Abstract
The ability to distinguish among the subspecies of Pasteurella multocida isolates is important epidemiologically; however, classification at the subspecies level based on the results of conventional biochemical tests (fermentation of sorbitol and dulcitol) is reportedly not accurate in all cases. Therefore, we developed a rapid, multiplex PCR assay to differentiate among the 3 subspecies of P. multocida. The PCR assay includes the P. multocida species–specific primers KMT1SP6 and KMT1T7 as an internal amplification control, with a newly designed gatD (galactitol-1-phosphate-5-dehydrogenase)-specific primer pair (unique for subsp. gallicida), and primers targeting a 16S rRNA gene region specific for subsp. septica. The subspecies specificity of the PCR was demonstrated by applying the test to a collection of 70 P. multocida isolates, including the Heddleston serovar reference strains; all isolates and strains were assigned correctly. The PCR assay is a sensitive, specific, and highly effective method for the identification of P. multocida subspecies, and an alternative to biochemical test–based differentiation. A possible relationship was noticed between P. multocida subspecies and lipopolysaccharide (LPS) genotype; all but one of the subsp. gallicida strains were isolated only from avian hosts and represented L1 LPS genotype. Subsp. multocida and subsp. septica isolates were classified into 5 and 4 different LPS genotypes, respectively, of which L3 was the only LPS genotype shared between these 2 subspecies.
Pasteurella multocida is a gram-negative bacterium that can infect many avian and mammalian species causing a variety of diseases. 18 The most common P. multocida–associated diseases are fowl cholera, atrophic rhinitis in pigs, and hemorrhagic septicemia of cattle and buffalo. The organism also causes pneumonia in ruminants, horses, and pigs, and is responsible for the development of pasteurellosis in rabbits. 18 Humans may also be infected, with wound infection injuries (bites or scratches) from pets 9 being the most frequent clinical manifestation of the disease. 18
Within P. multocida, 3 subspecies—P. multocida subsp. multocida, P. multocida subsp. septica, and P. multocida subsp. gallicida—have been distinguished by using DNA-DNA hybridization. 11 The subspecies also differ in their fermentation properties. Isolates of subsp. multocida can utilize sorbitol, but not dulcitol. In contrast, isolates of subsp. septica differ from subsp. multocida by not fermenting sorbitol; subsp. gallicida isolates differ by fermenting dulcitol. 11 Subspeciation of P. multocida strains is important for epidemiologic investigations. In previous studies, subsp. multocida was the subspecies found most frequently in pigs and poultry1,4; isolates of feline and canine origin generally belonged to subsp. septica. 10
Comparative sequence analysis of the 16S rRNA gene and key genes of basic cellular functions support the existence of only 2 taxonomic units within the species, a complex group comprising subsp. multocida/gallicida and subsp. septica.2,10,13,14 This phylogenetic divergence in the 16S rRNA gene sequence allows the identification of subsp. septica with molecular methods. 10 Subspecies classification based on the results of conventional biochemical tests (sorbitol and dulcitol) is reportedly not accurate in all cases,3,10,14,17 and 16S ribosomal RNA gene–based molecular methods are recommended for precise differentiation in addition to biochemical tests. 10 A PCR fingerprinting method has also been used to separate P. multocida subspecies. 5 However, inter-laboratory standardization and reproducibility of PCR-based fingerprinting methods are difficult.
Fermentation of dulcitol occurs only in subsp. gallicida.1,4,11 The genetic background of dulcitol (galactitol) utilization was studied in detail in Salmonella and revealed that the genes responsible for this metabolism are part of a gene cluster that encodes a phosphotransferase system. 12 This galactitol-specific phosphotransferase system was also identified in P. multocida reference strain X73 (subsp. gallicida), but not in P1059 (subsp. multocida). 8 A study found that genes responsible for galactitol metabolism are characteristic only of strains isolated from fowl cholera, and are not carried by strains isolated from other host species and diseases. 7 Another study on P. multocida pangenomic diversity also identified an association between subspecies and genes responsible for sorbitol and dulcitol metabolism. 15 We aimed to design a multiplex PCR scheme based on the detection of key genes associated with the P. multocida subspecies.
We included reference strains representing Heddleston serovars 1–16 (except 15), and a total of 55 field isolates of P. multocida from various host species (Table 1). The isolates were cultured on Columbia agar plates (Becton Dickinson) supplemented with 5% sheep blood under aerobic conditions at 37°C for 24 h. Their identity was confirmed by a species-specific PCR assay. 16 Dulcitol and sorbitol tests were performed as described before.1,4 The lipopolysaccharide (LPS) genotype was determined using the LPS multiplex PCR. 6
Origin of Pasteurella multocida strains and isolates used in our study.
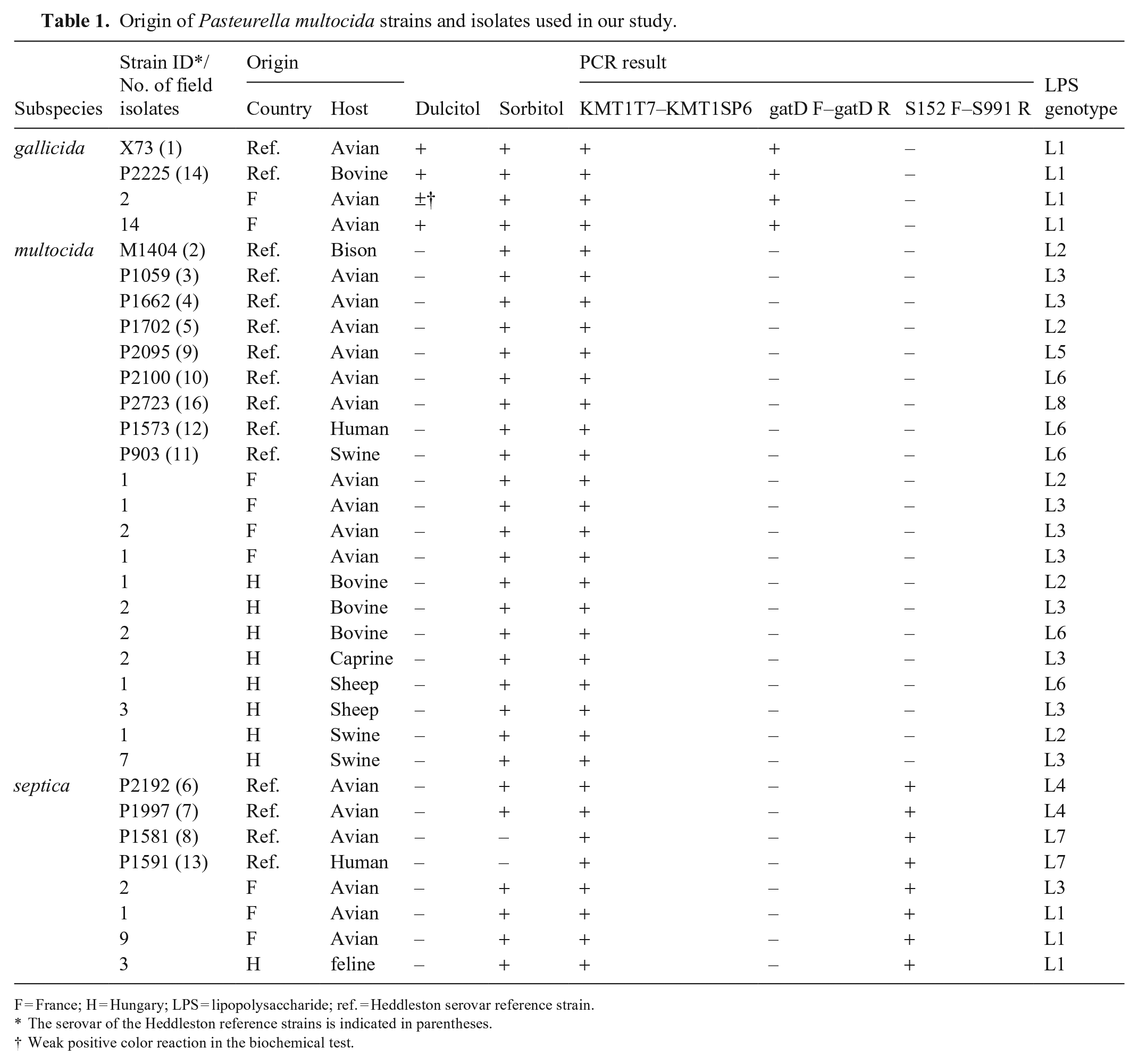
F = France; H = Hungary; LPS = lipopolysaccharide; ref. = Heddleston serovar reference strain.
The serovar of the Heddleston reference strains is indicated in parentheses.
Weak positive color reaction in the biochemical test.
Primers specific for the galactitol-1-phosphate-5-dehydrogenase gene (gatD) were designed (Table 2) from the complete genome sequence of X73 reference strain (GenBank NZ_CM001580). Primers specific for the 16S rRNA gene of P. multocida subsp. septica were designed from the sequences of reference strains available in GenBank (AF294410–AF294423). The primers were synthesized by Sigma-Aldrich. Primers gatD F and gatD R were designed to produce a 527-bp PCR fragment from the gatD gene, whereas primers S152 F and S991 R amplified an 840-bp fragment from the 16S rRNA gene of subsp. septica isolates. Amplicon sizes were calculated from the DNA sequences of subsp. gallicida and subsp. septica reference strains.
Sequences of the oligonucleotides used in the Pasteurella multocida subspecies multiplex PCR assay.
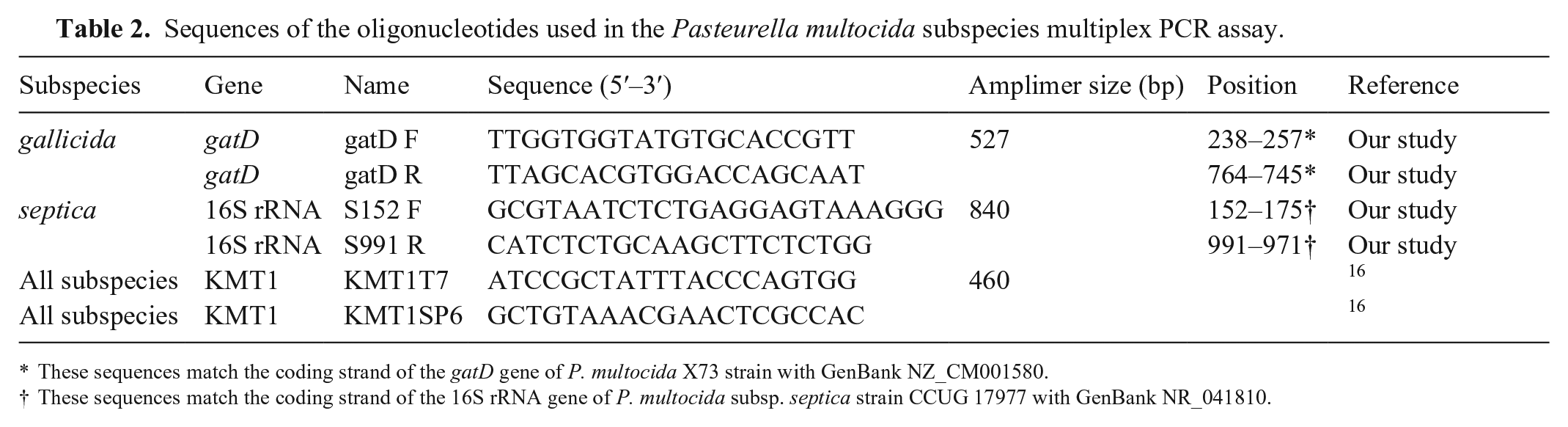

These sequences match the coding strand of the gatD gene of P. multocida X73 strain with GenBank NZ_CM001580.
These sequences match the coding strand of the 16S rRNA gene of P. multocida subsp. septica strain CCUG 17977 with GenBank NR_041810.
Lysates of pure cultures were prepared for all strains used in our study. A loopful of cultured bacterial growth was suspended in 100 µL of PCR-grade water (VWR) and heated in a thermocycler for 20 min at 99°C. Cellular debris was pelleted by centrifugation, and the supernatant was used as DNA template for PCR amplification. The lysates were stored at –20°C until use.
PCR was performed in a total volume of 25 µL containing 100 ng of template DNA, 2.5 µL of 10× DreamTaq buffer, 1.2 µL of 25 mM MgCl2, 0.5 µL of 10 mM dNTPs, 1 µL of forward and reverse primers (10 pmol/µL), and 1 U of DreamTaq polymerase (Thermo Scientific). Reactions were performed in a thermocycler (C1000; Bio-Rad). DNA was amplified for 25 cycles of denaturation at 95°C for 30 s, annealing at 58°C for 30 s, and extension at 72°C for 1 min; an initial denaturation step (95°C, 10 min) and a final extension step (72°C, 7 min) were also performed. Each reaction mixture was analyzed by electrophoresis in 1.5% agarose gel (SeaKem; FMC) stained with GelRed (Biotium) and visualized under UV light. The results were confirmed by sequencing of PCR products. Sequencing was performed by Macrogen Europe. Nucleotide sequences were analyzed using Geneious Prime software (v.2020.2.4, https://www.geneious.com) and confirmed by BLAST (https://blast.ncbi.nlm.nih.gov/Blast.cgi). The GenBank accessions for sequences obtained in our study are MZ476034–MZ476037.
Following sequence determination and analysis, a multiplex PCR assay was developed that contained the P. multocida species–specific primers KMT1SP6 and KMT1T7 as an internal amplification control, 16 with the newly designed gatD gene–specific primer pair (unique for subsp. gallicida), and S152 F and S991 R primers (targeting 16S rRNA gene regions specific for subsp. septica). Amplification and electrophoresis conditions were the same as described above. Amplimer lengths allowed clear size discrimination (Fig. 1).
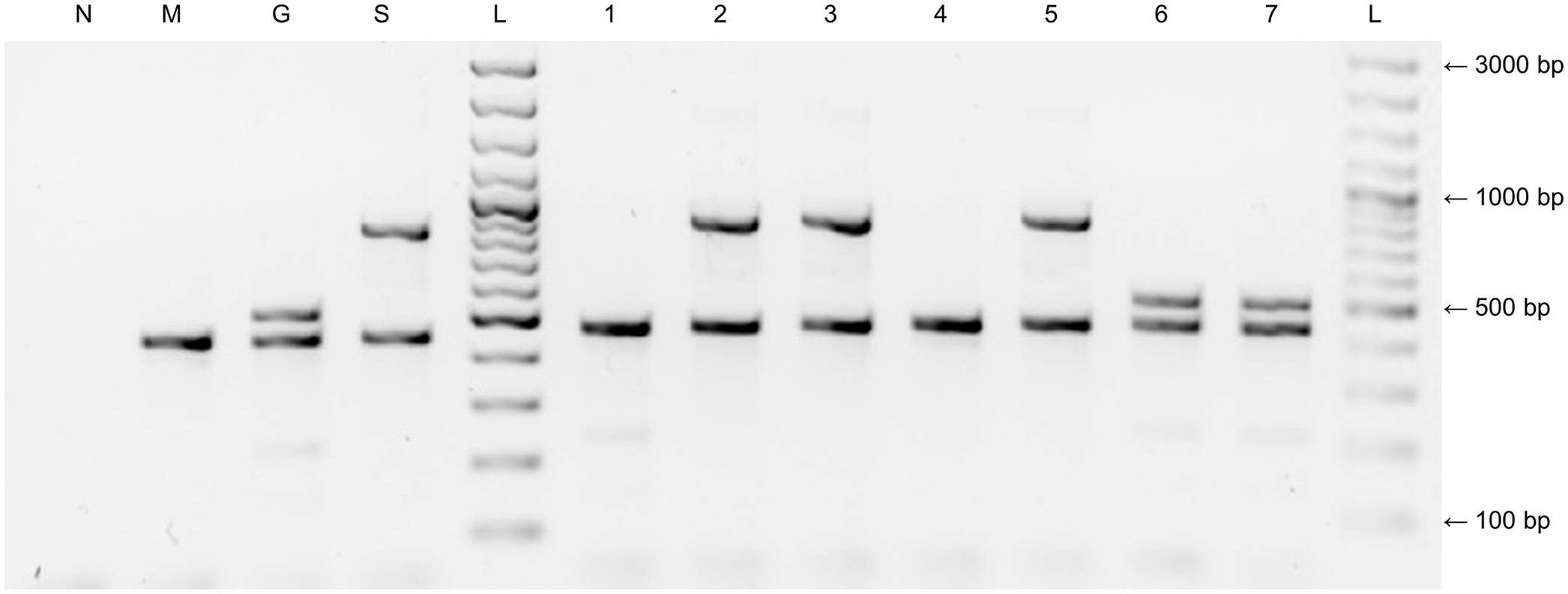
Multiplex subspecies PCR typing system for Pasteurella multocida.
To test the sensitivity of our assay, 10-fold dilutions of genomic DNA (106–100) of reference strains X73 (subsp. gallicida) and P1581 (subsp. septica) were used. DNA was quantified (NanoDrop spectrophotometer; Thermo Scientific), and template copy numbers per reaction were calculated using an online calculator (http://cels.uri.edu/gsc/cndna.html). The subspecies specificity of the PCR assay was determined by applying the test to a collection of 70 P. multocida isolates, including 15 of the 16 Heddleston serovar reference strains. For all strains or isolates belonging to subsp. gallicida or subsp. septica, a subspecies-specific fragment of the expected size and a 460-bp species-specific fragment were amplified. For all strains or isolates of subsp. multocida, only the species-specific fragment was produced (Fig. 1). Sensitivities of the assays were 105 genomic equivalents per reaction (data not shown).
All P. multocida field isolates typed as subsp. gallicida in the biochemical tests (dulcitol and sorbitol positive) were allocated to the corresponding subspecies by the PCR test. In accordance with previous studies,2,10,13,14 the ability to utilize sorbitol was found among strains belonging to subsp. septica. Thus, we confirmed the proposition that the 16S rRNA gene could be used to identify subsp. septica instead of sorbitol and dulcitol tests. Isolates belonging to subsp. multocida uniformly gave their characteristic phenotypic reactions (fermentation of sorbitol, but not dulcitol). Almost all subsp. gallicida strains had been isolated from avian hosts (with the only exception of P2225, the serovar 14 reference strain of bovine origin) and classified exclusively into L1 LPS outer core biosynthesis loci. On the other hand, subsp. multocida and subsp. septica isolates were classified into 5 and 4 different LPS genotypes, respectively. Interestingly, L3 was the only LPS genotype shared between the latter 2 subspecies.
The identification of P. multocida subspecies has been the subject of research for decades. Our results indicate that our developed PCR test is a sensitive, specific, and highly effective method for the differentiation of P. multocida subspecies. Problems related to the traditional biochemical tests (i.e., the existence of sorbitol-positive subsp. septica strains reported by others10,17) are avoided. This method could be an alternative to subspecies differentiation by biochemical tests in epidemiologic investigations.
Footnotes
Acknowledgements
The outstanding technical assistance of Éva Hegedűs and Katalin Oryszcsák is highly appreciated. We thank Dr. A. J. Lax for useful comments on the manuscript.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Our study was supported by the National Research, Development and Innovation Office (grant NKFIH K124457).
