Abstract
Cyclooxygenase-2 (COX-2) was detected and localized in 15 pigs with naturally occurring pleuropneumonia using a 437–base pair digoxigenin-labeled cDNA probe in an in situ hybridization protocol. Histopathologic changes in the acute stage were characterized by coagulative necrosis of lung parenchyma, hemorrhage, vascular thrombosis, edema, fibrin deposition, and infiltration of lung parenchyma by neutrophils and alveolar macrophages in nine pigs. In chronic lesions, a thick layer of granulation tissue surrounded foci of pulmonary necrosis in six pigs. All 15 pigs infected with
Porcine pleuropneumonia, caused by
Arachidonic acid–derived prostaglandins are potent inflammatory mediators, and they also modulate immune responses and physiologic processed such as bronchoconstriction, vasodilation, and mucus secretion in the lung.
12,28
In addition, prostaglandins increase transvascular hydrostatic pressure and promote increased microvascular permeability by synergizing with and stimulating the release of secondary mediators.
11,24
Prostaglandins are synthesized from arachidonic acid by two cyclooxygenase (COX) isoforms. COX-1 facilitates the synthesis of prostaglandins involved in homeostatic functions, whereas COX-2 is an inducible enzyme predominantly expressed at sites of inflammation. COX-2 expression is induced by LPS and inflammatory cytokines such as interleukin-1 (IL-1) and tumor necrosis factor-α (TNF-α).
16,21,25
The objective of this study was to determine whether COX-2 was expressed in the lungs of pigs naturally infected with
Materials and Methods
Animals
Samples were obtained at necropsy from pigs submitted to the Department of Veterinary Pathology of Seoul National University from January 1994 to December 1997. Fifteen pigs (Nos. 1–15) approximately 87–140 days old, from 15 different herds, were selected on the basis of clinical signs, characteristic lesions, bacteria isolation, and serotype. All 15 pigs were negative for
Positive control sections were from pigs that had been naturally infected with
Tissue processing
Samples of lung, liver, kidney, palatine tonsil, mediastinal lymph node, and large and small intestine specimens were collected from infected and noninfected animals, fixed in 10% (w/v) neutral buffered formalin for 24–48 hours, and embedded in paraffin according to standard laboratory procedures. Sections were cut 4-μm thick, floated on a water bath containing diethylpyrocarbonate-treated (DEPC) water and mounted on positively charged slides (Superfrost/Plus slide, Erie Scientific Co., Portsmouth, NH).
RNA extraction
LPS-stimulated alveolar macrophages, and pleuropneumonic lung specimens from 15 pigs naturally infected with
Primer
The primers were designed based on the porcine COX-2 cDNA sequence (GenBank accession number AF207824). The forward and reverse primers were 5′-GGAGAGACAGCATAAACTGC-3′ (nucleotides 413–432) and 5′-GTGTGTTAAACTCAGCAGCA-3′ (nucleotides 830–849), respectively. The primers amplified a 437–base pair (bp) cDNA fragment.
Reverse transcription-polymerase chain reaction
For the first-strand cDNA synthesis, 1 μl of the COX-2 RNA (5 ng/μl) was supplemented in a total reaction volume of 20 μl with 1× RT buffer (50 mM Tris-HCl, 8 mM MgCl2, 30 mM KCl, 1 mM dithiothreitol [pH 8.3]), 0.5 mM (each) deoxynucleotide triphosphates (dNTPs) (GeneClone, Suwon, Korea), 2.5 μM random hexanucleotide mixture (Applied Biosystems, Foster City, CA), 20 U of RNase inhibitor (Applied Biosystems), and 50 U of Moloney murine leukemia virus reverse transcriptase (Applied Biosystems). After incubation for 15 minutes at 42 C, the mixture was incubated for 5 minutes at 99 C to denature the products. The mixture was then chilled on ice.
The composition of the polymerase chain reaction (PCR) mixture (150 μl) was 30 μl of cDNA (5 ng/μl), 2 μl of each primer (250 nM), 15 μl of 10× PCR buffer (10 mM Tris-HCl, 40 mM KCl, 1.5 mM MgCl2 [pH 8.3]), 1.2 μl of each dNTP (0.2 mM), 2.5 unit of
Preparation of labeled probe
PCR products of COX-2 were purified using a 30-kD cutoff membrane ultrafiltration filter. The nucleotide sequences of the purified PCR products were determined by use of BigDye chemistry with the ABI Prism Sequencer (Applied Biosystems). Sequencing was performed in the purified PCR products before PCR products were labeled by random priming with digoxigenin-dUTP (Boehringer Mannheim, Indianapolis, IN) according to the manufacturer's instructions.
In situ hybridization
Sections were deparaffinized in xylene and rehydrated in phosphate-buffered saline (PBS) (pH 7.4, 0.01 M) for 5 minutes. Deproteinization was carried out in 0.2 N HCl for 20 minutes at room temperature. Tissues were then digested at 37 C for 20 minutes in 100 μg/ml proteinase K (Gibco BRL) in PBS (pH 7.4, 0.01 M). Serial sections of each tissue section examined were treated with RNase A (Boehringer Mannheim) at 100 μg/ml in 10 mM Tris-HCl (pH 7.4) for 30 minutes at 37 C to remove target RNA as a specificity control. After digestion, tissues were fixed in 4% paraformaldehyde in PBS for 10 minutes. After rinsing twice with PBS, the slides were acetylated in 300 ml of 0.1 mM triethanolamine-HCl buffer (pH 8.0) to which 0.75 ml of acetic anhydride (0.25%) had been added. After 5 minutes, an additional 0.75 ml of acetic anhydride was added, and 5 minutes later the slides were rinsed in 2× saline sodium citrate (SSC) (1× SSC contains 50 mM NaCl and 15 mM sodium citrate pH 7.0). The slides were allowed to equilibrate for 60 minutes in a standard hybridization buffer that consisted of 5× SSC with 50% deionized formamide, 10× 2% buffered blocking solution (Boehringer Mannheim), 0.1%
Hybridization was done overnight at 45 C for COX-2. The digoxigenin-labeled probe (0.1 ng/μl) was diluted in 300 μl of the standard hybridization buffer, heated for 10 minutes at 95 C on a heating block, and quenched on ice before being applied to the tissue sections. Approximately, 50 ng of the digoxigenin-labeled probe was added to the standard hybridization buffer (50 μl), which was then layered over the section. Fluid was held in place by a coverslip, and the edges were sealed with rubber cement. After hybridization, sections were thoroughly washed twice in 4× SSC for 5 minutes at room temperature, twice in 2× SSC for 10 minutes at 37 C, twice in 0.2× SSC for 5 minutes at room temperature, and once in maleic acid buffer (100 mM maleic acid and 150 mM NaCl, pH 7.5) for 5 minutes at room temperature.
For detection of hybridization, sections were incubated with anti-digoxigenin conjugated with alkaline phosphatase (Boehringer Mannheim) diluted 1:250 in 0.1 M Tris-HCl (pH 7.4), 0.15 M NaCl with 1% blocking reagent (Boehringer Mannheim). After three washes in buffer, substrate consisting of nitroblue tetrazolium and 5-bromocresyl-3-indolylphosphate was layered over the sections. Color was allowed to develop for 5–8 hours in the dark, and the reaction was stopped by dipping slides briefly in tri-ethylenediaminetetraacetic acid buffer (10 mM Tris-HCl and 1 mM EDTA, pH 8.0). Sections were counterstained with 0.5% methyl green, and the slides were then washed with distilled water for 1 minute, allowed to dry completely, dipped into the absolute xylene, and coverslipped with Canada balsam mounting medium (Hayashi Pure Chemical Industries Ltd., Osaka, Japan).
Results
Reverse transcription-polymerase chain reaction
Amplification of template cDNA with primers of COX-2 resulted in amplified products corresponding to those of the predicted size. PCR products were sequenced, and their identity was confirmed as COX-2. To investigate whether
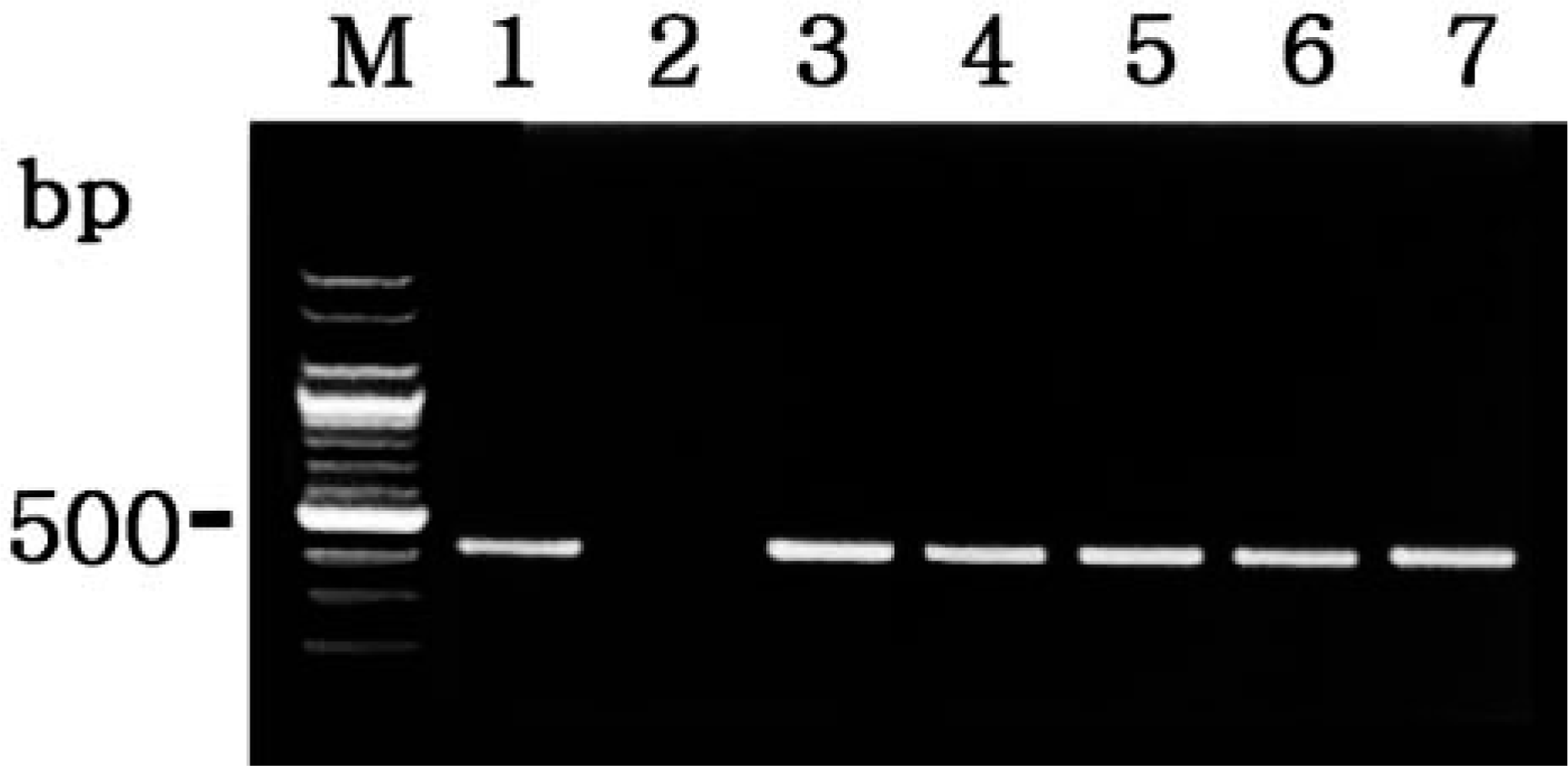
Agarose gel electrophoresis of PCR-amplified COX-2 cDNA products. From left to right: M = 100-bp DNA ladder; lane 1 = positive control from LPS-treated macrophages; lane 2 = negative control from normal macrophages; lane 3 = positive COX-2 cDNA from pig No. 1; lane 4 = positive COX-2 cDNA from pig No. 7; lane 5 = positive COX-2 cDNA from pig No. 8; lane 6 = positive COX-2 cDNA from pig No. 11; lane 7 = positive COX-2 cDNA from pig No. 14; lane 8 = positive COX-2 cDNA from pig No. 15.
Microscopic lesions
Histopathologic changes in the acute stage were characterized by coagulative necrosis of lung parenchyma, hemorrhage, vascular thrombosis, edema, fibrin deposition, and infiltration of lung parenchyma by neutrophils and alveolar macrophages in nine pigs (Nos. 1, 3, 7, 8, 11–15). In chronic lesions, a thick layer of granulation tissue surrounded foci of pulmonary necrosis in six pigs (Nos. 2, 4–6, 9, 10). There was no association between serotype and the characteristics of the histopathology.
In situ hybridization
The 15 pigs naturally infected with
In situ hybridization in 15 pigs naturally infected with
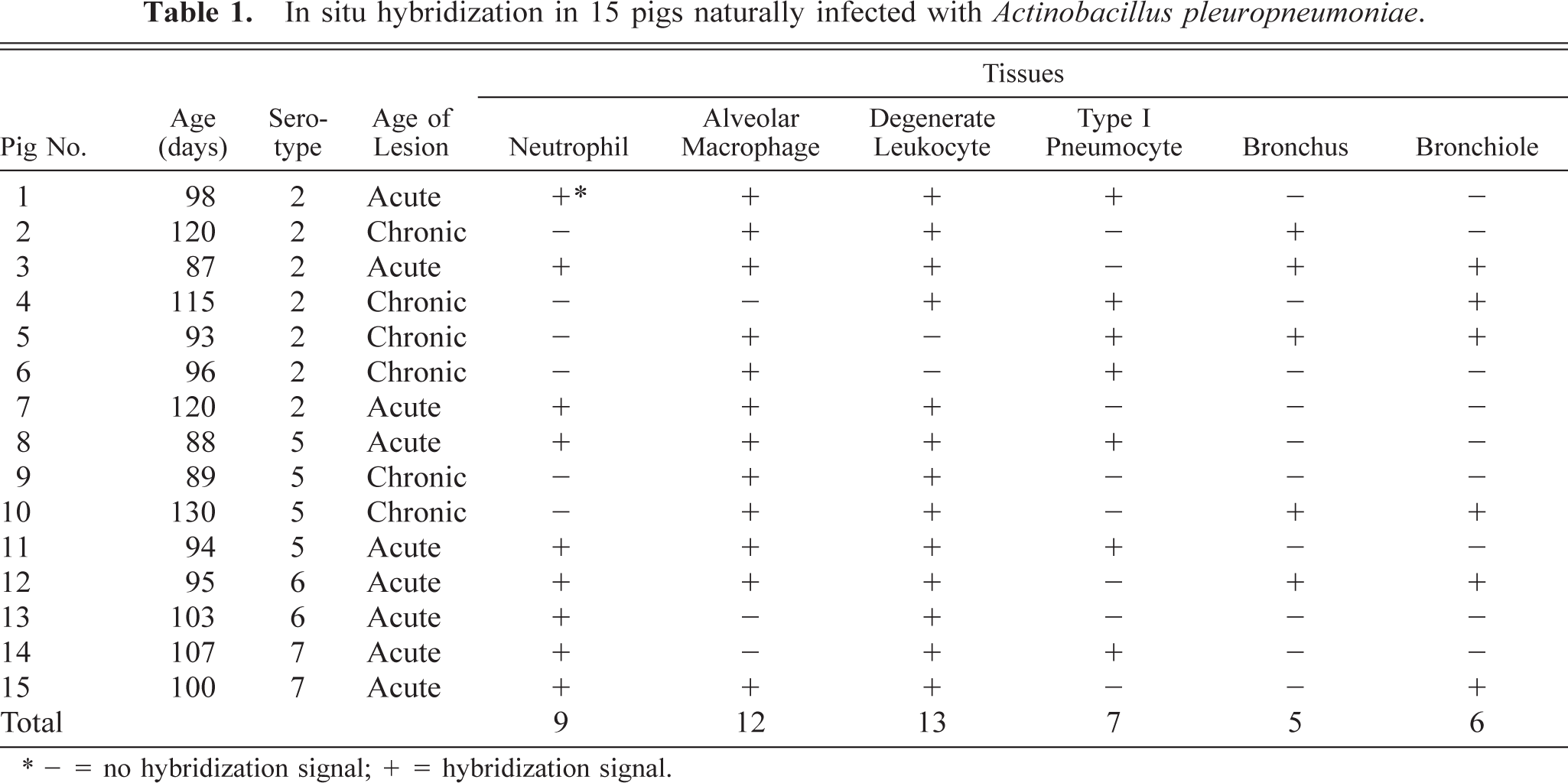
∗ - = no hybridization signal; + = hybridization signal
Bronchial, bronchiolar epithelial cells, alveolar macrophages, neutrophils, and type I pneumocytes had positive hybridization signals. A strong hybridization signal for COX-2 was seen in degenerate alveolar leukocytes (“oat cells”) adjacent to foci of coagulative necrosis, and in alveolar spaces (Fig. 2) and also in neutrophils (Fig. 3) and macrophages that had infiltrated alveolar spaces. Expression of COX-2 was consistently associated with lung lesions, but it was negligible in unaffected portions of lung from the
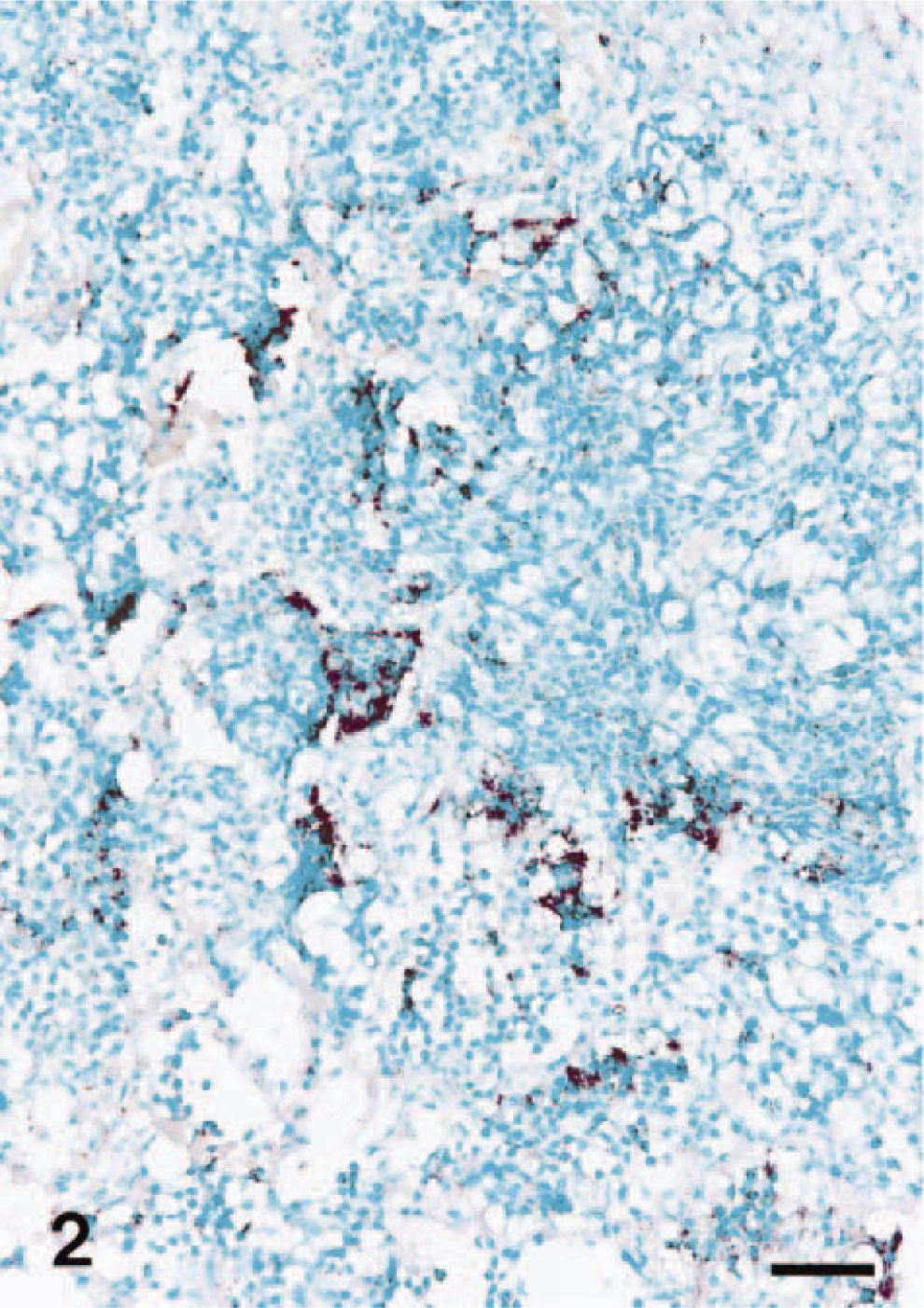
Lung; pig No. 1, naturally infected with
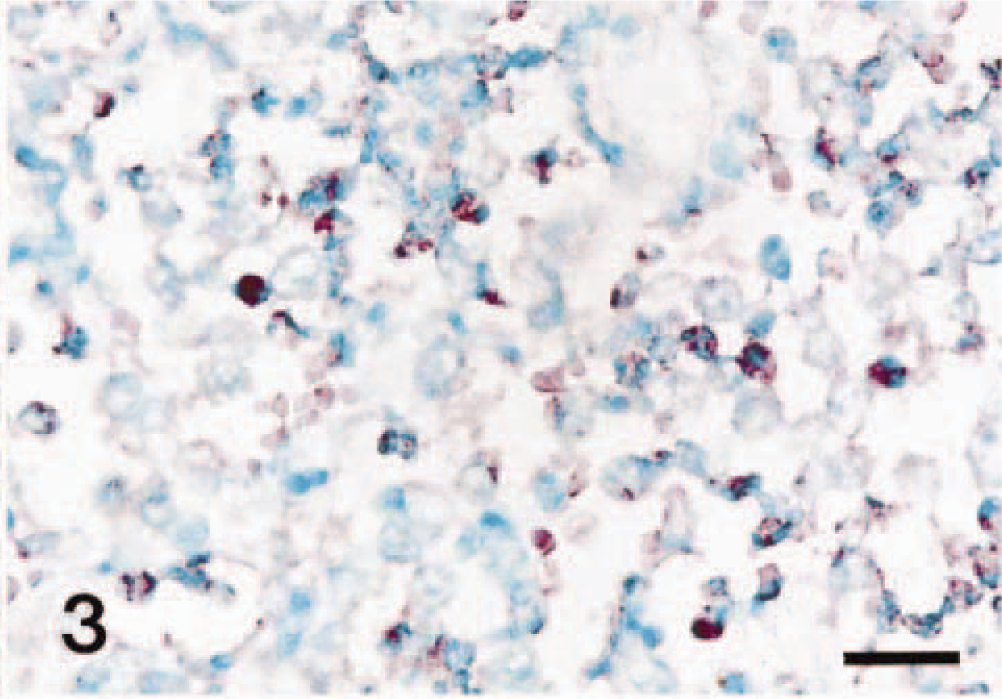
Lung; pig No. 3, naturally infected with
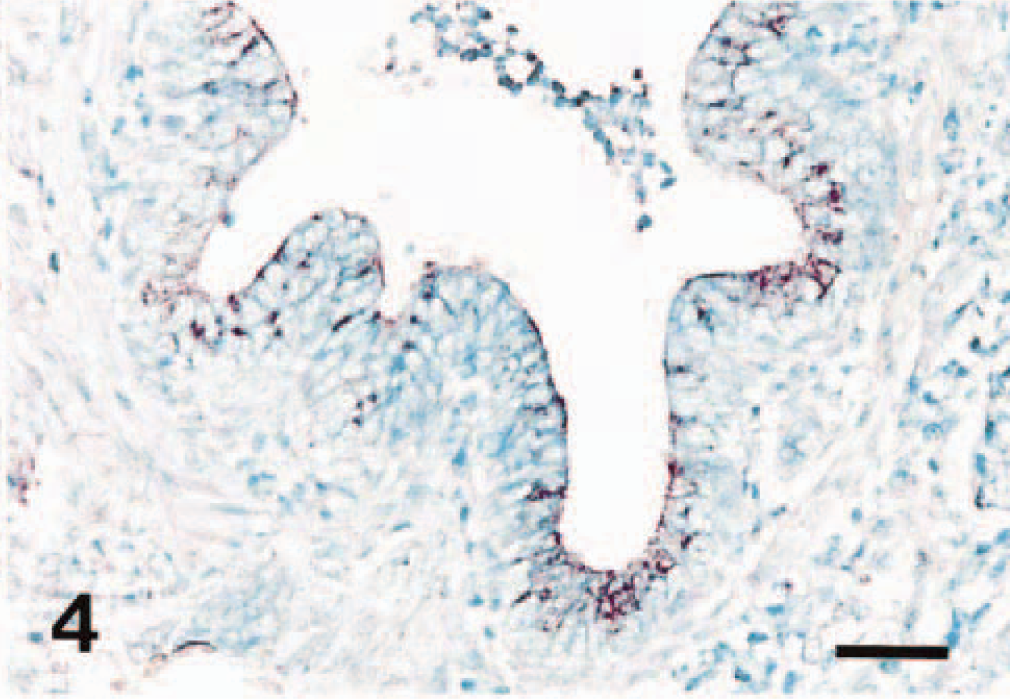
Lung; pig No. 10, naturally infected with
Pretreatment with RNase A eliminated hybridization signal from 15 pigs naturally infected with
Discussion
The major site of COX-2 expression in lungs of pigs naturally infected with
Traditionally, neutrophils have not been known to release products of the COX pathway of arachidonic acid metabolism. Although the COX isoforms have been described in a variety of cells and tissues, there have been few reports regarding the expression of these products in neutrophils. 26,29 Human neutrophils express COX-2 and release PGE2 after LPS stimulation. 14 Neutrophils that produce prostanoids such as PGE2 could modify the inflammatory process. PGE2 produced by stimulated neutrophils may be involved in the formation of edema at sites of acute inflammation. 18,24
COX-2 regulation may occur in a complex and cell-specific manner. COX-2 may be proinflammatory during the early stage of inflammation dominated by neutrophils, but it might also aid resolution in the later mononuclear cell–dominated stage by facilitating the synthesis of an alternative set of antiinflammatory prostaglandins.
16,17
Our results are in agreement with these previous reports in other species with COX-2 expression primarily in neutrophils of acute lesions and in alveolar macrophages of chronic lesions. It is well accepted that PGE2 formed by COX-2 contributes to lesions of acute inflammation by causing vasodilation, and increased vascular permeability.
10
In chronic infection where the macrophage is the dominant inflammatory cell, which contain COX-2 may exert an additional proinflammatory effect by contributing to angiogenesis.
15
Thus, a prostanoid product of COX-2 is an important component of the inflammatory response to acute and chronic
Two virulence factors of
The increased expression of COX-2 may be a direct response to
In this study, in situ hybridization with nonradioactive digoxigenin-labeled probes was successfully applied to
Footnotes
Acknowledgements
The research reported here was supported by a grant of the Korea Health 21 R&D Project (02-PJ1-PG3-20799-0001), the Ministry of Health and Welfare, Republic of Korea.
