Abstract
MLL (mixed-lineage-leukemia) gene rearrangements are typical for acute leukemia and are associated with an aggressive course of disease, with a worse outcome than comparable case, and thus require intensified treatment. Here we describe a 69-year-old female with adult B cell precursor acute lymphoblastic leukemia (BCP-ALL) with hyperleukocytosis and immunophenotype CD10- and CD19+ with cryptic MLL rearrangements. G-banding at the time of diagnosis showed a normal karyotype: 46,XX. Molecular cytogenetics using multitude multicolor banding (mMCB) revealed a complex rearrangement of the two copies of chromosome 11. However, a locus-specific probe additionally identified that the MLL gene at 11q23.3 was disrupted, and that the 5′ region was inserted into the chromosomal sub-band 4q21; thus the aberration involved three chromosomes and five break events. Unfortunately, the patient died six months after the initial diagnosis from serious infections and severe complications. Overall, the present findings confirm that, by far not all MLL aberrations are seen by routine chromosome banding techniques and that fluorescence in situ hybridization (FISH) should be regarded as standard tool to access MLL rearrangements in patients with BCP-ALL.
Keywords
Introduction
B cell acute lymphoblastic leukemia (B-ALL) is a heterogeneous disease accounting for approximately 20% of adult leukemia. B-ALL is also the most common leukemia in pediatrics, representing up to 80% of childhood leukemia, with a peak of prevalence between the ages of 1 and 6 years (Zuckerman and Rowe 2014; Pui et al. 2008).
One of the most common recurrent chromosomal rearrangements in B-ALL (observed in approximately 50% of the rearrangements) is the balanced translocation t(4;11)(q21;q23), which leads to fusion of the MLL (mixed-lineage-leukemia) gene on 11q23 to the AFF1 gene in 4q21 (Woo et al. 2014). MLL encodes for a protein with histone methyltransferase activity, which plays a critical role in the hematopoietic regulation of HOXA as well as embryonic development (Ansari and Mandal 2010). The translocation t(4;11) or MLL/AFF1 gene fusion is almost exclusively seen in infant B-ALL (<1 year of age) and in highest frequency in childhood B-ALL. Up to 93% of affected infants under the age of 90 days harbor MLL rearrangements such as translocations t(4;11), t(11;19), or t(1;11), and most of these children cannot be rescued with the currently available therapies. These MLL rearrangements are also approximately four times more common in children than in adults (Braoudaki and Tzortzatou-Stathopoulou 2012; van der Linden et al. 2009), and the most frequently observed translocation t(4;11), has a dismal prognosis (Pui et al. 2002; Biondi et al. 2000).
Cryptic structural abnormalities often remain undetected by routine chromosomal banding techniques in acute leukemia. However, molecular (cyto)genetics has been proven to be a reliable tool for identification of such cryptic aberrations. Well known examples are the recurrence of cryptic translocation t(12;21)(p13;q22), which is solely associated with childhood B-ALL, and the cryptic translocation t(5;14)(q35;q32), which is known to be present in children and adolescents with T-ALL (Lazic et al. 2010; Su et al. 2006). Overall, chromosomal translocations found in childhood and/or adult B-ALL may result in the production of chimeric fusion proteins with leukemogenic potential.
Here, we report the case of a patient with adult BCP-ALL with a novel cryptic submicroscopic balanced translocation and an additional cryptic insertion of 5’MLL region into the AFF1 locus at 4q21, with an unfavorable prognosis.
Materials & Methods
Clinical Description
A 69-year-old female presented in 2008 with hyperleukocytosis (white blood cell (WBC) count of 259.7×109/l; hemoglobin of 14.2 mmol/l and platelets of 103×109/l). The bone marrow (BM) aspiration showed hypercellularity, with 98% blasts. Immunophenotyping identified a variety of B-cell-specific antigens, with 96% of cells positive for CD15, CD19, CD22, CD34, CD45 and HLA-DR and all cells negative for CD10, CD13, CD20, and CD117. These findings were consistent with a diagnosis of BCP-ALL. It is noteworthy that the immunophenotypes CD10- and CD19+ as seen here are associated with MLL rearrangements in BCP-ALL. The patient was treated by induction therapy: Epi (4-epi-doxorubicin)/ VCR (vincristine)/ PEGAsp (polyethyleneglycole asparaginase)/ PDN (prednisone), two courses of consolidation and maintenance treatment (Mercaptopurin, Metotrexat). Unfortunately, she died six months after the initial diagnosis from serious infections and severe complications.
Diagnosis
Banding cytogenetic anaylsis was performed using an unstimulated bone marrow aspiration obtained at diagnosis and according to standard procedures (Claussen et al. 2002). A total of 20 metaphases were available for cytogenetic evaluation and analyzed on a level of 300 bands per haploid karyotype (Shaffer et al. 2013). Standard G-banding revealed a normal female karyotype as 46,XX and FISH test for a cryptic translocation t(9;22)(q34;q11.2) was negative.
Retrospective Analyses
Molecular Cytogenetics
FISH was performed according to standard procedures and/or to manufacturer’s instructions. The probes and probe sets were made in-house. FISH-banding probe-sets were created using genome-wide multitude multicolor banding (mMCB) and chromosome specific array-proven multicolor-banding (aMCB) (Weise et al. 2003, 2008; Liehr et al. 2002). BAC (bacterial artificial chromosome) clones of interest were identified through the Human Genome Browser Database of the Genome Bioinformatics Group at the University of California at Santa Cruz (http://genome.ucsc.edu/) and Ensembl Genome Data Resources of the Sanger Institute Genome Database (http://www.ensembl.org/). DNA probes (Table 1) obtained from the Resources Center (Oakland, USA) were labeled by PCR with SpectrumGreen, SpectrumOrange or TexasRed-dUTP and applied in two- or three-color FISH-approaches.
Results of Locus-Specific Probes Used for Breakpoint Characterization.
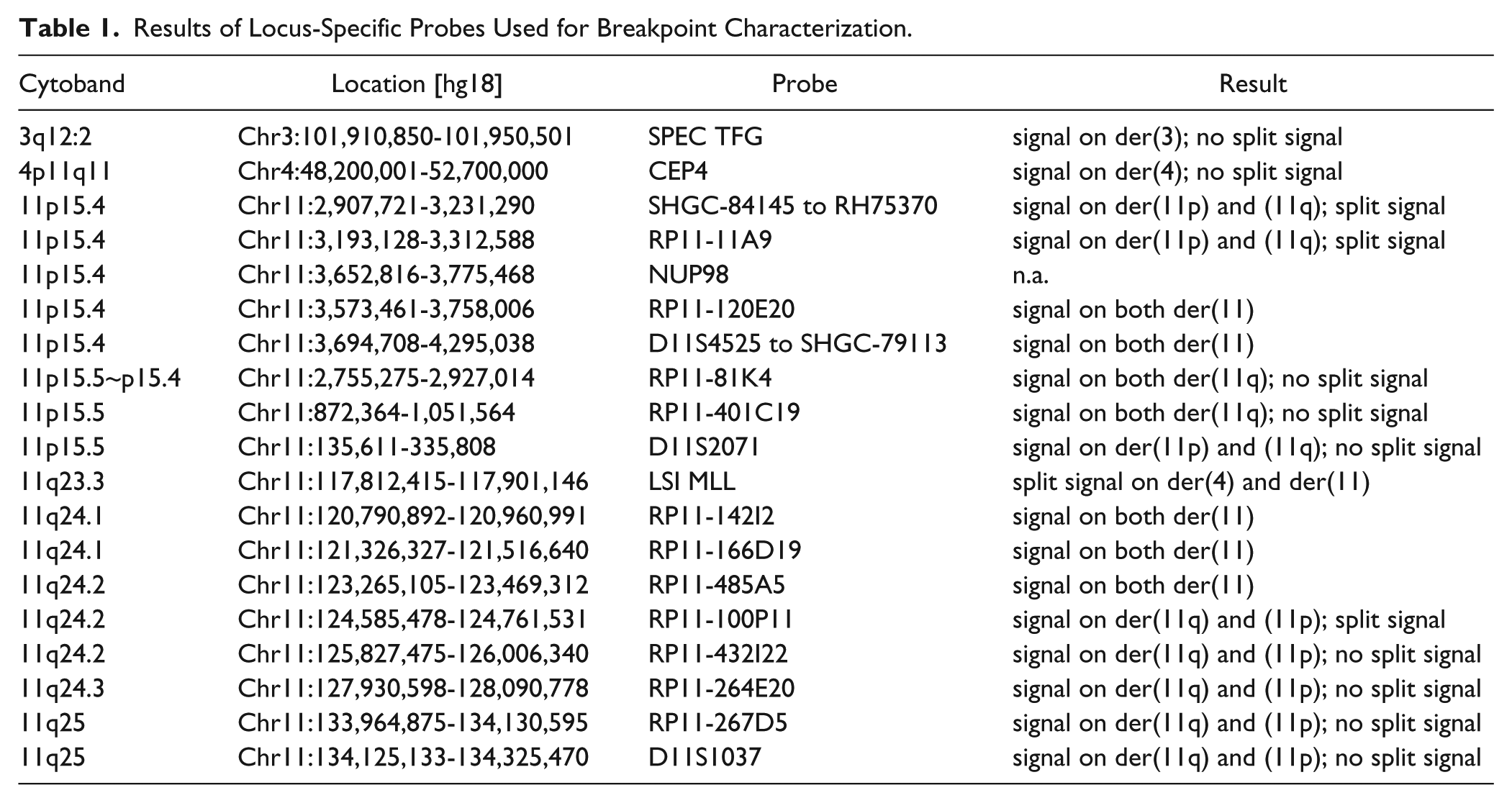
Additionally, the following commercially available probes were used: LSI MLL (11q23 Break probe, Abbott Molecular/Vysis, Mannheim, Germany), POSEIDON NUP98 (11p15 Break probe, Kreatech Diagnostics, Amsterdam, The Netherlands), SPEC TFG Break probe (TFG in 3q12.2, Zytovision, Bremerhaven, Germany), Centromere 4 (CEP4: 4p11-q11 Alpha Satellite DNA, Abbott Molecular/Vysis), and subtelomeric probes for 11p, and 11q (11p in D11S2071; 11q in D11S1037, Abbott Molecular/Vysis).
A total of 10–15 metaphase spreads were analyzed, using a fluorescence microscope (AxioImager.Z1 mot; Zeiss, Oberkochen, Germany) equipped with appropriate filter sets to discriminate between a maximum of five fluorochromes and the counterstain DAPI (Diaminophenylindol). Image capturing and processing were carried out using an ISIS imaging system (MetaSystems; Altlussheim, Germany).
DNA Isolation
Genomic DNA was extracted from cells fixed in acetic acid:methanol (1:3) by Puregene DNA Purification Kit (Gentra Systems; Minneapolis, MN). DNA concentration was determined using a Nanodrop spectrophotometer (NanoDrop Technologies, Inc., Thermo Scientific; Wilmington, DE). The quality of DNA was checked using agarose gel electrophoresis. DNA samples extracted from fixed cells of two healthy males and two healthy females by the same method were used as reference samples.
Multiplex Ligation-dependent Probe Amplification (MLPA)
The P377-A1 Hematologic malignancies probemix and SALSA reagents were used for this study (MRC-Holland; Amsterdam, The Netherlands). Amplified probes and Genescan 500 ROX standard were separated by capillary electrophoresis using a 4-capillary ABI-PRISM 3130XL Genetic Analyzer (Applied Biosystems; Foster City, CA). Sizing of peaks and quantification of peak areas and heights were performed using GeneMarker v1.9 software (Applied Biosystems). A minimum of four healthy control samples were included in each run.
Array-Comparative Genomic Hybridization (aCGH)
aCGH was performed using the Agilent SurePrint G3 Human Genome microarray 180 K (Agilent Technologies, Santa Clara, CA), an oligonucleotide microarray containing approximately 180,000 probes 60-mer with a 17 kb average probe spacing. Genomic DNA from the patient was co-hybridized with a male control DNA (Agilent Technologies). Labeling was performed using the Agilent Genomic DNA enzymatic labeling kit according to the manufacturer’s instructions. After hybridization, the aCGH slide was scanned on an Agilent scanner, processed with Feature Extraction software (v10.7) and results were analyzed using Cytogenomics (v2.9.1.3) using ADM2 as aberration algorithm.
Results
At diagnosis, banding cytogenetics at low resolution did not show any chromosomal aberrations. However, after subjecting the cytogenetic preparations in retrospective to FISH-banding probe-sets, mMCB identified a complex rearrangement for chromosome 11 involving reciprocal translocation and inversion (data not shown). The breakpoints were determined in more detail by further FISH experiments, such as aMCB, using a chromosome 11 specific probe set (Fig. 1) and by locus-specific FISH probes at 11p15.4 and 11q24.2 as shown in Table 1.
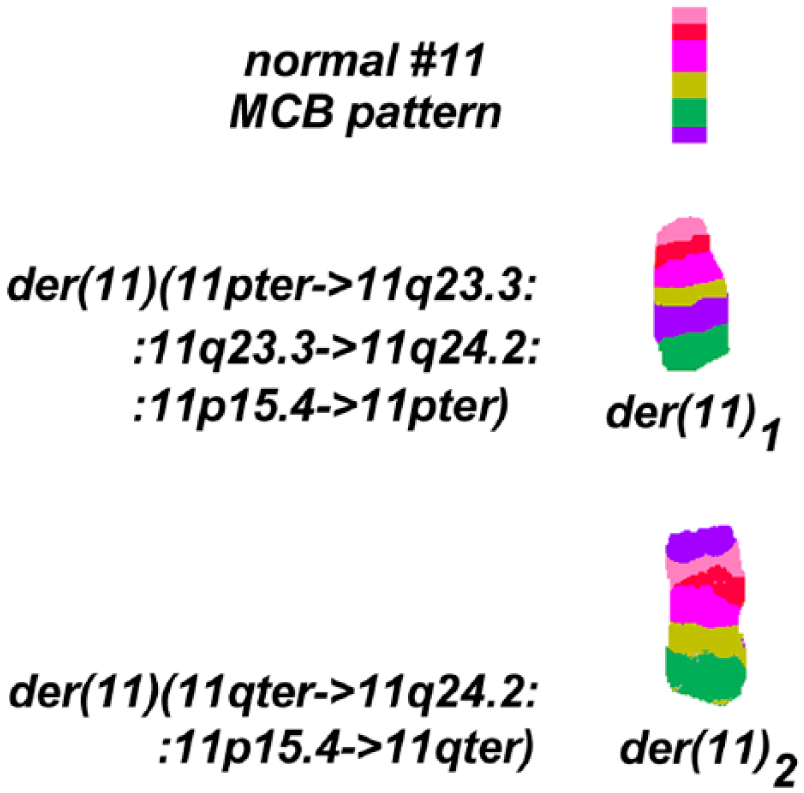
Result of the aMCB probesets for chromosome 11. Characterization of the complex rearrangements occurring in the derivative chromosomes. A normal chromosome 11 pattern (topmost) is provided as a comparison to the two derivative patterns of chromosome 11.
Additionally, dual-color FISH using a commercially available Break Apart Rearrangement probe specific for the MLL locus (LSI MLL) revealed an insertion of the 5′MLL gene into chromosome 4q21. According to the manufacturers of LSI MLL, a 350-kb portion (5’ region) centromeric of the MLL gene breakpoint cluster region was labeled in SpectrumGreen and includes exons 1–6, whereas the ~190-kb portion of the 3’ MLL region is labeled by SpectrumOrange; the latter remained on one of the two derivative chromosomes 11, while the green-labeled part of LSI MLL went to the der(4) (Fig. 2). This cryptic insertion was observed as signal splitting of the probe LSI MLL in 6/6 metaphases and 158/200 interphase-nuclei. Thus, the 5′ MLL region was inserted most likely into the AFF1 gene in chromosome 4q21.
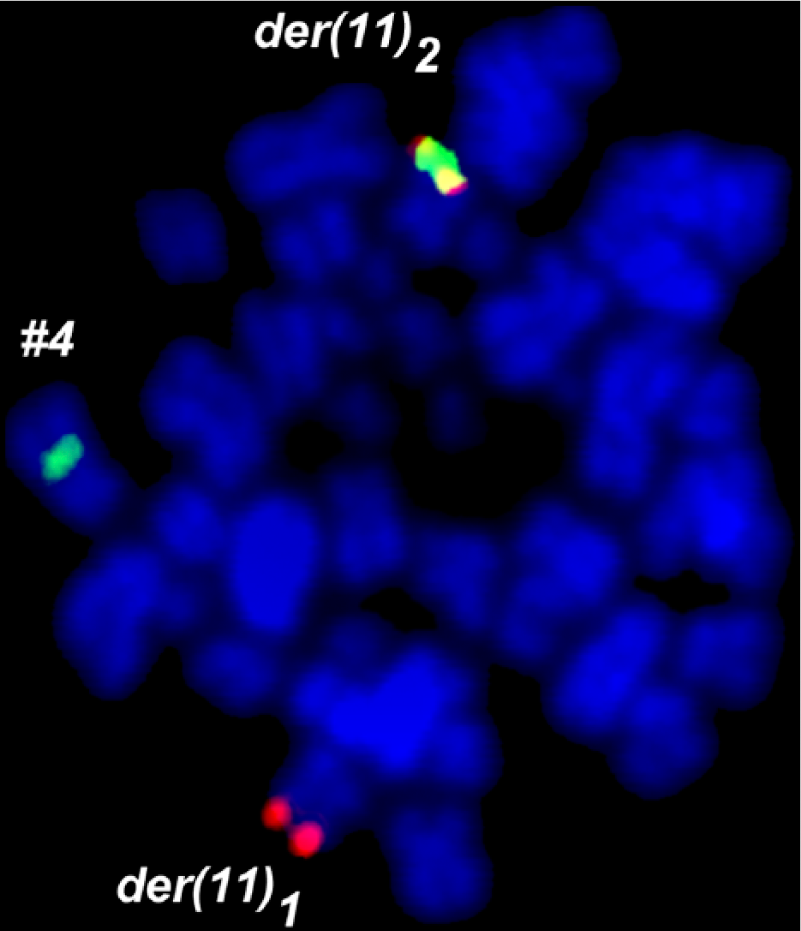
LSI MLL Break Apart probe showed one yellow fusion signal, and split of green signal and orange signal. Surprisingly the 5’MLL probe signal was inserted in a derivative chromosome 4.
The karyotype can be described as follows:
46,XX,der(4)(4pter->4q21.3::11q23.3->11q23.3::4q21.3->4qter), der(11)(11pter->11q23.3::11q23.3->11q24.2::11p15.4->11pter), der(11)(11qter->11q24.2::11p15.4->11qter).
In summary, the present case presents genetic changes involving three chromosomes and five break events.
The breakpoints in 11q24.2 and 11p15.4 were further delineated by locus-specific probes, as summarized in Table 1. The positions are given according to NCBI36/hg18, as a number of the used BAC-probes could not be found in later genomic browser versions.
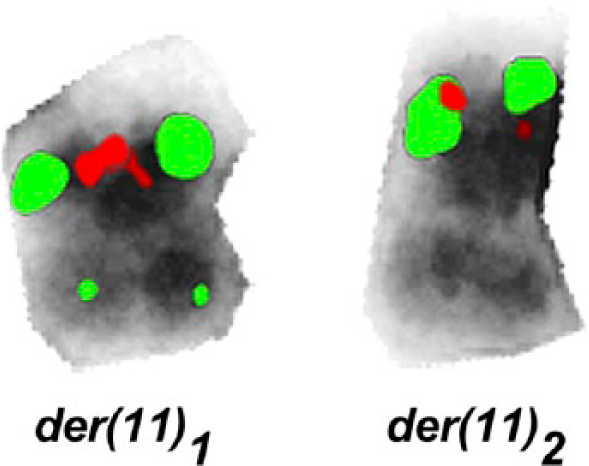
For 11q24.2, the break was narrowed down to lying between the positions 124,585,487 and 124,761,531; one OMIM gene is located there: PKNOX2 (PBX/KNOTTED 1 HOMEOBOX 2). The breakpoint in 11p15.4 was found to be spanned by a probe from locus SHGC-84145 to locus RH75370 that was part of a dual color/Break Apart probe from Kreatech (The Netherlands) flanking the NUP98 gene (Fig. 3). Additionally, BAC RP11-11A9 showed a split signal and the position of the break event can be given between 3,193,128 and 3,231,290; two OMIM genes are located there: MRGPRE and MRGPRG.
MLPA analysis showed no copy number variants; however, the array-CGH revealed an amplification of 83.4 Kb in the region of 3q12.2, which involves two genes, GPR128 and TFG; the latter result was confirmed using locus-specific FISH probes, which showed intrachromosomal amplification in 12% of the interphase nuclei (data not shown).
Discussion
Structural chromosomal abnormalities can be readily detected by metaphase analysis or FISH in B-ALL. The most common balanced or unbalanced translocations have been correlated with variable prognostic significance. Here, included aberrations such as translocations t(4;11) (MLL/AFF1), t(12;21) (ETV6/RUNX1), t(1;19) (E2A/PBX1), and t(9;22) (BCR/ABL) (Zhou et al. 2012; Pui et al. 2008). These alterations can be found in different incidences in childhood and adult B-ALL (Lazic et al. 2010).
In the present B-ALL case, a normal karyotype was initially reported, since the here-described translocation and insertion events were submicroscopic and only identifiable by a combination of different molecular (cyto)genetic approaches. The main problems hampering banding cytogenetics are the well-known difficulties in obtaining evaluable metaphases with well-spread chromosomes instead of clumsy ones or those that appear fuzzy with indistinct margins (Othman et al. 2014; De Braekeleer et al. 2011).
The patient whose case is presented here had high counts of WBC and blast cells, with a pre-B phenotype (CD19+, CD10-)-hallmarks of patients carrying a translocation t(4;11). Unfortunately, these hints were not further followed initially.
The MLL gene plays an important role in normal hematopoietic growth and differentiation. Abnormalities to this region can occur very early in hematopoietic stem cell development (Ansari and Mandal 2010; Ferrando et al. 2003). The translocation t(4;11)(q21;q23) is solely observed in B-ALL patients and presents in ~50% of MLL rearrangements as well as in the ins(4;11)(q21.3;q23.3) insertion as a typical variant of this translocation. In addition, an absence or low expression of CD10- in BCP-ALL and a very high WBC count are particularly common with the translocation t(4;11)(q21;q23) (Woo et al. 2014; De Braekeleer et al. 2011; Burmeister et al. 2009). MLL is well known to be rearranged in myeloid and lymphoid leukemia and can be classified into two groups. The first group includes MLL rearrangements, such as translocations or insertions, some of which are cryptic. These rearrangements result in the generation of in-frame fusion transcripts with various partner genes, with more than 120 loci already identified. The second group comprises amplification of 11q23, leading to the presence of multiple copies of the MLL gene located either intrachromosomally as a homogeneously staining region (hsr), or extrachromosomally in double minutes (dmin) (Meyer et al. 2013; De Braekeleer et al. 2011). The prognosis of MLL rearrangements in infants (<1 year of age) is extremely poor due to a high risk of treatment failure. Young children (1 to <10 years) have a better response to therapy than infants. Finally, for adults, event-free survival (EFS) is seen in 80% of cases. In general, the outcomes for adolescents and adults have improved significantly over time (van der Linden et al. 2009; Bassan 2005; de Bont et al. 2004; Pui et al. 2002; Morel et al. 2003). The present case, which involves 3 chromosomes and 5 break events in connection with an MLL gene rearrangement, is more complex than other comparable cases, but still belongs to the aforementioned first group.
Interestingly, it is considered that the fusion product of MLL-AFF1 is transcribed from the der(4) and not from the der(11), which supports the idea that the MLL-AFF1 is a protein with oncogenic potential. A review of the literature revealed that 10 cases with an insertion of chromosome 11 material in chromosome 4 have been identified in six children (all females) and four adult (3 elderly females and one male) B-ALL patients (Mitelman et al. 2014). Still, no other comparable cases have shown an additional reciprocal translocation between the two homologous chromosomes 11 and amplification in 3q12.2.
The chromosomal breakpoint 11p15 is recurrently involved in translocations in acute leukemia. The gene NUP98 can fuse with DOX10 in 11q22 or with MLL in 11q23 in acute myeloid leukemia (AML) (Kaltenbach et al. 2010; Romana et al. 2006). In the present case, the breakpoint at 11p15.4 involved two other genes MRGPRE and MRGPRG, which are related to the MAS1 oncogene and mainly expressed in sensory neurons. The proteins derived from the MRG gene contain transmembrane, extracellular, and cytoplasmic domains that regulate nociceptor function (Dong et al. 2001). In the second breakpoint observed here, 11q24.2, there is only one OMIM gene located: PKNOX2. PKNOX2 belongs to a homeodomain protein superfamily comprising a large number of sequence-specific transcription factors that share a highly conserved DNA-binding domain; they play fundamental roles in cell proliferation, differentiation, and death (Imoto et al. 2001). Thus, it can be speculated that MRGPRE and/or MRGPRG fused with PKNOX2 may lead to gene expression with oncogenic potential.
In the present case, it remains rather unclear which of the rearrangements—MLL with MRGPRE and/or MRGPRG, fusion of MLL with AFF1 or 3q12.2 amplification—were causative in the adverse outcome. In terms of the latter alteration, the TFG gene located at 3q12.2 is known to play a role in the NF-κB pathway and, thus, multiple copies of the gene may have contributed to oncogenic potential of the tumor cells. Indeed, translocations involving this gene have been observed in hematological malignancies (Chase et al. 2010).
Overall, this case shows that it is necessary to screen for further unbalanced submicroscopic abnormalities by molecular approaches such as MLPA and aCGH in acute leukemia. The present report highlights that MLL gene rearrangements should be considered and tested by molecular approaches in case of a normal cytogenetic result. This holds especially true for such patients with a BCP-ALL who are diagnosed as a result of high WBC counts and CD10-negative staining. However, if, in such cases, MLL rearrangements are detected, further cryptic aberrations with potential influence on the disease may be present. Overall, a normal routine chromosome banding karyotype in acute leukemia needs to be considered as a stimulus and reason for more detailed molecular (cyto)genetic analyses.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported in parts by DAAD.
