Abstract
Indoleamine 2,3-dioxygenase 1 (IDO1) catabolizes tryptophan to kynurenine at the first step of tryptophan metabolism. Recently, in addition to IDO1, a new isoform called IDO2 was identified. In this study, we examined the tissue expression pattern of IDO2 mRNA and the cellular localization of expressed IDO2 protein in mice. IDO1 mRNA expression was observed in the colon and epididymis, whereas IDO2 mRNA expression was found in the cerebral cortex, liver, kidney, and epididymis. Immunohistochemical analysis revealed that IDO2 protein was exclusively expressed on the hepatocytes, interlobular bile ducts, neuronal cells of the cerebrum cortex, Purkinje cells of the cerebellum cortex, lamina epithelialis, proximal convoluted tubule, and the collecting tubule of the kidney. In the epididymis, IDO1 protein expression was restricted to the caput, whereas IDO2 protein expression was observed on the caput, corpus, and cauda. Both IDO proteins were expressed on the caput, but both showed a different protein expression pattern in the segments. Immunohistochemical analysis in IDO1−/− mouse epididymis showed that IDO2 protein was extensively upregulated due to the loss of IDO1 expression.
Indoleamine 2,3-dioxygenase 1 (IDO1) is an intracellular enzyme that catalyzes indole compounds, including the essential amino acid tryptophan. IDO1 works at the first step of tryptophan metabolism via the kynurenine pathway. IDO1 plays a role in immune regulation. The enzyme suppresses effector T cell function and favors the differentiation of regulatory T cells (Tregs) through the capacity to degrade tryptophan to kynurenine and other downstream metabolites (Frumento et al. 2002; Grohmann et al. 2002; Uyttenhove et al. 2003).
Recently, an isoform of IDO1—called IDO2—was identified. The genomic structures of IDO1 and IDO2 genes are well conserved, and they are present via duplication on the same chromosome in mammals (Ball et al. 2007; Yuasa et al. 2007; Ball et al. 2009). IDO1 and IDO2 proteins have 43% similarity at the amino acid level in humans and mice. IDO1 and IDO2 catalyze the same substrate, but both enzymes have different efficiencies and distinct responses to their inhibitor (Metz et al. 2007; Loeb et al. 2009).
A recent study showed that IDO2 was predominantly expressed in the kidney and liver in humans and mice and that IDO2 expression did not change in murine malaria, a disease with high circulating levels of cytokines (Ball et al. 2009). However, there is no information available on the cellular distribution of IDO2.
In the present study, therefore, we examined the precise expression of IDO2 and the cellular localization of the enzyme in mouse tissues. We especially focused on the expression pattern of IDO2 in segments of epididymis in wild-type (WT) and IDO1−/− mice.
Materials and Methods
Mice
Six-week-old male mice were used in this study. IDO1 gene deficiency (IDO1−/−) mice of a C57BL/6J background were obtained from the Jackson Laboratory (Bar Harbor, ME). Mice that were WT (+/+) or homozygous null (−/−) for targeted disruption of the IDO1 gene were selected from the offspring of heterozygous/homozygous matings by using polymerase chain reaction (PCR) analysis of tail DNA. C57BL/6J mice obtained from Japan SHIMIZU Laboratory Supplies Co., Ltd. (Kyoto, Japan) were used as WT (IDO1+/+) controls. All experiments were performed in accordance with the Guidelines for Animal Care of Kyoto University.
RNA Extraction and RT-PCR Analysis
Total RNA was isolated from WT mouse tissue with Isogen (Nippon Gene; Tokyo, Japan), and the RNA concentration was determined spectrophotometrically at 260 nm. Reverse transcription PCR (RT-PCR) was performed by using the ReverTra Ace qPCR RT Kit (TOYOBO; Tokyo, Japan). The following PCR primers were synthesized: IDO1 sense: CCT ACA GAA GGT TCT AGA AGG, antisense: CTA CTG CAC TGG ATA CAG TG; IDO2 sense: CAC AGA GCC TCT GAA GTA CTC, antisense: CTA AGC ACC AGG ACA CAG GA.
Western Blot
Cell extracts were prepared by lysis buffer (100 mM Tris-HCL [pH 8.0], 100 mM NaCl, 0.5% NP40, and 0.2 mM phenylmethane sulfonylfluoride) and centrifuged for 5 min at 1800 × g at 4C. The supernatants collected for protein analysis were lysed with SDS-PAGE sample buffer, sonicated, and boiled. Protein samples were separated by one-dimensional SDS-PAGE, transferred onto a PVDF membrane, and blotted with the appropriate antibodies. The immunoblot signals were normalized to the corresponding β-actin (polyclonal antibody) from MBL (Tokyo, Japan). Antibody used for IDO1 was described in a previous report (Hoshi et al. 2010). For the detection of mouse IDO2 protein, we generated an IDO2 antibody. Two polypeptides, VPLSLGRYHIS and VATEPLK, corresponding to the mouse IDO2, were synthesized and coupled via cysteine to keyhole limpet hemocyanin. This conjugated peptide was used as the antigen to immunize rabbits to raise polyclonal antibodies (pAbs) against IDO2. This antibody specificity is mentioned in the results and shown in Supplemental Fig. S1.
Histopathology
Tissues were fixed in 10% formalin in phosphate-buffered saline (PBS) overnight and then embedded in paraffin. Sections (thickness, 4 µm) were used for hematoxylin and eosin (H&E) staining and immunohistochemical staining. The same antibodies for IDO1 and IDO2 were used as described in Western blotting. The preimmunized rabbit serum was used for negative controls. The results of one of several analyses with similar results are shown in each experiment.
Statistical Analysis
Values were expressed as means ± SD. Statistically significant differences (p<0.05) between two groups were determined using Student’s t-test. The numbers of mice are stated in the figure captions.
Results and Discussion
IDO1 and IDO2 Expression
First, we examined mRNA expression of IDO1 and IDO2 in various tissues of normal mice by RT-PCR. As shown in Fig. 1, IDO1 expression was detected in the colon and epididymis, whereas IDO2 was expressed in the brain, liver, kidney, and epididymis. The expression pattern of IDO2 was different from that of IDO1. However, both IDO1 and IDO2 were expressed in the epididymis. Next, to examine the protein expression and subcellular distribution of IDO2, we generated polyclonal antibodies (pAbs) in rabbits using a synthesized peptide corresponding to amino acids of IDO2 as an antigen. By immunoblotting, one of the affinity-purified pAbs specifically recognized the recombinant IDO2 protein but not the recombinant IDO1 (Suppl. Fig S1). We examined the protein expression levels of both enzymes in each organ by Western blot analysis using this antibody (Fig. 2). The quantitative protein expression of each gene was examined by normalization with β-actin. The highest IDO2 expression was found in the liver, followed by kidney, brain, and epididymis. In the epididymis, the protein expression of IDO1 was higher than that of IDO2. We detected no protein expression of IDO1 in the colon, although RT-PCR showed a clear expression of IDO1 mRNA in the tissue. Under normal conditions, no enzymatic activity of IDO1 has been detected in the mouse colon, although IDO1 is induced by inflammatory responses (Gupta et al. 2011). Immunohistochemical analysis (data not shown) showed that the epithelial cells of the colon did not express IDO1 protein. Some reports showed that IDO1-positive cells observed in various lymph tissues resemble various types of antigen-presenting cells (e.g., dendritic cells, macrophages, and reticular cells). Therefore, we speculated that IDO1 mRNA-positive signals come from the immune cells in the colon. As to the small population of positive cells that showed IDO1 mRNA expression, no protein expression of IDO1 could be detected in the colon.
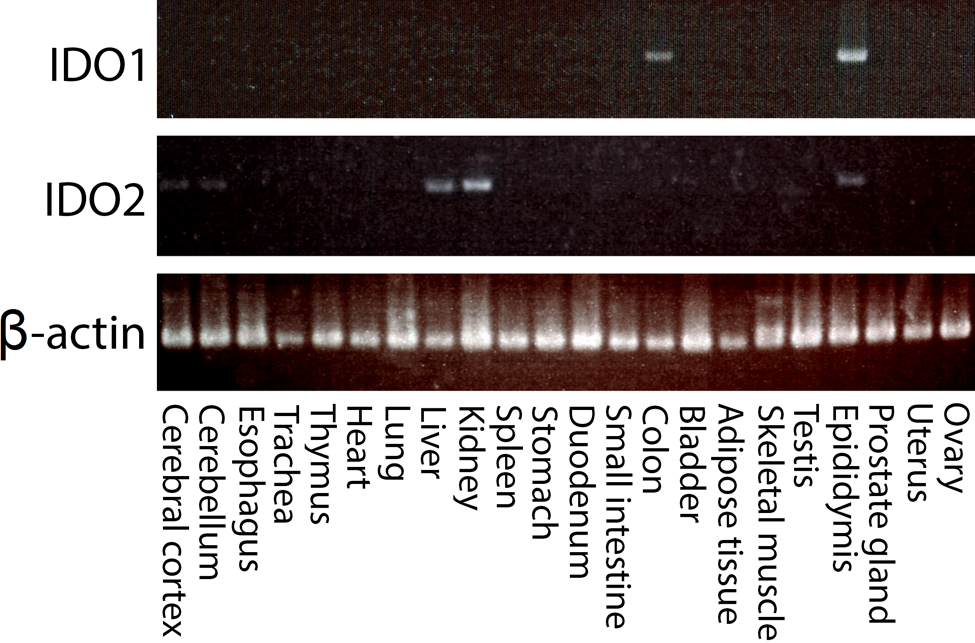
mRNA expression of IDO1 and IDO2. The mRNA expression of IDO1 and IDO2 in various tissues of a 6-week-old wild-type (WT) C57BL/6J mouse was examined by RT-PCR. IDO1 expression was observed in the colon and epididymis. IDO2 was expressed in the cerebral cortex, liver, kidney, and epididymis. Shown are the results of one of five mice with similar results.
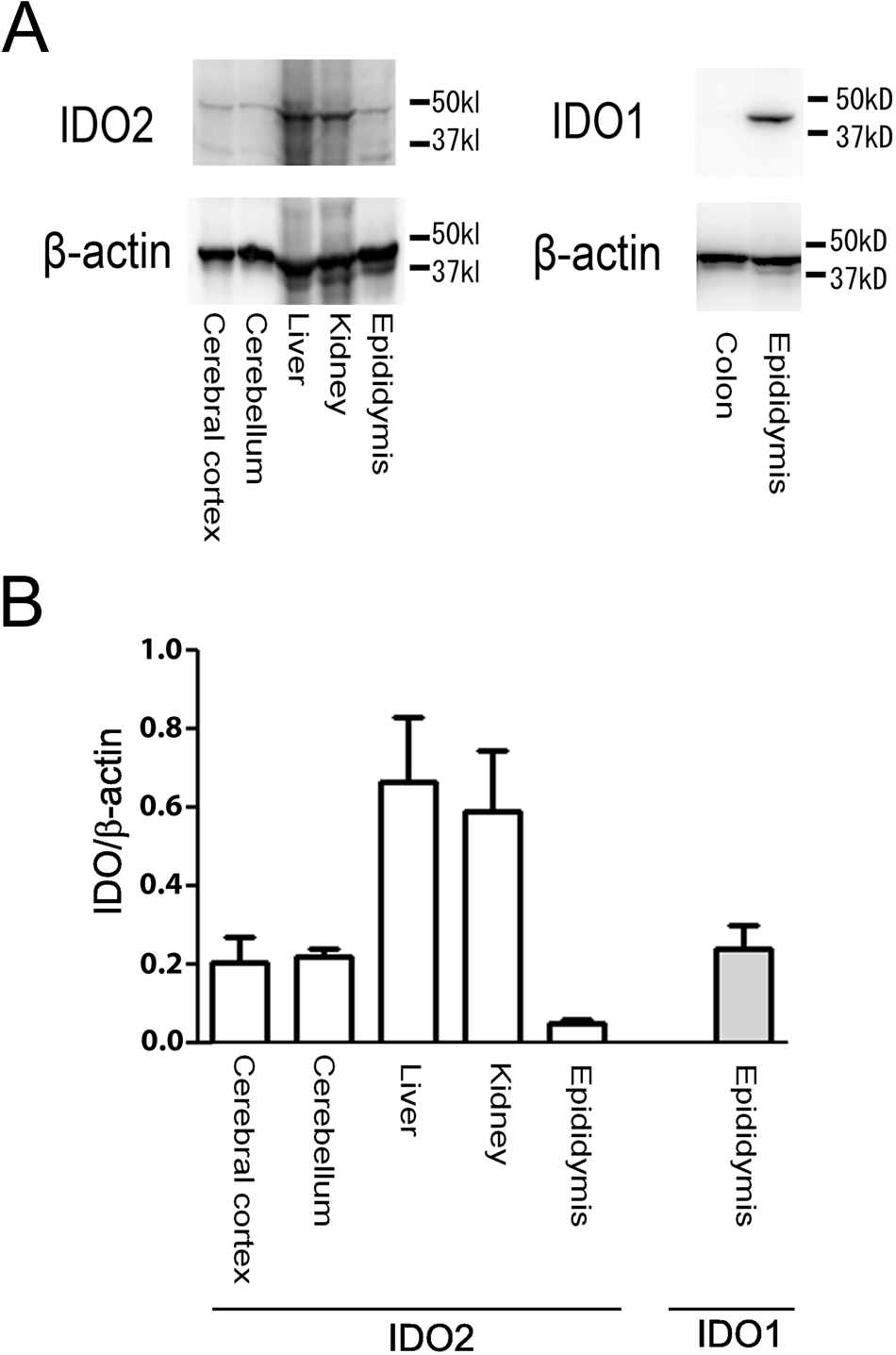
Protein expression of IDO1 and IDO2. (A) Western blot analysis showed that the results of protein expression were the same as the results of mRNA expression from Figure 1. (B) Quantitative expression of each gene was examined by normalization with β-actin (n=5).
Furthermore, we examined the subcellular distribution of IDO2 in mouse tissues by using the specific IDO2 pAb. When the formalin-fixed paraffin sections of mouse tissues were immunostained with anti–IDO2 pAb, positive signals were seen in the brain, liver, kidney, and epididymis (Fig. 3). Immunohistochemical microscopy analysis revealed that IDO2 was exclusively expressed on the neuronal cells of the cerebrum cortex, Purkinje cells of the cerebellum cortex, lamina epithelialis, hepatocytes, interlobular bile ducts, the proximal convoluted tubule, and the collecting tubule of the kidney (Fig. 3).
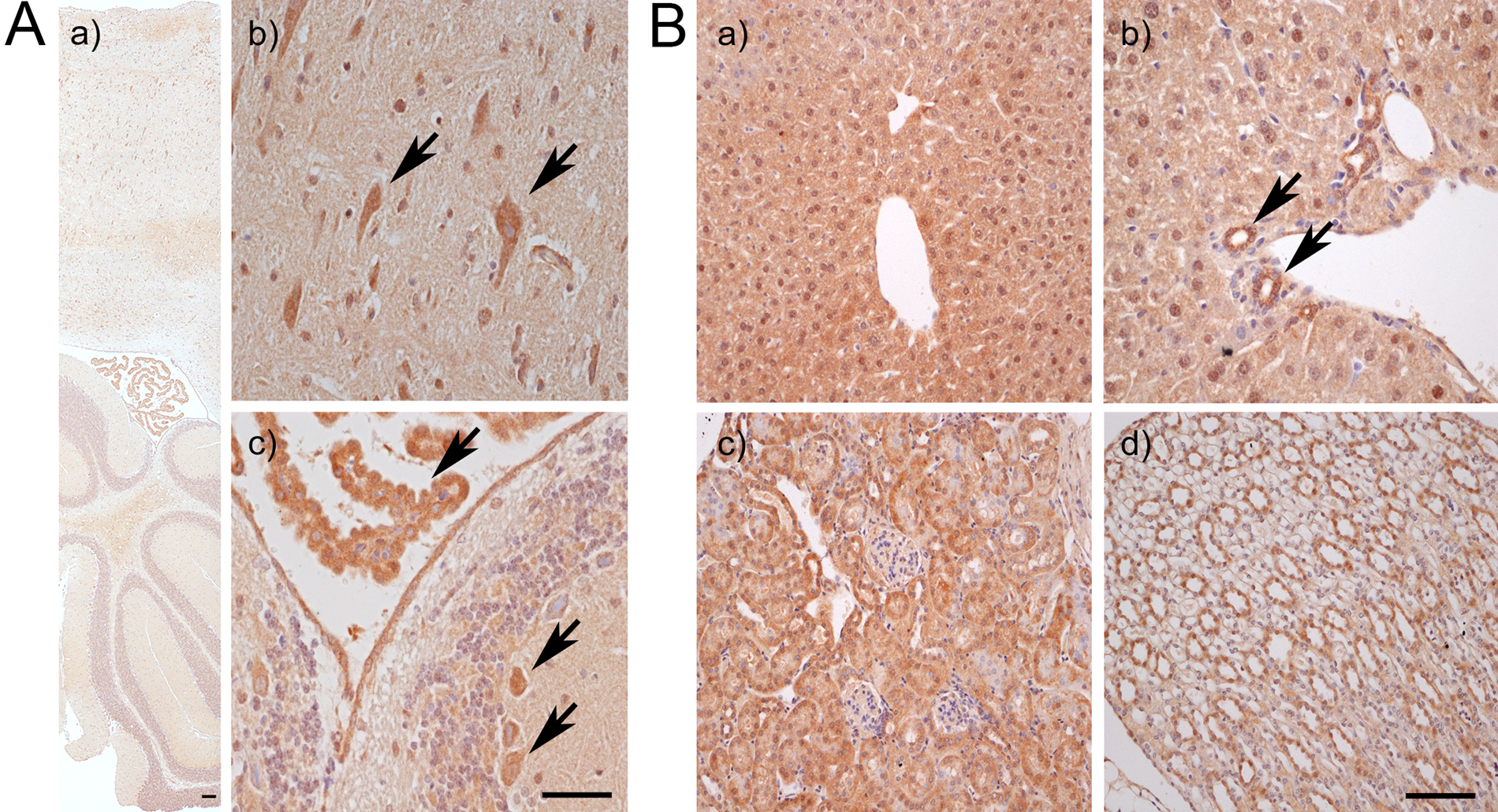
Localization of IDO2 in brain, liver, and kidney of wild-type (WT) mouse tissues. The “A” panels show IDO2 expression in the brain. (a) Lower magnifications of the brain. Scale bar: 500 µm. (b, c) Higher magnifications of the brain. Scale bar: 100 µm. IDO2-positive signal was detected in neuronal cells (arrows) of the cerebrum cortex (b) and Purkinje cells (arrows) of the cerebellum cortex and lamina epithelialis (c). The “B” panels show IDO2 expression in the liver and kidney. IDO2 was exclusively expressed on hepatocytes (a) and interlobular bile ducts (b) in the liver (arrows), as well as the proximal convoluted tubule (c) and collecting tubule (d) in the kidney. Scale bar: 100 µm.
IDO2 protein was predominantly expressed in the brain, liver, and kidney. IDO2 exists not only in mammals but also in lower vertebrates, but no gene corresponding to mammalian IDO1 has been found in the chicken and zebrafish genomes (Ball et al. 2007; Yuasa et al. 2007). Based on the gene expression evolution of IDO2 and our results, one can speculate that IDO2 contributes to the maintenance of tryptophan under physiological conditions. Some reports showed that IDO2 enzyme activity was much lower than that of IDO1 (Ball et al. 2007; Austin et al. 2010). Although the pathophysiological role of IDO2 is not clear at present, the enzymatic activity of IDO2 may be very important as a homeostatic control to maintain tryptophan concentration at basal levels.
IDO1 and IDO2 Expression in the Epididymis
The epididymis is a component of the testicular excurrent duct system, which is divided into the caput, corpus, and cauda. The luminal fluid microenvironment of the epididymis differs from segment to segment and is created by the epididymal epithelium, which has both secretory and absorptive activities (Turner 1991, 1995).
The epididymis is a unique organ that expresses both IDO1 and IDO2 in normal conditions. Immunohistochemical analysis revealed that IDO1 expression was restricted to the caput of the epididymis, whereas IDO2 expression was observed in the caput, corpus, and cauda (Fig. 4, Suppl. Fig. S2). These observations were in agreement with previous reports on IDO1 expression (Britan et al. 2006; Dai and Zhu 2010; Jrad-Lamine et al. 2011).
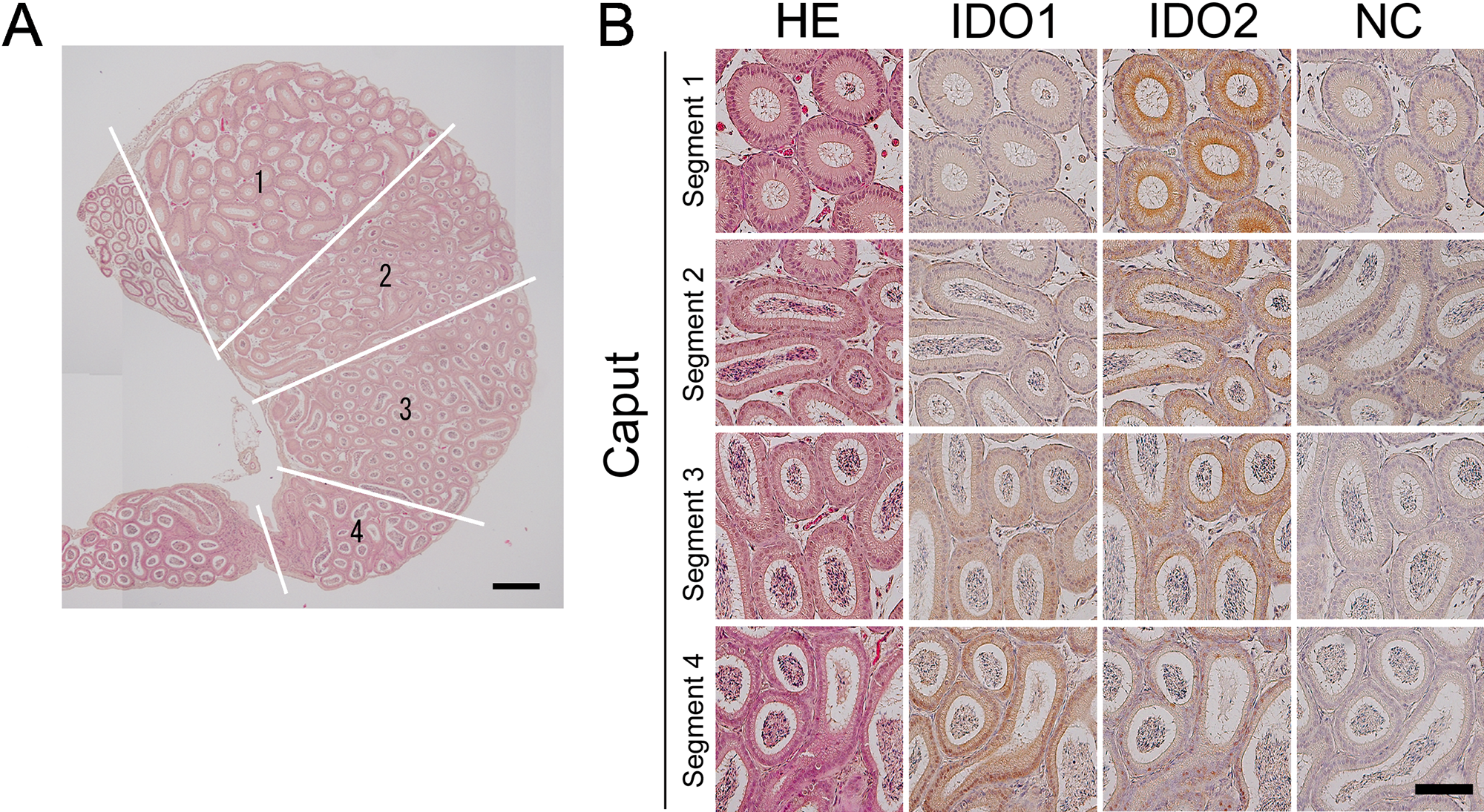
Segment-specific expression of IDO1 and IDO2 in the caput of the epididymis. Immunohistochemical analysis showed that IDO1 and IDO2 were expressed in different segments in the epididymis. (A) Longitudinal sections of the entire caput (hematoxylin and eosin stain). The left panel schematizes various segments separated from the connective septa found in the caput. Scale bar: 500 µm. (B) Higher magnifications of caput epididymal tubules in various segments. Scale bar: 100 µm.
In the caput of the epididymis, IDO1 and IDO2 showed different expression patterns in the segments. As shown in Fig. 4, segment 1 had the highest positive signal of IDO2, and segment 4 had high expression of IDO1. Both IDO1 and IDO2 work at the first step of tryptophan metabolism via the kynurenine pathway. Therefore, the difference in the distribution of IDO1 and IDO2 in the epididymis may be correlated with a functional difference between the genes, which is related to sperm maturation. Our data may prove useful in revealing the functional difference between IDO1 and IDO2.
Difference in IDO2 Expression between WT and IDO1−/−
It has been shown that the highest IDO1 activity is seen in the mouse epididymis without inflammatory responses (Yoshida et al. 1980). IDO1 KO mouse have decreased sperm numbers. These findings indicate that IDO1 in spermatozoa helps to control immunity. Therefore, the lack of IDO1 expression had no noticeable impact on overall male fertility but did induce a highly significant increase in both the number and the percentage of abnormal spermatozoa (Jrad-Lamine et al. 2011).
Immunohistochemical analysis in this study showed that IDO1−/− mice had the highest IDO2-positive signal in the segments of the caput (Fig. 5). Furthermore, the protein expression of IDO2 on the caput of each segment was compared in WT and IDO1−/− mice using Western blotting. IDO1−/− mice had the highest expression of IDO2 in segments 3 and 4 when compared with WT (Fig. 6). These results indicate that IDO2 expression was upregulated because of the loss of IDO1 expression. We measured kynurenine concentration in the epididymis of IDO1−/−. Although IDO2 expression was upregulated, the kynurenin concentration in IDO1−/− decreased when compared with the WT (data not shown) (Jrad-Lamine et al. 2011). Thus, the increased expression of IDO2 protein in the epididymis of IDO1−/− was not sufficient to compensate for the production of kynurenine in the tissue. However, these data might allow us to speculate that IDO2 protein expression can change depending on the environmental conditions. Furthermore, a recent study by Metz et al. (2007) has shown that IDO2 overexpression generates a stable LIP signal that presents independently of tryptophan availability. Although IDO2 has a very low enzymatic activity, its protein expression may control the signaling pathway. This function of IDO2 is quite distinct from that of IDO1, even if IDO2 and IDO1 cause a similar outcome to elicit immune tolerance.
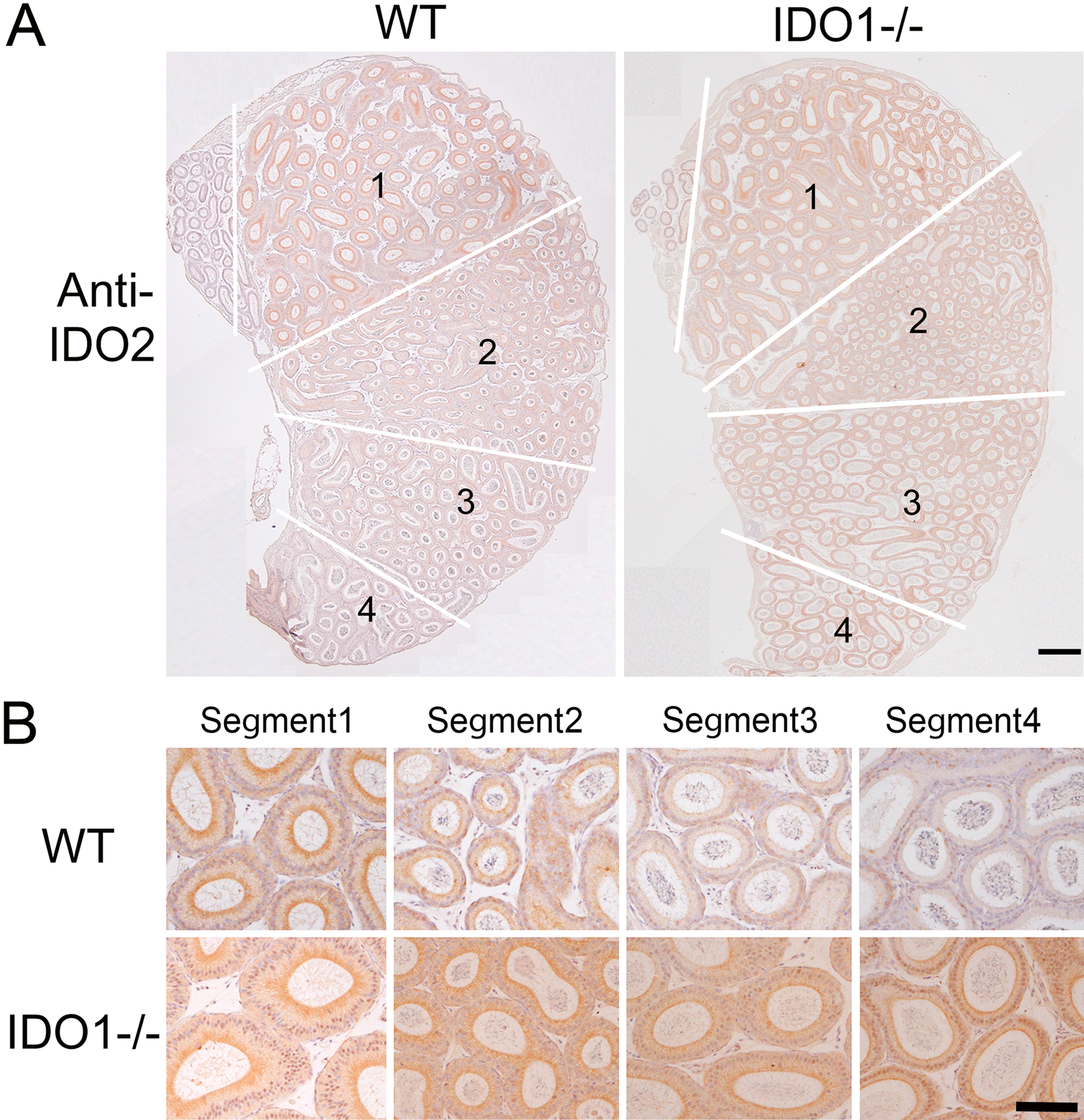
Localization of IDO2 in various caput segments of wild-type (WT) and IDO1−/− mice. (A) Longitudinal caput epididymal sections of WT and IDO1−/− mice aged 6 weeks. In both panels, connective septa separating the various caput segments are highlighted (white lines). Scale bar: 500 µm. (B) Higher magnifications of caput epididymal tubules at various segments from WT and IDO1−/− male mice. Scale bar: 100 µm.
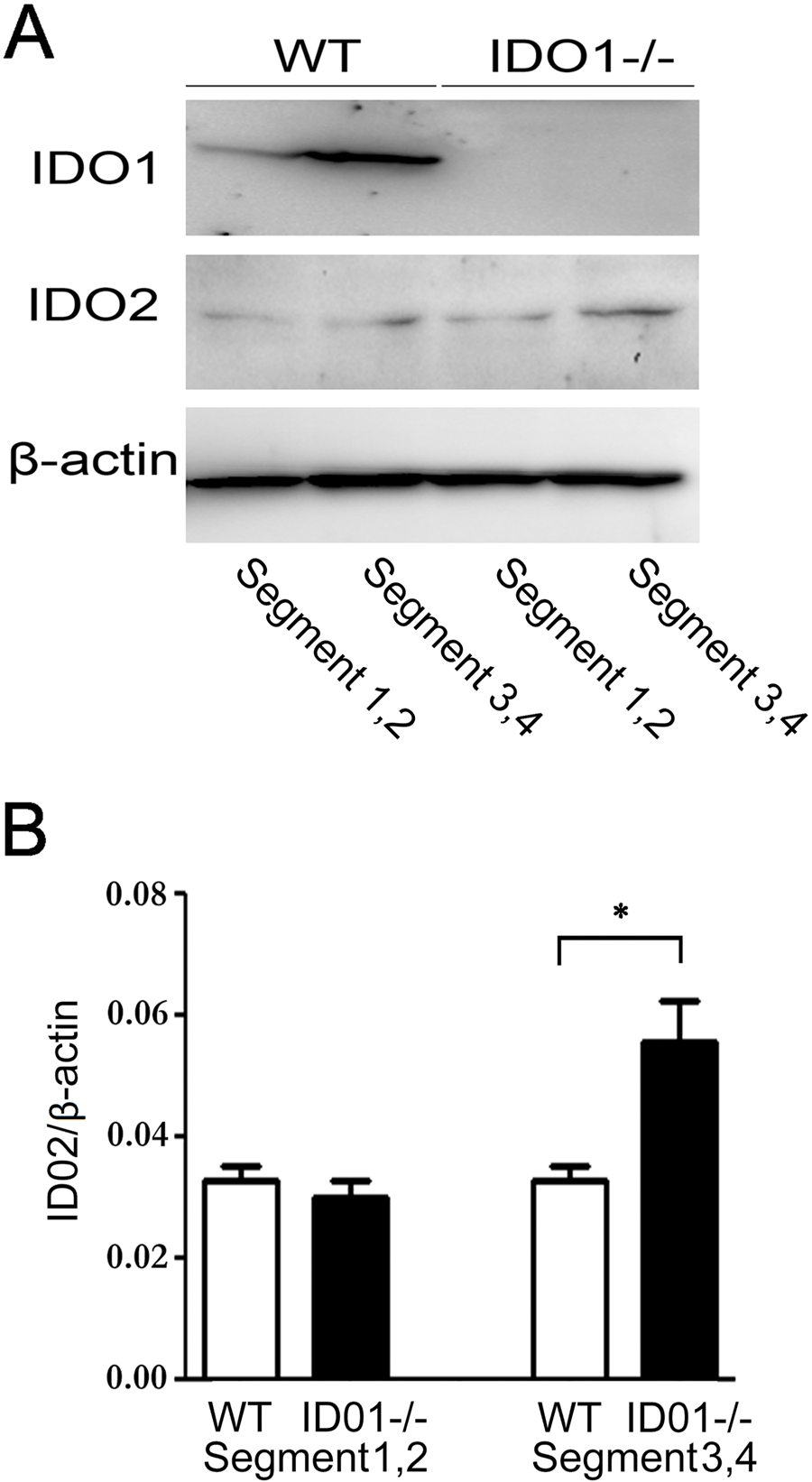
IDO2 expression in each segment of the caput was observed by Western blotting. (A) Caput segment was divided into segments 1,2 and segments 3,4. (B) Quantitative expression of IDO2 in each segment was examined (n=5). *p<0.05.
Spermatogenesis is a process that involves an array of cellular and biochemical events, collectively culminating in the formation of haploid spermatids from diploid precursor cells known as spermatogonia. Therefore, cellular and biochemical events in spermatogenesis are potently immunogenic (Robair and Hermo 1988). In the epididymis, high IDO expression induces an immunotolerant microenvironment to protect spermatozoa from the autoimmune response (Itoh et al. 2005).
In this study, we have clarified the distribution of IDO2 in various organs of mice. Furthermore, IDO2 protein expression has been found to increase in the epididymis of IDO1−/−. Further study is needed, but from the present results, it is speculated that IDO2 may contribute to immunoregulation and maintenance of physiological tryptophan metabolism.
Footnotes
Acknowledgements
We thank Dr. Y. Ohta for scientific discussion and proofreading the manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported in part by a Grant-in-Aid from the Ministry of Education, Science and Culture of Japan (to K. Saito [23390149] and Y. Yamamoto [23790621]).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
