Abstract
As recommendations for specific pathogen-free housing change, mouse facilities need to re-derive their colonies repeatedly in order to eliminate specified bacteria or viruses. This paper describes the establishment of a new mouse facility using as starting point a small colony of CD-1 mice colonized with the Charles River altered Schaedler flora (CRASF®) housed in individually ventilated cages (IVCs). The import of new strains was performed exclusively via embryo transfer using CD-1 mice as recipients. The integrity of the CRASF® in caecum samples of the original CD-1 colony and of three inbred mouse lines imported into the colony was proven by a quantitative realtime polymerase chain reaction approach. Furthermore, we searched for bacterial contaminants in the gut flora using non-specific 16S rRNA primers. The bacterial sequences found were closely related to but not exclusively sequences of altered Schaedler flora (ASF) members, suggesting that the ASF is heterogeneous rather than restricted to the eight defined bacteria. Moreover, no pathogens were found, neither using the non-specific 16S rRNA primers nor in routine quarterly health monitoring. As one effect of this defined gut flora, interleukin-10 knockout mice are devoid of colitis in our facility. In conclusion, our approach building up a mouse facility using foster mothers and embryo transfer as well as a strict barrier system and IVCs is suitable to maintain a colony free from contaminating bacteria over the long term. CRASF® remained stable for seven mouse generations and was efficiently transferred to the imported mouse strains.
In the modern husbandry of mice, the recommendations for specific pathogen-free (SPF) housing tend to change over time. In Europe, the Federation of European Laboratory Animal Science Associations (FELASA) sets the standards for SPF status in laboratory animal colonies. 1 When newly found organisms are deemed significant by the research community, facilities must clean up their colonies by embryo transfer in order to fulfil the newer standard. Furthermore, differences in the enteric flora in mice can have a great impact on some experiments as well as on the outcome of mouse phenotyping; for example, the differing severity of colitis in interleukin-10 knockout (IL-10−/−) mice. 2 In order to avoid periodic re-derivations of mouse colonies and to constantly have a comparable enteric flora in all mouse strains over longer periods of time, a small colony of germfree CD-1 mice that were colonized by the Charles River altered Schaedler flora (CRASF®) was the starting point of our new mouse facility. New strains were imported exclusively via embryo transfer. With this approach, we assured that in all likelihood none of the microorganisms that our research community considers significant to their projects would be present in our colony.
The gastrointestinal system is the organ containing the largest number of microorganisms of the entire body. 3 About 400–1000 different bacterial species colonize the adult human intestinal tract. 4,5 It has been estimated that only 20–40% of the intestinal bacteria can be cultured with current methods 6 and novel species were found using 16S rRNA libraries, 7 meaning that the intestinal flora contains about 60–80% undefined bacteria. The composition of the bacterial flora is dependent on environmental factors such as the diet 8 and the stress level of the animals. 9 Additionally, the genetic background of the mice is influencing the intestinal flora. 10 Newborn mice are colonized through the faeces of their parents by coprophagia. This can lead to a high variability of the intestinal flora between different mouse colonies as well as between groups housed in different cages in the same mouse facility. 11 In attempts to reduce this variability, defined microflora with known ingredients have been applied. The first defined mouse intestinal flora was developed by Schaedler in 1965. It consists of eight commensal bacterial strains naturally occurring in mice. 12 In 1987, Orcutt et al. 13 developed an ‘altered Schaedler flora’ (ASF) that has become the standardized flora most commonly used. Charles River Laboratories provide mice associated with two CRASF®, one consists of the eight ASF strains and CRASF®plus contains in addition non-haemolytic Escherichia coli and Enterococcus faecalis. The mice at the Helmholtz Centre for Infection Research (HZI) were associated with CRASF®. An overview of the defined microflora mentioned is given in Table 1.
Composition of the most frequently used defined microflora for mice
OSF: original Schaedler flora; ASF: altered Schaedler flora; CRASF®: Charles River altered Schaedler flora; G–C: guanine–cytosine
The IL-10−/− mouse is one example of how the microflora can influence a phenotype. Housed in a conventional facility these mice develop a chronic enterocolitis similar to human inflammatory bowel disease (IBD). 14 In SPF mouse facilities, the onset of the inflammation is delayed and the severity is variable due to unknown environmental factors. 2
To our knowledge, this study describes for the first time (a) the colonization of an entire facility, that to date encompasses over 150 mouse lines, with mice carrying the defined CRASF® microflora, (b) the housing of these mice exclusively in individually ventilated cages (IVCs) under controlled environmental conditions, (c) the analysis of the stability of this defined flora for four years or seven mouse generations after the initial association and (d) the absence of defined murine pathogens in this mouse facility.
Material and methods
The animal facilities
The animal housing facility at the HZI encompasses three buildings: a small quarantine; a main, dedicated building for housing SPF stocks as well as experimental breeding colonies; and an infection unit for performing infection challenge experiments at biosafety level 2 under defined environmental conditions. The last building is the only one to which researchers are allowed access. The animal rooms in all three buildings are protected by barriers and supplied with high efficiency particulate air-filtered air with a temperature of 22°C and a relative humidity of 55% at an exchange rate of 15–17 room volumes per hour. Access to the animal rooms in the main building is via three-chamber air locks consisting of a central air shower flanked by gowning rooms on the external and internal sides. Personnel are required to wear sterile surgical protective clothing including bonnet, face mask, gloves and clogs at all times in the animal rooms. All materials (food, cages, bedding material, etc.) entering the animal rooms are autoclaved (118°C, 20 min). Drinking water is acidified with sulphuric acid (pH 3), then filled into bottles and autoclaved. Electrical and other equipment that is temperature sensitive is sterilized with vaporized hydrogen peroxide (VHP) (Steris, Cologne, Germany) when taken into the animal rooms. All rooms within the barriers of the main building and the infection unit are sterilized with VHP before being put into operation. Mice are housed exclusively in IVCs (Sealsafe®, 1284L, Tecniplast, Hohenpeißenberg, Germany). IVC racks are exchanged on a regular basis (every 3 months), cleaned and autoclaved. Operations requiring that cages be opened, both experimental procedures and routine cage changing, are always carried out under laminar flow safety cabinets or changing stations. Lids of dirty cages are replaced and closed before removing cages from the safety cabinet for transport out of the barrier to the washing facility. Following disposal of dirty bedding, cages are washed in a tunnel washer (Tecniplast) using cycles of prerinse, alkaline wash, acidic wash, final rinse and drying. Fresh bedding is added to the clean cages, and the lids are replaced for transport and sterilization back into the barrier. The cage washing room has two entrances at the dirty and clean ends of the tunnel washer, respectively, ensuring a unidirectional work flow. All mouse lines imported into the facilities are quarantined until they can be re-derived by embryo transfer using CD-1 recipients with CRASF® (see below) and subsequently transferred into the SPF areas. Health monitoring of mouse colonies is performed on a quarterly basis using CD-1 mice from our stock colony as sentinels. During the three-month monitoring period, sentinels are placed in dirty cages from a different mouse line weekly using a systematic, rotating protocol. Monitoring is performed according to FELASA recommendations 1 with the addition that Helicobacter species are tested every quarter.
The mice
CD-1 mice (Crl:CD1[ICR]) associated with the CRASF® via transfer into a CRASF® associated gnotobiotic colony by Caesarean section were acquired from Charles River Laboratories (Sulzfeld, Germany) in March 2001. The analysed individuals were out of the sixth to seventh generation bred in our mouse facility. All other mouse strains were imported via embryo transfer using CD-1 foster mice. The mouse strains C57BL/6J and BALB/c (BALB/cByJ) were acquired from The Jackson Laboratory (Bar Harbor, ME, USA) in April 2004. The individuals were out of the second generation bred in our mouse facility after embryo transfer. 129P2 (129P2/OlaHsd) mice were acquired from Harlan-Winkelmann (Borchen, Germany) in November 2004 and were out of the first generation bred in our mouse facility after embryo transfer. All mice were at least six months of age (range: 6–21 months, average: 11 months). Animals from at least two different litters of each strain were analysed. All inbred strains were fed with rodent maintenance diet Ssniff® V1534-3 R/M-H, whereas rodent breeding diet Ssniff® V1124-3 M-Z (Ssniff, Soest, Germany) was the diet for the CD-1 foster mothers. Mice were anaesthetized by a rising CO2 concentration and killed by organ excision, the entire caecum was excised and stored at −20°C until further processing.
Composition of CRASF®
According to Charles River Laboratories, CRASF® contains ASF519 (Bacteroides sp.), ASF360 (Lactobacillus sp.), ASF361 (Lactobacillus murinus), ASF356 (Clostridium sp.), ASF492 (Eubacterium plexicaudatum), ASF500 (low guanine–cytosine [G–C] content Gram + group), ASF502 (Clostridium sp.), ASF457 (Flexistipes group). Further available is the flora CRASF®plus containing E. faecalis (Enterococcus spp. #ATCC 10541) and E. coli (non-haemolytic # ICO-IM 1803) in addition (Table 1). The CD-1 mice used to build up the HZI mouse colony were associated with a CRASF®. CRASF® control flora was a kind gift from Charles River Laboratories (Wilmington, MA, USA).
DNA extraction from mouse caecum
DNA extraction from mouse caecum was carried out using the QIAamp® DNA Stool Kit (Qiagen, Hilden, Germany), according to the manufacturer's instructions with minor modifications. The DNA concentration was determined with a GeneQuant RNA/DNA Calculator (GE Healthcare, Freiburg, Germany) at 260 and 280 nm.
Amplification of 16S rRNA genes
The qualitative determination of ASF strains was done using specific primers for the amplification of 16S rRNA genes as described in Sarma-Rupavtarm et al. 15 Primers were commercially synthesized (Invitrogen, Karlsruhe, Germany).
Sample preparation for DNA sequencing
Optimal annealing temperature was determined using a PTC-200 gradient thermal cycler (MJ Research, Waltham, MA, USA). Amplification was performed with the following conditions: an initial step of 3 min at 98°C and 40 cycles, with one cycle consisting of one minute at 96°C, one minute at annealing temperature, and one minute at 72°C and one final step for one minute at 72°C.
A product from a 100 µL polymerase chain reaction (PCR) mixture was run on a preparative 2% agarose gel. The DNA was stained with SYBR Safe® (Invitrogen) and visualized on a Dark Reader® transilluminator (Clare Chemical Research, Dolores, CO, USA). The bands corresponding to the amplification products were excised and directly purified with the QIAquick Gel Extraction kit (Qiagen).
Confirmation of 16S rRNA genes by sequencing
Purified DNA samples from preparative PCRs were sequenced using the specific primers for each of the expected 16S rRNA genes and the ABI PRISM® BigDye Terminator v 3.1. Ready Reaction Cycle Sequencing Kit (Applied Biosystems, Foster City, CA, USA). The reactions were analysed with an ABI 3730xI 96-capillary automatic DNA Analyzer (Applied Biosystems) at the Department of Genome Analysis, HZI.
Realtime PCR conditions
Realtime PCR amplification was performed using a LightCycler 1.0 (Roche, Mannheim, Germany). Amplification was carried out with a QuantiTect SYBR Green PCR kit (Qiagen), according to the manufacturer's instructions.
16S rRNA gene clone library construction
For the 16S rRNA clone library, rRNA genes were amplified using the universal forward oligonucleotide primer 5′-AGTTTGATYMTGGCTCAG-3′ and the universal reverse primer 5′-AAGGAGGTGWTCCARCC-3′(Y = C/T, M = A/C, R = A/G, W = A/T). PCR was carried out with Taq polymerase (Qiagen). The following programme was used: a 8 min degeneration step, 35 cycles consisting of 95°C for 45 s, 60°C for 45 s and 72°C for 90 s with an increase of 5 s per cycle. A final extension step was carried out at 72°C for 5 min. Approximately 1.5 kb PCR products were excised from the gel and cloned with a Qiagen PCR Cloning Kit (Qiagen) in accordance with the manufacturer's instructions. Ligation products were transformed into chemically competent E. coli TOP 10F' cells (Invitrogen) according to the manufacturer's instructions and standard techniques. 16 Transformants were picked with a Qbot robot (Genetix, New Milton, UK).
High-throughput plasmid purification for capillary sequencing
Plasmids were isolated by alkaline lysis 16 using MADVN6550 and MSNUPSD50 multiscreen plates (Millipore, Schwalbach/Ts, Germany) with a Perfect Vac Manifold (Eppendorf, Hamburg, Germany). The plasmid inserts were sequenced using the standard T7 and SP6 promotor primers with standard protocols for the ABI 3730xI 96-capillary automatic DNA analyser.
Analysis of the 16S rRNA gene sequences
The 16S rRNA sequences were blasted against the 16S rRNA (Prokaryotes) database (
Cultivation of E. coli and E. faecalis
The cultivation of E. coli and E. faecalis was performed at the Institute for Laboratory Animal Science and Central Animal Facility of the Hannover Medical School. Faecal samples were collected from two CD-1 mice and one BALB/c female mouse, six months of age. After 24 h of cultivation in thioglycolate at 37°C, samples were plated on blood agar and Gassner agar plates. Plates were further incubated overnight at 37°C. E. faecalis was identified by morphology, Gram staining and biochemical analysis using rapid ID 32 STREP from bioMérieux® SA, Marcy-l'Étoile, France.
Histomorphological analysis of IL-10−/− mice
C57BL/6-Il10tm1Cgn J (IL-10−/−) mice were obtained from The Jackson Laboratory in May 2003 and introduced to our facility via embryo transfer. In total, 16 IL-10−/− mice (10 males, 13 months of age, 6 females, 10 and 13 months of age) and 13 C57BL/6J mice (5 males, 6 months of age, 8 females, 9 months of age) bred and maintained at the HZI animal facility were analysed. Mice were killed by organ excision after anaesthesia with rising CO2 concentrations. Intestines were immediately excised rolled up to ‘Swiss rolls’ and fixed in 4% neutrally buffered formaldehyde. Paraffin-embedded tissues were obtained by standard laboratory procedures in order to analyse the mucosa throughout the entire length of the intestine in haematoxylin–eosin (HE) stained sections.
Results
Qualitative analysis of CRASF® after four years
In order to test whether the CRASF® is efficiently transmitted to mouse strains via embryo transfer and is stable over several mouse generations, we used a qualitative PCR approach to detect the original ASF members. The gut flora of the original CD-1 colony and of three inbred mouse strains imported into the colony (C57BL/6J, BALB/c, 129P2) were analysed. Bacteria were isolated from the caecum. To evaluate the method, original CRASF® obtained from Charles River Laboratories was used as positive control. As shown in Figure 1, all 16S rRNA genes could be detected after amplification by using the specific primer pairs. Each PCR with a specific primer pair showed only one band in the SYBR Safe®-stained agarose gel, corresponding to the expected size of the 16S rRNA gene. Using the qualitative PCR approach, we were able to detect all members of the original CRASF® in C57BL/6J male mice (Table 2). In the other mouse strains, only an incomplete spectrum of the CRASF® members could be identified. ASF360, ASF457 and ASF519 were the bacteria constantly found throughout all individuals of the mouse strains. ASF360 could be detected in samples including the caecal wall, but not on samples containing caecal content only (data not shown).
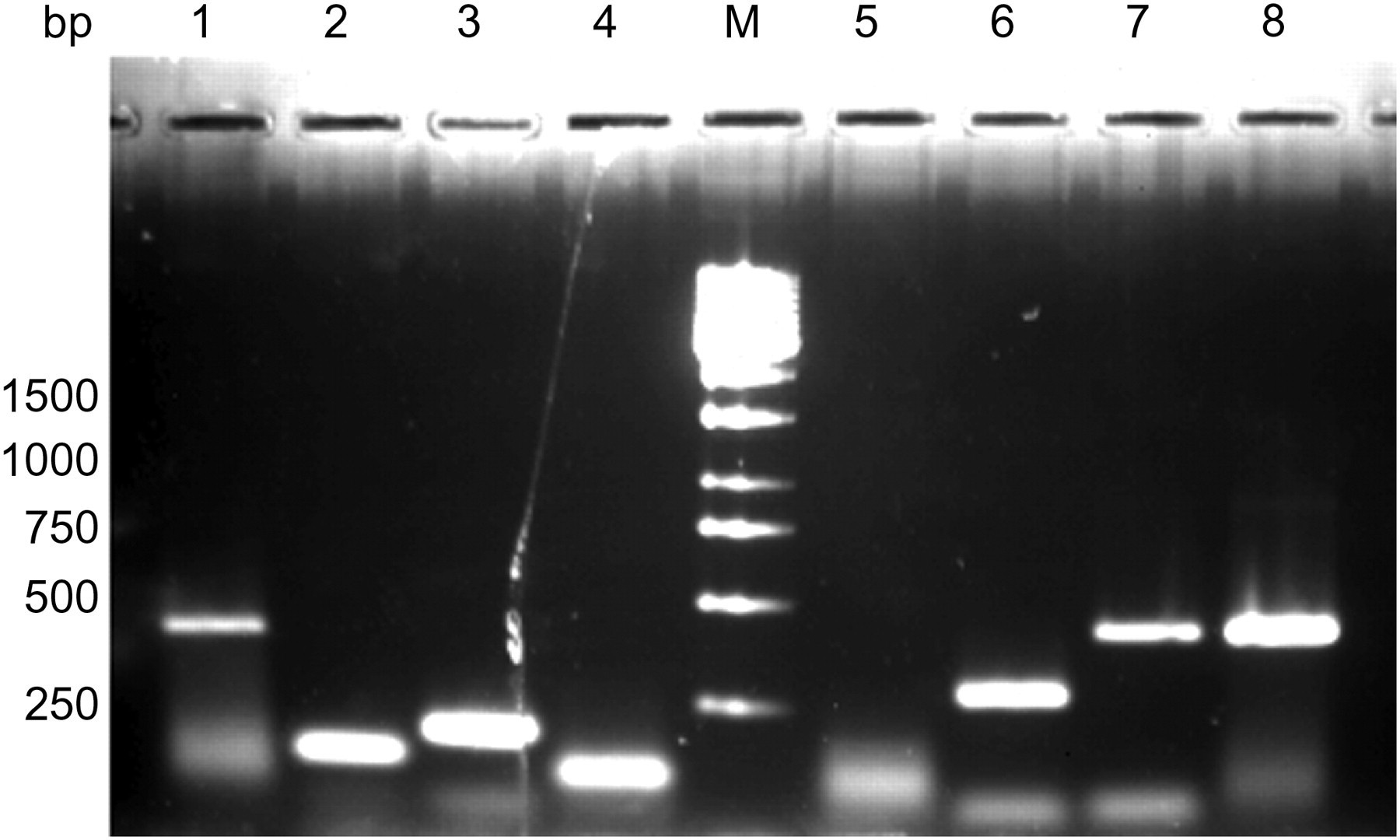
Specific polymerase chain reaction products obtained by amplification of original Schaedler Flora obtained from Charles River Laboratories (CRASF®) on a 2% agarose gel stained with SYBR Safe®. Lanes 1–4: ASF356 (417 bp), ASF360 (131 bp), ASF361 (182 bp), ASF457 (95 bp). Lane ‘M’: 1 kb DNA ladder (MBI). Lanes 5–8: ASF492 (167 bp), ASF500 (285 bp), ASF502 (427 bp), ASF519 (429 bp)
Distribution of ASF in mouse individuals (qualitative PCR analysis)
Numbers of positive-tested mice for a given ASF strain divided by the number of mice tested (‘ASF ratio’). The ‘ASF score’ is the addition of ‘ASF ratios’. The ‘ASF score’ represents the fraction of 23 mouse individuals that have been tested positive for a given ASF strain on a 0–8 grading scale
M: male; F: female; ASF: altered Schaedler flora; PCR: polymerase chain reaction
Quantitative realtime PCR
All the CRASF® members were detected using the realtime PCR (Figure 2). In all four mouse strains ASF457 (Flexistipes group) (23–42%), ASF502 (Clostridium sp.) (21–35%) and ASF519 (Bacteroides sp.) (14–34%) displayed the highest copy number and therefore the highest amount of bacterial DNA in the caecum. ASF500 and ASF492 were found in intermediate concentrations (0.5–11%). ASF360 (Lactobacillus sp.) was least represented in the caecum content (0.06–0.4%). Further, bacteria found in low amounts were ASF356 (Clostridium sp.) and ASF361 (L. murinus) (0.2–3.3%) (Figure 2). The overall distribution of ASF strains was very similar in all mouse strains analysed. No significant interstrain variation could be found. Interindividual differences were very high.
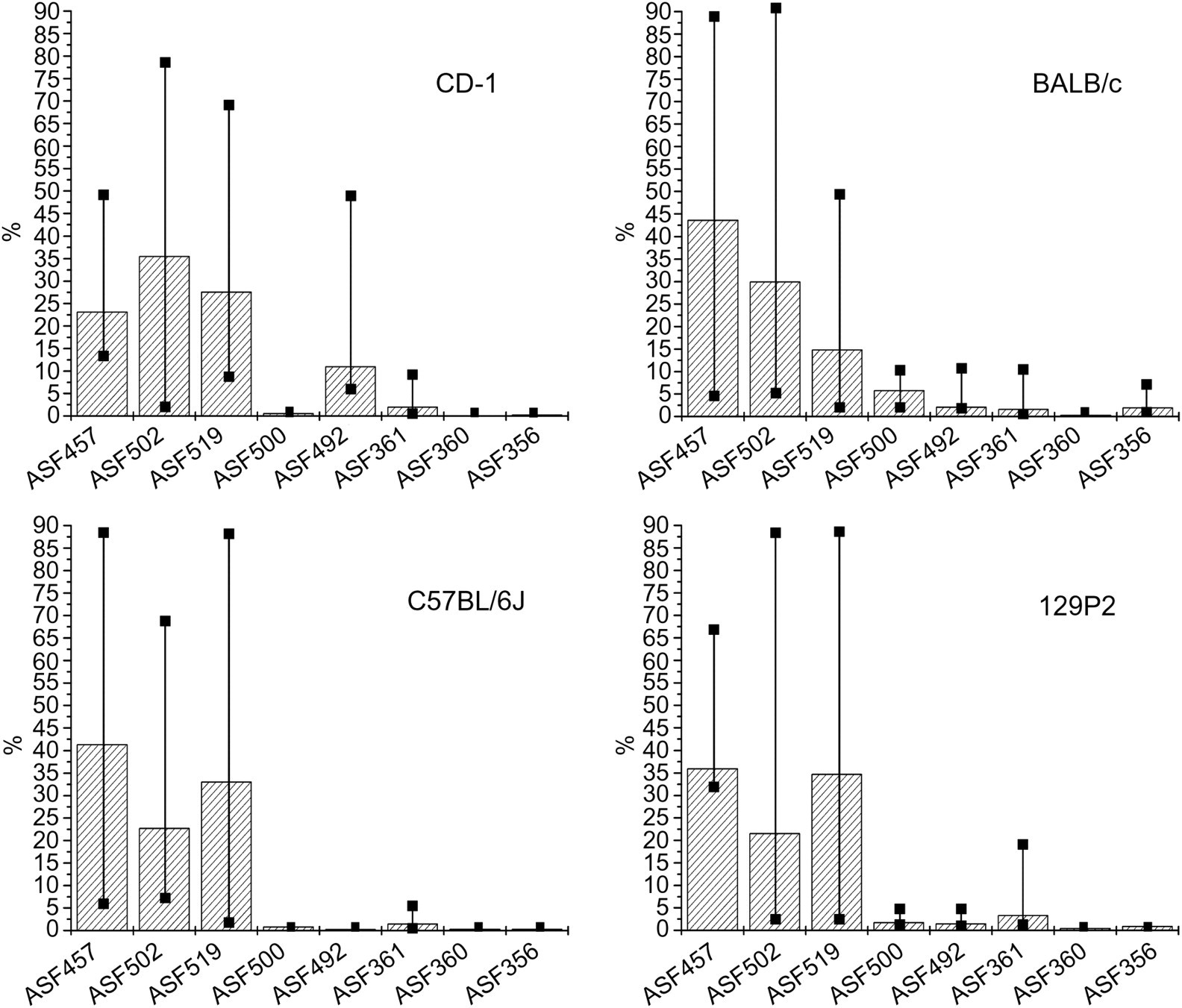
Average percentage of altered Schaedler flora strains in four mouse strains determined by realtime polymerase chain reaction. The number of individuals were nine (CD-1), eight (C57BL/6J), ten (BALB/c) and seven (129P2). The error bars indicate the maximal and minimal percentage found
Cultivation and biochemical analysis revealed E. faecalis but no E. coli
Faecal samples of two CD-1 mice and one BALB/c mouse were cultured as described in materials and methods. E. faecalis was detected by morphological and biochemical analysis (99.9% confidence interval) in all samples, while E. coli could not be cultured.
Analysis for bacterial contaminants
In order to determine possible bacterial contaminants, we used degenerated primers for the amplification of the complete genomic 16S rRNA genes, cloned the amplified fragments and determined the bacterial content by sequencing and database analysis. The caecum content of one C57BL/6J and one CD-1 male mouse was analysed. In total, 96 clones from caecum samples of C57BL/6J and CD-1 were sequenced. Eleven clones from the caecum sample of CD-1 and five clones from the caecum sample of C57BL/6J contained 16S rRNA sequences (Table 3). Several clones contained mouse DNA sequences (37 for CD-1, 21 for C57BL/6J) or no insert (49 for CD-1, 66 for C57BL/6J). In both mice, 16S rRNA genes clustering with Clostridium sp. ASF502 and Bacteroides sp. ASF519 were identified. The sequence identity to the closest relative ranged from 88% to 99%. From the defined ASF strains only the 16S rRNA gene of Bacteroides sp. ASF519 was clearly identified in two clones of each mouse (Table 3). All other 16S rRNA genes clustered with the ASF strains, but were not identical (Figure 3). We identified 16S rRNA genes closely related to Clostridium saccharolyticum DSM 2544, Clostridium sp. CCUG 52336 and Clostridium indolis, which belong to the genus Clostridium as the defined ASF strain Clostridium sp. ASF502. We detected 16S rRNA sequences, closely related to Lachnospiraceae bacterium A4 and butyrate-producing bacterium T2-145, which belongs to the order Clostridiales. The identified 16S rRNA gene of Eubacterium eligens, which is closely related to E. plexicaudatum (ASF492), belongs to the genus Eubacterium. The phylogenetic trees show the identified sequences along with the sequences of their closest relatives and the sequences of the ASF members (Figure 3). No 16S rRNA gene sequences of pathogenic bacteria were found.
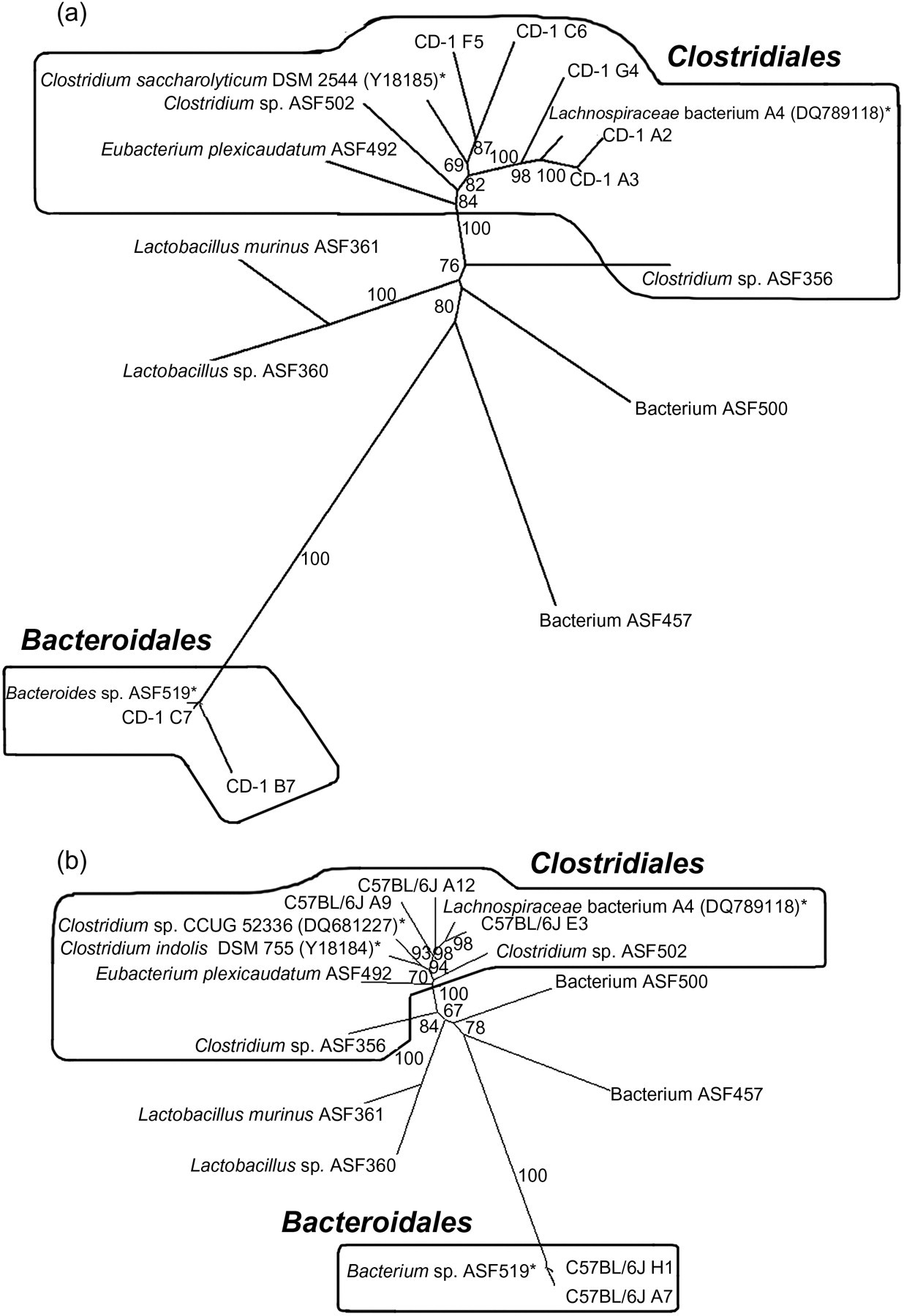
Phylogenetic tree showing the relationships of 16S rRNA gene sequences obtained from 16S rRNA clone library from caecum samples with their closest relative sequences and altered Schaedler flora strains. The closest related sequences based on a multiple-sequence alignment are marked with an asterisk. The closest related sequences are named as in Table 3 (almost-complete sequences). Bootstrap values based on 1000 replications display the significance of the nodes. Accession numbers for each of the published sequences are given. The bacterial orders are written in bold letters
Summary of sequenced clones from the 16S rRNA clone library
*16S rRNA database (
IL-10−/− mice do not develop IBD at the HZI mouse facility
As an example of the phenotypic consequences of the CRASF®, the IBD phenotype of IL-10−/− mice housed in our facility is described below. Clinical examination and necropsy did not reveal any macroscopical abnormalities. HE-stained sections of the intestine in its entire length revealed small single foci of inflammatory infiltrations in the lamina propria in seven individuals out of 16 IL-10−/− animals. Less than 0.01% of the intestine was affected by the inflammation. The inflammatory infiltrate consisted of granulocytes, macrophages and lymphocytes (data not shown). Compared with 10–30% inflamed intestine in IL-10−/− mice described in Mähler and Leiter, 2 the inflammation was very mild and, most likely, clinically irrelevant. C57BL/6J animals did not show alterations in histology.
Discussion
The set-up of our mouse facility is suitable to maintain a mouse colony housed in IVCs associated with the defined CRASF® for four years or seven mouse generations. Differences in the amount of organisms present for each bacterial strain might account for the discrepancy between the mouse strains observed in the qualitative PCR. The sensitivity of the qualitative PCR might not be high enough to detect low numbers of bacteria. In contrast, using the quantitative realtime PCR approach, all ASF members could be detected in every mouse strain analysed, indicating that realtime PCR is more sensitive than the qualitative PCR. For all mouse strains tested, ASF457, ASF502 and ASF519 were the most prominent bacteria, while the other ASF strains were found only in small numbers. Similarly, Sarma-Rupavtarm et al. 15 described a high number of ASF502, ASF457 and ASF500 in the caecum, while Deloris Alexander et al. 11 found a dominating population of ASF492, ASF500 and ASF502 in the caecal bacterial flora. Genetic background and exogenous factors, including the use of acidified water, nutrition and environmental stress, may be responsible for the variances. 9,11 Remarkably, the composition of the gut flora was equivalent between the analysed mouse strains. High interindividual differences might have masked differences between mouse strains. Significant differences in the composition of the gut flora between individual mice have been described before i.e. by Deloris Alexander et al., 11 Sarma-Rupavtarm et al. 15 and Dubos and Schaedler. 8
The CRASF® is named a ‘defined microflora’. We have evidence that the 16S rRNA genes, identified in the caecum of the C57BL/6J and CD-1 mice, were not identical to the defined ASF strains in the National Center for Biotechnology Information nucleotide database. The 16S rRNA gene sequences rather match to bacterial species related to the defined ASF. Only for the ASF strain Bacteroides sp. ASF519 we found highly identical 16S rRNA genes. All other identified 16S rRNA genes were only related at genus or order level to the expected ASF sequences. In 1999, Dewhirst et al. 19 characterized ASF strains, grown on agar media for the first time by 16S rRNA sequence analysis. Since only 60–80% of microorganisms of the intestine can be cultured, the isolates might show lower species diversity compared with identification by 16S rRNA sequence analysis of the intestinal content. 6,30 The strength of the 16S rRNA sequence analysis method is that even non-culturable anaerobe bacteria are detected. 31 Furthermore, the ASF strains were originally only identified based on phenotypic criteria, the definite identification at the genus and species levels might be uncertain. 13,32 Therefore, we assume that the ASF might contain additional species or phylotypes, which are closely related to the defined ASF strains. We exclude sequencing errors because of the good quality of the obtained sequences. Furthermore, we were able to clearly identify the 16S rRNA gene sequence of Bacteroides sp. ASF519 with an identity of 96–99%, which confirms the validity of our method. However, we cannot exclude certain minor changes of the particular flora composition of the HZI mouse facility over the time and generations, since the flora was not analysed on arrival. Moreover, the mouse colony used to associate our founder mice with CRASF® was initially associated 10–20 years ago (Charles River Laboratories, personal communication). Even though the presence of the eight ASF members in the CRASF® is regularly verified by sequencing at Charles River Laboratories (Charles River Laboratories, personal communication), the mouse colony at HZI might contain further closely related bacteria. Therefore, we assume that the identified 16S rRNA sequences belong to bacteria that are not contaminants, but rather unidentified components of the CRASF®. With our approach, we could only get a first impression of the complexity of the flora in the mouse caecum. A further metagenomic approach of the mouse intestinal tract, associated with the ASF, may enlighten the full species abundance in the ASF cocktail. An important observation is that no pathogenic bacteria were found using the described method. This is in line with the quarterly health monitoring results performed by analysis of sentinels that did not reveal any bacteria listed in the FELASA recommendations up to now (data not shown). E. faecalis could not be detected using non-specific primers for 16S rRNA genes, but was clearly identified with classical cultivation methods. Although these bacteria are most frequently found by cultivation because they are facultative aerobes, they might not represent the predominant gut flora in mice but be outnumbered by anaerobes that are not culturable with standard techniques. For human gut flora it has been shown that rather than the frequently used indicator organisms for faecal contaminations such as E. faecalis, the strict anaerobes like Bacteroides sp. represent the highest numbers of intestinal bacteria. 33 Recently, Stecher et al. 34 confirmed that, also in the mouse intestine, Proteobacteria are only present in low numbers. E. faecalis was the only aerobic contaminant observed during routine health monitoring. A flora containing exclusively anaerobic bacteria might leave an ecological niche for a common facultative aerobe like E. faecalis. In order to avoid such contaminations, the association with the CRASF®plus inculding E. faecalis and E. coli might be advantageous.
The present data show that the bacterial strains of the CRASF® do not induce colitis in IL-10−/− mice. The status of our mouse facility allows us to test for specific bacteria responsible for the induction of colitis in IL-10−/− mice. It is known that the severity of the IBD phenotype in IL-10−/− mice is strongly dependent on the intestinal flora: germfree mice do not show intestinal inflammation. 35 In addition, several bacteria induce colitis of distinct characteristics in IL-10−/− mice: mono-association with E. faecalis for instance induces severe colitis. 36 In our setting, E. faecalis is not able to induce intestinal inflammation. This effect might be due to a protective effect of the Lactobacilli present in the CRASF®. It has been demonstrated that Lactobacilli can improve the barrier function of the epithelium, 37,38 ameliorate chemically induced colitis 39 and prevent apoptosis of intestinal epithelial cells. 40
In conclusion, we were able to show that the CRASF® remained stable for four years or seven mouse generations in our mouse facility. Furthermore, the colonization of newly introduced mouse strains via embryo transfer by the foster mothers is efficient. By exclusively importing mouse strains via embryo transfer into a mouse facility equipped with a strict barrier system and IVCs, we were as yet able to avoid contaminations with pathogenic agents listed on the FELASA recommendations. 1 No pathogenic contaminant was detected. The advantage of such a pathogen-free mouse facility is that sensitive mouse strains, such as the IL-10-deficient mouse mutants, can be maintained without apparent phenotype. Moreover, it might provide the basis to understand which bacterial agent is necessary to induce colitis in these animals. Our approach further ensures a stable and comparable enteric flora in all our mouse strains, thereby reducing the variation due to differences in gut flora between mouse strains housed in the same facility.
Footnotes
ACKNOWLEDGEMENTS
We thank André Bleich for performing the cultural and biochemical analysis and Karin Lammert and Martina Hoffmann for invaluable assistance.
M C Greweling was supported by the German Research Council through the GRK705II.
