Abstract
Histochemistry—chemistry in the context of biological tissue—is an invaluable set of techniques used to visualize biological structures. This field lies at the interface of organic chemistry, biochemistry, and biology. Integration of these disciplines over the past century has permitted the imaging of cells and tissues using microscopy. Today, by exploiting the unique chemical environments within cells, heterologous expression techniques, and enzymatic activity, histochemical methods can be used to visualize structures in living matter. This review focuses on the labeling techniques and organic fluorophores used in live cells.
The goal of histochemistry is to provide color and contrast to microscopic images. The field uses disparate techniques to accomplish the specific labeling of biological structures. Histochemists pioneered the use of small-molecule cellular stains, labeled molecules such as antibodies, and enzyme-mediated detection and signal amplification (Kiernan 2008). Historically, however, histochemistry has been synonymous with the imaging of fixed (i.e., dead) cells and tissues. The advent of genetic manipulation techniques has greatly expanded histochemical methods to living cells. This review examines the current collection of labeling strategies and discusses the correlating fluorescent dyes that allow biologists to add color to live systems.
Fluorescence and Fluorophores
Fluorescence occurs when a dye absorbs one or more photons, yielding a molecule in an excited state (Valeur 2002; Lakowicz 2006). This excited molecule can relax through a variety of processes, including the emission of a photon of another, typically longer, wavelength. It is this property—emission of a photon at a different wavelength—that allows detection of fluorophores in the presence of billions of other molecules in a biological sample. Thus, fluorescence-based techniques find widespread utility in bioresearch as they provide exquisite sensitivity and contrast in imaging experiments (Lichtman and Conchello 2005).
Fluorescent molecules come in various forms, including small-molecule dyes accessed by organic chemistry (Haugland et al. 2005; Lavis and Raines 2008), the genetically encoded fluorescent proteins (Giepmans et al. 2006; Chudakov et al. 2010), or the inorganic “quantum dots” (Smith A and Nie 2009). Here, we focus on the organic fluorophore classes, which encompass both synthetic small-molecule dyes and the fluorescent proteins. A key principle to understanding organic fluorophores is modularity. Building different substituents in and around the molecular structure of the dye allows tuning of the excitation maxima (λex) and emission maxima (λem) across the entire visible spectrum and into both the ultraviolet and infrared (Lavis and Raines 2008). Figure 1 summarizes the strategies used to label cells with organic fluorophores and shows specific examples of different fluorescent dye structures (compounds
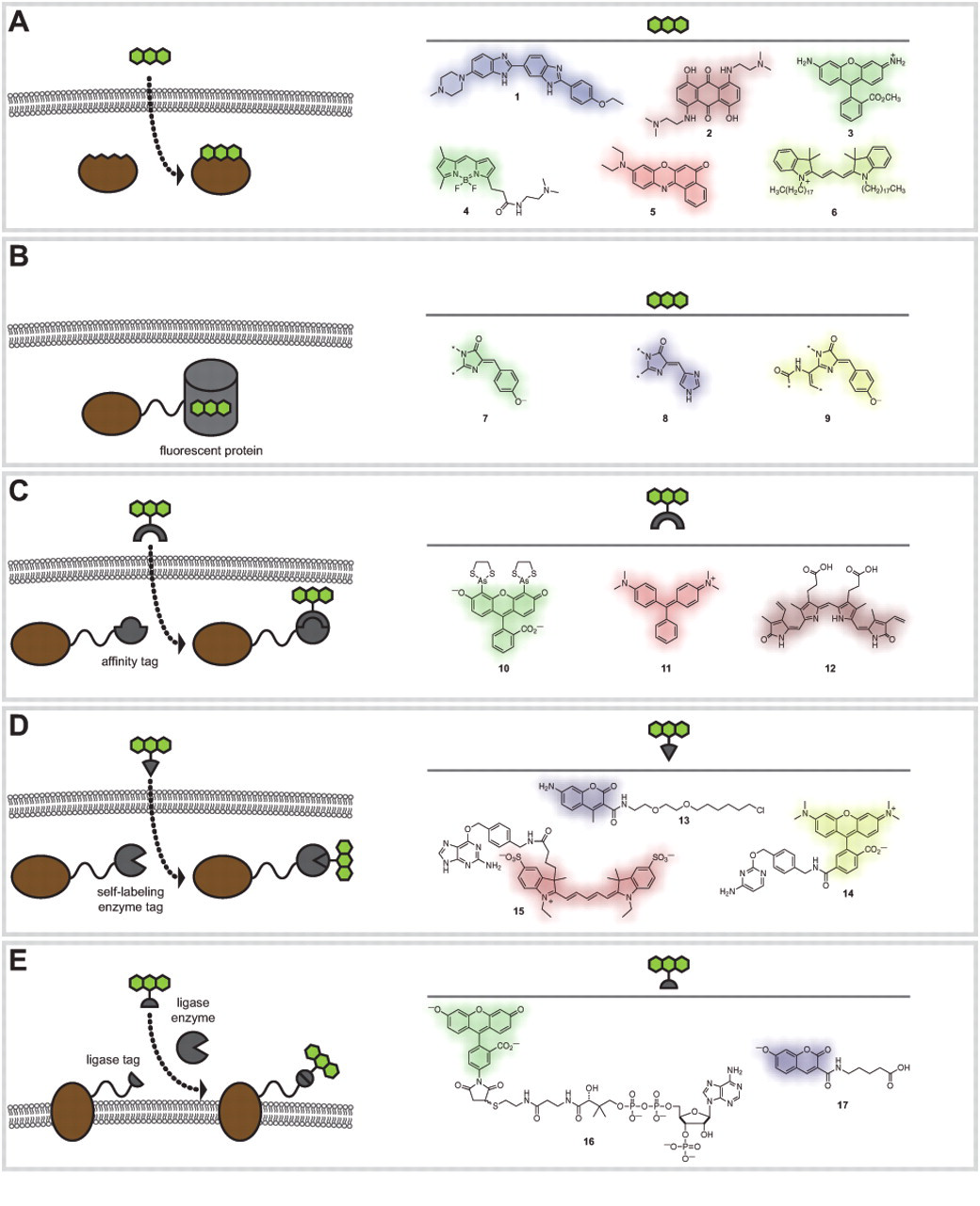
Labeling strategies and representative fluorescent labels
Intrinsic Labeling
Small fluorescent molecules with intrinsic affinity for the target can be used to stain regions of interest (Fig. 1A). This concept harkens back to the genesis of the histochemical discipline. Painstaking application of large collections of colored molecules allowed the discovery of compounds with affinity for different biological structures. The idea of the “magic bullet” put forth by Paul Ehrlich, a noted histochemist and Nobel laureate, was inspired by the affinity of dyes such as methylene blue and trypan red for particular types of cells (Strebhardt and Ullrich 2008). Today, this concept is being revisited as large libraries of fluorescent dyes are generated via diversity-oriented synthesis and screened for utility in live-cell imaging experiments (Vendrell et al. 2010).
An important subset of fluorescent stains includes molecules with intrinsic affinity for nucleic acids. For example, the dibenzimidazole Hoechst 33342 (
In addition to the screening approach, rational design of small-molecule fluorophores can furnish stains for native cellular structures. The labeling of organelles can be accomplished by exploiting different chemical environments, including electric potential, pH, and lipophilicity within a specific subcellular region (Haugland et al. 2005). For example, positively charged molecules can label mitochondria; this selective concentration is due to the potential across the mitochondrial inner membrane. Thus, dyes bearing a fixed positive charge such as rhodamine 123 (
Lipophilic regions inside cells can be stained with non-polar dyes. Compounds that exhibit increased fluorescence in hydrophobic environments are particularly advantageous for this application. One example of an environmentally sensitive dye is the oxazine-based Nile Red (
In addition to exploiting environmental differences to label subcellular locations, appending binding motifs for native proteins allows specific staining of cellular structures such as the endoplasmic reticulum (Zünkler et al. 2004). An interesting and useful extension of this strategy involves fashioning a suicide substrate for an enzyme where the enzymatic reaction elicits a large change in fluorescence. This fluorogenic reaction yields a labeled protein. Although used extensively for proteome investigations (Sadaghiani et al. 2007), these “activity-based probes” also can enable the imaging of native enzymes inside live cells (Blum et al. 2007).
Genetically Encoded Fluorophores
Perhaps the most straightforward method to label specific proteins in living cells is to use the genetically encoded fluorescent proteins (FPs; Fig. 1B) (Chudakov et al. 2010). These compounds are essentially small-molecule fluorophores encased in a protein shell. The chromophores form spontaneously, requiring only molecular oxygen. The ability to express many proteins as fusions with a fluorescent protein tag allows a myriad of biological imaging experiments. In addition to protein labeling, these fluorophores can be directed to various parts of the cell such as the nucleus, endoplasmic reticulum, cellular membrane, or mitochondria by expression of variants with different localization motifs (Chudakov et al. 2010).
The relative ease of labeling with fluorescent proteins has led to the development of numerous variants displaying a broad range of colors and properties. Encased in the protein structure, the green fluorescent protein (GFP) chromophore (
Genetically encoded fluorophores with larger chromophore structures have been discovered and further engineered to open up imaging in the red region of the visible spectrum using fluorescent proteins (Piatkevich and Verkhusha 2009). Such imaging agents are known collectively (and imprecisely) as red fluorescent proteins (RFPs). The chromophore of DsRed (
Despite the unquestionable utility of fluorescent proteins, this labeling strategy has some limitations. The relatively large size of the FP can disrupt the activity and location of the resulting fusion protein. These systems can use only the 20 proteogenic amino acids and thus suffer from a finite (albeit still growing) number of accessible fluorophore structures. Most FP systems express a protein variant with a single color. Given the larger diversity of small-molecule fluorescent dyes, it would be useful to combine the precise genetic targeting available with FPs and the structural flexibility of small-molecule fluorophores. Incorporation of non-natural amino acids is one possible route to this goal (Young and Schultz 2010), although the ultimate utility in mammalian systems might be limited (Uttamapinant et al. 2010). Alternatively, intersectional strategies that use a genetically expressed “tag” that interacts specifically with a small-molecule “ligand” can be advantageous. Appending the ligand motif to a fluorescent dye can allow the labeling of a single tagged protein with fluorophores of different colors or properties, depending on experimental requirements.
Affinity-Based Tag–Ligand Systems
Numerous genetically encoded tags have been discovered or engineered (O’Hare et al. 2007). The first type involves expression of a protein bearing an affinity tag that can bind tightly and specifically to a small-molecule ligand (Fig. 1C). Examples of tags include small-molecule binding proteins (Marks et al. 2004; Krusemark and Belshaw 2007; Wombacher et al. 2010), peptides (Kelly et al. 2007), antibody fragments (Szent-Gyorgyi et al. 2007), streptavidin for biotinylated probes (Wu et al. 2000), tetra-aspartate to bind Zn2+ complexes (Nonaka et al. 2010), and polyhistidine to bind nitrilotriacetic acid (NTA) motifs via a Ni2+ bridge (Goldsmith et al. 2006). Further engineering can introduce a nucleophilic residue on the tag near the ligand-binding site that can react with a corresponding electrophilic motif installed in the small-molecule ligand to form a covalent conjugate (Krusemark and Belshaw 2007; Gallagher et al. 2009; Nonaka et al. 2010).
A classic small-molecule binding tag is the tetracysteine-containing helix described by Griffin and coworkers (1998). The fluorescein arsenical helix binder (FlAsH,
Endogenous molecules can also serve as ligands, negating the need for delivery of exogenous compounds. The engineering of bacteriophytochrome proteins that bind the naturally occurring biliverdin
Self-Labeling Tag–Ligand Systems
In addition to labeling based on expressed affinity tags, exogenous enzymes can be used to label structures inside cells. Exploiting enzymatic activity can increase the specificity and speed of labeling in bioconjugation experiments (Kalia and Raines 2010). Enzymes in which the mechanism uses a covalent intermediate can be engineered to stop at that step of the reaction. Small proteins have been modified to serve as tags with substrate moiety–fluorophore conjugates as ligands (Fig. 1D). Examples of these “selflabeling tags” include a halogenase (HaloTag) (Los et al. 2008; Watkins et al. 2009) and variants of the O 6alkylguanine-DNA alkyltransferase (SNAPTag and CLIPTag) (Maurel et al. 2010). In addition, substrates for β-galactosidase have been constructed that label the protein with a concomitant change in the spectral properties of the dye (Komatsu et al. 2006). Such fluorogenic systems eliminate the need for washing unbound ligand out of the cell.
These self-labeling tags have been engineered to accommodate disparate dye structures. Derivatives of the pH-insensitive 7-aminocoumarin have established utility as the basis for fluorescent ligands for many labeling strategies, including the HaloTag (e.g., compound
Ligase-Based Tag–Ligand Systems
Enzymatic activity can be applied another way by exploiting ligases that facilitate posttranslational modifications on particular protein motifs (Fig. 1E). Here the expressed peptide tag can be significantly smaller than fluorescent proteins, affinity tags, and enzyme variants while facilitating the formation of a specific covalent bond. Examples include a formyl glycine-generating enzyme to install aldehydes (Carrico et al. 2007), the acyltransferase protein (ACPTag) (George et al. 2004), and biotin ligase from Escherichia coli that can attach biotin and biotin mimics to a short peptide sequence on the N-terminus of proteins (Chen et al. 2005).
It must be noted that these ligase-based strategies require two proteins and efficacious delivery of the small-molecule label. Therefore, the utility of these labeling systems is often confined to the exterior of cells. For example, coenzyme A derivative
Future Directions
Numerous systems are available to allow the extension of histochemical techniques inside living cells. The modularity of organic fluorophores allows multiple colors to be used to label structures for imaging experiments. Each labeling strategy has distinct advantages and weaknesses. Molecules with intrinsic affinity do not require genetic manipulation and could be useful for imaging in living organisms where genetic access is limited or unethical (e.g., humans). Fluorescent proteins will likely remain the dominant imaging modality in cell biology and model organisms, as addition of exogenous compounds is not required. Limitations of fluorescent protein labels can be circumvented by intersectional strategies employing both small-molecule fluorophore ligands and genetically encoded tags. These strategies do, however, add complexity to the experiment as they require expression of one (or more) proteins, delivery of the small-molecule ligand, and removal of any excess dye.
The established histochemical methods were developed and refined through collaborative efforts between chemists, biochemists, and biologists. Further improvements in live-cell labeling strategies will also require interdisciplinary research. Development of efficient synthetic methods for constructing fluorescent molecules along with improved screens is required to discover new stains with intrinsic affinity for biological structures. New fluorescent proteins with lower toxicity and enhanced optical properties are needed to perform long-term imaging experiments in vivo. Novel strategies for the efficient delivery and targeting of small molecules are also essential to expand the utility of labeling in complex cellular environments. Collectively, these exciting and useful tools will allow increasingly intricate histochemical experiments to illuminate biological processes in living cells.
Footnotes
Acknowledgements
I thank L. L. Looger and L. M. Wysocki for critical reading of the manuscript.
The author(s) declared no potential conflicts of interest with respect to the authorship and/or publication of this article.
Authorship of this article was supported by the Howard Hughes Medical Institute.
