Abstract
The ability of chondroitin/dermatan sulfate (CS/DS) to convey biological information is enriched by the presence of iduronic acid. DS-epimerases 1 and 2 (DS-epi1 and 2), in conjunction with DS-4-O-sulfotransferase 1, are the enzymes responsible for iduronic acid biosynthesis and will be the major focus of this review. CS/DS proteoglycans (CS/DS-PGs) are ubiquitously found in connective tissues, basement membranes, and cell surfaces or are stored intracellularly. Such wide distribution reflects the variety of biological roles in which they are involved, from extracellular matrix organization to regulation of processes such as proliferation, migration, adhesion, and differentiation. They play roles in inflammation, angiogenesis, coagulation, immunity, and wound healing. Such versatility is achieved thanks to their variable composition, both in terms of protein core and the fine structure of the CS/DS chains. Excellent reviews have been published on the collective and individual functions of each CS/DS-PG. This short review presents the biosynthesis and functions of iduronic acid-containing structures, also as revealed by the analysis of the DS-epi1- and 2-deficient mouse models.
Keywords
Structure and Biosynthesis of Chondroitin/Dermatan Sulfate
Chondroitin (CS) is a long, linear acidic polysaccharide consisting of the repeating disaccharide unit -4GlcAβ1–3GalNAcβ1- (Silbert and Sugumaran 2002). During assembly, dermatan sulfate (DS) can be generated via epimerization of glucuronic acid (GlcA) into iduronic acid (IdoA) by two DS-epimerases (DS-epi1 and DS-epi2). Subsequently, O-sulfation in various positions is carried out by specific sulfotransferases: Four add a sulfate to the 4-OH position of GalNAc (C4ST1–3 and D4ST1), three are specific for sulfation on the 6-OH position of GalNAc (C6ST1/2 and GalNAc4–6ST, the latter adding a 6-O sulfate to an already 4-O–sulfated GalNAc), and a single CS2ST/UST adds a sulfate to the 2-OH position of both GlcA and IdoA. The reactions described above do not go to completion (i.e., only a fraction of available substrate is converted by a specific enzyme). Highly variable structures are therefore present in CS/DS chains (Fig. 1). In this review, we will indicate the CS/DS structures with the nomenclature O, C, A, B, D, E and iO, iC, iA, iB, iD, iE for the iduronic-containing counterpart, respectively. However, the disaccharide structures depicted in Fig. 1 are just the building blocks of longer protein binding domains, which are usually five to eight monosaccharides long. The exact sequencing of these protein binding domains is still a major challenge and has only been successfully completed in some instances (Li F et al. 2010; Purushothaman et al. 2012). Noteworthy, the disulfated structures B/iB, D/iD, and E/iE, which usually constitute no more than 10% of the chains, often stand out as being part of protein binding domains in CS/DS. Presumably, their double sulfation favors protein interaction. Moreover, a single CS/DS may contain multiple protein binding domains and potentially have multiple functions. In addition to protein binding, CS/DS chains have self-adhesion properties (Fransson et al. 1982).
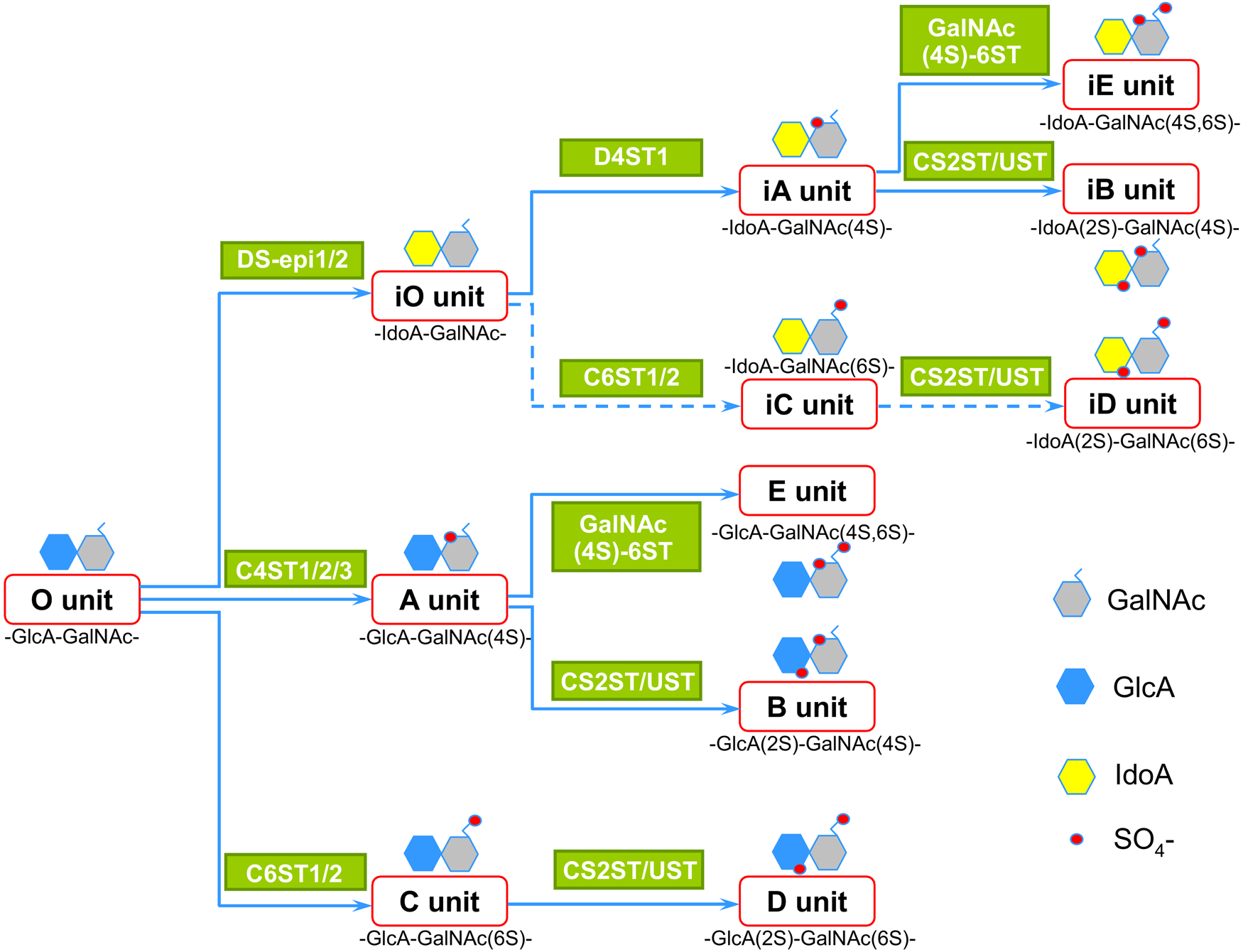
Structures and biosynthetic pathways of chondroitin/dermatan sulfate (CS/DS) disaccharide units. Studies on the disaccharide composition of CS/DS chains and enzyme specificities are the basis of this scheme. In iduronic acid (IdoA)-containing units, epimerization always occurs before sulfation. In disulfated structures, 2-O-sulfation follows both 4-O and 6-O sulfation, whereas 4-O sulfation precedes 6-O sulfation when a 4S, 6-di-O–sulfated unit is generated. Dotted lines indicate putative pathways because iC and iD unit formation has not been clarified yet.
The conversion of GlcA into IdoA is variable and can range from one IdoA residue per chain to almost 100% IdoA. The term CS/DS indicates the glucuronic/iduronic acid hybrid nature of the galactosaminoglycan chain. IdoA can be found in blocks (stretch of ≥6 IdoA residues), in alternating IdoA/GlcA structures, or as isolated IdoA interspersed in unmodified GlcA residues (Fig. 2) (Malmstrom et al. 1975; Maccarana et al. 2009). High content of DS epimerases, especially of DS-epi1 in vivo, and a concomitant high content of the DS-specific 4-O-sulfotransferase D4ST1 are required for formation of IdoA blocks (Maccarana et al. 2009; Pacheco, Maccarana, and Malmstrom 2009; Pacheco, Malmstrom, and Maccarana 2009). Indeed, DS-epi1 and D4ST1 are co-localized in the Golgi apparatus, as seen by confocal staining (unpublished observation). The distribution of IdoA governs some of the subsequent O-sulfation reactions. For instance, the IdoA blocks are never found containing 6-O-sulfated GalNAc and are instead 4-O sulfated and subsequently good substrates for the 2-O sulfation reaction, giving rise to consecutive iB structures. Downregulation of DS-epi1, DS-epi2, or D4ST1 all resulted in a reduced amount of iduronic acid blocks and iB residues. In addition, downregulation of D4ST1 using siRNA decreased the E/iE structures. The amount and distribution of IdoA within a single chain are cell/tissue specific (Cheng et al. 1994). Furthermore, a cell can produce different CS/DS chains depending on the core protein. For example, human skin fibroblasts produce decorin/biglycan with 60% IdoA, mostly present in blocks, and versican with 7% IdoA, mostly as isolated residues (Pacheco, Malmstrom, and Maccarana et al. 2009). Both the amount and distribution of IdoA are subjected to physiological regulation—for example, transforming growth factor (TGF)-β1 considerably decreases IdoA blocks in decorin and biglycan produced by fibroblasts (Tiedemann et al. 2005).
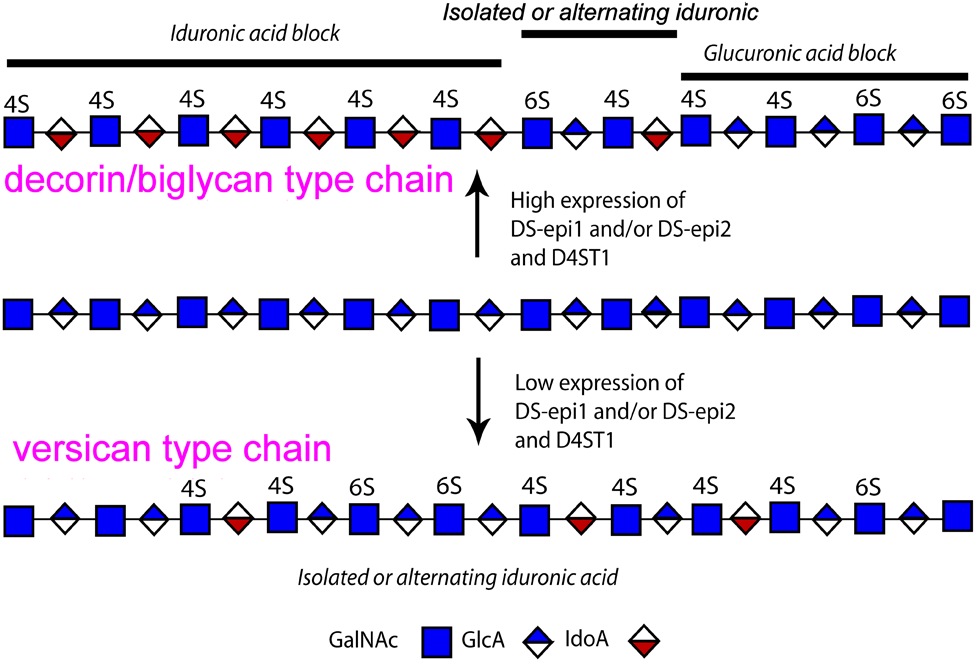
Hybrid structure of chondroitin/dermatan sulfate (CS/DS) and distribution of iduronic acid (IdoA). In vivo, IdoA is commonly found in clusters (IdoA blocks) or as isolated or alternating glucuronic acid (GlcA)/IdoA residues. High expression of DS-epimerases, in close collaboration with the dermatan-specific 4-O-sulfotransferase 1 (D4ST1), is necessary for IdoA block formation.
DS-Epimerase 1 and 2 Structure and Catalytic Mechanism
Only three groups of enzymes catalyze the stereochemical inversion of the C5 carboxyl group of a hexuronic acid at the polymer level (i.e., the conversion takes place after the monosaccharide has been incorporated in the polysaccharide chain). In vertebrates, two DS epimerases and a single HS epimerase convert GlcA into IdoA, whereas in algae and some bacteria, alginate epimerases convert mannuronic into guluronic acid (Valla et al. 2001). No primary sequence or three-dimensional (3D) similarities have been detected between the DS-epimerases and the HS epimerase, which therefore seem to be the result of convergent evolution. DS-epi1 is coded by the DSE gene on chromosome 6, whereas DS-epi2 is coded by the DSEL(-like) gene on chromosome 18 (Maccarana et al. 2006). Interestingly, in DSE, the protein coding sequence is divided into five exons, whereas in DSEL, a single exon contains all the protein coding sequence. The two enzymes show apparent domain similarities (Fig. 3). Both share an N-terminus epimerase domain (51% amino acid identity, identical secondary and 3D predicted structure). DS-epi1 has a C-terminal domain (amino acids 691–958), terminating with two membrane-spanning domains, whose function is currently unknown. No homolog of known structure has been found in 3D databases for this particular domain. Likewise, DS-epi2 has a central domain (amino acids 720–823, including the two transmembrane regions) with unknown function. These domains in DS-epi1 and 2 do not share sequence similarities.
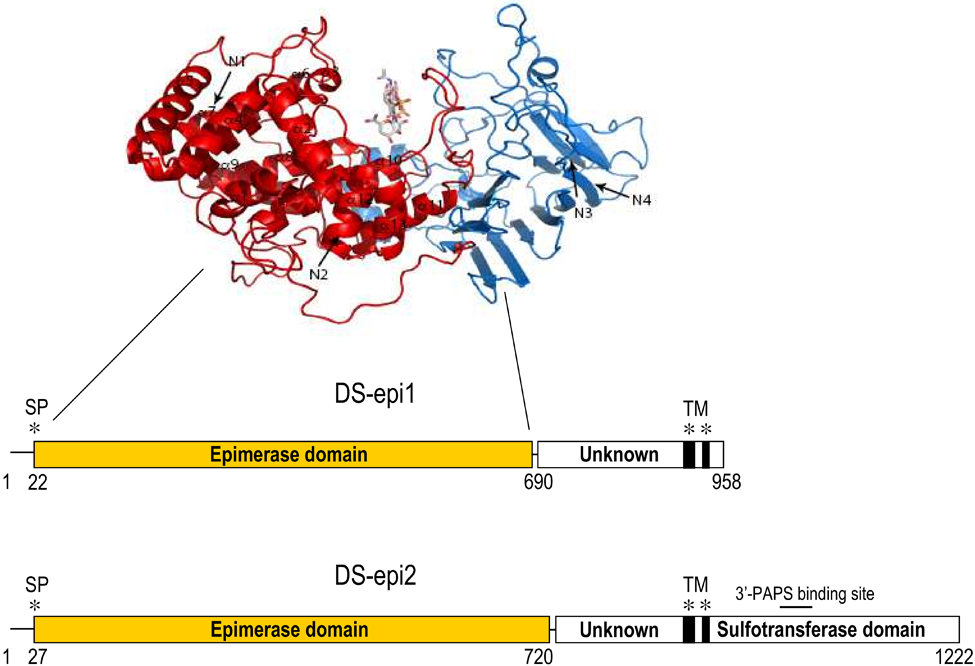
Three-dimensional structure of the DS-epi1 epimerase domain (above) and domain structure of DS-epimerases (below). A tetrasaccharide substrate is positioned in the groove formed by the two subdomains. The four N-glycosylation sites are indicated with arrows and labeled N1 to N4. The locations of signal peptides (SPs) and predicted transmembrane regions (TMs) are shown with asterisks.
Interestingly, to get enzymatic activity, the epimerase domain alone is not sufficient. In fact, recombinant 1–755-DS-epi1 is inactive, whereas 1–775-DS-epi1 is active (E. Tykesson, personal communication 2012). This is mirrored by the fact that 1–734-DS-epi2 is inactive, whereas the full size 1–1222-DS-epi2 is active (Pacheco, Malmstrom, and Maccarana 2009). In addition, DS-epi2 has a C-terminal sulfotransferase domain homologous to several CS/DS O-sulfotransferases. Recombinant DS-epi2 indeed has detectable sulfotransferase activity (unpublished results), but the position of the added sulfate has yet to be defined. The epimerase domains of both enzymes carry four conserved N-glycans, which are required for activity (Pacheco et al. 2009). Based on structural predictions, the epimerase domain of DS-epi1 can be further subdivided into an N-terminal domain composed of alpha-helices (amino acids 1–390; SCOP superfamily a.102.2) and a C-terminally located domain predominantly composed of beta-sheets (amino acids 391–690; SCOP superfamily b.30.5). Together, the two domains form a groove where the substrate is positioned during catalysis. Several HA-, HS-, CS-, and alginate-lyases show structural similarities to these two DS-epi1 domains. Modeling of DS-epi1 into the structure of heparinase II, as determined by crystal X-ray diffraction, showed conservation and striking similarities of the position of key amino acids shown to be essential in the catalytic mechanism of heparinase II. The importance of these amino acids has been validated through site-directed mutagenesis in recombinant DS-epi1. From these data, a mechanism of epimerization has been proposed where the C5 stereochemical inversion of the carboxyl group is achieved by the removal of the C5 proton from one side of the sugar followed by the readdition of another proton from the opposite side of the sugar plane. Four steps have been detailed: (1) His-450 removes the C5 proton from GlcA, and (2) the glycosidic linkage between the attacked GlcA and the GalNAc at the non-reducing side is broken by Tyr-261 with the subsequent formation of a C4–C5 double bond containing intermediate hexuronic acid. In (3), the intermediate hexuronic acid is protonated from the opposite side by His-205, and in (4), the glycosidic linkage is re-created by an unidentified residue. Steps 1 and 2 are in common with the mechanism of lyases, whereas subsequent steps 3 and 4 are DS-epimerase specific. In summary, due to structural similarities and catalytic mechanism, the two DS epimerases appear to have evolved from bacterial lyases.
Dermatan Sulfate Epimerase 1- and 2-Deficient Mice
DS-epi1- and 2-deficient mouse models have been generated, and the double DS-epi1/2 KO mouse, which should be a DS-null model, is under way (Maccarana et al. 2009; Bartolini et al. 2012). DS-epi1 and -2 are ubiquitously expressed in the body (Nakao et al. 2000; Goossens et al. 2003). Brain has the lowest expression of DS-epi1 and the highest expression of DS-epi2. DS-epi1 is the major epimerase in vivo and contributes to the major part of enzyme activity in extracts of most tissues. Ablation of DS-epi1 led to an approximate 80% reduction of IdoA in the entire animal. This reduction affected both decorin chains (73% reduction in skin decorin) and versican chains (89% reduction in skin versican). DS-epi1 is mainly responsible for the biosynthesis of the iduronic acid blocks, which decreased 90% in DS-epi1-deficient mice. A remaining single functional allele of DS-epi1 (i.e., in the condition of heterozygosity) results in half of the total protein amount and enzyme activity but interestingly leaves the CS/DS structure unaltered. DS-epi2 is the major epimerase in brain, accounting for approximately 90% of the epimerase activity. These data are in agreement with in situ hybridization expression patterns in the developing mouse brain (Akatsu et al. 2011). Nevertheless, brains in DS-epi2-deficient mice had a consistent, but not a dramatic, 38% reduction of IdoA in CS/DS, which in wild-type mice accounts for only 2% of total hexuronic acid residues. No phenotype in unchallenged DS-epi2-deficient mice could be observed, and brain extracellular matrix (ECM) features were normal. It thus remains to be explained why there are two DS-epimerases.
Although no apparent phenotype could be observed in DS-epi2-deficient mice, the phenotypes observed in DS-epi1-deficient mice are multiple. Mice lacking DS-epi1 having a mixed C57BL6/Sv129 genetic background were vital and fertile (Maccarana et al. 2009). However, in pure C57BL6 background, they died at a late stage of embryonic development and perinatally (unpublished observation), and in a pure NFR genetic background, 21% of them presented a birth defect called gastroschisis, consisting of a protrusion of the abdominal content due to a defect in the abdominal wall closure (R. Gustafsson, personal communication 2012). These mice are smaller, with a lower body weight at birth compared with wild-type littermates, and this phenotype persists in adulthood. This condition becomes aggravated when DS-epi2 is also inactivated (unpublished results). DS-epi1-deficient mice have a defect in skin collagen fibril formation, leading to skin fragility. This finding links the presence of high IdoA content in skin decorin CS/DS with its capacity to modulate collagen fibrils and deepens the previous knowledge implicating the decorin proteoglycan (Danielson et al. 1997) and its CS/DS chain (Ruhland et al. 2007) in this function. Also, iB structures, which decrease 70% in DS-epi1-deficient skin decorin, could take part in this function. Interactions between antiparallel CS/DS chains carried by two decorins bound to adjacent collagen fibrils could be altered in DS-epi1-deficient skin (Scott 1991). Another phenotype observed in DS-epi1-deficient mice is the alteration of the epidermal basement membrane, as seen by electron microscopy (S. Kalamajski, personal communication 2012). There is an abnormal distance between the basement membrane, which appears more convoluted, and the keratinocyte cellular membrane. This phenotype is reminiscent of alterations observed in the dermal-epidermal basement membrane of double nidogen-1 and 2-deficient mice (Mokkapati et al. 2008). Possibly linked to the altered basement membrane, the epidermal basal layer is also thicker (R. Gustafsson, personal communication 2012).
Genetic Diseases Related to DS-Specific Enzymes
A recently discovered genetic defect in the D4ST1 gene has been identified in 20 patients with a new type of Ehlers–Danlos syndrome (Zhang et al. 2010). The disorder affects various connective tissues and is the result of impaired assembly of collagen fibrils. The main features are joint dislocations and deformities, skin hyperextensibility and fragility, and recurrent large subcutaneous hematomas. The mutations lead to a loss of D4ST1 activity, and decorin produced by affected fibroblasts was devoid of IdoA (Miyake et al. 2010). Not surprisingly, given the fact that D4ST1 and the DS-epimerases are functionally closely linked in the biosynthesis of the IdoA blocks, the DS-epi1-deficient mouse presents similar defects in collagen fibril formation.
Also DS-epi2 could be the cause of a human disease. In fact, single-nucleotide polymorphism variants of DS-epi2 could be detected at very low frequency in a group of patients with type II bipolar disorder but not in the control group (Goossens et al. 2003).
Functions of Iduronic Acid in CS/DS Proteoglycans
Table 1 lists 29 CS/DS-PGs, 19 of which have not been analyzed in detail for the presence of IdoA. For instance, in studies of CS/DS in the nervous system, most research has analyzed only total CS/DS extracted from whole tissue and not from individual PGs. Further work comprising single PG purification and identification with specific antibodies followed by analysis of its chains with multiple chondroitinase ABC, AC, and B digestions is required.
CS/DS-PGs

BM, basement membrane; CS/DS, chondroitin/dermatan sulfate; ECM, extracellular matrix; FGF, fibroblast growth factor; HB-GAM, heparin-binding growth-associated molecule; HCII, heparin cofactor II; HGF/SF, hepatocyte growth factor/scatter factor; IdoA, iduronic acid; NA, not analyzed; N-CAM, neural cell adhesion molecule; PDGF, platelet-derived growth factor; PG, proteoglycan; TGF-β, transforming growth factor–β; VEGF, vascular endothelial growth factor; VWF, von Willebrand factor.
Some are part-time CS/DS-PGs because they can be unglycanated or carry HS chains. The division in groups is arbitrary because some CS/DS-PGs can belong to more than one group.
For the symbols of the units, refer to Figure 1.
Each PG has a reference reporting the experimental evidence of CS or eventually the CS/DS chain.
Indirect evidence was reported.
One classic way through which CS/DS-PGs can regulate biological processes at or near the cell surface is their ability to bind several growth factors. The flexibility of iduronic acid, which can be found as 1C4 and 4C1 chairs and 2S0 skew-boat conformers as opposed to GlcA found only in 4C1 chair conformation (Ferro et al. 1990), is thought to facilitate protein binding. For instance, hepatocyte growth factor/scatter factor (HGF/SC) requires IdoA residue flanked by sulfated hexosamine (Deakin et al. 2009). IdoA-containing domains have also been shown to interact with the fibroblast growth factor (FGF) family, thereby regulating wound healing and cell migration (Trowbridge and Gallo 2002).
Basement membrane CS/DS-PGs mainly regulate the architecture of the structure they are part of. There is no clear evidence of IdoA presence in these PGs.
In brain, the abundant CS/DS-PGs regulate plasticity during development and regeneration after injury. Most of the described CS/DS-PGs interact with specific brain molecules of the tenascin family, midkine, pleiotrophin, and neural cell adhesion molecules, in addition to growth factors (Yamada and Sugahara 2008).
The representative CS/DS-PG secreted in the extracellular space is decorin. It regulates collagen fibril formation and acts as a growth factor reservoir and as an anticoagulant factor, thanks to the ability of its CS/DS chain to interact with von Willebrand factor (VWF) (Guidetti et al. 2004) and with heparin cofactor II (HCII), the latter via iB structures (Tollefsen 2010). Decorin is therefore believed to be part of a complex acting at the subendothelial matrix regulating the clotting cascade and platelet activation.
The cell membrane CS/DS-PGs are co-receptors for several growth factors, and because of their transmembrane localization, they are involved in adhesion to substrates, migration, and intracellular signaling by connecting the outside environment with intracellular responses. IdoA presence is documented for CD44 after PDGF stimulation (Clark et al. 2004) and for syndecan-1 produced by fibroblasts grown on a 3D collagen I gel and during in vivo wound healing (Lee et al. 2004).
Endocan and SRPX2 have been recently described as IdoA-containing PGs. They show different functions, although they have been both connected with tumor progression. Endocan is a soluble PG bearing a single CS/DS chain expressed by vascular endothelium and released in the bloodstream. It is capable of binding HGF/SF via the CS/DS chain (Bechard et al. 2001), regulating the mitogenic activity of cells. In a recent work, its chains have been analyzed both by NMR and by chondroitinase ACI split products, showing 30% of IdoA present as isolated or two to four consecutive residues (Sarrazin et al. 2010). SRPX2 is upregulated in gastric cancer cells and affects cell adhesion, migration, and angiogenesis, and its IdoA content is evident given the partial digestion with chondroitinase B (Tanaka et al. 2012).
Finally, some members of the collagen family can also be decorated with CS/DS chains, although so far they are described as CS-PGs since a proper analysis for IdoA content is lacking. Collagen XV connects the basement membrane with underlying structures, and skin collagen XII and XIV have been described in specialized sites called anchoring plaques that stabilize the cohesion between the upper dermis and the basement membrane zone (Agarwal et al. 2012).
Dermatan Sulfate and Cancer
CS/DS proteoglycans have been extensively studied in cancer (Wegrowski and Maquart 2006). Focus in the past years has been on the role of the cancer stroma whose structure affects proliferation and migration of tumor cells (Mueller and Fusenig 2004). There are several mechanisms through which CS/DS could influence tumor biology, among which are cell proliferation and migration via binding of growth factors (Villena and Brandan 2004; Taylor et al. 2005; Radek et al. 2009) and hiding cancer cells from T-cell cytotoxic surveillance in blood through a selectin-mediated coating by platelets (Kozlowski et al. 2011), to mention a few. Cell surface CS/DS-PGs on human metastatic breast cancer cell lines have been suggested as receptors for P-selectin (Monzavi-Karbassi et al. 2007). Furthermore, removal of CS/DS chains by chondroitinase B has been shown to inhibit endothelial and melanoma proliferation and invasion (Denholm et al. 2001). DS-epi1 has been found upregulated in almost all cancers examined (Nakao et al. 2000). In a recent study, the hypothesis that IdoA-carrying proteoglycans affect tumorigenesis was directly tested (Thelin et al. 2012). DS-epi1 was downregulated in the esophageal cancer cell line TE-1. The consequent 80% to 90% reduction in IdoA in CS/DS chains hampered cellular invasion and migration. This effect was pronounced in the presence of hepatocyte growth factor (HGF) and mediated through the c-Met receptor. Moreover, DS-epi1-downregulated TE-1 cells have malfunctioning assembly of adhesion complexes and abnormal cytoskeleton architecture. In conclusion, iduronic acid in esophageal cancer cells is important for invasion, migration, and signaling through the c-Met receptor.
Future Directions
Biosynthesis of CS/DS still awaits the answer to several basic questions: How is the machinery producing structurally diverse chains controlled? Does it exist as a single macromolecular biosynthetic complex, or are there several specialized biosynthetic workstations made up of a few biosynthetic enzymes (e.g., DS-epi1 with D4ST1) to make the iduronic acid blocks? Where in the Golgi does the CS/DS biosynthesis take place? How does the core protein govern the variable structure of CS/DS chains? Which transcriptional/translational (and posttranslational) controls are exerted on the biosynthetic enzymes? Which proteoglycans contain iduronic acid? The answer to the latter question would also help to explain the largely unexplored phenotype of the DS-epi1-deficient mouse and the even more profound effect of the double ablation of DS-epi1 and -2. In fact, these models run longitudinally across the functions of each single CS/DS-PG. Finally, can the biological effects of iduronic acid be pharmacologically modulated by inhibition of DS-specific enzymes? For sure, future research on such a basic life component as CS/DS will surprise curious minds for years to come.
Footnotes
Acknowledgements
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from the Swedish Science Research Council; the Medical Faculty of Lund University; the Greta and Johan Kock, the Albert Österlund, the Anna-Greta Crafoord to [B.B.] foundations; the Mizutani Foundation for Glycoscience to [M.M. and A.M.].
