Abstract
A flock of Rambouillet sheep was examined because of increased lamb mortality due to ineffective hemostasis at parturition. Decreased activities of coagulation factors II, VII, IX, and X, and severely reduced hepatic γ-glutamyl carboxylase activity with adequate vitamin K 2,3 epoxide reductase activity was determined.1,21 Parenteral vitamin K1 supplementation did not improve vitamin K-dependent coagulation factor activities in 3 affected lambs. Affected lamb γ-glutamyl carboxylase deoxyribonucleic acid was sequenced, and 4 single nucleotide polymorphisms (SNPs 2-5) of the γ-glutamyl carboxylase gene were identified. Single nucleotide polymorphism-4 results in an arginine to stop codon (UGA) substitution, which prematurely terminates the peptide at residue 686 (R686Stop). This genotype (GATT/GATT) has a strong association with the coagulopathy observed in clinically affected lambs, P < 0.001. The frequency of SNP-3 in exon 11 (R486H) within the MARC 1.1 database is high in the US sheep population overall. Gamma-glutamyl carboxylase activity in hepatic microsomes from a SNP-3 homozygous lamb lacking the SNP-4 mutation (GACC/GACC) was similar to control sheep homozygous for arginine at 486 and also lacking SNP-4 (TGCC/TGCC), indicating that the R486H does not measurably impact γ-glutamyl carboxylase activity. The remaining two SNPs (2 and 5) are located within non-coding intron sequences. These 4 SNPs allowed for determining the genotype associated with the observed fatal coagulopathy. Screening for the premature truncation (SNP-4) based on the presence of a Bbv I restriction site in clinically normal lambs but not in the homozygous affected lambs allows for detection of the heterozygous state (GATT/GACC), because carrier animals are clinically normal.
Introduction
Vitamin K is a required cofactor for the post-translational enzymatic conversion of varying numbers of glutamic acid residues (Glu) in the protein amino acid peptide sequence to γ-carboxylated glutamic acid (Gla). 36 Proteins that share this processing before being secreted are referred to as vitamin K-dependent (VKD) proteins. Post-translational modification occurs for VKD proteins involved in hemostasis, bone metabolism, mineralization, growth control, signal transduction, and cell survival. 3, 13, 34 Carboxylation allows calcium binding and tertiary protein folding for normal activity of VKD proteins II, VII, IX, X; protein C; protein S 16 ; and osteocalcin-hydroxyapatite binding in bone. This unique gamma carboxylation is catalyzed by γ-glutamyl carboxylase (GGCX) in multiple tissues.
Gamma carboxylation is the addition of a carboxyl group to the gamma carbon of glutamic acid 9 and occurs on the inner surface of the endoplasmic reticulum. 6 Energy is provided through the oxidation of vitamin K hydroquinone (K1H2) subsequent to binding the propeptide region of VKD proteins. 3, 9 Calcium-binding Gla residues are formed, and K1H2 is converted to vitamin K1 2,3 epoxide, which is recycled to K1H2, in a 2-step process by vitamin K epoxide reductase, an enzyme sensitive to inhibition by warfarin. This complex process allows for coagulation proteins to fold, bind receptors, and participate in a sequential reaction process culminating in the formation of a stable fibrin clot. 9
We reported a coagulopathy in Rambouillet Sheep similar to man. 1, 5, 8, 20, 21, 29 Breeding data suggest that the coagulopathy is inherited as an autosomal recessive trait. Affected lambs are born alive but lack the ability to achieve hemostasis of the transected umbilical artery and vein. Without intervention, newborn lambs continually bleed from the umbilicus and develop extensive subcutaneous and body-cavity hemorrhage resulting in death.
We have previously demonstrated that the underlying defect in affected lambs from this sheep flock is markedly reduced hepatic GGCX activity. To determine the underlying genetic and functional mechanism of reduced enzymatic activity, we determined the genetic sequence of the normal ovine GGCX and the genetic defect associated with the observed coagulopathy. We also developed an assay for detection of the heterozygous condition, because hemostasis and coagulation factor activities are normal in heterozygous carrier animals.
Materials and Methods
Three Rambouillet lambs were born with an activated clotting time of >7 minutes (normal, 2–3 minutes). Blood was collected for coagulation studies before transfusion with thawed citrate anticoagulated ovine plasma. Phenotypically affected lambs, prothrombin, activated partial thromboplastin times, and coagulation factor activities were determined with an MLA Electra 700 coagulation analyzer (Pleasantville, NY). Coagulation factor activity was reported as percent activity compared with pooled plasma (100% activity) from 20 unrelated adult sheep by using human factor deficient plasma (George King Biomedical, Overland Park, KS). Gamma-glutamyl carboxylase activity was measured to confirm the phenotype from hepatic microsomes prepared as outlined below. The sequence of GGCX gene was determined for all the members of the flock where the three affected lambs originated.
Hepatocellular microsomes
Microsomes were prepared according to Kotkow et al. 23 by using fresh liver obtained from all phenotypically affected lambs, adult sheep homozygous for single nucleotide polymorphism (SNP)-3 and lacking SNP-4 mutations, and an age/sex matched control lamb from a separate sheep flock and breed. Gamma-glutamyl carboxylase activity was determined from standard reaction mixtures (125 µl) containing 250 µg of microsomal protein, 0.8 M (NH4)2SO4, 28 mM morpholine propanesulfonic acid at pH 7.5, 0.5 M NaCl, 20 µl 1% [3-(cholamidopropyl) dimethylammonio]-1-propanesulfonate, 8 mM dithiothreitol, 10 µCi NaH14CO3, 220 µM of vitamin K1H2. Exogenous substrates added were either the pentapeptide phenylalanine-leucine-glutamic acid-glutamic acid-leucine (FLEEL) (3.6 mM) or FLEEL + bovine proII (propeptide sequence of bovine coagulation factor II, 8 µm), a peptide based on amino acids −18 to −1. The mixture was incubated at 25°C for 30 minutes in sealed tubes. Adding 1 ml of 10% trichloroacetic acid to the reaction mixture stopped reactions. Unbound 14CO2 was removed by gently boiling the mixture for 10 minutes, under a hood vented to the exterior. Total incorporation was determined by using a Beckman LS1801 liquid scintillation counter (Beckman Coulter, Fullerton, CA). Background counts per minute for assays lacking vitamin K1H2 were subtracted from all data points.
Populations and sampling
The sheep populations used for SNP discovery and validation consisted of 24 individuals from the affected flock, including the 3 phenotypically affected lambs, and the Meat Animal Research Center (MARC) Sheep Diversity Panel 1.1 (MSDP 1.1). 12 The MSDP 1.1 was designed to sample the breadth of genetic diversity in US breeds of sheep and consisted of 90 individuals from 9 popular breeds. This panel provides sufficient sequence information to estimate the frequency of sheep SNPs and haplotypes in US sheep populations.
Primer design, Polymerase Chain Reaction amplification and DNA sequencing
Primers were designed from the ovine GGCX cDNA sequence (AF312035, http://www.ncbi.nlm.nih.gov/), by using OLIGO 6.0 (National Biosciences Inc., Plymouth, MN). Primers generated a set of 7 overlapping amplicons spanning the entire ovine GGCX cDNA (15 exons). Human genomic sequence for GGCX was used to predict intron-exon junctions in ovine cDNA sequence (HSU65896 and NM 000821). 25 Amplicons spanning introns were targeted because these regions tend to be the most variable. 19, 38 Amplification and sequencing reactions were performed by using standard procedures. 15, 18
SNP identification
DNA segments from animals in the affected flock and MSDP 1.1 were amplified, sequenced, and compared to identify candidate SNPs. Sequences were edited and aligned by using Phred and Phrap software. 11 Semi-automated SNP identification was performed by using PolyPhred 3.0 and Consed 8.0. 28 Deviation of SNP genotype frequencies from the Hardy-Weinberg equilibrium was tested by using GENEPOP 3.1. 33 Relations between haplotypes were determined by median-joining-network analysis by using the method of Bandelt et al. 2 and Network 3.1.0.1 software. Comparisons of allele frequencies were performed by Pearson's χ2 test using system 9.01 (SPSS, Inc., Chicago, IL, USA).
RFLP of R686Stop
We designed a specific polymerase chain reaction (PCR) approach for the analysis of R686Stop to determine the carrier state in newborn lambs. The mutant allele of SNP-4 results in loss of a Bbv I restriction site. First, we amplified a 218-bp fragment, which included part of exon 14 and intron 14 of the ovine GGCX by using primers RFLP (5′-AAGGCTCCAAGAGATTGAAC-3′) and (5′-AGGGAAAAGTTAGCACTGG-3′). The PCR product was cut into fragments with Bbv I (New England Biolabs, Beverly, MA, USA) and subjected to electrophoresis in 3% agarose.
Results
Coagulation parameters
The 3 phenotypically affected bleeding lambs had prolonged prothrombin and activated partial thromboplastin times and decreased vitamin K-dependent factor activities (Table 1), verifying the GGCX deficiency phenotype.
Summary of affected lambs' coagulation data, vitamin K-dependent protein levels, and enzyme activity.∗
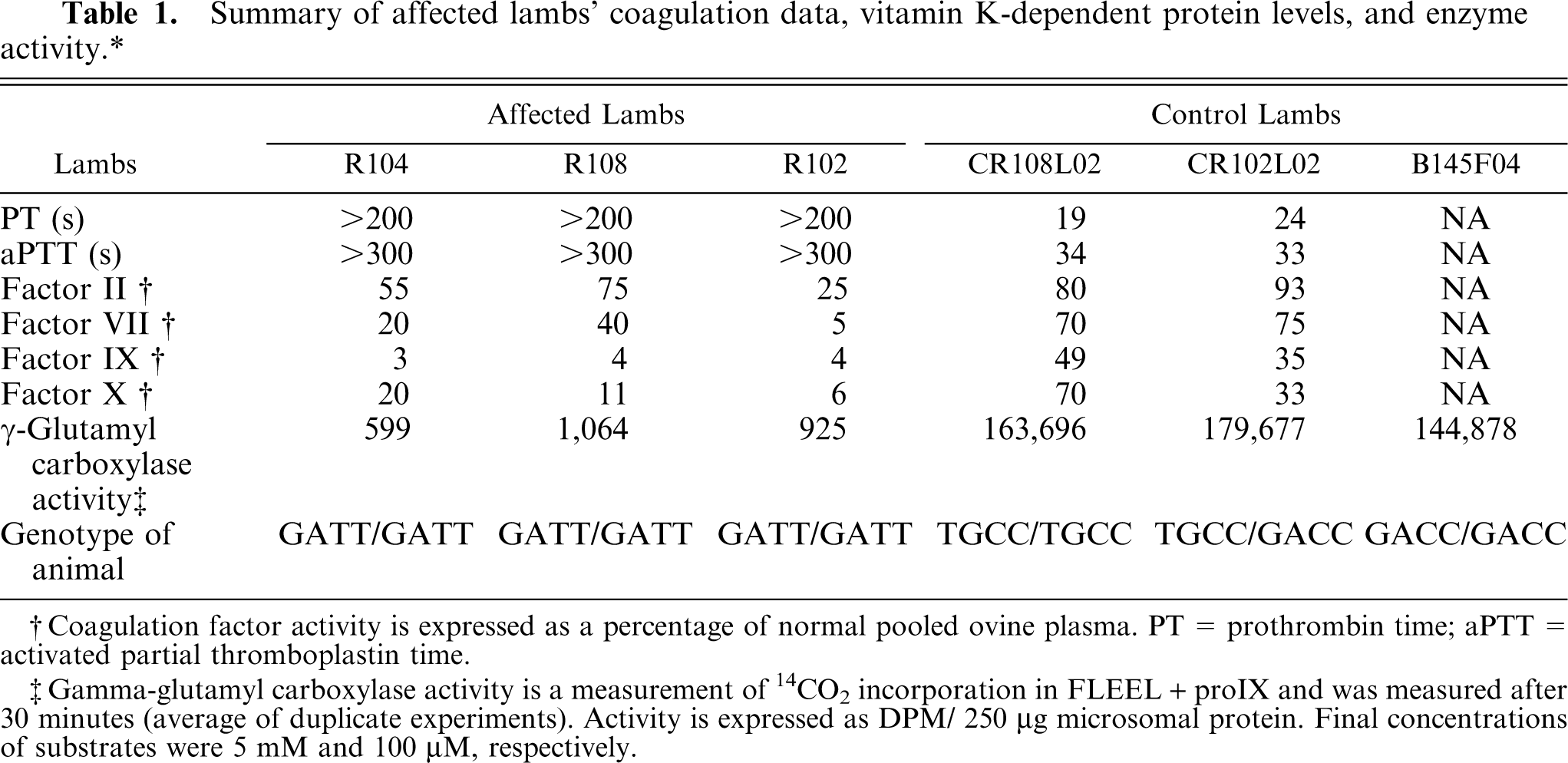
Coagulation factor activity is expressed as a percentage of normal pooled ovine plasma. PT = prothrombin time; aPTT = activated partial thromboplastin time.
Gamma-glutamyl carboxylase activity is a measurement of 14CO2 incorporation in FLEEL + proIX and was measured after 30 minutes (average of duplicate experiments). Activity is expressed as DPM/ 250 μg microsomal protein. Final concentrations of substrates were 5 mM and 100 μM, respectively.
Gamma-glutamyl carboxylase SNP and haplotype identification
The entire GGCX cDNA (2345 bp, less 29 bp of the 5′ end and 34 bp of the 3′ end that were subject to primer remodeling) was sequenced from 2 affected lambs, 1 carrier ewe, and 1 unrelated ewe. Two SNPs, SNP-3 and −4, were identified. Two GGCX genomic DNA segments, OARGGCXDS6 (AY330326) and OARGGCXDS7 (AY330325), which included these SNPS, were chosen for characterization based on cDNA sequencing results and robust PCR amplification (Fig. 1). The OARGGCXDS6 DNA sequence was 100% identical with the 3′ 28 nucleotides of exon 10 and the 5′ 70 nucleotides of exon 11 of ovine GGCX (AF312035). OARGGCXDS7 had 100% DNA sequence homology with the 3′ 148 nucleotides of exon 14 and the 5′ 150 nucleotides of exon 15 of ovine GGCX, indicating the DNA segments were amplified from the ovine GGCX gene locus.
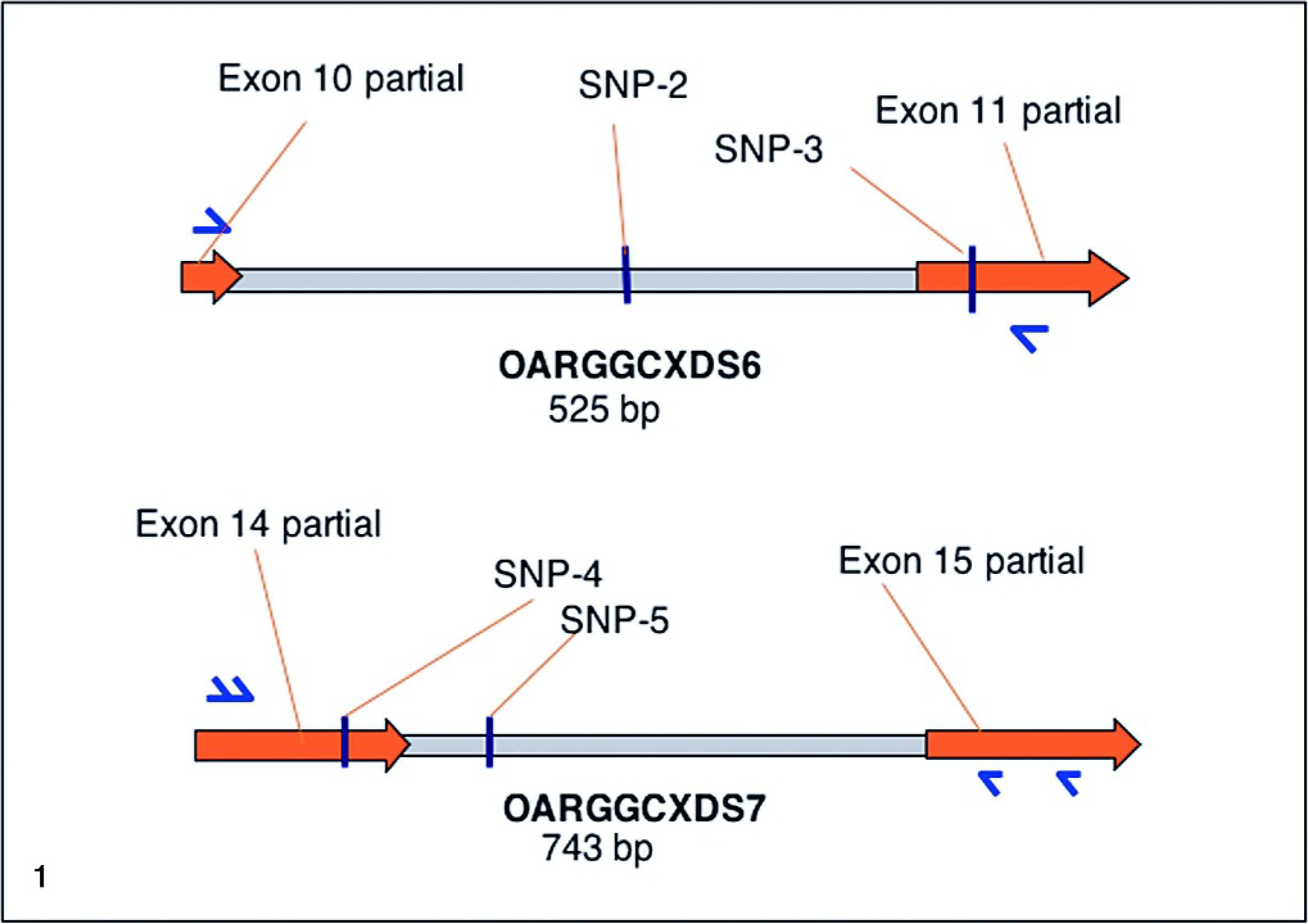
Sheep γ-glutamyl carboxylase SNPs. Individual SNPs and their locations within sheep γ-glutamyl carboxylase DNA.
OARGGCXDS6 and OARGGCXDS7, totaling 1245 bp of ovine GGCX genomic DNA, were sequenced from MSDP1.1 and 24 members of the affected flock, revealing 1 SNP in intron 10 (SNP-2), 1 in exon 11 (SNP-3), 1 in exon 14 (SNP-4), and 1 in intron 14 (SNP-5, Fig. 1). The allele of SNP-3 results in an A/G, R486H substitution (exon 11). The rare allele of SNP-4 results in a C/T, R686Stop substitution (exon 14). The homozygous and heterozygous state of SNP-2, −3, and −5 were present in the MSDP 1.1 panel (Table 2). The rare allele of SNP-4, resulting in the haplotype GATT, was only detected in the affected flock (Fig. 2 and Table 2).
SNP frequency in MARC sheep diversity panel 1.1.∗
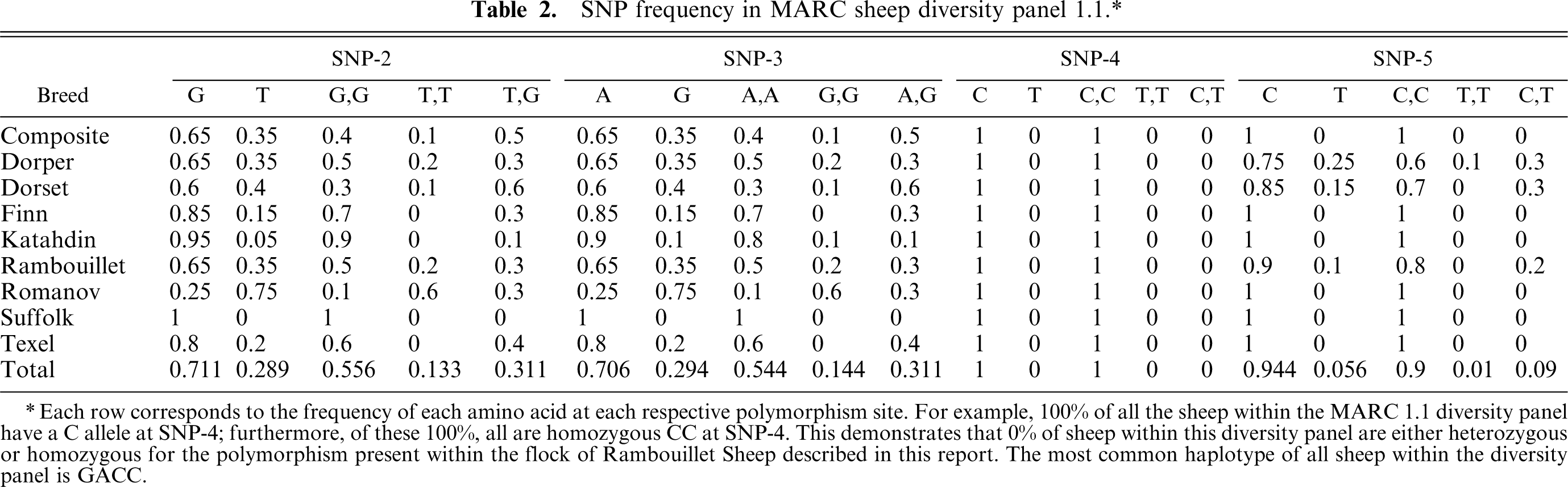
Each row corresponds to the frequency of each amino acid at each respective polymorphism site. For example, 100% of all the sheep within the MARC 1.1 diversity panel have a C allele at SNP-4; furthermore, of these 100%, all are homozygous CC at SNP-4. This demonstrates that 0% of sheep within this diversity panel are either heterozygous or homozygous for the polymorphism present within the flock of Rambouillet Sheep described in this report. The most common haplotype of all sheep within the diversity panel is GACC.
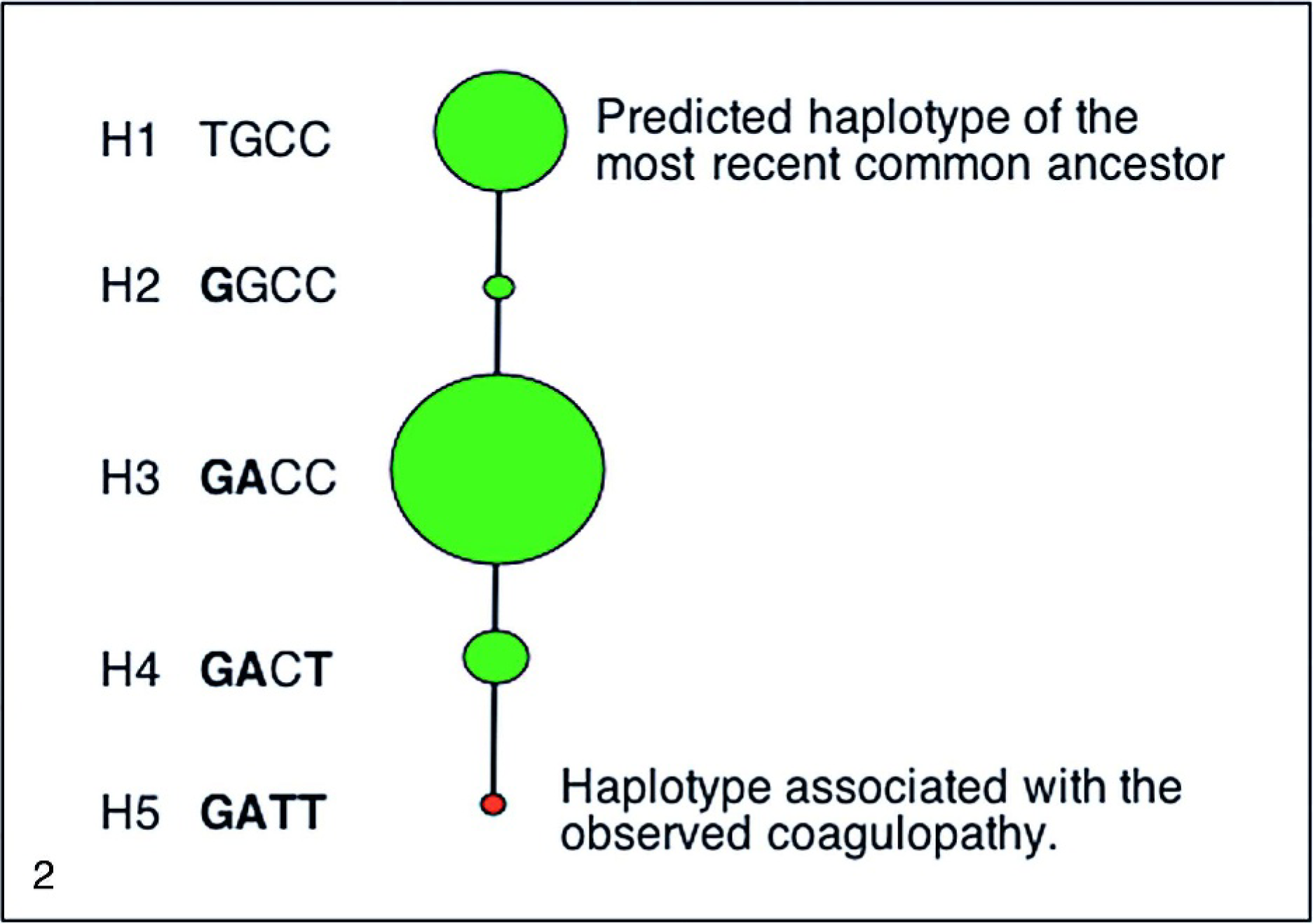
Sheep γ-glutamyl carboxylase haplotypes. Individual haplotypes of the most recent common ancestor (H1), haplotypes present within the sheep flock (H2–H5), and the haplotype associated with the heritable coagulopathy (H5). The diameter of each node reflects the frequency of that haplotype in the MARC sheep diversity panel 1.1. H1 corresponds with the ancestral haplotype, H2 corresponds with the ancestral haplotype with SNP-2, H3 corresponds with the ancestral haplotype with SNP-2 and SNP-3, H4 corresponds with the ancestral haplotype with SNP-2, −3, and −5, H5 corresponds to the ancestral haplotype plus SNP-2, −3, −4, and −5.
Enzyme activity
The GGCX activity of liver microsomes prepared from sheep liver homozygous for the identified SNP-3 R486H mutation was equivalent to control lamb homozygous for arginine at 486 (Table 1). Both animals were homozygous normal for arginine at 686.
RFLP of R686Stop
The gel electrophoresis of the 218-bp fragment amplified from normal, heterozygous, and affected lamb DNA is depicted in Fig. 3. The resulting gel electrophoresis of the affected animal had 1 dense band of amplified DNA fragment (lane 3) that was not significantly digested, whereas the normal unaffected sheep amplified DNA had 2 bands from the cleaved products of the amplified DNA fragment sequence (lane 4) retaining the Bbv I cleavage site. The heterozygous animal has 3 bands, 1 uncleaved larger sequence from the amplified DNA having the R686Stop and losing the Bbv 1 cleavage site, and the 2 cleaved products from the amplified normal sequence coding for the normal arginine at 686.
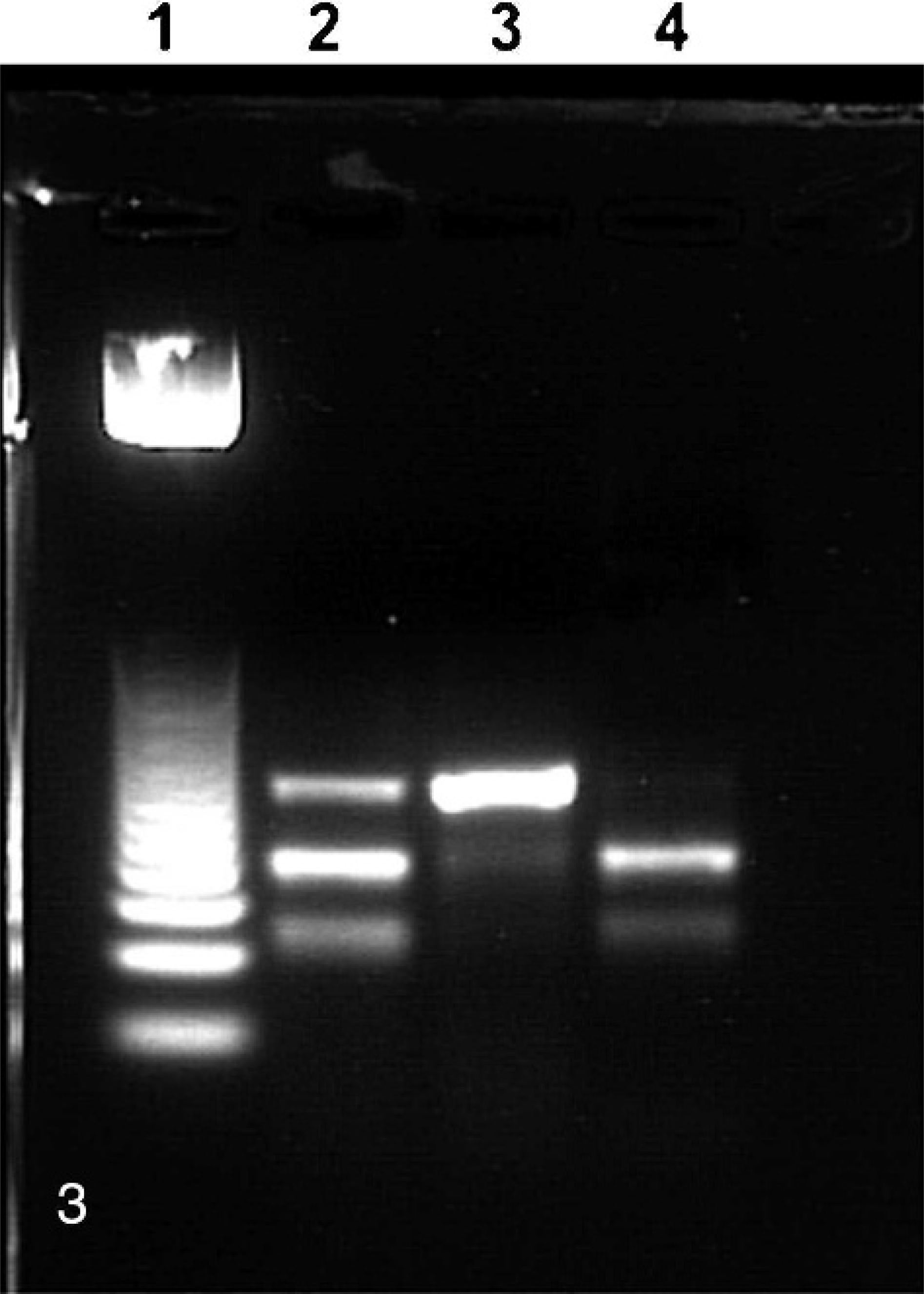
Restriction fragment length polymorphism analysis of R686Stop. Lane 1, 25 bp ladder; lane 2, DNA from a lamb that is heterozygous for SNP-4 mutation in GGCX genes; lane 3, DNA from affected sheep that is homozygous for the SNP-4 mutation on each gene; lane 4, DNA from normal control sheep lacking the SNP-4 mutation in either GGCX gene. 3% agarose gel.
Discussion
The 4 observed SNPs defined 5 haplotypes, all of which were observed in this study (Fig. 2). Haplotype-5 was only observed in the affected flock and none of the sheep in the diversity panel of genetic material. Homozygosity for haplotype-5 was strongly associated with the fatal coagulopathy, P < 0.001, and only animals homozygous for haplotype-5 were clinically affected.
These findings are similar to those reported by Roth et al. 32 in which truncation of the peptide to 676 residues resulted in a 28-fold decrease in the carboxylase Vmax . The theory that the vitamin K epoxidase domain also resides within this truncation 32 is supported by the lack of increased vitamin K-dependent coagulation factor activity (factor II, VII, IX, and X) to vitamin K1 administration in affected lambs. 21
Based upon the frequency of homozygosity (54.4%, Table 2) of SNP-3 (R486H) in the MARC 1.1 sheep diversity panel, we conclude it is not associated with the bleeding phenotype. The GGCX activity from a sheep homozygous for R486H (homozygous for histidine at residue 486) and lacking the truncation at 686 mutation was equivalent to the GGCX activity of a control sheep homozygous for arginine at 486 and lacking the R686Stop mutation, further supporting the contention that this variation does not influence enzyme activity.
The variable coagulation factor activities of factors II, VII, IX, and X at the initial examination of these affected lambs pose an interesting dilemma. Propeptide binding regions for GGCX are variable between coagulation factors and their influence on GGCX activity is somewhat controversial. According to Wu et al., 40 the propeptide binding site lies somewhere COOH-terminal to residue 438, while other investigators suggest it is either within the first 314 amino acids of peptide 24, 40 or involves 2 NH2-terminal cysteine residues. 31 Recent evidence suggests that residues 495–513 are involved in propeptide binding. 26 These reported studies suggest the possibility that multiple, noncontiguous regions of the carboxylase may form functionally important propeptide binding sites.
The various propeptide affinities have been determined for human recombinant carboxylase. 35 Evidence suggests that the propeptide release from GGCX after fully carboxylating all Glu residues is likely to be the rate-limiting step in factor IX turnover in vitro. 7, 16, 27, 30 Propeptide II binds 10-fold more loosely to the carboxylase than proIX, and studies evaluating expression of recombinant prothrombin and FIX showed a greater percentage of fully carboxylated prothrombin presumably because of the higher off rate of the propeptide from the carboxylase. 21 Wallin et al. 39 found that while prothrombin and proX were tightly bound to the carboxylase (though not as tight as proIX), in vitro carboxylation of these substrates resulted in release of prothrombin precursors but not FX precursors. These findings support the notion that propeptides with lower affinity for GGCX have higher substrate turnover, more efficient carboxylation, and suggest one explanation of the variable coagulation factor activities observed in affected lambs.
We have successfully identified the genotype associated with the coagulopathy in this sheep flock, a premature stop codon at 686. The affected lamb genotype, GATT, represents the homozygous state of each individual SNP. For example, all affected lambs are homozygous for the amino acid adenine (A) at SNP-3 (residue 486). This unique flock represents the only known animal model of defective GGCX activity and allows for in vivo investigation of the function and importance of various VKD proteins.
Footnotes
Acknowledgements
This research was supported by NIH Grant 1R24 RR17569-01A1.
