Abstract
Cell-free fetal DNA (cffDNA) is a highly accurate screening test for major aneuploidies. Despite high detection rates and low false-positive rates compared to current screening methods, cffDNA testing is approved for use in only the high-risk population and is expensive for low-risk women because of a lack of insurance coverage. As the technology improves and research demonstrates the efficacy of cffDNA testing in the general population, insurance coverage of this test likely will be extended to all women. At that time, cffDNA testing in most cases will replace current screening methods. Furthermore, cffDNA testing may become a routine diagnostic test, effectively rendering most invasive diagnostic procedures obsolete.
Chromosomal abnormalities (aneuploidies) occur in 0.1% to 0.2% of live births, the most common among live-born infants being Down syndrome (trisomy 21), occurring in approximately 1 of every 690 live births. Other sonographically detectable aneuploidies include trisomy 18 (Edwards syndrome, 1 out of 6000 live births), trisomy 13 (Patau syndrome, 1 out of 20 000 live births), and monosomy X (Turner syndrome, 1 out of 2500 live female births). 1 Prenatal diagnosis of aneuploidy is traditionally achieved by either chorionic villus sampling or amniocentesis, as both methods have very high detection rates (99.25% for chorionic villus sampling and 99.4% for amniocentesis). However, chorionic villus sampling and amniocentesis carry an associated risk of spontaneous abortion of approximately 0.5% to 1%. 2 Because of the potential complications associated with these procedures, researchers and obstetricians have long sought noninvasive screening methods that would assess a woman’s risk of carrying a fetus with one or more chromosomal disorders, effectively reducing the number of women requiring invasive testing.
Three major screening methods currently used in obstetrical practice include the quad screen, combined/first-trimester screen, and the sequential/integrated screen. The combined screen measures maternal age, presence or absence of nasal bone, nuchal translucency, free β-human chorionic gonadotropin, and pregnancy-associated plasma protein A in the first trimester to calculate the risk of aneuploidy. The quad screen measures maternal age along with the four markers α-fetoprotein, estriol, dimeric inhibin A, and β-human chorionic gonadotropin in the second trimester to calculate the risk of aneuploidy. The sequential/integrated screen calculates a patient’s individualized risk based on the combination of both the first-trimester (combined) and the second-trimester (quad) screenings. While these screening methods have high detection rates (78%−95%), the screening tests are also associated with false-positive rates between 5% and 11%.2–8 Furthermore, these screening tests can be done within only a limited gestational age window. For example, quad screening can be performed only between approximately 15 and 20 weeks of gestation. 9
Ideally, a noninvasive diagnostic test would have a much lower false-positive rate. Initial research for such a test focused on the isolation of intact fetal nucleated cells in maternal blood for the prenatal diagnosis of aneuploidies. These cells were not ideal for use in prenatal diagnosis, as concentrations of such cells are very low, typically of the order of one or a few fetal nucleated cells per milliliter of maternal blood. Consequently, prenatal testing carried out with circulating fetal cells has been found to have a relatively low sensitivity and specificity.4,10 In addition, fetal cells remain in maternal circulation long after delivery, making it impossible to distinguish new fetal cells from those of prior pregnancies. In 1997, Lo and colleagues reported the presence of cell-free fetal DNA (cffDNA) in maternal plasma and serum, indicating its potential for use in prenatal diagnosis of aneuploidies. 11 This review article discusses the benefits and limitations of aneuploidy testing based on cffDNA.
What Is cffDNA?
Circulating cffDNA makes up 3% to 10% of the total maternal cell-free DNA. It is derived from placental cells and is cleared from the maternal blood within hours after childbirth. 10 Fetal cells enter the maternal circulation where they undergo apoptosis, the normal process of cell death and elimination. This is a necessary step in the development of fetal tissues and organs. In the process, fetal DNA is cleaved into small 150– to 200–base pair fragments. These cffDNA fragments can be detected as early as the fourth week of gestation and reliably after the seventh week of gestation.4,10
Validation of Aneuploidy Detection With cffDNA
Studies have shown the ability of cffDNA to detect aneuploidy with a high sensitivity and specificity (Table 1). Since the discovery of cffDNA, researchers have used various methods to isolate and analyze fetal DNA in a maternal blood sample. The first method utilized was massive parallel sequencing, a technique that utilizes the amplification of all fetal DNA molecules in maternal plasma. Individual sequence tags are aligned to the human genome to determine its chromosome of origin. 10 A known euploid reference sample can be used to calculate the percentage of each chromosome represented in the sample. If the percentage for a particular chromosome is significantly higher in the fetal sample as compared to the euploid sample, the fetal sample is classified as having a trisomy for that chromosome. For example, trisomy 21 makes up about 1.5% of a euploid sample. For a fetal sample to be classified as trisomy 21, the percentage of chromosome 21 would show an increase from 1.5% to approximately 1.575%. 4
Testing Sensitivity and Specificity for T21, T18, T13, and MX a .
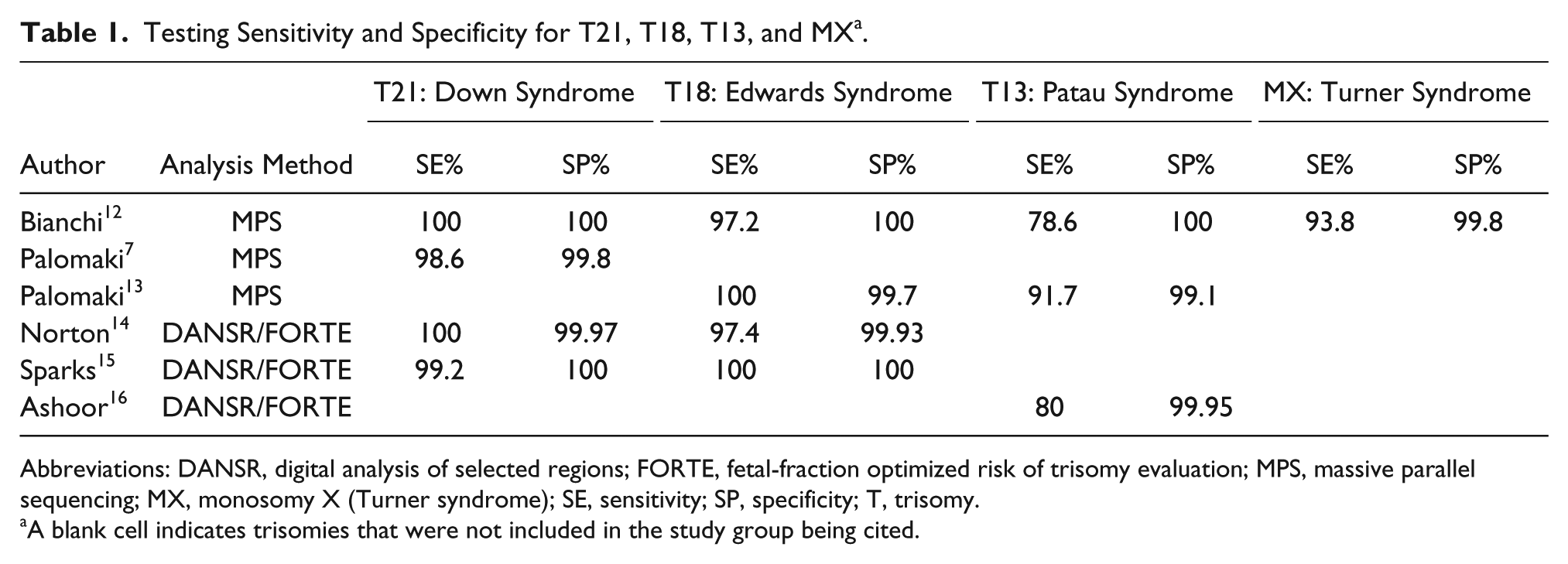
Abbreviations: DANSR, digital analysis of selected regions; FORTE, fetal-fraction optimized risk of trisomy evaluation; MPS, massive parallel sequencing; MX, monosomy X (Turner syndrome); SE, sensitivity; SP, specificity; T, trisomy.
A blank cell indicates trisomies that were not included in the study group being cited.
In recent years, researchers have looked for more cost-effective methods to isolate and analyze cffDNA. This has been achieved to a certain extent by targeting and amplifying only the genes of interest in the sample. One method of targeted sequencing is digital analysis of selected regions, which selectively evaluates specific genomic fragments from cffDNA and is used in combination with an algorithm called fetal-fraction optimized risk of trisomy—a mathematical formula that calculates the risk of trisomy for each chromosome studied in the sample. 15 While targeted sequencing appears to be an effective method to detect aneuploidy, its primary disadvantage is the inability to rule out chromosomal conditions that have not been selected for. 4
Commercial Applications
Clinical research validation studies showed that cffDNA could be offered in obstetrical practice. In October 2011, Sequenom (San Diego, CA) offered the first commercially available prenatal diagnostic test based on cffDNA (MaterniT21), which detected trisomy 21. Sequenom also currently offers testing for trisomy 13 and 18 under the name MaterniT21 Plus.17,18 Three other companies now offer aneuploidy testing (Verinata Health, Redwood City, CA; Ariosa Diagnostics, San Jose, CA; and Natera, San Carlos, CA), and all these companies offer testing to determine the sex of the fetus as well (Table 2).
Companies That Offer Free-Fetal DNA Testing.
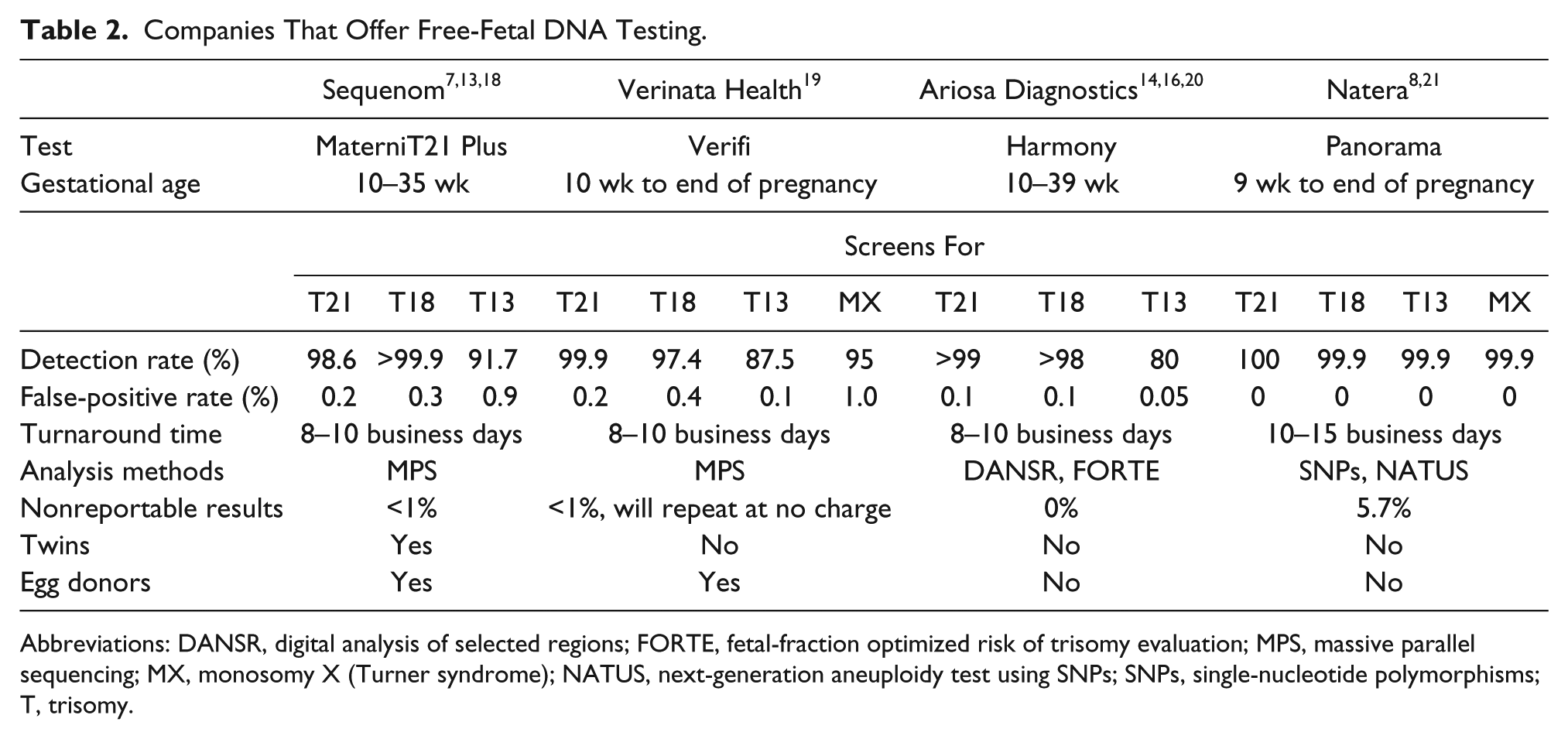
Abbreviations: DANSR, digital analysis of selected regions; FORTE, fetal-fraction optimized risk of trisomy evaluation; MPS, massive parallel sequencing; MX, monosomy X (Turner syndrome); NATUS, next-generation aneuploidy test using SNPs; SNPs, single-nucleotide polymorphisms; T, trisomy.
Natera was the latest of the four companies to offer prenatal cffDNA testing. This company offers testing based on a new method of DNA analysis called NATUS—next-generation aneuploidy test using single-nucleotide polymorphisms—a mathematical calculation that determines the number of copies and the accuracy of detecting each chromosome studied in a tested sample of DNA. Single-nucleotide polymorphisms are variations in DNA among the human population that can be used to distinguish fetal from maternal DNA. A proof-of-concept study showed that NATUS can detect trisomy 18, trisomy 13, and monosomy X with an accuracy of greater than 99.9%. In addition, it can detect trisomy 21 with 100% sensitivity and specificity. 8
Application to Obstetrical Practice
As most of the research to date has focused on a selective group of high-risk women, both the National Society of Genetic Counselors and the American Congress of Obstetricians and Gynecologists recommend cffDNA testing only as a highly specific screening test and recommend a follow-up diagnostic test for any positive screen results. Accordingly, further studies showing the efficacy of cffDNA testing in the general population (low-risk women) must be carried out to facilitate universal third-party pay or coverage for these tests. High-risk women who are currently recommended for cffDNA screening include women older than 35 years, women having a prior child affected by a trisomy, women with a positive serum screening result, and/or women with prior sonograms indicating that their fetuses had increased risk of a chromosomal abnormality.22,23
According to Palomaki and colleagues, the use of the cffDNA screening test following a screen-positive result from traditional screening methods has the potential to reduce invasive diagnosis-related fetal losses in the United States by approximately 95%. This can be attributed to higher false-positive rates for traditional screening methods compared to cffDNA screening. Women with a false positive by traditional screening would receive reassurance due to the low risk of carrying a fetus with aneuploidy given a negative cffDNA result. 7
cffDNA is also an option following a sonographic finding indicating that the fetus has one or more “soft markers” for aneuploidy. The presence of one or more of these markers increases the risk of aneuploidy. Soft markers detectable during the first trimester scan (gestational age, 11–14 weeks) include the absence of the nasal bone (Figure 1) and an increased nuchal lucency. The major soft signs that can be detected in the second trimester are an increased nuchal fold thickness (nuchal edema), choroid plexus cysts, an echogenic focus or foci in the heart, echogenic bowel, and mild renal pyelectasis (Figures 2–7). In addition, shortening of the femur and humerus is considered a soft sign that increases the risk of trisomy 21, trisomy 18, and Turner syndrome. 1
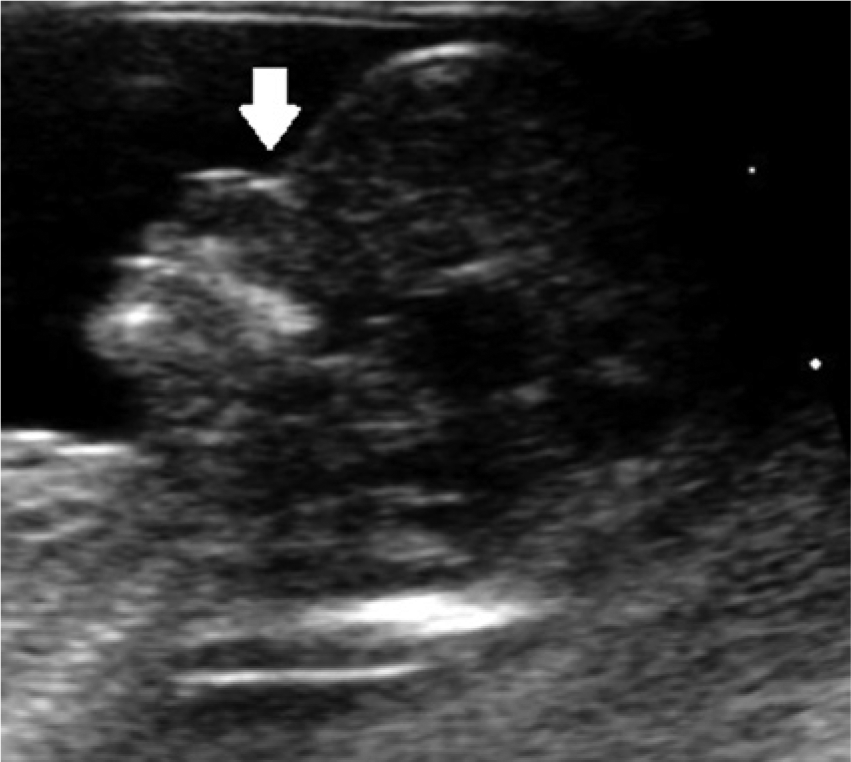
Midline sagittal image of the fetal head in the first trimester showing an absent nasal bone, an early “soft” marker of aneuploidy.
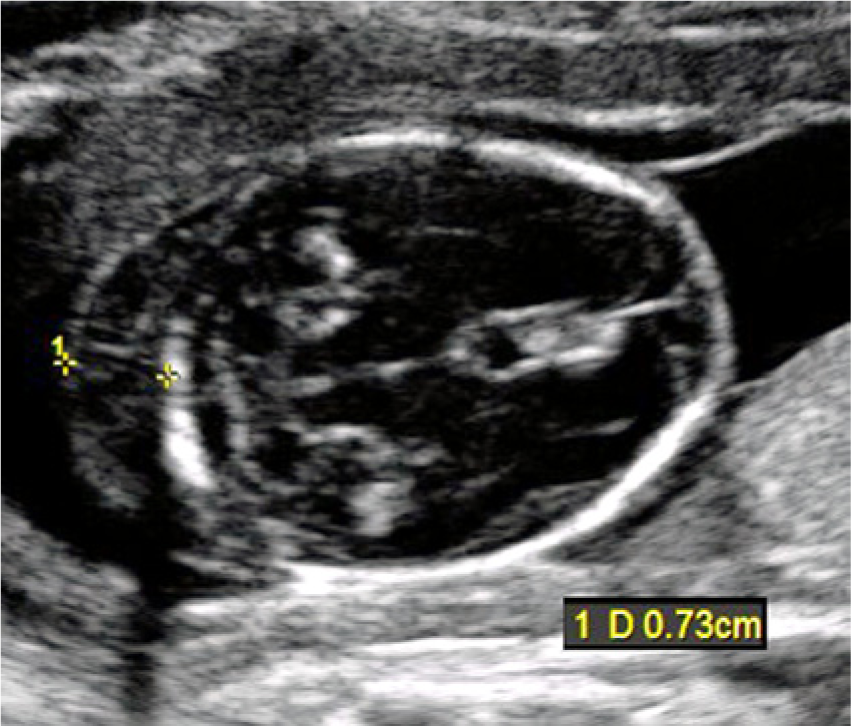
Axial image of the fetal head in the second trimester showing thickening of the nuchal fold to 7.3 mm (> 6 mm), a predictive sonographic marker for aneuploidy. The anatomic markers of cerebellum, cisterna magna, and occipital bone are all seen.
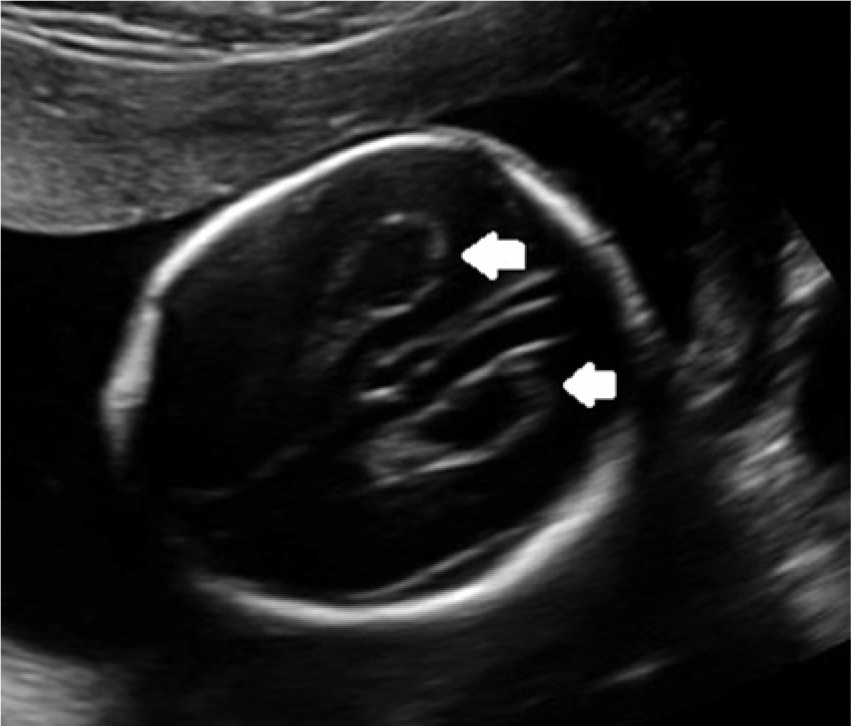
Axial image of the fetal head during the second trimester showing bilateral choroid plexus cysts (arrows).
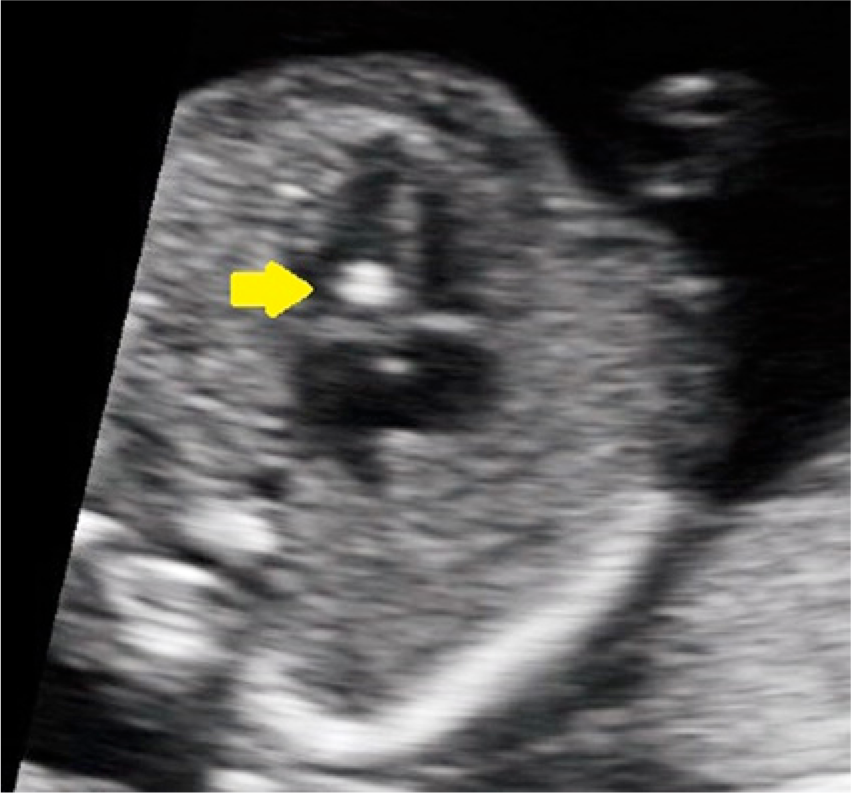
Four-chamber view of the fetal heart during the second trimester showing an echogenic intracardiac focus (arrow).
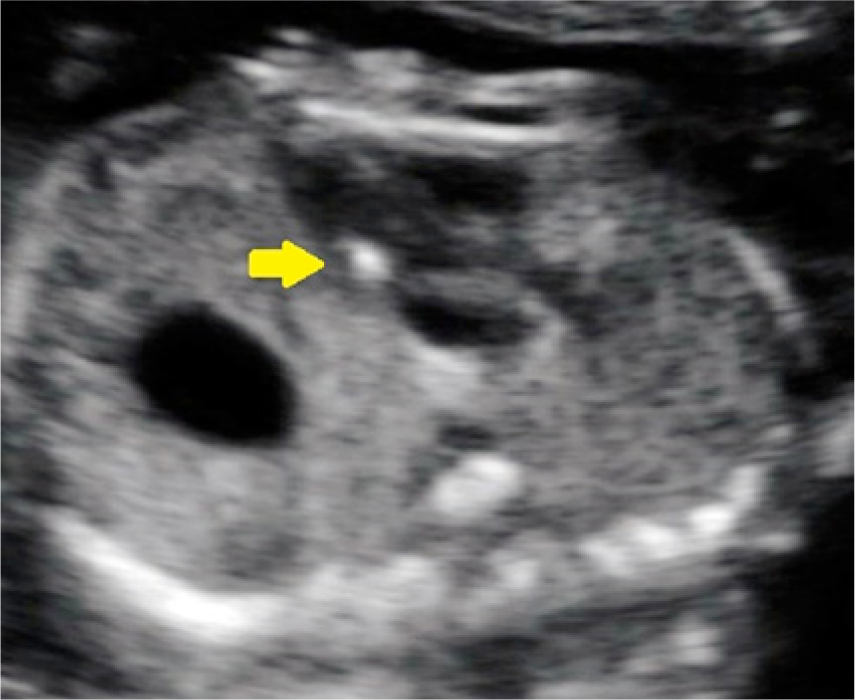
Left outflow tract view of the heart during the second trimester showing an echogenic intracardiac focus (arrow).
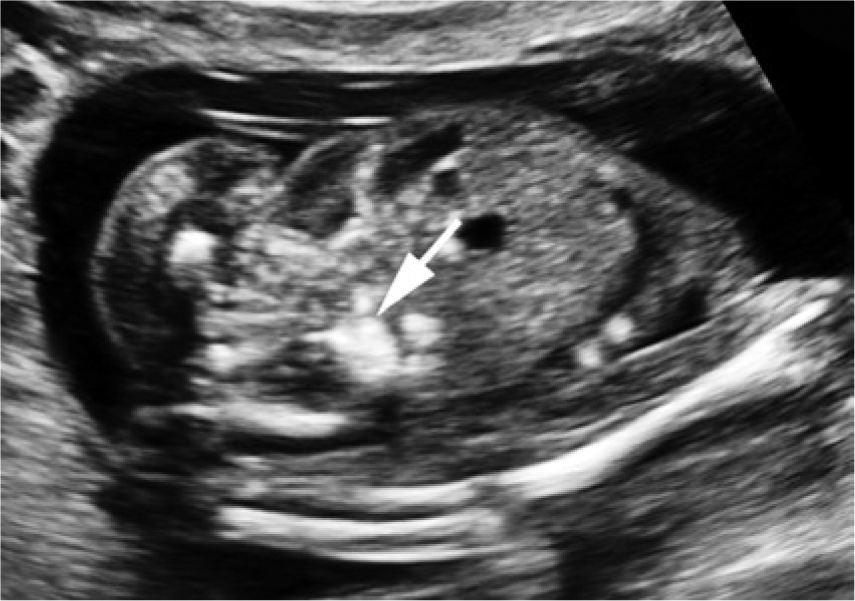
Coronal view of the fetal abdomen during the second trimester showing echogenic bowel (arrow). Note that the bowel is isoechoic or hyperechoic relative to the fetal bone.
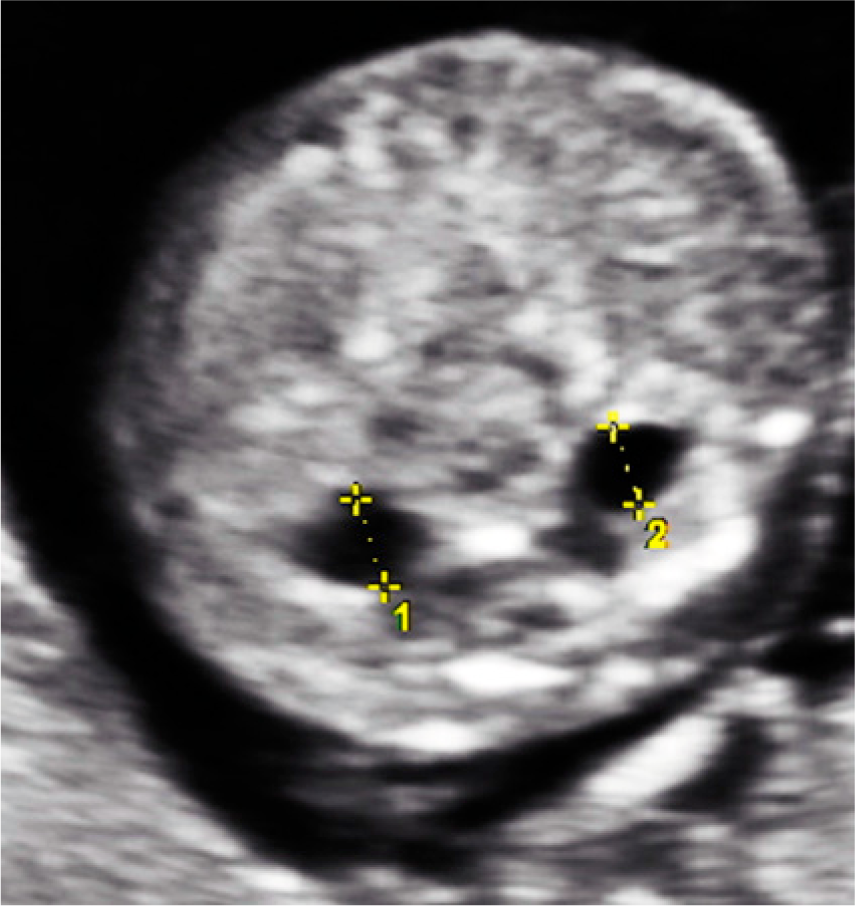
Cross-sectional image of the fetal abdomen during the second trimester showing bilateral dilation of the renal pelvis (pyelectasis).
These soft markers may be seen in the normal fetus but have an increased incidence in infants with chromosomal abnormalities. A negative cffDNA result would most likely indicate that these soft markers have no clinical significance. A positive cffDNA result would still require invasive testing for definitive diagnosis of aneuploidy.
Conclusion / Future Directions
cffDNA is currently used in obstetrical practice as an effective screening test for major aneuploidies. However, cffDNA testing is still quite expensive, and most insurance companies will not cover this test in the general (low-risk) population. As the technology used to isolate cffDNA improves, cffDNA testing will become less expensive and faster. 10 In addition, studies to assess clinical validity in the general population are currently underway. As these studies are completed and reported, it is likely that their efficacy will be demonstrated and insurance companies will cover this test for all women. 22 In the future, cffDNA testing could very well replace conventional screening methods due to its comparatively high aneuploidy detection rates and low false-positive rates.
Studies indicate that there is promise in the near future for detecting sex-linked aneuploidies. Bianchi et al used massive parallel sequencing to correctly identify 8 of 10 aneuploidies for karyotypes 47,XXX, 47,XXY, and 47,XYY. 12 A study submitted for publication in 2013 by Sequenom indicates a high specificity and sensitivity for detection of these sex aneuploidies. 18 A proof-of-concept study by Lo has shown that a genomewide genetic and mutational map of a fetus can be constructed from the maternal plasma DNA-sequencing data. 10 These studies demonstrate the possibility that cffDNA testing will become a routine diagnostic test for all fetal chromosomal abnormalities, effectively replacing amniocentesis and chorionic villus sampling.
Footnotes
Acknowledgements
I thank Bryann Bromley, MD, for obtaining the sonographic images presented in this article and for reviewing the article. I also thank Patricia Bullock, RDMS, the genetic counselors, and the staff of the Ultrasound and Prenatal Diagnostics Center at Massachusetts General Hospital for assisting in the preparation of this review article. Special thanks to Jane Morrison, RDMS and Nada Almoujahed, MD, RDMS, professors at Middlesex Community College in Bedford, Massachusetts, for encouraging me to submit this article.
Declaration of Conflicting Interests
The author declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author received no financial support for the research, authorship, and/or publication of this article.
