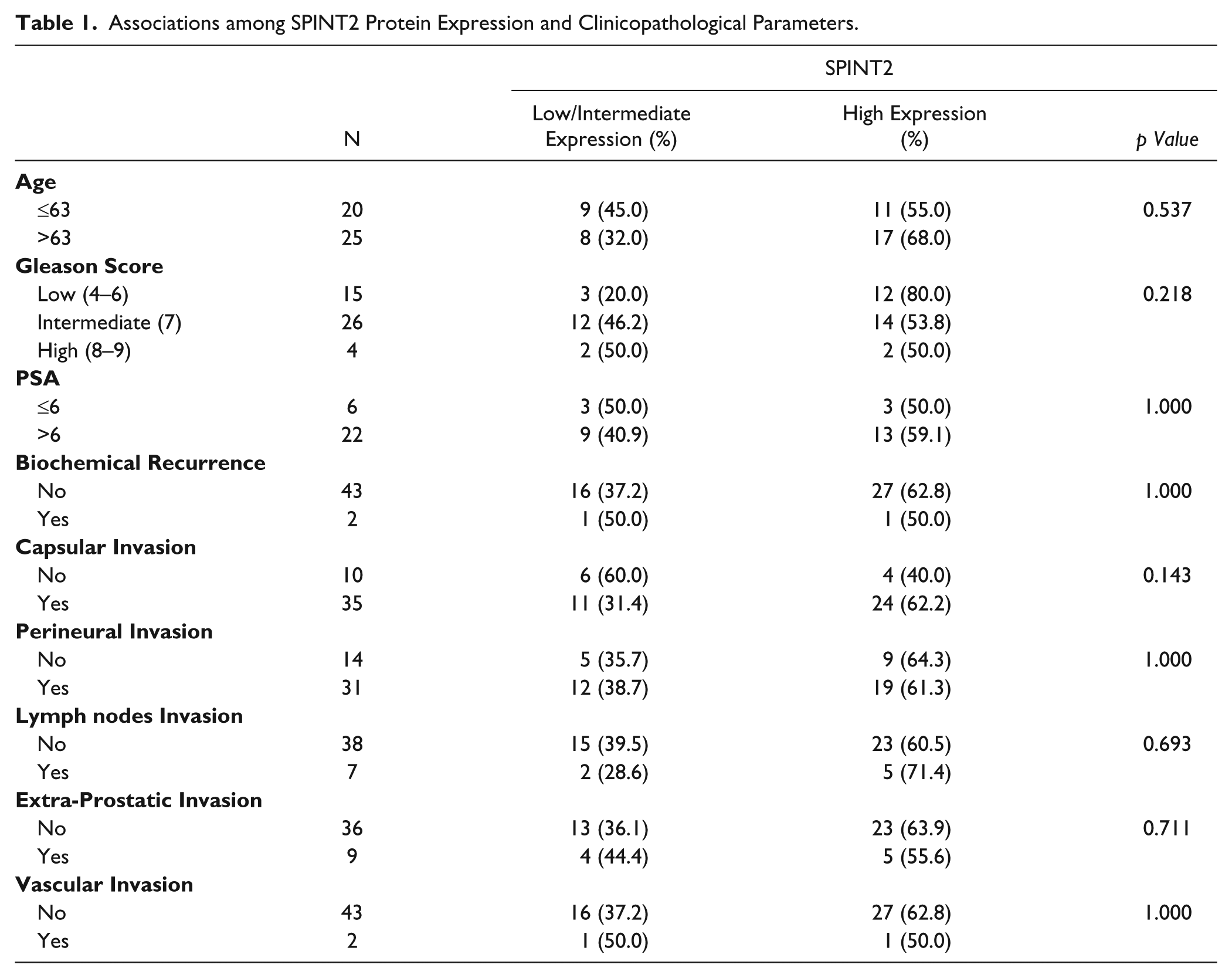
Associations among SPINT2 Protein Expression and Clinicopathological Parameters.
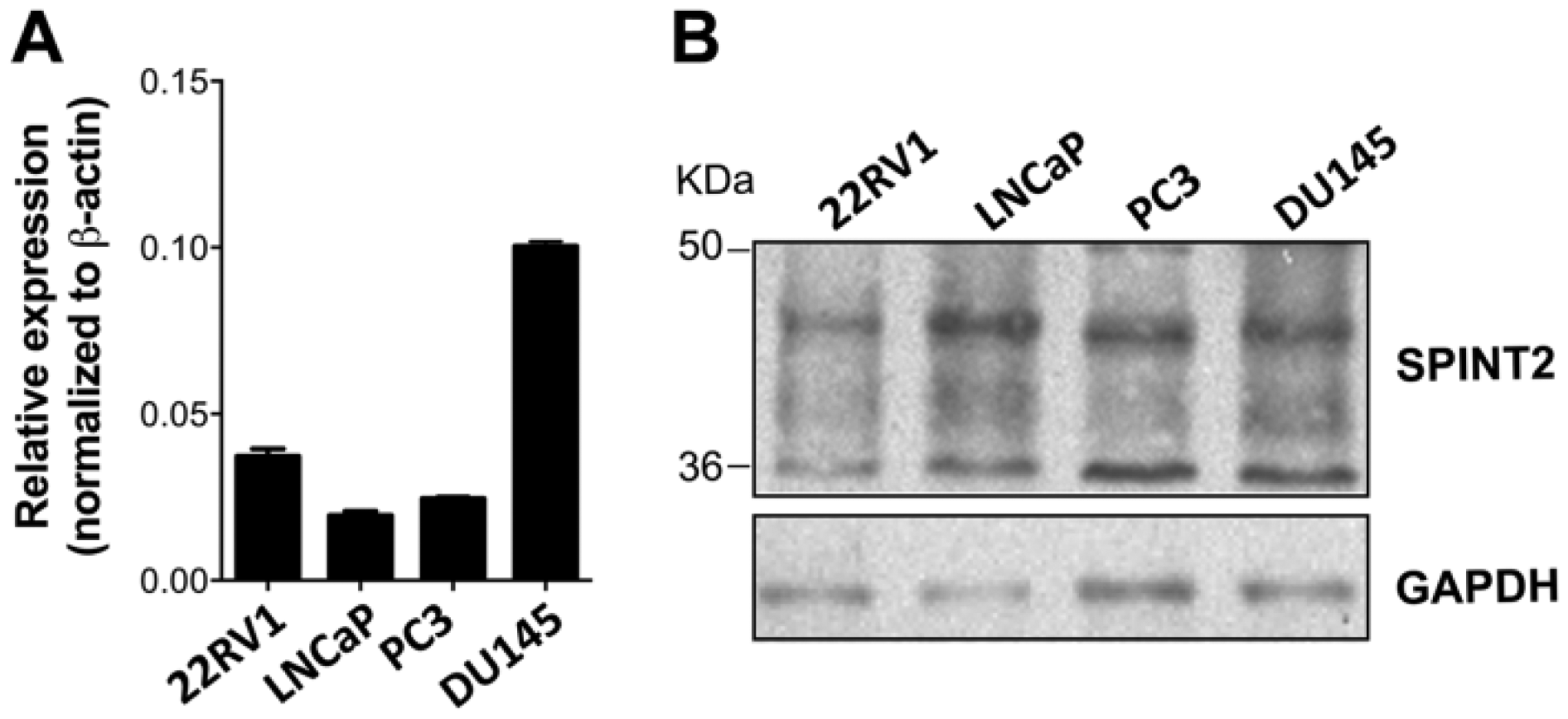
SPINT2 expression characterization in prostate cancer cell lines. (A) Detection of mRNA expression by RT-qPCR. (B) Detection of protein expression by western blot.
SPINT2 Promoter Hypermethylation Analysis in Prostate Carcinoma
We were able to evaluate the methylation status of SPINT2 by MSP in 55 of 57 primary PCa samples using specific primers for CpG island 5’ of the transcriptional start of SPINT2. In this analysis, we found that none of the cases presented SPINT2 promoter methylation (Fig. 3). Similar findings were observed in PCa cell lines (Fig. 3).
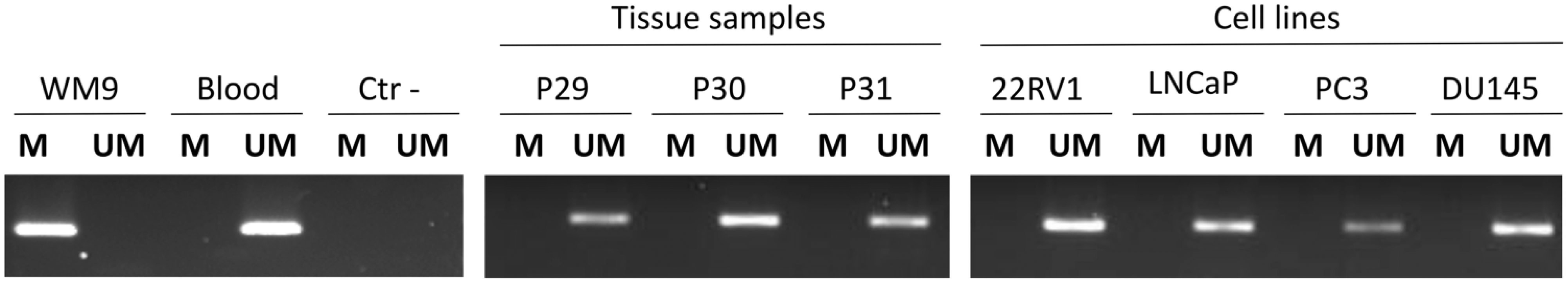
SPINT2 promoter methylation analysis. Representative example of methylation-specific PCR for SPINT2 in prostate carcinoma tissue and cell lines. WM9 cell line and blood were used as SPINT2 -methylated and -unmethylated controls, respectively. M, product from methylated DNA (162 bp) and UM, from unmethylated DNA (168 bp).
In order to clarify whether the methylation status was absent in the SPINT2 promoter, we performed bisulfite sequencing using the 22RV1 cell line, together with the SPINT2 methylated control cell line (UW228). We verified that all of the 46 CpG sites sequenced in the UW228 cell line SPINT2 promoter were methylated (data not shown). In contrast, in the 22RV1 cell line, none of the 60 CpG sites sequenced in SPINT2 promoter region were methylated (data not shown). In addition, to confirm that methylation status does not affect SPINT2 expression, we treated the four PCa cell lines, together with a melanoma cell line known to have promoter methylation (WM9), with a demethylating agent 5-Aza-2’-Deoxycytidine (5 µM). After treatment, we verified that SPINT2 promoter remains unmethylated in the PCa cells (Fig. 4A) and that the inhibition of DNA methyltransferases had no significant effect on the SPINT2 gene expression in the PCa cell lines, in contrast to the control-methylated cell line (Fig. 4B).
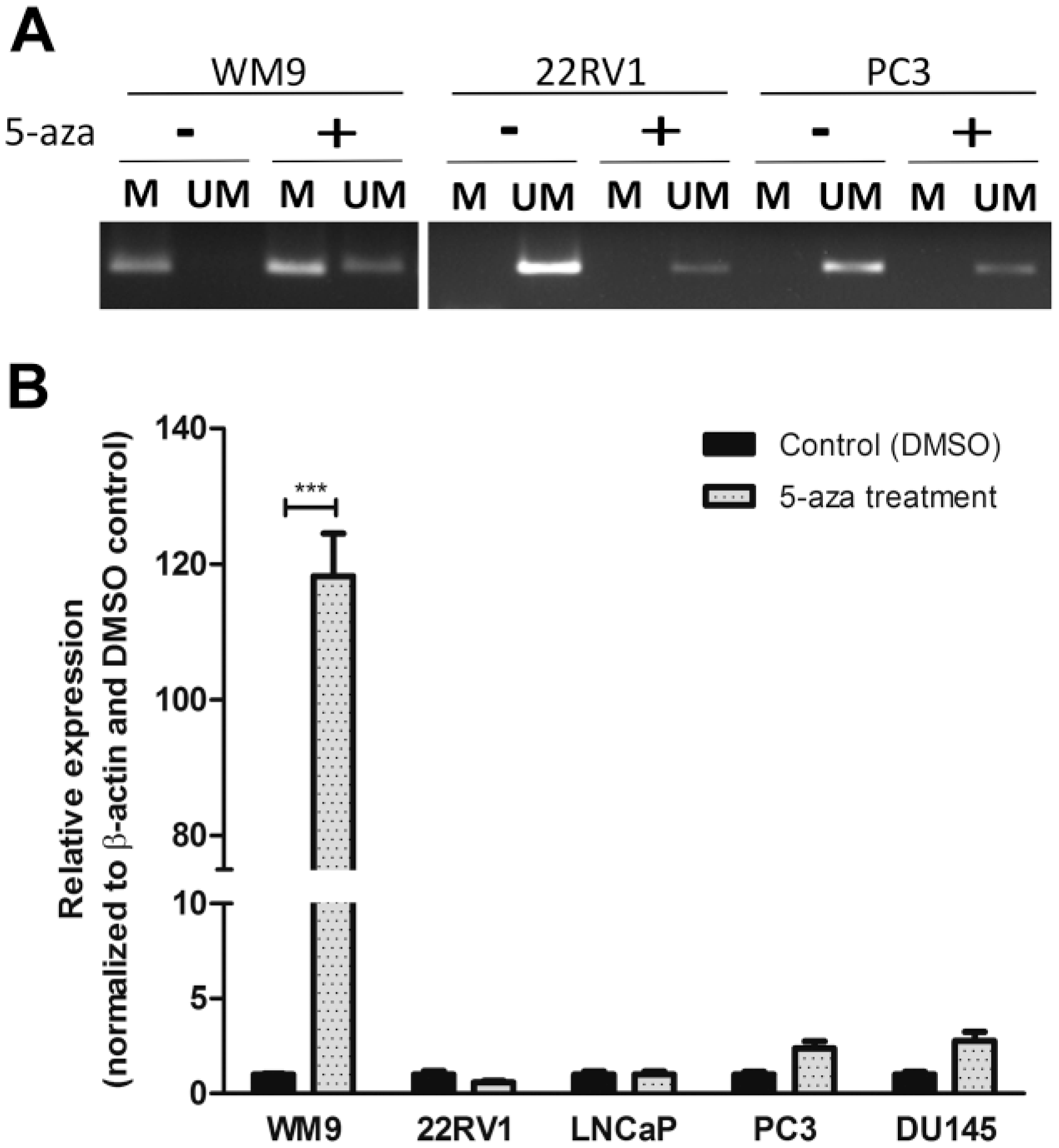
SPINT2 expression analysis after treatment with the demethylation agent 5-aza. 5-aza treatment was performed in PCa cell lines: 22RV1, LNCap, PC3 and DU145. (A) SPINT2 methylation status before and after 5-aza demethylation treatment, as determined using MSP. (B) Detection of the SPINT2 expression levels by qPCR before and after 5-aza treatment. WM9 cell line was used as positive control for the methylated SPINT2 promoter. *p<0.05; **p<0.01; ***p<0.001.
In silico Analysis of SPINT2 in Prostate Tissue
Using the bioinformatics tool Oncomine (Rhodes et al. 2007; www.oncomine.org), we explored the expression of SPINT2 mRNA in 885 cases from ten different datasets. From Grasso (122 samples), Tomlins (101 samples), Taylor (179 samples) and Holzbeierlein (53 samples) datasets, we observed no statistical differences in SPINT2 mRNA expression among the normal, primary PCa and metastatic tissues (Fig. 5). Moreover, Yu (111 samples), Lapointe (111 samples), Singh (102 samples) and LaTulippe (35 samples) datasets demonstrated an increase in SPINT2 mRNA levels from normal prostate gland to PCa and metastatic samples (Fig. 5). Also in Yu and Lapointe datasets, the highest levels of SPINT2 were associated with the presence of metastasis (Fig. 5). In contrast, the Chandran (31 samples) and Vanaja (40 samples) datasets showed that the lowest levels of SPINT2 were associated with the presence of metastasis (Fig. 5).
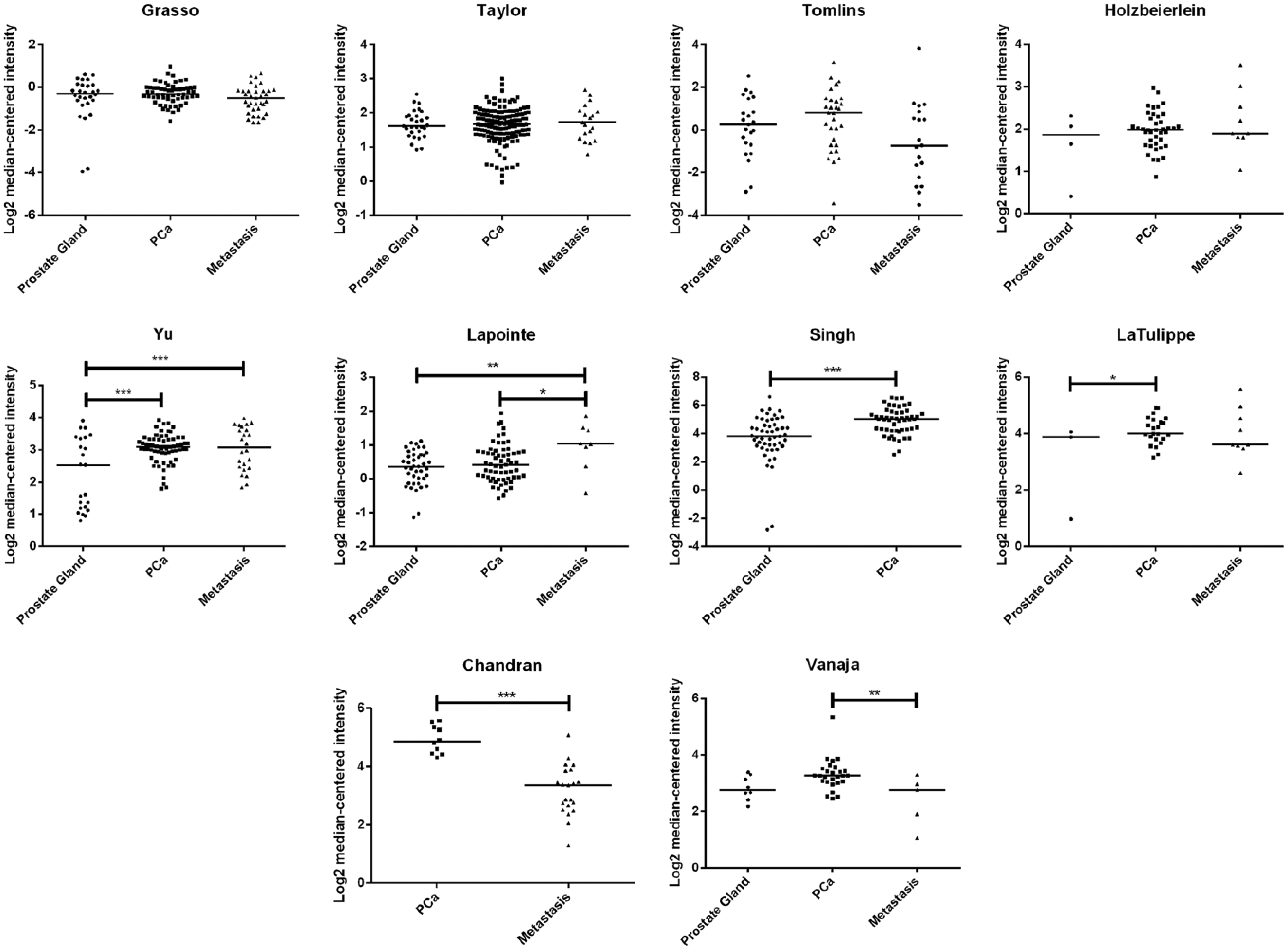
In silico analysis of SPINT2 levels from the Oncomine database. SPINT2 mRNA level distribution among prostate gland, primary carcinoma and metastatic tissues in ten Oncomine datasets (885 samples). *p<0.05; **p<0.01; ***p<0.001.
Additionally, we further characterized the methylation status of SPINT2. For this, we analyzed the TCGA (www.tcga-data.nci.nih.dov) dataset comprising 201 patients diagnosed with PCa. All seven probes that recognize the SPINT2 promoter region (cg10154122, cg27176870, cg18354742, cg15375239, cg19337852, cg16655905 and cg23090870) showed an absence of SPINT2 promoter hypermethylation (Fig. 6).
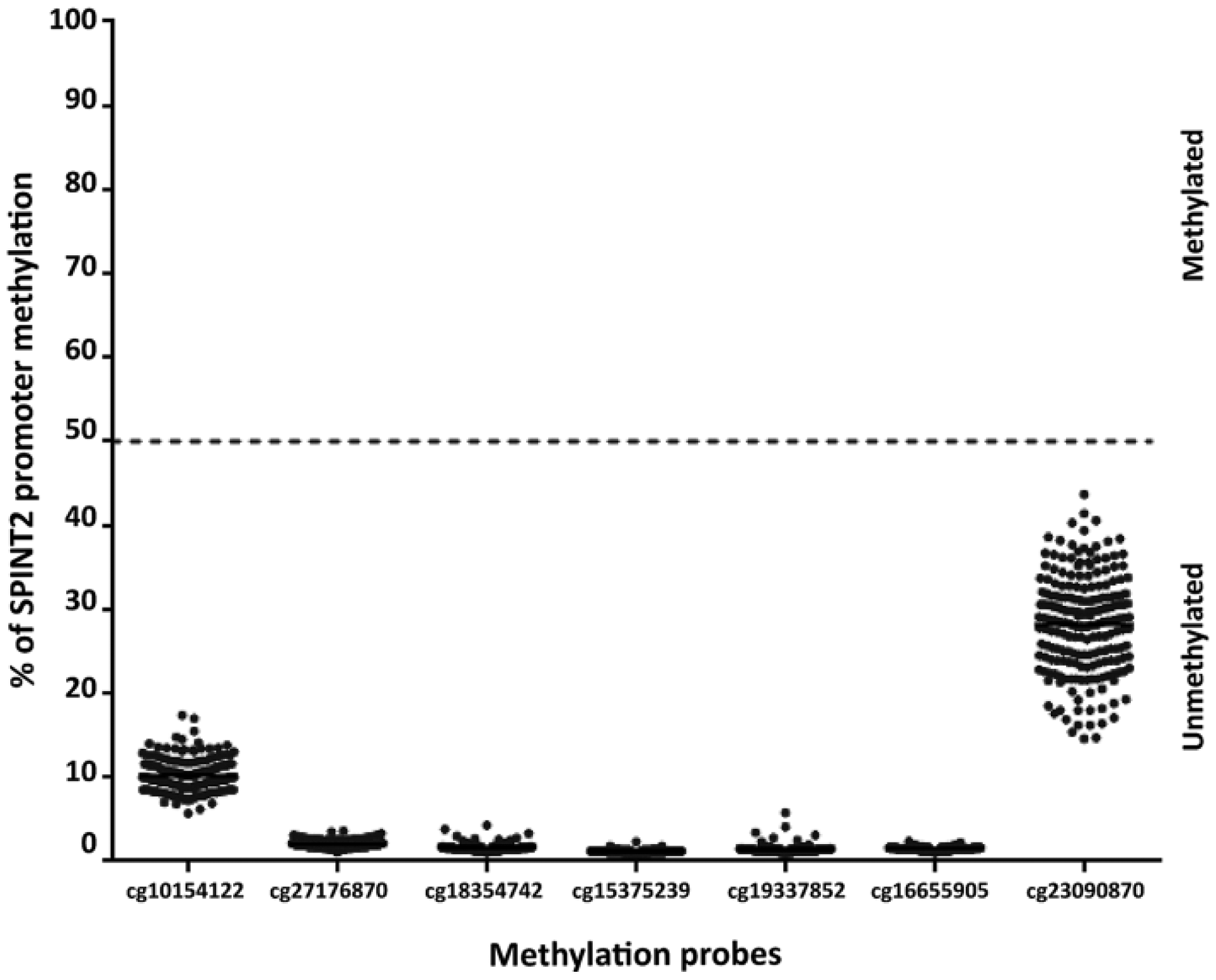
SPINT2 promoter methylation analysis using TCGA database. Methylation of SPINT2 promoter in 201 prostate carcinomas using TCGA database. Seven specific methylation probes (cg10154122, cg27176870, cg18354742, cg15375239, cg19337852, cg16655905 and cg23090870) were used to detect SPINT2 promoter methylation.
Discussion
The ability of serine proteases to degrade the extracellular matrix is essential for PCa progression (Bok et al. 2003; Klezovitch et al. 2004). SPINT2 is a kunitz-type serine protease inhibitor, in which the Kunitz domains are responsible for its inhibitory activity. The SPINT2 protein is thought to be first produced in a membrane-associated form and then released from cells by ectodomain shedding through proteolytic cleavage in a COOH-terminally truncated form (Kawaguchi et al. 1997; Shimomura et al. 1997; Qin et al. 1998). SPINT2 displays an inhibitory activity in secreted proteases (HGFA and plasmin), as well as in membrane serine proteases (hepsin and matriptase) (Delaria et al. 1997; Marlor et al. 1997; Qin et al, 1998; Szabo et al. 2008). Tsai et al. (2013) demonstrated that SPINT2 losses are associated with PCa cell invasion and metastatic capacity. Moreover, the authors showed that SPINT2 regulates matriptase activity, in that its presence is associated with the inhibition of matriptase activation and the consequent reduction in MET signaling, PCa cell migration and invasion (Tsai et al. 2013).
In the current study, we observed a decrease in immunohistochemistry intensity of SPINT2 between non-neoplastic and neoplastic prostate tissues. This result is in accordance with previous studies that showed a significant decrease in SPINT2 protein expression in malignant prostate tissues as compared with normal glands, with the lowest levels of expression found in the most undifferentiated of the prostate tumors (with higher Gleason score) (Bergum et al. 2010; Tsai et al. 2013). The lack of an association between SPINT2 and Gleason score in our cohort might be justified for the limited number of evaluated patients. Our immunohistochemical results were in agreement with Chandran (Chandran et al. 2007) and Vanaja (Vanaja et al. 2003) datasets, which reported that the loss of SPINT2 mRNA levels is associated with metastasis formation. The mRNA expression profiles among the other eight Oncomine datasets are not in agreement.
In several tumor types, the mechanism underlying SPINT2 downregulation is gene promoter hypermethylation (Fukai et al. 2003; Chiba et al. 2005; Morris et al. 2005; Kongkham et al. 2008; Morris et al. 2008; Schuster et al. 2003; Tung et al. 2009; Dong et al. 2010). Therefore, we analyzed, for the first time, the possibility of this epigenetic regulation in PCa. We verified that none of the 55 cases in our cohort or in the 4 PCa cell lines we studied presented SPINT2 promoter methylation. In addition, none of the 201 cases in the TCGA cohort showed SPINT2 promoter methylation. In the cell lines, we validated these findings by distinct methodologies, namely bisulfite sequencing and following treatment with the demethylating agent 5-Aza-2’-Deoxycytidine. These two techniques showed the absence of SPINT2 promoter hypermethylation in prostate cancer. In our PCa cohort, we observed SPINT2 expression in all of the cases (only one PCa sample with weak staining), which is in accordance with the observed unmethylated status of the SPINT2 promoter. However, the mechanism involved in the different expression intensity between normal and neoplastic tissues remains to be determined. Morris and collaborators demonstrated that, in addition to the frequent methylation of SPINT2 promoter in renal cell carcinoma, a nonconservative missense substitution was detected. However, they showed that this mutation did not lead to a reduction in SPINT2 expression but led to reduced tumor suppressor function (Morris et al. 2005). We have confirmed that SPINT2 protein is decreased in PCa but the levels of mRNA were not consistent neither among the various datasets nor compared to protein levels. Therefore, it is plausible to consider that post-translational alterations may occur. SPINT2 is a glycosylated protein, potentially containing 2 N- and 5 O-glycosylation sites (Kawaguchi et al. 1997; Steentoft et al. 2013). Glycosylation alterations are very frequent in cancer, and have been shown to be involved with cancer development and progression (Stowell et al. 2015). A possible deregulation of the mechanism of glycosylation may lead to a SPINT2 misfolding, which can undergo ubiquitination and posterior degradation of the protein.
In conclusion, our results suggest a downregulation of SPINT2 in PCa tumorigenesis, potentially associated with post-translational mechanisms.
Footnotes
Acknowledgements
The authors acknowledge the collaboration of Dr. Pedro Monteiro in the anatomo-pathological evaluation of tumors.
Author Contributions
MSP performed the SPINT2 expression (immunohistochemistry and western blot), methylation and in silico analysis; collected and analyzed the data and wrote the manuscript; GCA evaluated the staining of the SPINT2 immunohistochemistry; FP performed the real-time PCR for SPINT2 expression and the silico analysis; MVP supervised the experiment and edited the manuscript; RMR designed the experiment, supervised its analysis and edited the manuscript.
Competing Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the ICVS internal research funds of participating authors and by FCT project, ref. PTDC/SAU-ONC/115513/2009. F.P. received fellowship from the FCT, ref. SFRH/BD/81369/2011 and M.VP from the ON.2 SR&TD Integrated Program (N-01-01-01-24-01-07), ref. UMINHO/BPD/36/2013.
