Abstract
Nine juvenile ferrets (Mustela putorius furo) with a history of diarrhea were severely dehydrated and had distended abdomens and thin-walled small intestines that contained gas and fluid. Histologically, small intestines exhibited acute superficial atrophic enteritis. Transmission electron microscopy of the small intestine showed rotavirus-like particles within apical vacuoles. Reverse transcription polymerase chain reaction (RT-PCR) was negative for group A rotavirus. A group C rotavirus-specific RT-PCR assay was developed using consensus primers designed from the alignment of VP6 gene sequences of porcine, bovine, and human strains. A 182-bp product of the VP6 gene was sequenced and showed significant similarity to group C rotavirus VP6 sequences. This strain was designated “Ferret Rota C-MSU.” The entire coding sequence of VP6 was determined and compared with other rotaviruses. Ferret Rota C-MSU virus was found to be most closely related to Shintoku group C rotavirus. This is the first definitive identification of a group C rotavirus in ferrets, based upon RT-PCR, sequencing, and genetic analysis.
Rotaviruses are classified in the family Reoviridae. They are 55–70-nm icosahedral particles that are nonenveloped with a triple-layered viral capsid made up of outer, intermediate, and inner layers. Their genome consists of 11 segments of dsRNA, which encode 6 structural and 5 nonstructural proteins. 4,9,10 Seven major serogroups have been identified and are designated A–G based upon antigenic properties and RNA migration patterns in polyacrylamide gels. 9,10,14,19,25 Group A rotaviruses are “typical” rotaviruses and share a common antigen within the inner virion capsid layer. 14,22,33 All other groups (B–G) are designated as “atypical” rotaviruses and do not contain the common antigen present in typical rotaviruses. 20,22
We report the first definitive identification of group C rotavirus in ferrets, based upon reverse transcription polymerase chain reaction (RT-PCR), sequencing, and genetic analysis. This strain was designated “Ferret Rota C-MSU.”
Nine ferrets (Mustela putorius furo) were presented dead for necropsy. All 9 ferrets had been euthanatized and shipped on ice over night to the Diagnostic Center for Population and Animal Health in Lansing, Michigan. These ferrets were approximately 1 week old and of undetermined sex and had a history of diarrhea. Gross necropsy examination of each ferret was performed, and sections of all major organs were examined microscopically. For ultrastructural examination, sections of formalin-fixed small intestine were dehydrated in graded alcohols and embedded in Epon resin by routine methods for transmission electron microscopy. Ultrathin sections were cut and stained with uranyl acetate and lead citrate and examined using an electron microscope.
Sterile sections of small intestine from each ferret were pooled and submitted for routine bacterial culture, including aerobic culture on blood agar, and enrichment followed by aerobic culture on special media for Enterobacteriaceae, including Salmonella spp.
Sterile samples of small intestine were collected individually from each ferret for virology testing. RNA from small intestinal sections from each of the 9 ferrets was extracted using the RNeasy Mini Kit (QIAGEN Inc., Valencia, CA) according to the manufacturer's guidelines. An RT-PCR assay targeting a 294-bp fragment of the VP6 gene of mammalian group A rotavirus was performed on sections of small intestine from each ferret as previously described. 13
A group C rotavirus–specific RT-PCR assay was developed using primers designed from the consensus sequence of aligned VP6 gene sequences (including 5′ and 3′ untranslated region sequences) of porcine, bovine, and human group C rotavirus strains reported in GenBank. The accession numbers for these sequences are D00326, AY86571, and M88768, respectively. The selected primers, Rota C fwd (5′-TTCCATACTGGTGGCATTGG-3′) and Rota C rev (5′-CTTGAAGCTTCTCTAACCATTTC-3′) (nucleotide positions 157–176 and 338–316, respectively, in bovine group C rotavirus VP6 gene, GenBank accession no. M88768), target a 182-bp region of the gene. RT-PCR was performed with the Superscript One-Step RT-PCR System (Life Technologies, Rockville, MD). Five microliters of extracted RNA were added to the reaction mix containing the 2× amplification buffer, primers (at 0.4 µM final concentration), and water, giving a volume of 49 µl. This mixture was heated at 95°C for 4 minutes then quickly cooled to 4°C before the addition of 1 µl of the RT-Taq enzyme mix. RT was at 48°C for 30 minutes, followed by 40 PCR cycles of 94°C for 30 seconds, 53°C for 30 seconds, and 72°C for 45 seconds. A final extension was performed at 72°C for 7 minutes. Products were analyzed by agarose gel electrophoresis and ultraviolet transillumination.
Primers were designed from the VP6 consensus sequence derived previously from the alignment of porcine, bovine, and human rotavirus sequences. RT-PCR was performed using the One-Step RT-PCR Kit (QIAGEN) with 7 µl of sample RNA and primers at a final concentration of 0.6 µM each in a 50-µl reaction volume. Again, template RNA was denatured in the reaction mixture before the addition of the RT-Taq enzyme mix. Primers VP6 5′ end (5′-GGCTTTAAAAATCTCATTCAC-3′) (nucleotide positions 1–20 in bovine group C rotavirus VP6 gene, GenBank accession no. M88768) and Rota C rev were used to amplify the 5′ end of the gene (∼300-bp product). Primers Rota C fwd and VP6 3′ end (5′-CACATAGTTCACATTTCATCCT-3′) (nucleotide positions 1,349–1,328 in bovine group C rotavirus VP6 gene, GenBank accession no. M88768) were used to amplify the approximately 1.1-kb 3′ end of the gene. RT was carried out at 45°C for 30 minutes followed by a predenaturation step of 95°C for 15 minutes for both sets of primers. PCR cycling conditions for the VP6 5′ end amplification consisted of 45 cycles of 94°C for 30 seconds, 51°C for 1 minute, and 72°C for 1 minute. For the VP6 3′ end amplification, conditions were 45 cycles of 94°C for 30 seconds, 52°C for 1 minute, and 72°C for 2 minutes. A final extension of 72°C for 7 minutes was employed for both PCR assays.
Amplicons of the expected size were cut from the gel and purified using the QIAquick Gel Extraction Kit (QIAGEN). PCR products were sent, with both forward and reverse primers, to the Research Technology Support Facility of Michigan State University for automated DNA sequencing. The facility uses Applied Biosystems (Foster City, CA) cycle sequencing technology, and sequences are determined on an ABI Prism 3100 Genetic Analyzer. Basic local alignment search tool analysis of sequence data was performed to search for significant similarities to sequences in the GenBank database. 2 Sequence assembly and analyses, including multiple alignments of nucleotide and predicted amino acid sequences using the Clustal W method and construction of phylogeny trees, were done with Lasergene biocomputing software (DNASTAR, Inc., Madison, WI). 26
All 9 ferrets were in a good state of preservation with minimal autolysis. Gross examination findings were similar for all and included dehydration, distended abdomens (Fig. 1), and thin-walled small intestines that were distended with gas and fluid (Fig. 2). There were no other significant gross abnormalities. Microscopically, the small intestines from each ferret exhibited acute superficial atrophic enteritis characterized by degeneration and necrosis of epithelial cells at the villous tips with loss or sloughing of affected epithelial cells into the lumen (Fig. 3). These histologic findings were similar to lesions of rotaviral enteritis in other species. Calcified casts were present within renal tubules, consistent with dehydration. All other organs appeared microscopically unremarkable. Transmission electron microscopy of sections of small intestine showed viral particles within apical vacuoles (Fig. 4). These viral particles had a viral capsid and were between 55 and 70 nm in diameter, consistent with rotavirus-like particles.
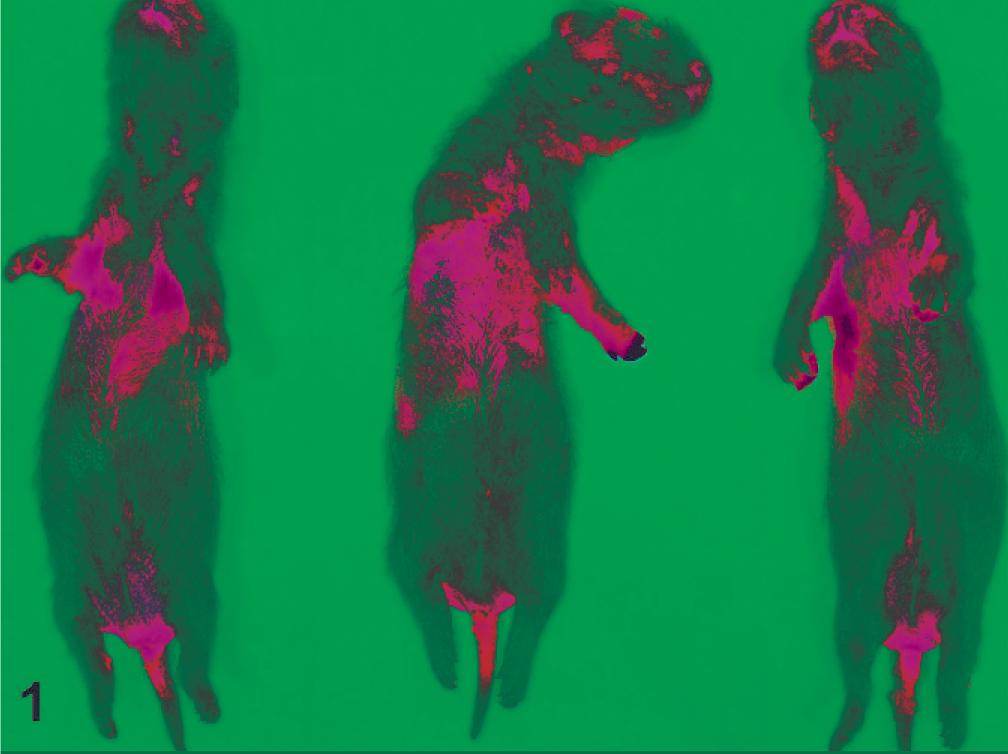

Ferret, kits. The bodies are thin and dehydrated with distended abdomens.
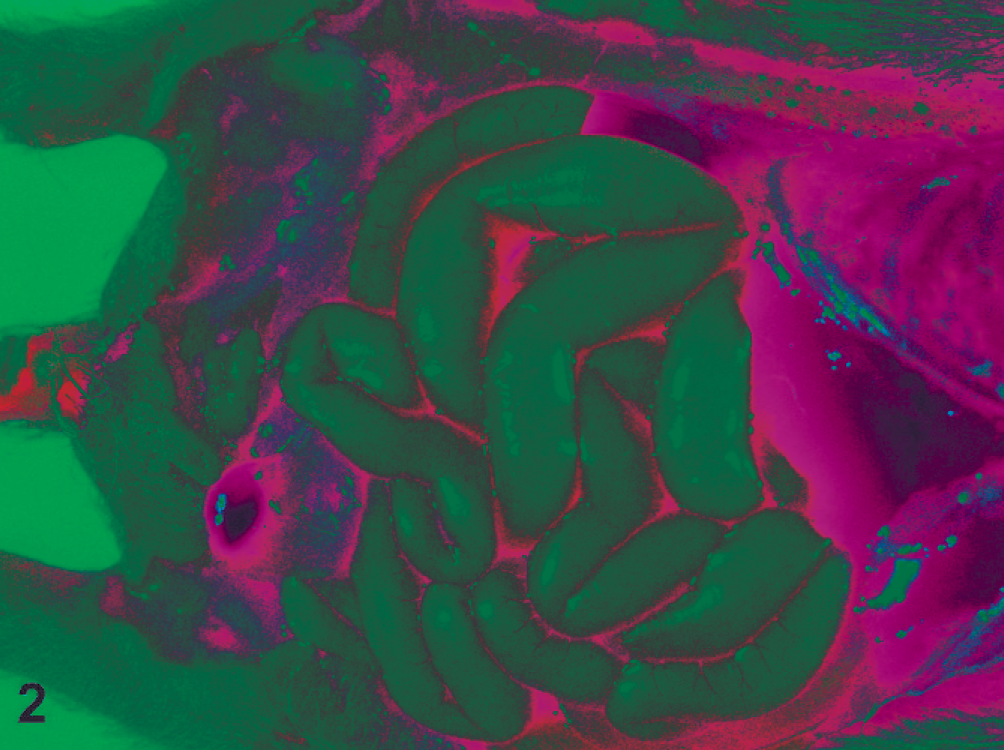
Ferret, intestines. The small intestines are thin walled and distended with gas and fluid.
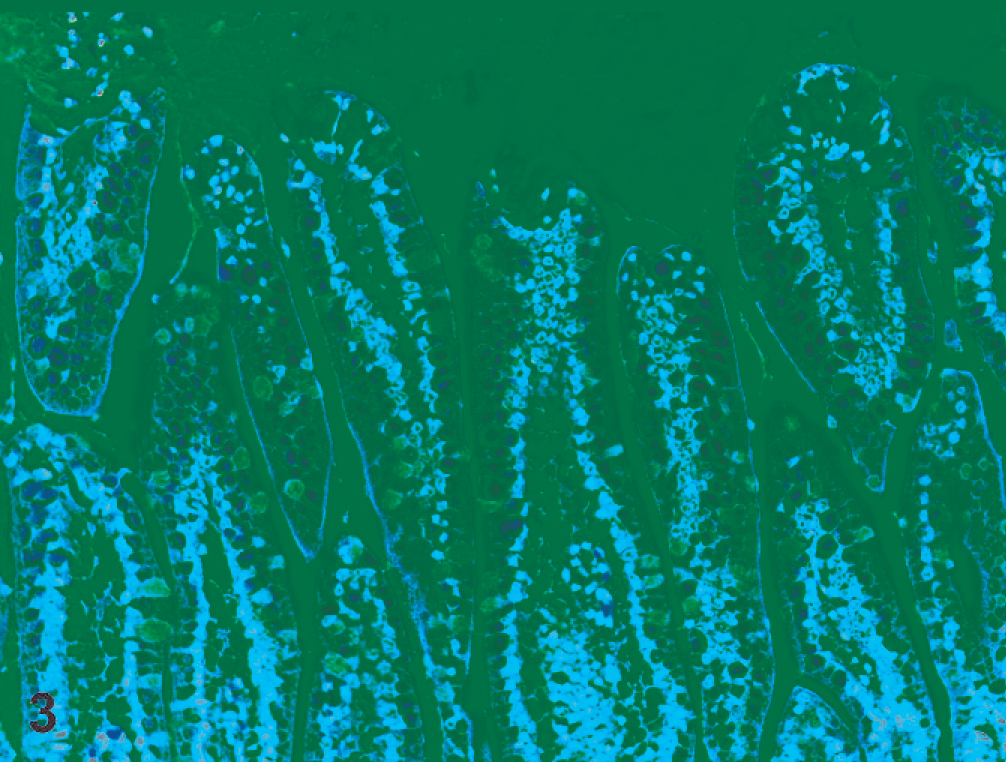

Ferret, small intestine. Microscopically, there was degeneration and necrosis of epithelial cells at the villous tips with loss or sloughing of affected epithelial cells into the lumen. HE.
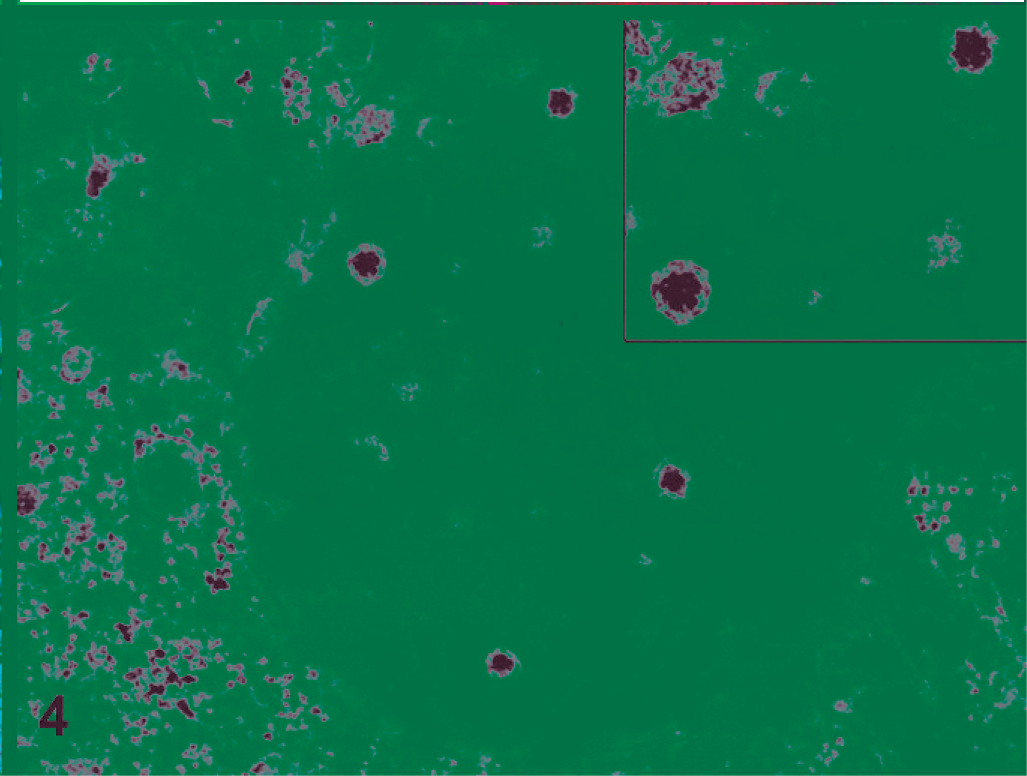

Ferret, small intestine epithelial cell. Viral particles are within apical vacuoles, have a viral capsid, and are between 55 and 70 nm in diameter. Transmission electron microscopy.
Bacterial culture of the pooled sections of small intestine from each of the 9 ferrets was negative for Salmonella spp. and other pathogenic bacteria.
RT-PCR tests for bovine group A rotavirus, using sections of small intestine from the ferrets, were negative. The group C rotavirus–specific RT-PCR assay that was developed used primers that target a 182-bp region of the VP6 gene. With this assay, RNA extracted from small intestine samples from each ferret was positive for the 182-bp amplicon (Fig. 5). The product was directly sequenced, and it showed significant similarities to group C rotavirus VP6 sequences in GenBank by basic local alignment search tool analysis. This strain was designated “Ferret Rota C-MSU.”
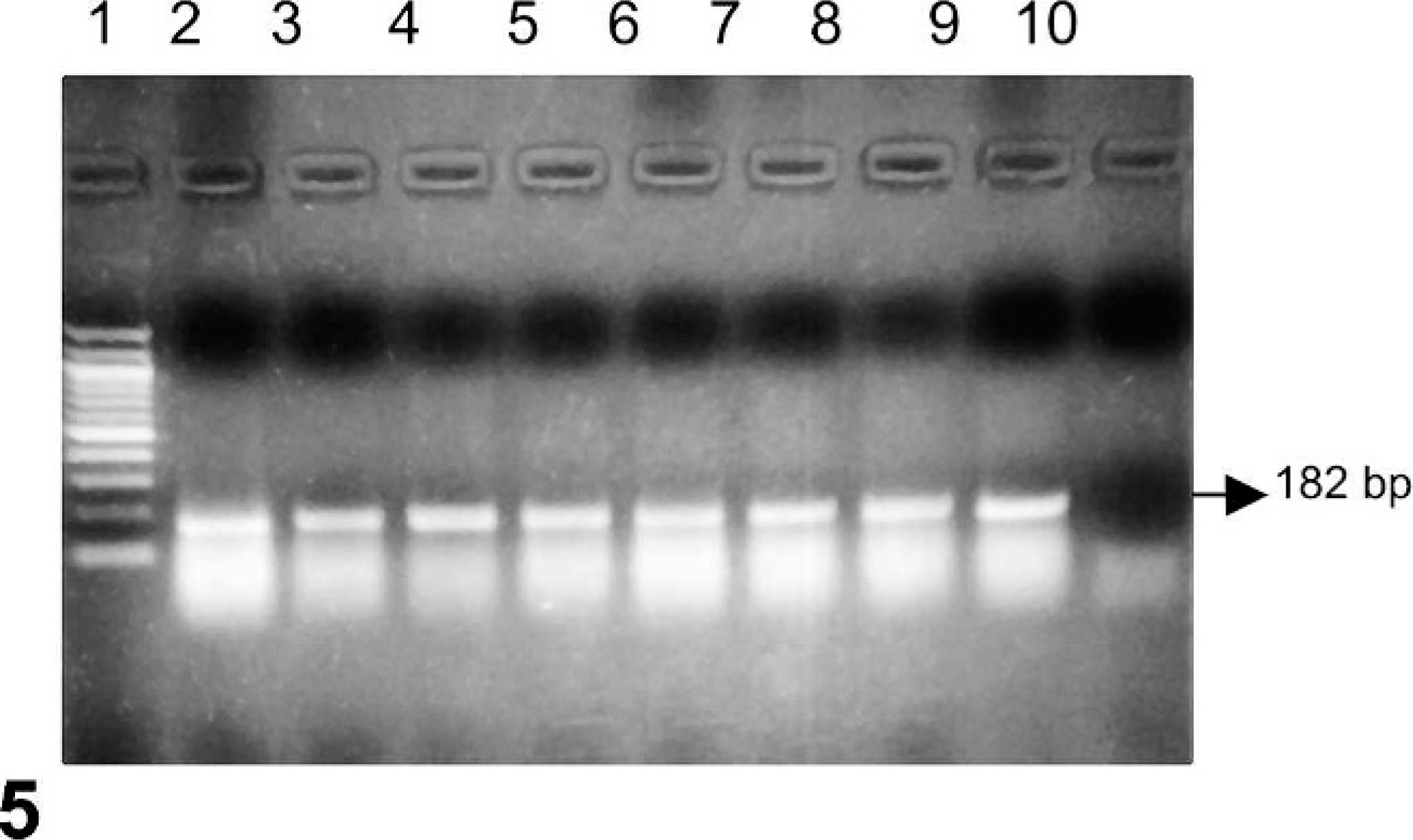
Reverse transcription polymerase chain reaction assay for the detection of group C rotavirus. Expected amplicon size is 182 bp. Lane 1, 100-bp DNA ladder; lanes 2–9, RNA from 8 ferret small intestinal samples; lane 10, no template control.
The entire coding sequence of VP6 was sequenced, and this VP6 gene was found to have 1 open reading frame of 1,185 nucleotides, which translates into a 395–amino acid protein with a predicted molecular weight of 44.5 kDa. Basic local alignment search tool (BLAST analysis), sequence alignment, and phylogenetic analyses were performed. The VP6 sequence of Ferret Rota C-MSU was compared with corresponding sequences of known group C and A rotaviruses and was found to be most closely related (81.1% nucleotide similarity and 92.2% amino acid similarity) to the Shintoku (bovine strain) group C rotavirus (Table 1). Ferret Rota C-MSU was also compared with Cowden (porcine strain), Jajeri (human strain), and Preston (human strain) group C rotaviruses, as well as with avian rotavirus A and bovine rotavirus A. The ferret rotavirus appears to be more similar to the porcine strain than to the 2 human-derived strains. The group A rotaviruses showed less than 45% sequence identities to the ferret strain and the other group C viruses. Nucleotide and amino acid similarities between these rotavirus strains are shown in Table 1. Multiple sequence alignment for the deduced amino acid sequence of the VP6 protein of Ferret Rota C-MSU and other rotavirus strains is presented in Fig. 6. Ferret Rota C-MSU and the rotavirus C strains showed a high degree of amino acid sequence conservation within their VP6 proteins, including a conserved deletion of an arginine residue at position 276, in contrast to the 2 rotavirus A strains. The phylogenetic tree corresponding to the VP6 protein alignment is shown in Fig. 7. The ferret sequence was observed to form a tight cluster with the 2 animal-derived group C rotavirus strains, Shintoku and Cowden.
Percent amino acid and nucleotide sequence identities between rotavirus group C and A VP6 proteins/genes.
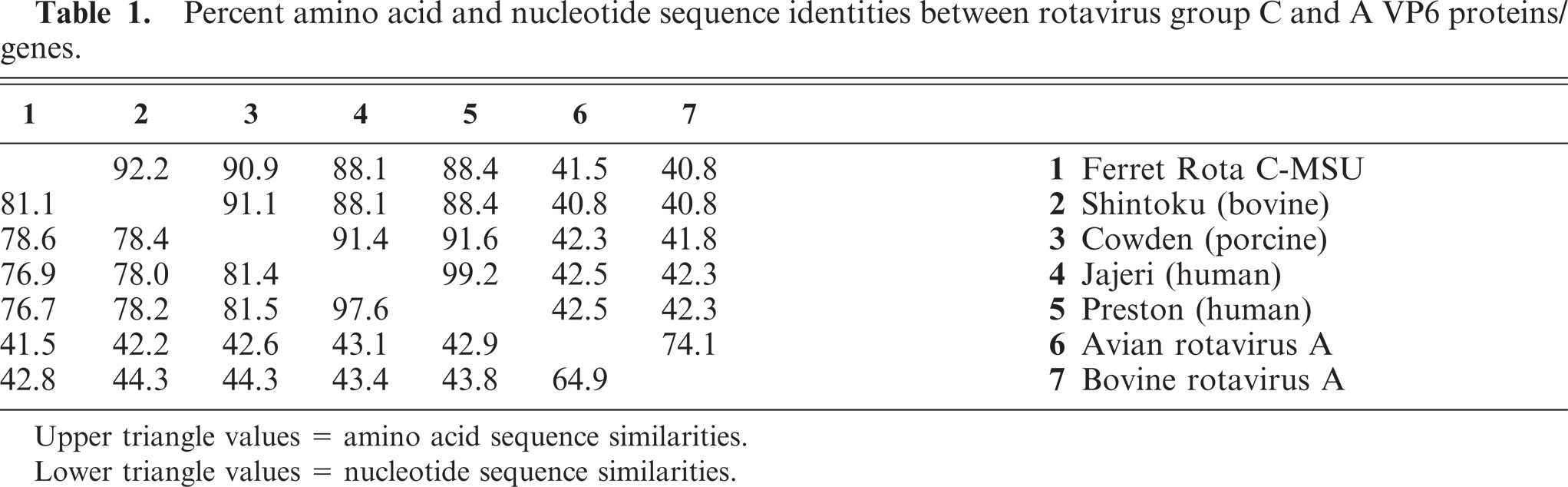
Upper triangle values = amino acid sequence similarities.
Lower triangle values = nucleotide sequence similarities.
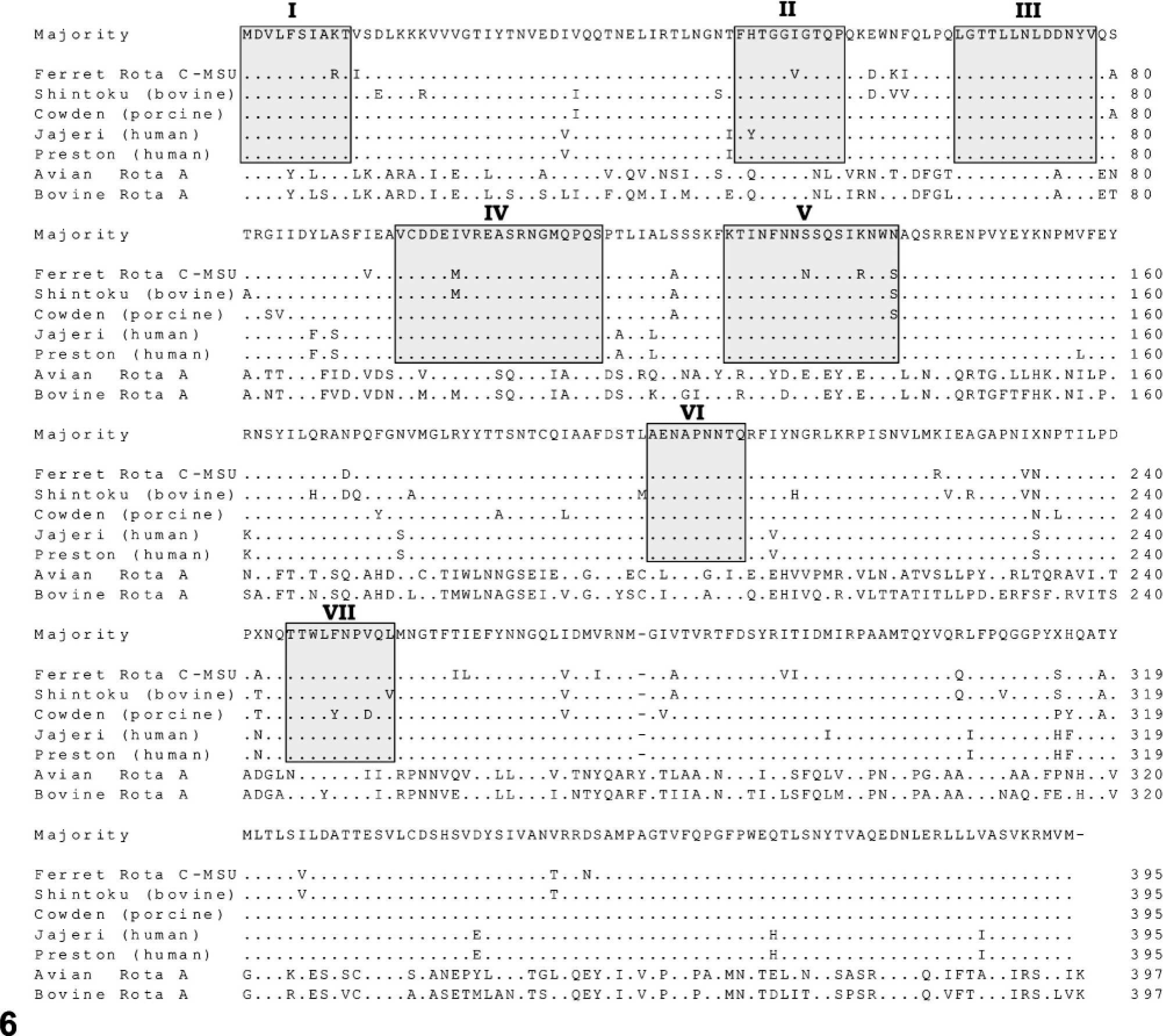
Alignment of the deduced amino acid sequences of the VP6 proteins of Ferret Rota C-MSU (this study); rotavirus group C strains, Shintoku (bovine) (M88768), Cowden (porcine) (M94157), Jajeri (human) (AF325805), and Preston (human) (M94156); and rotavirus group A strains, avian rotavirus A (D16329) and bovine rotavirus A (K02254). Residues that match the majority (consensus) sequence are shown in dots. A deleted residue is indicated by a minus sign. The 7 conserved regions, I–VII, are boxed. GenBank accession numbers are in parentheses.
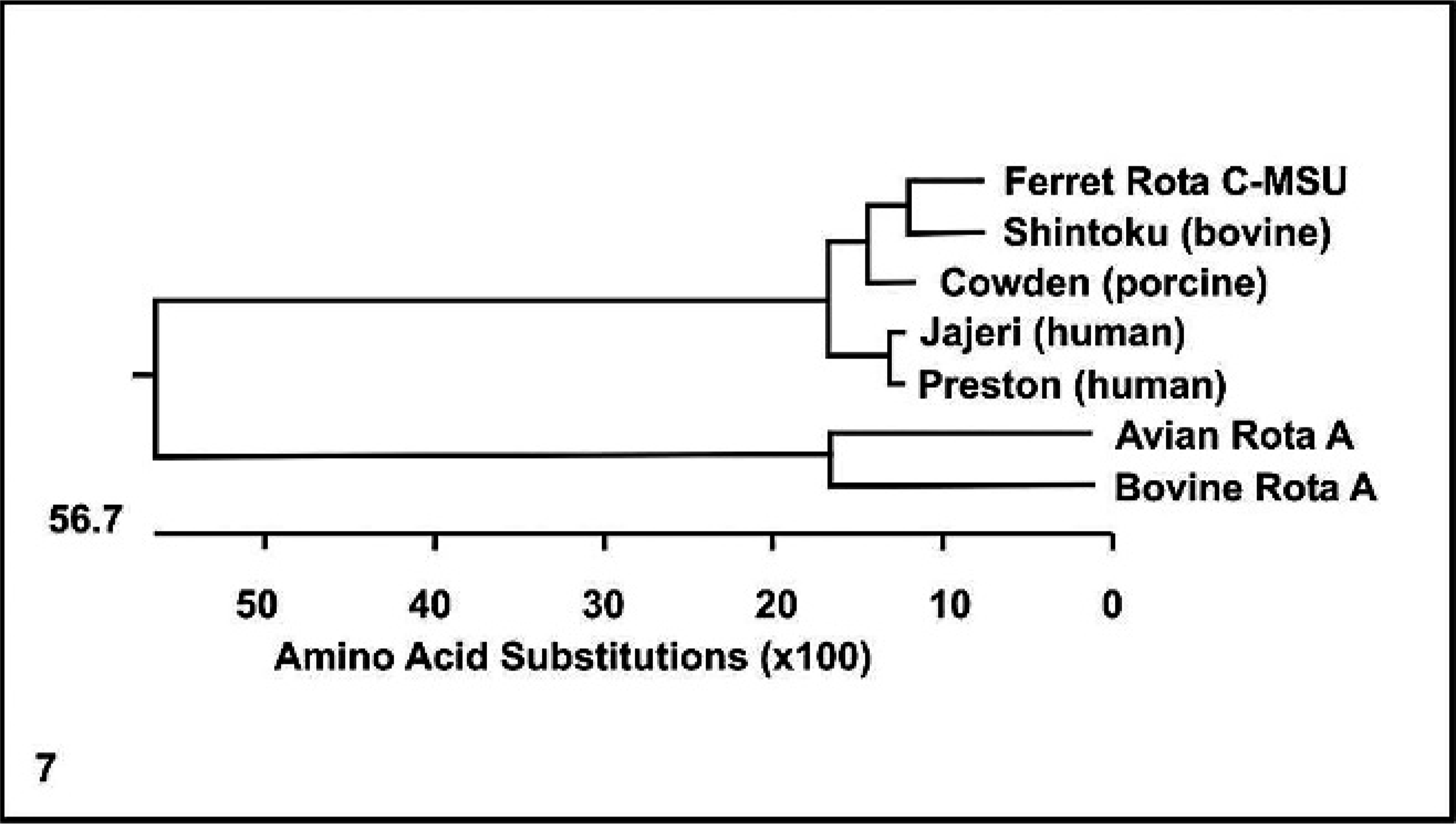
Phylogenetic tree based upon deduced amino acid sequences of the VP6 proteins of Ferret Rota C-MSU (this study); rotavirus group C strains, Shintoku (bovine) (M88768), Cowden (porcine) (M94157), Jajeri (human) (AF325805), and Preston (human) (M94156); and rotavirus group A strains, avian rotavirus A (D16329) and bovine rotavirus A (K02254). GenBank accession numbers are in parentheses.
This paper describes the first definitive identification of group C rotavirus in ferrets, based upon RT-PCR, sequencing, and genetic analysis. Rotaviruses have been reported to cause diarrhea worldwide in humans and a wide range of animal species, including calves, piglets, rats, mice, foals, sheep, goats, rabbits, monkeys, poultry, dogs, and cats. 10,14–16,18,19,22,33,34 Young animals and children are most commonly affected; infections in older animals and adults can occur but are usually clinically mild. 12,14,34 The first isolate of group C rotavirus from any species was found in nursing piglets in 1980. 23 Since then, group C rotaviruses have been reported in cattle, humans, and dogs. 3,5,19,28,29 The gross and microscopic lesions caused by enteric rotaviruses are very similar across species. Dilated, thin-walled intestines filled with gas and clear to yellow fluid are typical. Enterocytes near the distal tips of small intestinal villi are most commonly affected. 22,24 Destruction of these enterocytes leads to a malabsorptive diarrhea. 24
There are relatively few known causes of enteritis in ferrets. Previously reported viral causes of enteritis in ferrets include ferret enteric coronavirus (FECV), anecdotal reports of group A rotavirus, and a single report of an atypical rotavirus isolated from neonatal ferrets. 27,31,32 FECV has been associated with epizootic catarrhal enteritis in ferrets. 32 Clinical signs associated with epizootic catarrhal enteritis include lethargy, inappetence, vomiting, and distinct green mucoid diarrhea. 31 Epizootic catarrhal enteritis is usually most severe in older ferrets, whereas young ferrets tend to exhibit only mild or subclinical disease. 31 This is in contrast to Ferret Rota C-MSU, which affected young kits. Typically, high morbidity and low mortality are seen with FECV. 31,32 There have been few anecdotal reports of group A rotavirus causing diarrhea in juvenile ferrets. However, an extensive literature review did not identify any confirmed published cases. There is a single report of isolation of an atypical rotavirus causing diarrhea in neonatal ferrets, which most likely represents a group C rotavirus based upon the lack of the rotavirus type A common antigen and an electrophoretic dsRNA migration pattern of 4-3-2-2 that is characteristic of atypical rotaviruses. 27 However, this suspicion was not confirmed by sequencing. 27
PCR and sequence analysis are important diagnostic tools for identifying the cause of viral enteritis in ferrets. The microscopic lesions seen with ferret coronavirus and rotavirus enteritis are identical and are consistent with an atrophic enteritis. The age of affected ferrets, clinical history, and character of the diarrhea (FECV usually causes green mucoid diarrhea) can help to differentiate between FECV and rotavirus infection. However, additional diagnostic testing is needed to identify the group and strain of rotavirus present in cases suggestive of viral enteritis. Standard ELISAs and immunofluorescence tests only identify group A rotaviruses, and standard negative staining methods used for electron microscopy cannot differentiate between different groups or strains of rotaviruses. 9,22 PCR and sequence analysis are necessary to confirm rotavirus infection and to identify which group of rotavirus is present.
In this report, a group C rotavirus–specific RT-PCR assay was developed using primers that target a 182 bp-region of the VP6 gene. VP6 is a major structural inner capsid protein located on the outer surface of single-shelled particles. 9,11,24 It is encoded by genome segment 6 in group A rotaviruses and by gene 5 in group C rotaviruses. 7,9,11,21 This protein is commonly targeted in diagnostic assays used to detect virus particles, and it is highly immunogenic and antigenic. 9 RNA extracted from the small intestine from each of the 9 affected ferrets was positive for this 182-bp amplicon of the VP6 gene. Based upon sequencing of the VP6 gene, this virus, designated Ferret Rota C-MSU, was found to be most closely related to the Shintoku (bovine strain) group C rotavirus. The VP6 gene was found to have 1 open reading frame of 1,185 nucleotides, which translates into a 395–amino acid protein with a predicted molecular weight of 44.5 kDa. As previously reported, gene 5 of group C rotaviruses Shintoku, human Bristol and 88-220 strains, and a porcine Cowden strain also have 1 long open reading frame that is 1,185 nucleotides (395 amino acids). 7,11 Similarly, the VP6 proteins of the two group C human strains, Jajeri and Preston, included in the alignment in Figure 6, each had 395 amino acid residues with a predicted molecular weight of 44.7 kDa. The predicted molecular mass of the Shintoku strain has been reported as 44.5 kDa, which is the same as Ferret Rota C-MSU. Cowden, Bristol, and 88-220 strains have been reported to have a molecular weight of 44.7 kDa. 7,11 In one study, VP6 sequences of Shintoku, Cowden, Bristol, and 88-220 strains were compared, and 7 highly conserved regions (I–VII) were identified. 11 These 7 conserved regions were also observed in the sequences of Ferret Rota C-MSU and the 2 human group C strains, Jajeri and Preston, in the amino acid alignment in Fig. 6. Among the group C rotavirus strains in this figure, there were 21 positions with amino acid substitutions identical between either the 3 animal strains or the 2 human strains. Based on similar observations when comparing Shintoku and Cowden animal strains with the 2 human strains, Bristol and 88-220, Jiang et al. 11 suggested that animal- or human-specific sequences may exist.
The group C rotavirus identified in this report appears to be highly prevalent, and has been detected in additional pooled fecal samples from ferrets with diarrhea. Transmission of rotaviruses occurs via contact with infected individuals or a contaminated environment. Infected individuals shed large amounts of the virus in their feces. In general, most rotaviruses are very stable within feces, and common disinfectants often do not kill the virus, thus making it difficult to eliminate it from a housing facility once animals become infected. 14 Group C rotaviruses, however, are not shed in high amounts and are unstable in feces. 5 Colostral antibodies can protect neonates from rotaviral diarrhea but not from rotavirus infection. 14 Most rotaviruses are described as being species specific. However, experimental cross-species transmission of naturally occurring rotaviruses (both group A and some nongroup A strains) has been reported. 8,22,24 There have also been reports of naturally occurring rotavirus reassortants that may be able to cross species and cause disease. 1,6,8,17,30 There has been no evidence of cross-species transmission from ferrets to other species or vice versa thus far. However, the possibility of such transmission must be considered.
Footnotes
Acknowledgements
We thank members of the virology section at the Diagnostic Center for Population and Animal Health for technical assistance.
