Abstract
Clostridia dominate the rodent intestinal bacterial community and play an important role in physiological functions of the host. However, their ecology and diversity are still unclear. In our previous report, we showed that phylogenetically novel groups of Clostridia inhabit the mouse intestine and contribute to the normalization of germfree mice. In this study, five new oligonucleotide probes were designed and applied to detect these clostridial groups that are essential for the normalization of germfree mice. Faecal microbiota of conventional mouse strains and specific pathogen-free mice from different breeding colonies were analysed by fluorescence in situ hybridization using these five probes. Our results showed that the composition of Clostridia differed among mouse strains and also among mouse groups of the same inbred strain from different breeding colonies. These five new probes for mouse Clostridia were able to detect the difference in clostridial diversity in each mouse group. In addition to Clostridium, we also analysed Bacteroides and Lactobacillus using previously described probes and the number or the frequency of occurrence of Bacteroides was shown to be different among mouse groups analysed. The oligonucleotide probe set including our newly developed and previously described probes used in this study can be applied to monitoring of significant groups of mouse intestinal microbiota.
Some genetically-modified animals have been reported to show no or fewer lesions under germfree (GF) and specific pathogen-free (SPF) conditions. 1 It has also been reported that the same inbred strains raised in different SPF colonies possess different compositions of intestinal microbiota resulting in different metabolic responses to oligosaccharides and different development of lymphocyte populations in the intestinal and systemic organs of the mice. 2 These findings suggest that intestinal microbiota is strongly related to and determines host responses. Therefore, exact information on intestinal microbiota is important when we use experimental animals in various studies. It is also necessary to monitor the composition of intestinal microbiota of SPF mice, and focus not only on pathogenic bacteria.
Clostridia, which are defined as spore-forming Gram-positive anaerobes, form a dominant population in the intestinal microbiota of mice. Most of them are extremely oxygen sensitive and have been known as fusiform-shaped bacteria based on their characteristic morphology for 40 years.3,4 Studies have shown that antibiotic administration to mice results in disappearance of fusiform-shaped bacteria and enlargement of the caecum5,6 and a lowering of colonization resistance to pathogens. 7 Their numbers are decreased markedly by dietary and environmental stress. 8 They also play a significant role in the regulation of the ratio of CD4-CD8+/CD4+CD8- cell numbers in αβ T-cell receptor-bearing intraepithelial lymphocytes in mouse large intestine. 9 These findings indicate that mouse Clostridia play an important role in physiological functions of the host. However, the ecology and diversity of mouse Clostridia are still unclear.
In earlier studies, Itoh and Mitsuoka prepared a mixture of clostridial spores obtained by chloroform treatment of a faecal suspension of conventional (CV) CF#1 mice and produced ex-GF mice that were administered the mixture. They reported that the caecum was reduced to normal size in these ex-GF mice. 10 GF mice should be associated with ‘normal’ microbiota for the expression of ‘normal’ physiological functions. This is called ‘normalization’ of GF mice and is indexed especially by caecal size. In our previous study, we selected Clostridia by chloroform treatment of mouse faeces and obtained Culture strains that can be cultured only in an anaerobic chamber and Culture Jar strains that can grow in both an anaerobic chamber and an anaerobic jar. DNA was extracted from these strains. DNA was also extracted directly from the chloroform-treated faecal suspension, and cloned 16S rRNA genes were obtained. The 16S rRNA gene sequences of the clostridial isolates and the cloned sequences were analysed and they were not found to be identical to known identified Clostridia. 11 The majority of them were members of Clostridium clusters XlVa and IV. Others belonged to the Acinetobacter cluster. Mouse Clostridia formed novel groups within these three clusters and it was revealed that these novel clostridial groups play an essential role in the normalization of GF mice. 11
The genus Clostridium belongs to the phylum Firmicutes and Clostridium is the most phylogenetically diverse group.12,13 Oligonucleotide probes have been designed to quantify the predominant clostridial clusters XlVa and IV in the human gut ecosystem by fluorescence in situ hybridization (FISH)14–16 and one of these probes (Erec482) has been applied to the analysis of mouse intestinal bacteria. 17 However, this Erec482 probe could only detect the diversified cluster XlVa as a whole group. It is important to focus on the significant (sub) groups to understand the functions of mouse Clostridia in the intestinal ecosystem that contribute to the normalization of GF mice. The aim of this study was to design oligonucleotide probes for rapid detection of the mouse clostridial groups, and to apply them to investigate the ecology in the mouse intestine by FISH.
Materials and Methods
Animals
Three strains of CV male mice at 12–13 weeks of age were used in this study. CF-1 and BALB/cA strains were originally obtained from The Institute of Medical Science, The University of Tokyo and CLEA Japan, respectively, and maintained in our laboratory as CF#1 and BALB/cA strains since 23 years. Interleukin-1 receptor antagonist knockout (IL-1Ra KO) 18 mice were provided from The Institute of Medical Science, The University of Tokyo and maintained under conventional conditions in our laboratory for three years. They were given a pelletted commercial diet (CMF, Oriental Yeast, Tokyo, Japan) and water ad libitum. They were housed in a room on a 12:12 light cycle at 24 ± 1°C with a relative humidity of 55 ± 5%. SPF BALB/c mice (male, 11–13 weeks old) were purchased from three commercial laboratory animal breeders in Japan: Japan SLC (Shizuoka), CLEA Japan (Tokyo) and Charles River Japan (Kanagawa) and faecal samples were collected on receipt to examine the composition of microbiota maintained in each breeder. All experiments were ethically reviewed and performed in accordance with the guidelines for the care and use of laboratory animals of The University of Tokyo.
Design of new oligonucleotide probes and dot blot hybridization
The sequences of mouse Clostridia obtained in our previous study 11 and their close relatives were aligned and group-specific probes were designed using the ARB software package. 19 The 16S rRNA gene sequence data of known bacteria were obtained from the DDBJ/EMBL/GenBank database. The XIVa401 probe was designed to detect mouse clostridial groups clustering in Clostridium cluster XlVa and the XIVa217 probe was designed for one of the groups within cluster XlVa that consists of cloned sequences (Figure 1). The IV1011 and IV481 probes were designed to detect each of the two mouse clostridial groups within Clostridium cluster IV The Aci211 probe was designed to detect one clostridial group within the Acinetobacter cluster. The specificity of the probes was tested by dot blot hybridization. For dot blotting, the 16S rRNA genes were amplified by polymerase chain reaction (PCR) using 16S rRNA gene specific primer pairs 8F (5′-GAGTTTGATCMTGGCTCAG-3’) and 15R (5′-AGGAGGTGATCCARCCGCA-3’). 20 Bacterial strains tested are listed in Table 2. The amplification program consisted of one cycle of 94°C for 3 min; 35 cycles of 94°C for 30 s, 55°C for 30 s and 72°C for 2 min; and finally one cycle of 72°C for 2 min. The PCR products were purified with Microspin S-400 HR Columns (GE Healthcare Bio-Sciences, Buckinghamshire, UK) and denatured in 0.4 mol/L NaOH and 10 mmol/L EDTA at 100°C for 10 min. They were applied to Hybond-N+ nylon membranes (GE Healthcare Bio-Sciences) and were fixed with ultraviolet light for one minute. Synthetic oligonucleotides were obtained from Greiner Bio-One (Tokyo, Japan) and labelled using a Dig Oligonucleotide 3′-End Labeling Kit (Roche, Mannheim, Germany) to probe the dot blots. Dot blot hybridization was carried out as described previously. 21 Hybridization and the washing process were performed at 50°C. Hybrids were detected by a Dig Nucleic Acid Detection kit (Roche) according to the manufacturer's instructions.
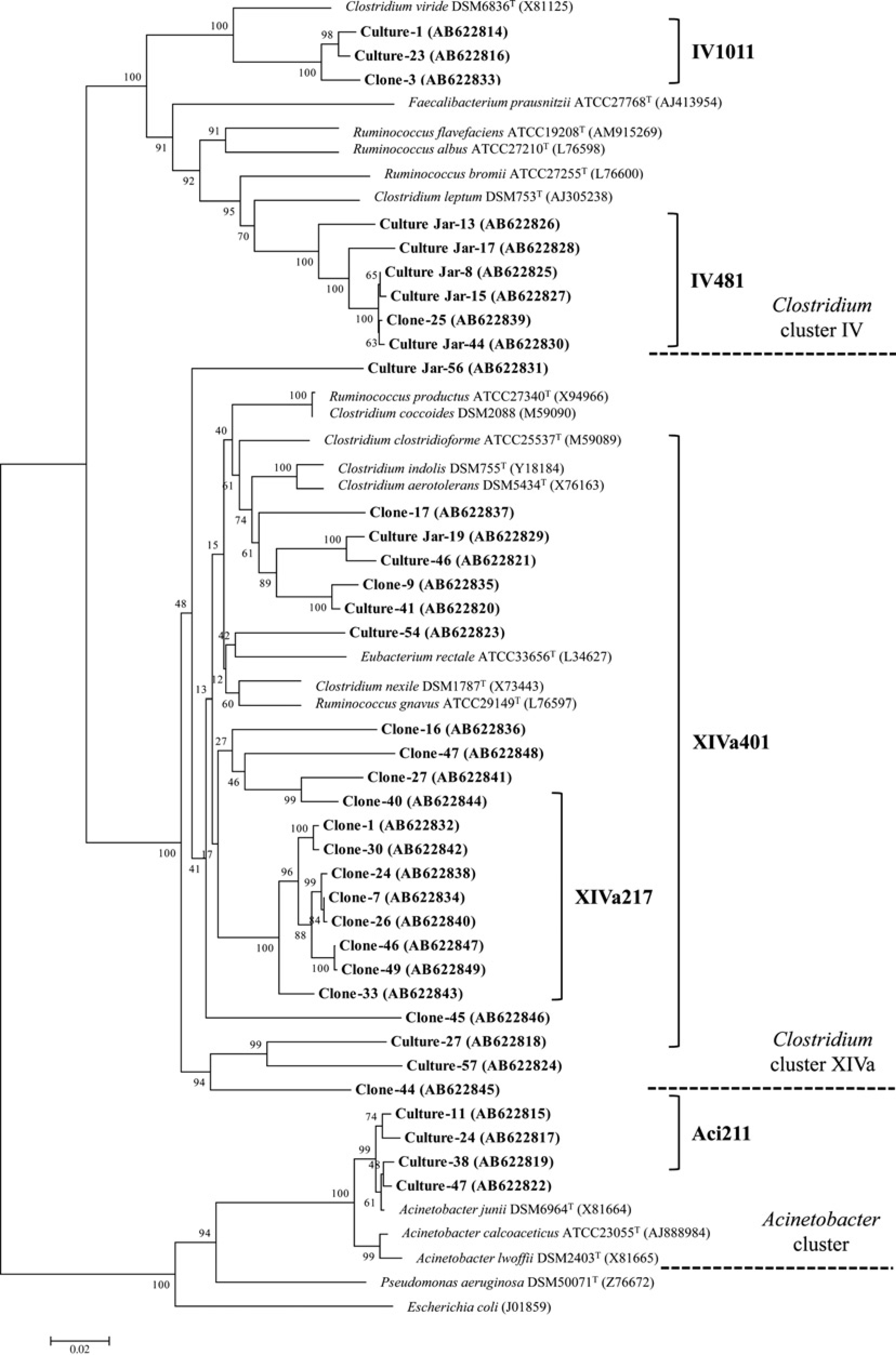
Phylogenetic tree based on 16S rRNA sequences illustrating the target groups of the newly developed probes. The tree shows the relationship among the sequences of mouse Clostridia with close relatives. The oligonucleotide probes, shown on the right of the figure, were designed to identify the organisms in the corresponding branches indicated by the brackets. Bootstrap values, based on 1000 replications, at the nodes of the tree indicate the significance of these nodes. The accession number for each of the published sequences is given in parentheses. Horizontal bars represent sequence divergence
Sequences and target groups of the probes
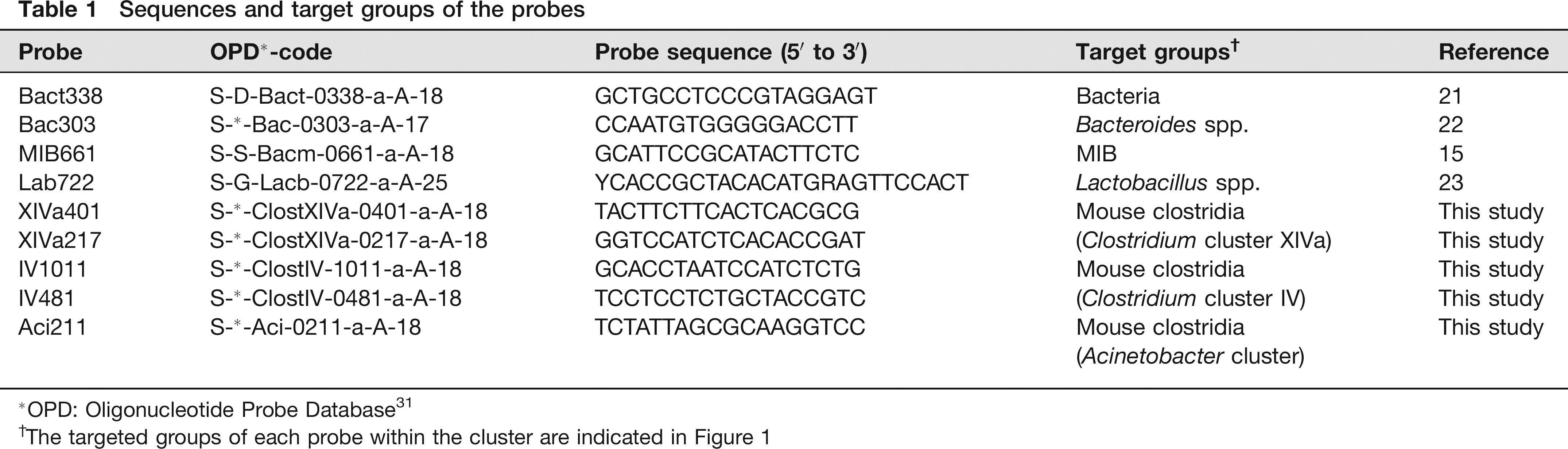

OPD: Oligonucleotide Probe Database 31
The targeted groups of each probe within the cluster are indicated in Figure 1
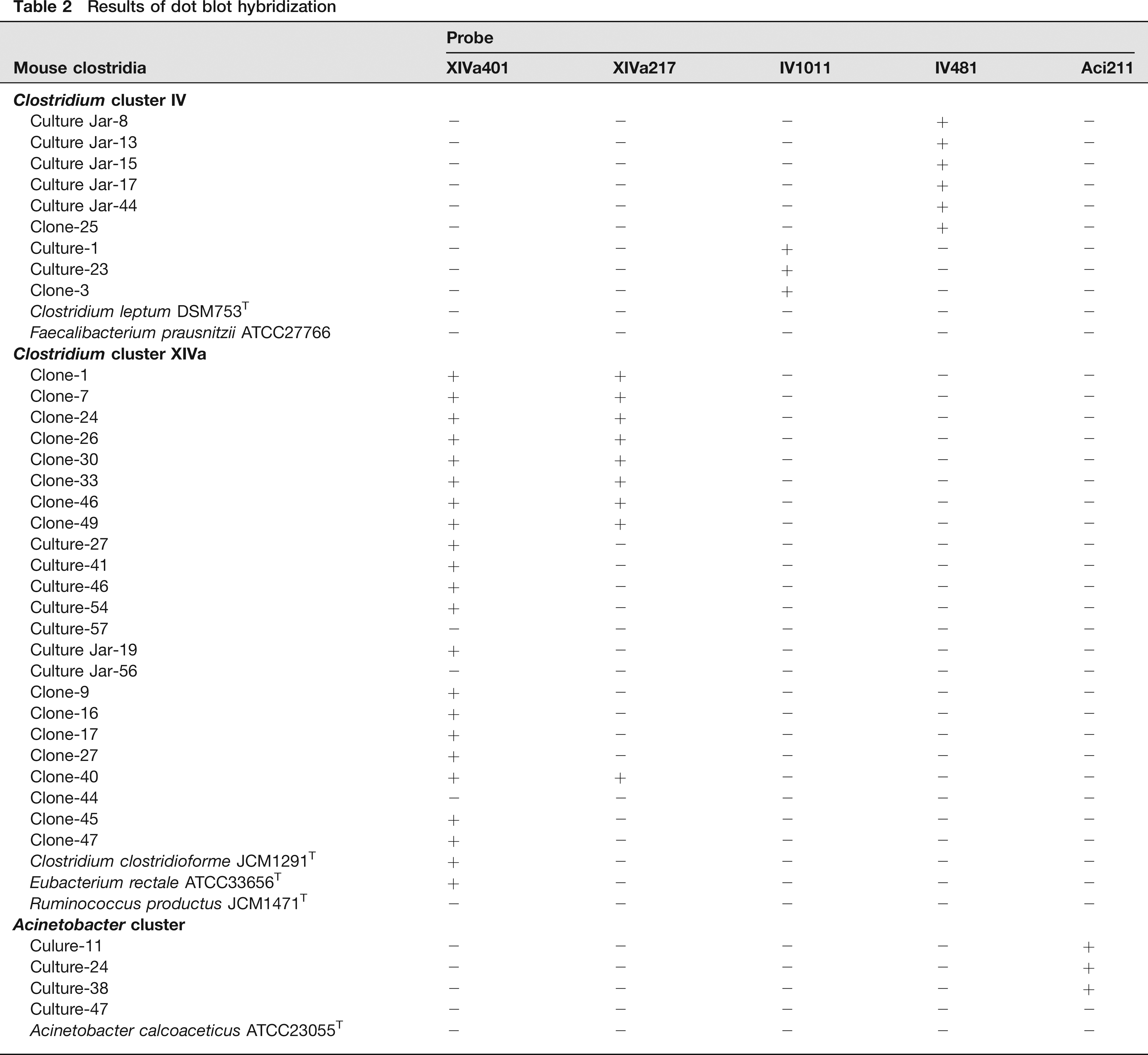
Results of dot blot hybridization
Preparation of faecal samples
Sample preparation and subsequent FISH were carried out according to the protocol described previously 22 with some modifications. Specifically, a buffer of five-fold volume was used to prepare the faecal suspensions because mouse faeces contain more debris and particles than human samples. Freshly-voided faeces were collected from four mice in IL-1Ra KO group and six mice each in other mouse groups, and a 1:50 faecal suspension was prepared immediately with PBS. After centrifugation (50 × g, 5 min), each supernatant was mixed with three volumes of 4% (w/v) paraformaldehyde in PBS and incubated at 4°C for 3 or 6 h to fix cells for analysis by Bac303 and MIB661 probes or for analysis by other probes, respectively. Serial two-fold dilutions were prepared with PBS and a 10 μL aliquot of each dilution was spread over the surface of a 1 × 1 cm well of a Matsumani Adhesive Slide Glass slide (Matsunami, Osaka, Japan). The slides were fixed for 10 min using 96% (v/v) ethanol. For analysis by the Lab722 probe, LABMIX treatment was carried out at 37°C for 40 min prior to hybridization. LABMIX is a mixture of 0.2% (w/v) lysozyme (chicken egg white), 0.01% (w/v) lipase (type II, porcine pancreas), 0.03% (w/v) sodium taurocholate and 0.07% (w/v) CaCl2 dissolved in 100 mmol/L Tris-HCl (pH 8.0).
Fluorescence in situ hybridization
Along with the five new probes, four previously described probes (Bact338, Bac303, MIB661 and Lab722) were used to detect the bacterial kingdom, Bacteroides, mouse intestinal bacteria (MIB) and Lactobacillus, respectively.17,23–25 MIB is an operational taxonomic unit within the Cytophaga–Flavobacter–Bacteroides phylum belonging to a separate branch of the Bacteroides group and a large population has been reported to inhabit the mouse gastrointestinal tract. 17 Hybridization buffer (20 mmol/L Tris-HCl [pH 7.5], 0.9mol/L NaCl and 0.1% SDS) containing 5 ng of oligonucleotide probe μL-1 (listed in Table 1) was applied to the slide surface and incubated at 50°C for 16 h. Oligonucleotide probes with fluorescein isothiocyanate (FITC) or Cy3 incorporated into their 5’ end were synthesized by Greiner Bio-One (Japan). Cy3 was used for Bac303 and FITC for all other probes. The slides were rinsed with washing buffer containing 20 mmol/L Tris-HCl (pH 7.5) and 0.9 mol/L NaCl at 50°C for 20 min. For analysis by the Bac303 and MIB661 probes, hybridization was performed for 3 h and washing for 10 min. After mounting 6 μL of VECTASHIELD Mounting Medium (Vector Laboratories, Burlingame, CA, USA) on each well, slides were viewed under oil immersion using an Axio Imager A1 epifluorescence microscope (Carl Zeiss, Oberkochen, Germany) equipped with FITC, Cy3 and 4’,6-diamidino-2-phenylindole (DAPI) filters. For the total cell count, VECTASHIELD Mounting Medium with DAPI was used.
Statistical analysis
The numbers of DAPI-stained bacteria and probe-positive bacteria in the faecal samples were expressed as mean values and standard deviations. Differences in bacterial counts were analysed using Student's t-test. Differences in the frequency of occurrence of microbes were determined by the chi-square test (df = 1). A significant difference was defined as a P value of <0.05.
Results
Design of new probes
The newly designed probes were confirmed for specificity by using the ARB software package (http://www.arb-home.de). The specificity was subsequently checked by dot blot hybridization using target clostridial groups and close relatives (Table 2). The hybridization was carried out at 50°C in a buffer not containing formamide and washing was also carried out at 50°C. The XIVa401 probe was shown to detect major groups of mouse Clostridia in Clostridium cluster XlVa and also some other known species. This XIVa401 probe did not hybridize with Ruminococcus productus JCM1471, one of the typical species of cluster XlVa. The XIVa217 probe detected only one group that consists of cloned sequences within cluster XlVa. The IV1011 and IV481 probes each specifically detected one of two mouse clostridial groups within cluster IV and did not hybridize with known species tested. The Aci211 probe detected a mouse clostridial group clustering within the Acinetobacter cluster. This Aci211 probe did not hybridize with Acinetobacter calcoaceticus ATCC23055, the type species of Acinetobacter. The corresponding branches that were detected by these five probes are shown in Figure 1.
Comparison of faecal microbiota among CV mouse strains
Faecal microbiota of CV CF#1, BALB/cA and IL-1Ra KO mice raised in our laboratory was analysed by FISH using a combination of five newly designed and four previously described probes (Table 1). The results are shown in Figure 2. The number of DAPI-stained bacteria in BALB/cA mice was statistically lower than that in CF#1 and IL-1Ra KO mice. The percentage of Bact338 positive/DAPI-stained cells was approximately 100% in CF#1 and IL-1Ra KO mice, whereas that in BALB/cA mice was 90.2%. The number of MIB-positive bacteria was higher in CF#1 and IL-1Ra KO mice than in BALB/cA mice. Bac303 and Lab722-positive bacteria were not significantly different among these three mouse strains.
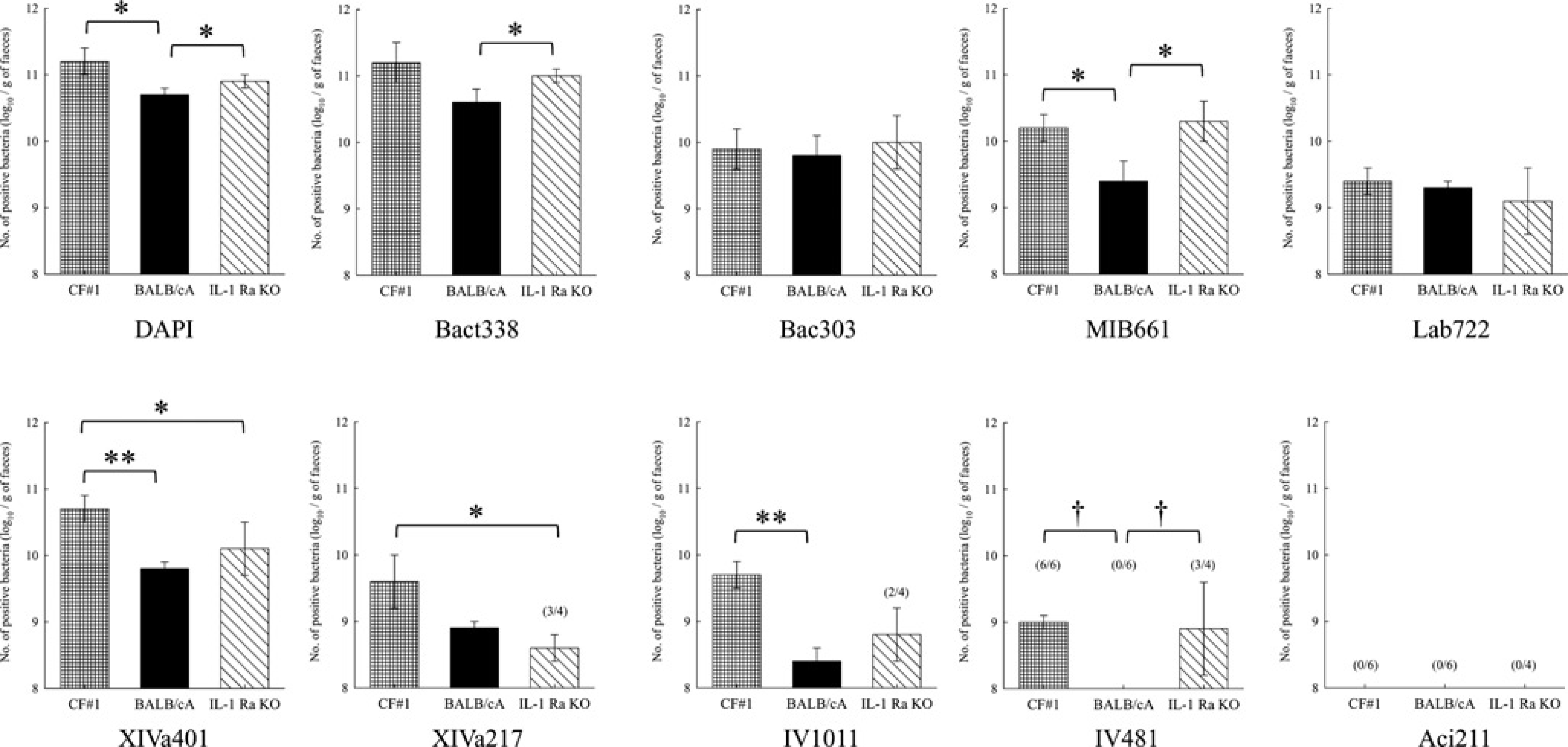
Comparison of faecal microbiota among CV mouse strains. Means and standard deviations of logarithmic counts of each probe-stained bacterium. Figures in parentheses refer to frequency of occurrence (no. positive/no. tested). The number in parentheses not in the column indicates that the frequency is 100%. **P < 0.01; *P < 0.05 for Student's t-test and †P < 0.05 for the chi-square test. CV: conventional; DAPI: 4’,6-diamidino-2-phenylindole
XIVa401-positive bacteria were present in significantly higher numbers in CF#1 mice than in BALB/cA and IL-1Ra KO mice. The numbers of XIVa217 and IV1011-positive bacteria in CF#1 mice were higher than those in IL-1Ra KO mice and BALB/cA mice, respectively. Bacteria labelled with IV481 were below the detection level (<108 g-1) in BALB/cA mice. Aci211-positive bacteria were not detected in any CV mouse strains.
Comparison of faecal microbiota among SPF mice bred in different colonies
Faecal microbiota of BALB/c mice from Japan SLC (SLC mice), CLEA Japan (CLEA mice) and Charles River Japan (CR mice) are shown in Figure 3. The numbers of both DAPI-stained and Bact338-positive bacteria in CR mice were significantly lower than in other colonies. The percentage of Bact338-positive/DAPI-stained cells was approximately 100% in SLC and CLEA mice, whereas it was 71.1% in CR mice. Bac303-positive bacteria in CR mice were also present in smaller numbers than in other colonies of animals. The number of MIB661-positive bacteria in SLC mice was higher than that in CLEA mice and below the detection level in CR mice. Lab722-positive bacteria were not significantly different among these three mouse colonies.
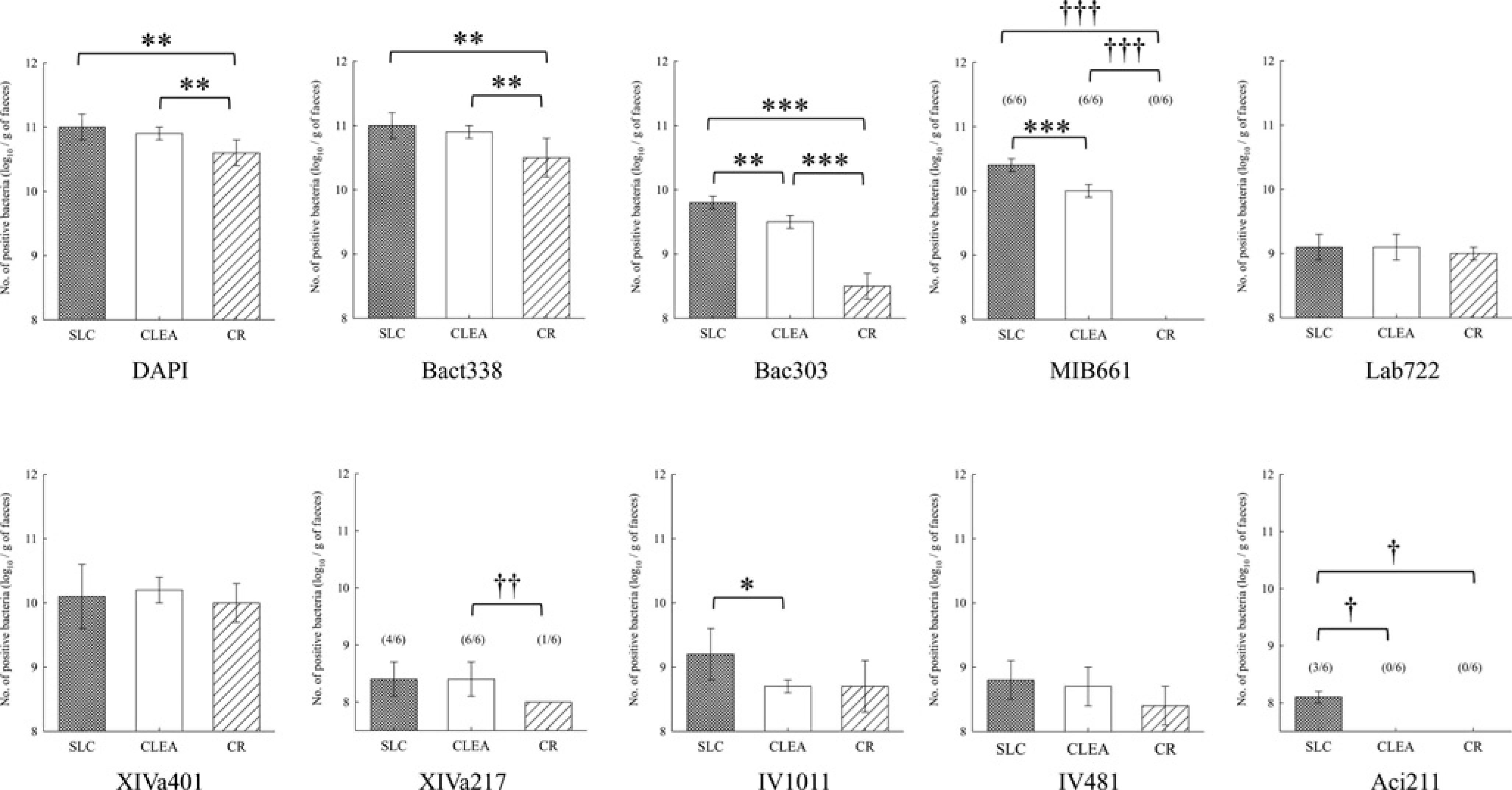
Comparison of faecal microbiota among SPF BALB/c mouse colonies. Means and standard deviations of logarithmic counts of each probe-stained bacterium. Figures in parentheses refer to frequency of occurrence (no. positive/no. tested). The number in parentheses not in the column indicates that the frequency is 100%. ***P < 0.001; **P < 0.01; *P < 0.05 for Student's t-test and †††P < 0.001; ††P < 0.01; †P < 0.05 for the chi-square test. DAPI: 4’,6-diamidino-2-phenylindole
XIVa217-positive bacteria were present in all CLEA mice, whereas they were detected in four and one out of six faecal samples each of SLC and CR mice, respectively. The number of IV1011-positive bacteria in SLC mice was higher than that in CLEA mice. Aci211-positive bacteria were detected in three out of six SLC mice, whereas they were never detected in CLEA and CR mice. The numbers of XIVa401- and IV481-positive bacteria were not significantly different among these three mouse colonies.
Comparison of faecal microbiota between CV and SPF BALB/cA mice
The results for CV BALB/cA mice and SPF CLEA mice, both of which have the same BALB/cA background, were also compared and are shown in Figure 4. The number of DAPI-stained bacteria in SPF BALB/cA mice was higher than that in CV mice. Bact338, Bac303, MIB661 and Lab722-positive bacteria were not significantly different between these two groups.
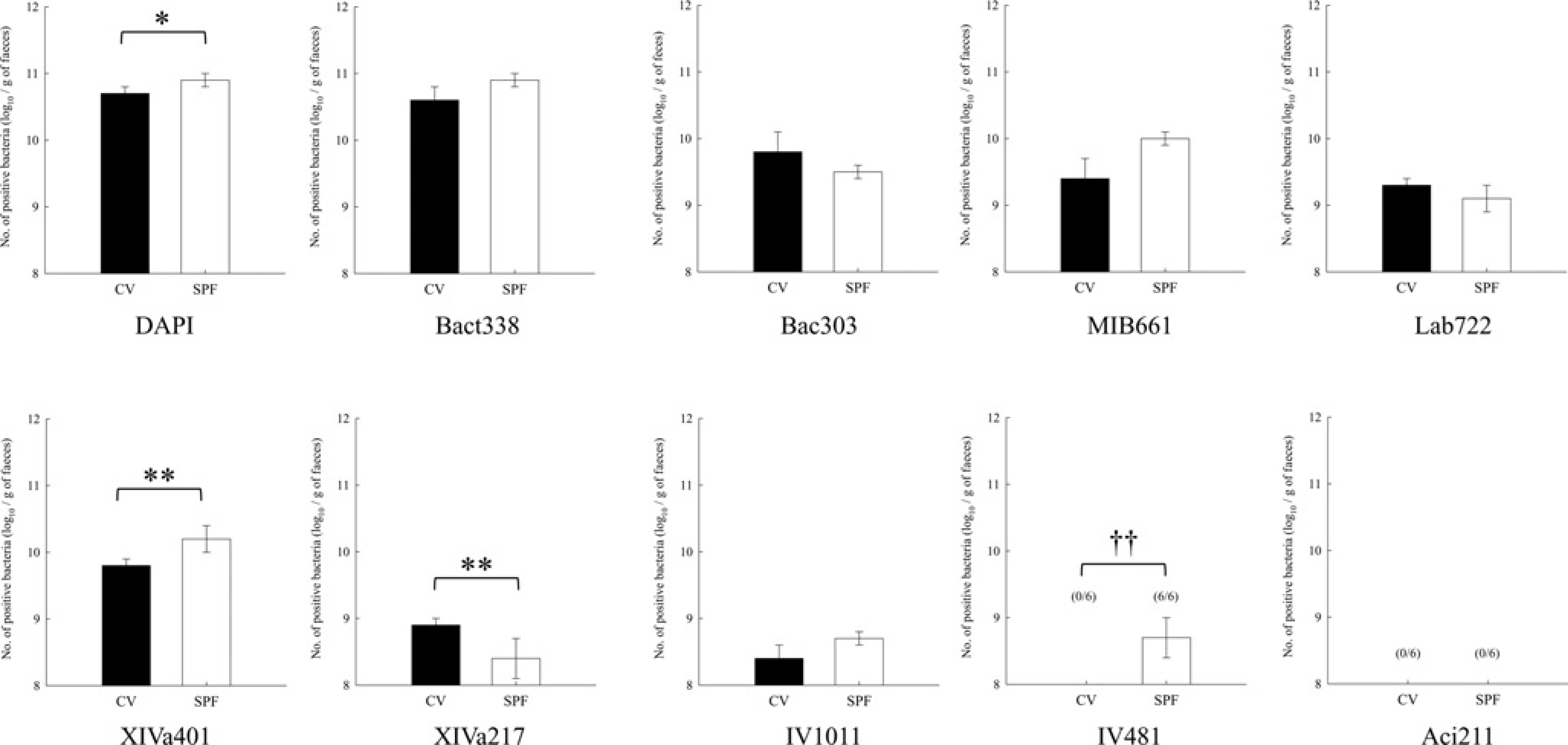
Comparison of faecal microbiota between CV and SPF BALB/cA mice. Means and standard deviations of logarithmic counts of each probe-stained bacterium. Figures in parentheses refer to frequency of occurrence (no. positive/no. tested). The number in parentheses not in the column indicates that the frequency is 100%. **P < 0.01; *P < 0.05 for Student's t-test and ††P < 0.01 for the chi-square test. CV: conventional; SPF: specific pathogen-free; DAPI: 4’,6-diamidino-2-phenylindole
XIVa401-positive bacteria in SPF BALB/cA mice were present in higher numbers than in CV mice. A larger number of XIVa217-positive bacteria was observed in CV mice. There was no significant difference in numbers of IV1011-positive bacteria between the two groups. The number of IV481-positive bacteria was below the detection level in CV mice. Aci211-positive bacteria were never detected in CV and SPF BALB/cA mice.
Discussion
Detailed information on faecal microbiota is important when we use experimental animals. One of the important points of this study is that five groups of mouse Clostridia, which previously were recognized as a single dominant population from their characteristic spore-forming properties and microscopic observations, can be detected quantitatively using newly developed oligonucleotide probes. Another feature is that the oligonucleotide probe sets used in this study can be hybridized under the same conditions, which is important for rapid detection of each bacterial group harboured in the mouse intestine. These five probes were designed specifically to detect mouse clostridial groups essential for the normalization of GF mice. Among them, specificity of the XIVa401 probe was relatively broad and included some other known species within Clostridium cluster XlVa, but target bacteria could be distinguished by their specific fusiform-shaped morphology. Although the target bacterial groups of the XIVa401 probe overlapped with the previously published Erec482 probe, one feature of our XIVa401 probe was that it excluded R. productus, one of the typical species of cluster XlVa. It would, therefore, be better to use the XIVa401 probe to detect and monitor the kinetics of clostridial groups of cluster XIVa in the mouse intestine. One strain, Culture-47, could not be detected by the Aci211 probe. This strain has higher sequence similarity with Acinetobacter junii DSM6964T than with the other three strains of mouse Clostridia 11 and it was difficult to design a group-specific probe including Culture-47. Among mouse groups tested, Aci211-positive bacteria were detected only in SLC mice of low frequency and low bacterial counts near the detection limit, indicating that this clostridial group occupied a minor niche in the mouse clostridial community.
It has been reported that different strains of mice have different intestinal microbiota based on culture methods,26,27 but detailed information on the composition of mouse Clostridia is still unclear. In this study we designed five oligonucleotide probes to detect significant mouse clostridial groups that contribute to the normalization of GF mice, and examined the faecal composition of various mouse groups by FISH. In addition to our five probes, four previously described probes were also used to detect the bacterial kingdom, Bacteroides and Lactobacillus. Clostridium, Bacteroides and Lactobacillus are the three key bacterial groups that have marked effects on mouse physiological conditions and stable composition of mouse intestinal microbiota.10,28
The count of Lactobacillus was not significantly different among any mouse groups, whereas the count or the frequency of occurrence of other probe-positive bacteria including total counts did differ. The composition of mouse clostridial groups and the count of MIB were different between CV CF#1 and BALB/cA mice, and between BALB/cA and IL-1Ra KO mice. IL-1Ra KO mice were first produced by Horai et al. 29 and found to develop chronic inflammatory polyarthropathy spontaneously on a BALB/cA background. 18 The faecal microbiota could be changed by genetic or host physiological factors even if the mice were reared in the same CV environment. The numbers or frequencies of mouse clostridial groups, and Bac303 and MIB661-positive bacteria differed among SPF BALB/c mice bred in different colonies. Faecal samples from CV BALB/cA mice and SPF CLEA mice had different compositions of mouse clostridial groups although these mice were derived from the same GF BALB/cA colony. These results suggested that the environmental factors where the mice were raised could also affect the composition of faecal microbiota even if they belonged to the same strain or had the same background. The probe set used in this study can be used to detect these differences and changes. Not only inter-mouse group difference, intra-mouse group difference was also observed; XIVa217-positive bacteria were detected in four out of six SLC mice and one out of six CR mice. The observation that the composition of faecal microbiota had some variation even within the same strain or the same colony could be one of the reasons why not all the animals show the same responses in the experiments in vivo.
The bacterial kingdom specific Bact338 probe was reported to detect only 60.9% of the total DAPI-stained cells in human faeces, 15 whereas this probe was able to account for 95% of the DAPI-stained bacteria in the large intestine and caecum of mice. 17 In this study, approximately 100% of DAPI-stained mouse faecal bacteria could be detected by Bact338 except in CV BALB/cA and SPF CR mice where 90.2% and 71.1% of the DAPI-stained bacteria, respectively, were detected. The reasons why some DAPI-stained bacteria were not detected by the Bact338 probe might be based on the permeability of their cell wall. The ratio of Bact338 positive/DAPI-stained cells differed among mouse groups, suggesting that both DAPI and Bact338 probe should be used for the monitoring of mouse intestinal microbiota.
Hirayama et al. 30 and Nagura et al. 2 reported that SPF mice bred in different colonies possessed different compositions of culturable microbiota at the genus level. In this study, we demonstrated, by FISH, that the composition of mouse clostridial groups and MIB also differed among SPF mouse colonies. As mentioned above, various factors affect mouse faecal microbiota. Therefore, the exact monitoring of mouse intestinal microbiota in SPF facilities is important. The monitoring of specific pathogens is carried out regularly in each SPF facility. However, monitoring of intestinal microbiota, especially the composition of mouse Clostridia, is not performed even though they account for a significant part of the rodent faecal microbiota. The oligonucleotide probes designed in this study are a useful tool for rapid detection of mouse clostridial groups inhabiting the mouse intestine. These newly developed probes combined with previously reported probes used in this study can be applied for the monitoring of mouse intestinal microbiota, not only as a regular monitoring system in SPF facilities but also when establishing and maintaining genetically-modified mice. They will also help to provide important new information on the ecology and functions of mouse intestinal microbiota.
Footnotes
Acknowledgement
This work was supported by Grant-in-aid No. 14380379 from the Ministry of Education, Culture, Sports, Science and Technology of Japan.
