Abstract
Objective
This study aimed to comprehensively analyze the detection capacity of non-invasive prenatal testing (NIPT) for chromosomal abnormalities of all 24 chromosomes, as well as high-risk indications for pregnancy and the fetal fraction, in a large cohort.
Methods
We retrospectively enrolled 118,969 pregnant women who underwent NIPT at Sichuan Provincial Maternity and Child Health Care Hospital from March 2019 to June 2022. The sensitivity, specificity, positive predictive value, negative predictive value, and positive chromosomal abnormality rate were calculated. The fetal fraction based on gestational age, maternal body mass index, and number was examined.
Results
NIPT demonstrated > 99% sensitivity and specificity for almost all of the common trisomies (T21, T18, and T13), sex chromosomal aneuploidies, rare autosomal trisomies, and microdeletion/microduplication syndromes. Positive predictive values varied from 12.0% to 89.6%. Advanced maternal age was associated with an increased risk of three major aneuploidies. The fetal fraction was positively correlated with gestational age and negatively correlated with the maternal body mass index.
Conclusions
NIPT can be used to effectively screen for chromosomal abnormalities across all 24 chromosomes. Advanced maternal age is a risk factor for high-risk pregnancy, and careful consideration of the fetal fraction is essential during NIPT.
Keywords
Introduction
Chromosomal abnormalities such as aneuploidies are common causes of genetic disorders. 1 Aneuploidies typically result in embryo growth arrest, implantation failure, early miscarriage, congenital abnormalities, or perinatal death.2,3 To diagnose chromosomal abnormalities, invasive amniocentesis or chorionic villus sampling has been used.3,4 However, these procedures carry the risk of procedure-related miscarriage and infection.
The discovery of fetal DNA in the cell-free plasma of pregnant women and the development of next-generation sequencing methods have enabled more accurate screening of fetal chromosomal abnormalities than traditional screening methods, such as biochemical analytes and nuchal translucency. 5 This circulating cell-free DNA, primarily derived from apoptotic placental trophoblasts, comprises 3% to 20% of DNA fragments in maternal blood during pregnancy. 5 Non-invasive prenatal testing (NIPT) for fetal aneuploidies using cell-free DNA has been widely adopted in clinical practice because of its greater sensitivity and specificity than maternal serum screening. While the detection efficiency of NIPT for trisomy 21 (T21), T18, and T13 has been widely studied overseas and in some regions of China, it has not been fully investigated in large cohorts, especially for other chromosomal abnormalities. These chromosomal abnormalities include sex chromosomal aneuploidy (SCA), rare autosomal trisomy (RAT), and microdeletion/microduplication syndrome (MMS). Additionally, the efficiency of NIPT for these chromosomal syndromes in people with a general risk and specific-risk pregnancies still needs to be validated and analyzed to guide its clinical application.
This study aimed to evaluate the performance of NIPT for common trisomies (T21, T18, and T13), SCAs, RATs, and MMSs in pregnant women in relation to various indicators (e.g., advanced maternal age, serum screening of high risk or borderline risk of fetal aneuploidies, abnormal ultrasound findings, and in vitro fertilization and embryo transfer [IVF-ET]) in Sichuan located in southwest China. We also aimed to investigate variation in the fetal fraction along with different clinical characteristics to highlight its importance in NIPT. Additionally, we aimed to analyze false positives in NIPT to identify areas for technological improvement. Overall, our goal was to gain insight into the clinical application of NIPT in a large cohort.
Methods
Patients
Between March 2019 and July 2022, we retrospectively recruited 118,969 pregnant women who underwent routine obstetrical care at Sichuan Provincial Maternity and Child Health Care Hospital. The study was approved by the institutional review board of Sichuan Provincial Maternity and Child Health Care Hospital (No. 20240607-220). All procedures were performed in accordance with relevant regulations and guidelines. We obtained written informed consent from all of the participants for testing and anonymized research. Qualified professionals ensured that the participants fully understood the consent form before signing. All experiments were conducted in the prenatal testing laboratory of Sichuan Provincial Maternity and Child Health Care Hospital. The patients details have been de-identified. The reporting of this study conforms to the STARD guidelines. 6
Assay optimization for NIPT
A 10-mL peripheral venous blood sample was drawn using an EDTA blood collection tube. Within 6 hours of collection, the blood samples were processed using two-step centrifugation (1,600 × g for 10 minutes and 16,000 × g for 10 minutes at room temperature). Cell-free DNA was extracted with the MagPure Circulating DNA Midi Kit (Magen, Guangzhou, China). Libraries were constructed and quantified using the Rapid CM DNA Lib Prep Kit Set (BGI; Shenzhen, China) and sequenced on the MGI-2000 platform (MGI Tech; Shenzhen, China) to generate approximately 10M 50-base pair single-end reads.
Polymerase chain reaction duplicates were removed from the sequencing reads. A minimum of 70% of the reads were required to achieve Q30 quality. Reads containing more than five “N” bases were removed. Subsequently, the reads were mapped to the human reference genome hg19, allowing for at most one mismatch, and allocated to continuous 100-kb bins with 50-kb overlapping. Bins with abnormal GC content (<30% or >70%) and low coverage were filtered out. Coverage from each bin was counted and normalized. GC bias was corrected using a smooth spline method and weighed linear regression as previously described. 7 Maternal copy number variations (CNVs) were removed using the hidden Markov model to enhance the detection of fetal chromosomal abnormalities. Aneuploidies were detected by comparing them with the reference genome using the Z-test method. The fetal fraction of male fetuses was determined using reads from the Y chromosome, while that of female fetuses was calculated on the basis of single nucleotide polymorphism heterozygosity as previously reported. 8 The resolution of the NIPT technique in this study was 2 Mb for maternal CNV and 10 Mb for fetal CNV.
Follow-up and NIPT performance assessment
In our study, all patients who were positive for NIPT were advised to have a prenatal diagnosis performed, which served as the gold standard for evaluating the performance of NIPT. In positive cases of MMS in NIPT, we recommended additional diagnostic tests, such as chromosomal microarray and CNV-sequencing for prenatal confirmation. Patients who were negative for NIPT received routine prenatal care. A subsequent ultrasound assessment, clinical examination, and postnatal follow-up were performed to confirm negative NIPT.
Statistical analysis
We calculated the sensitivity, specificity, and positive predictive value (PPV) to assess the performance of NIPT. We compared continuous variables between different groups using the Wilcox Rank Sum test. We generated plots using R (version 4.2.2) software (www.r-project.org).
Results
Clinical characteristics of the pregnant women
A total of 118,969 pregnant women underwent NIPT (Figure 1). Their median age was 29 years old (range: 14–52 years old). The median gestational age was 17 weeks (range: 11–33 weeks). The enrolled pregnant women had a median body mass index (BMI) of 22.16 kg/m2 (range: 13.20–53.91 kg/m2). Most pregnancies were spontaneous (94.13%, 111,980/118,969) and singleton (96.77%, 115,121/118,969). Among the 6,989 IVF-ET pregnancies, 60.81% (4,250) were singletons and 39.19% (2,739) were twins. Among the whole cohort, 6.7% (7,935) of all pregnant women had an abnormal ultrasound soft index (ultrasound findings generally did not represent structural abnormalities, but indicated an increased age-adjusted risk of underlying fetal aneuploidy or certain non-chromosomal abnormalities), 4.4% (5,239) were at high risk and 13.1% (15,572) were at borderline risk in serum screening, and 10.14% (12,069) had an advanced maternal age without other high-risk indicators. Furthermore, 59.8% (71,145/118,969) of pregnant women voluntarily requested NIPT despite lacking specific clinical indicators.
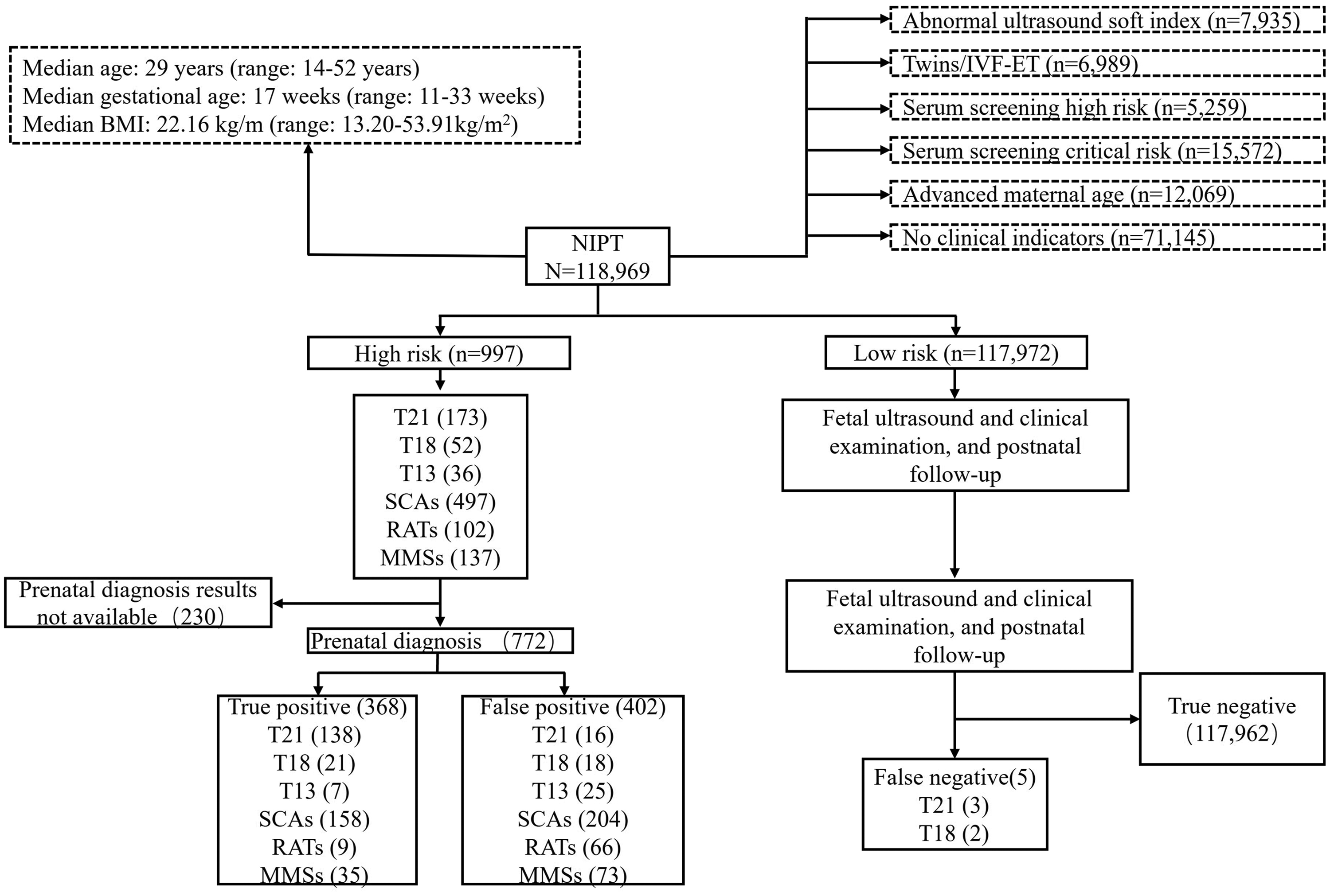
Results of NIPT and prenatal diagnosis for 118,969 pregnant women. BMI, body mass index; IVF-ET, in vitro fertilization and embryo transfer; NIPT, non-invasive prenatal testing; T21, trisomy 21; T18, trisomy 18; T13, trisomy 13; SCAs, sex chromosomal aneuploidies; RATs, rare autosomal trisomies; MMS, microdeletion/microduplication syndromes.
Performance of NIPT for common trisomies and SCAs
Among the 118,969 patients, NIPT detected T21 in 173, T18 in 52, and T13 in 36 (Figure 1 and Table 1). Prenatal diagnosis results were available for 772 pregnant women, and they confirmed 138 with T21, 21 with T18, and 7 with T13 as true positives. There were five false negative samples including three with T21 and two with T18. The sensitivity, specificity, and PPV for T21, T18, and T13 are shown in Table 1. Regarding SCAs, NIPT detected 497 positive cases, with prenatal diagnosis results available for 362. Of these, 158 were confirmed as true positives: 42 (26.58%) were Turner syndrome (45, X), 27 (17.09%) were triple X syndrome (47, XXX), 32 (20.25%) were Jacob syndrome (47, XYY), 56 (35.44%) were Klinefelter syndrome (47, XXY), and one was 48, XXYY. Among the false positives, 110 were 45, X, 28 were 47, XXX, 7 were 47, XYY, and 59 were 47, XXY. The overall PPV, sensitivity, and specificity for SCAs were 42.65%, 100%, and 99.83%, respectively.
Performance parameters of NIPT for detecting chromosomal abnormalities in 118,969 pregnancies.

TP, true positive; FP, false positive; TN, true negative; FN, false negative; PPV, positive predictive value; T21, trisomy 21; T18, trisomy 18; T13, trisomy 13; SCAs, sex chromosomal aneuploidies; RATs, rare autosomal trisomies; MMSs, microdeletion/microduplication syndromes.
NIPT for RATs and mosaic interference
In our study, NIPT detected 102 cases of RATs. Supplementary Figure 1 shows a bar graph of the absolute number of RATs detected. The most frequently identified RATs in this cohort were trisomies 22, 20, 15, 16, and 9, in descending order.
Prenatal diagnosis results were available for 75 of the 102 cases of RATs. Among these, nine cases were confirmed as true positives: three were T9, three were T16, two were T15, and one was T2. The PPV, sensitivity, and specificity of all RATs are shown in Figure 1 and Table 1. Of the true positives, six were confirmed as mosaicism by prenatal diagnosis, while the mosaicism status for the other three was unavailable.
We then assessed the accuracy of NIPT detection for RATs regarding the mosaicism status in the nine true positive cases. We used the trisomic fraction and fetal fraction from NIPT data to calculate the mosaic fraction. The trisomic fraction refers to the proportion of cell-free fetal DNA that indicates the presence of an extra chromosome compared with the expected amount in a typical euploid (normal) sample. Ideally, the trisomic fraction is approximately equivalent to the fetal fraction. By comparing this fraction with the fetal fraction, we can evaluate potential mosaicism. Supplementary Figure S2 shows a dot plot of both fractions to indicate the mosaic status of these true positive RATs. We found that in most cases, the trisomic fraction was notably lower than the approximated fetal fraction (Supplementary Figure 2). Specifically, in six of nine pregnancies with true positive RAT results, the trisomic fraction was less than 80% of the fetal fraction. The mosaic cases calculated from NIPT data were consistent with the prenatal diagnosis.
Evaluation of NIPT for MMSs
We also assessed the performance of NIPT in identifying MMSs. We selectively reported high-confidence MMS cases for clinical reference, including those with maternal inheritance or a strong fetal mutation signal. NIPT predicted 137 cases of MMS across the whole genome, of which 35 were confirmed as true positives by prenatal diagnosis and 73 as false positives (Table 1 and Figure 1). Prenatal diagnosis results were unavailable for the remaining 29 samples. The PPV, sensitivity, and specificity of MMSs are shown in Table 1.
Among the 35 true positive MMS cases, 30 were maternally inherited, and the remaining 5 originated from the fetuses, including 3 samples with a fetal fraction >10% and 2 with large fragments (>10 Mb). Of the true positive MMS cases, 20 involved 22q11 microduplication syndrome, and 3 involved 22q11 microdeletion syndrome, which meant that they were the most common. These cases were not necessarily pathogenic because of the variable penetrance of 22q11 microduplication syndrome. Our results are consistent with the prevalence of 22q11 microdeletion syndrome as the most common MMS.
Chromosomal abnormalities within populations with specific clinical indicators
A proportion of pregnant women in our study had high-risk pregnancy indicators. Therefore, we analyzed the positive rates of different chromosomal abnormalities in subpopulations in relation to different clinical indicators (Table 2). The positive rate represents the proportion of true positives confirmed through prenatal diagnosis within a group showing specific indications. We found that T21, T18, T13, and SCAs had the highest positive rates among the population with advanced maternal age. This finding indicated that advanced age is an important factor to focus on for these syndromes. In this study, advanced maternal age refers to age being the only high-risk factor for pregnancies. Among the identified MMSs, the highest rate (0.050%) was due to ultrasound abnormalities compared with other indications. Overall, the highest positive rate of chromosomal abnormalities was found for advanced maternal age, followed by positive serological screening. IVF-ET had the lowest positive rate of chromosomal abnormalities, followed by an abnormal ultrasound soft index. The paired t-test was conducted to assess aneuploidy positive rates within each of two of the six distinct groups characterized by varying clinical indications. There was no significant difference in the positive rate between any two groups.
Positive rates of chromosomal abnormalities in 118,969 pregnant women with different clinical indications.

IVF-ET, in vitro fertilization and embryo transfer; T21, trisomy 21; T18, trisomy 18; T13, trisomy 13; SCAs, sex chromosomal aneuploidies; RATs, rare autosomal trisomies; MMSs, microdeletion/microduplication syndromes.
Association between the positive rates of chromosomal abnormalities and maternal age
We analyzed the positive rates of various types of chromosomal abnormalities in relation to maternal age because age is a risk factor for high-risk pregnancies. Our study included 73,144 pregnant women aged 18 to 30 years, 35,520 aged 31 to 35 years, 9,172 aged between 36 and 40 years, and 632 aged older than 40 years. The positive rates of T21, T18, T13, and SCAs increased with age (Figure 2). T21 showed the largest increase, reaching a 1.27% positive rate in women aged older than 40 years. SCAs had the highest positive rate in pregnant women younger than 35 years, while T21 was highest in those older than 35 years. RATs and MMSs remained relatively stable with increasing age. Notably, in young women, the positive rate of MMSs was higher than that of T18 and T13.

Positive rates of chromosomal aneuploidies in 118,969 pregnant women of different maternal ages. T21, trisomy 21; T18, trisomy 18; T13, trisomy 13; SCAs, sex chromosomal aneuploidies; RATs, rare autosomal trisomies; MMSs, microdeletion/microduplication syndromes.
Fetal fraction analysis of this cohort
The fetal fraction plays an important role in NIPT because a sufficient fetal fraction ensures reliable results. Gestational age and BMI affect the fetal fraction. We investigated how the fetal fraction varies with gestational age and maternal BMI, and compared the fetal fraction between singletons and twins. We found that the fetal fraction remained relatively stable at a gestational age <20 weeks and significantly increased as gestational age advanced in singletons and twins (both p < 0.001) (Figure 3(a)). The fetal fraction was higher in twins than in singletons (p < 0.001) (Figure 3(a)). There was no difference in the maternal age of mothers with twins and those with singletons. The fetal fraction was negatively correlated with maternal BMI (r2 = 0.038, p < 0.001) (Figure 3(b)), and linearly decreased as maternal BMI increased, and it dropped below 10% when maternal BMI reached approximately 35 kg/m2. We compared the fetal fraction between false positive and true positive cases because the fetal fraction can cause false positives. We found that the false positive cases had a significantly lower fetal fraction than the true positive cases (p < 0.001) (Figure 3(c)). These findings indicated variation in the fetal fraction across our cohort.

Variation in the fetal fraction with gestational age and maternal BMI, and comparison of FPs and TPs. (a) Fetal fraction in singleton and twin pregnancies in relation to gestational age. (b) Fetal fraction in relation to maternal BMI and (c) comparison of false positive cases and true positive cases in relation to the fetal fraction. BMI, body mass index; FP, false positive; TP, true positive.
Discussion
This study showed that NIPT had a high sensitivity and specificity for detecting fetal trisomies 21, 18, and 13, SCAs, RATs, and MMSs in pregnant women, which is a similar finding to that in previous studies.9–15 However, PPVs for T18 and T13 were relatively low compared with the PPV for T21. This variation in PPV might be due to the differing prevalence of each syndrome. T21 has a higher prevalence than other chromosomal abnormalities, leading to a higher PPV for NIPT. In contrast, rarer aneuploidies with lower prevalence rates might result in lower PPVs because of limited sample sizes or challenges in accurate identification. Understanding this variation in PPVs is crucial for healthcare professionals when counseling expectant parents because it helps provide accurate information on the likelihood of a positive result, truly indicating a specific chromosomal abnormality.
Five false negatives were identified in the NIPT process, and occurred in three T21 samples and two T18 samples. Notably, the reason for the missed detection of false negative cases could not be attributed to a low fetal fraction because the observed range was 8.1% to 17.1%. Plausible explanations for the false negative samples include the presence of fetal or placental mosaicism, as suggested by previous studies. 16 Importantly, one false negative case of T18 showed a Z-score of 2.8 in NIPT, with a trisomic fraction and fetal fraction of 2.08% and 17.1%, respectively. Despite indications of a low rate of mosaicism in NIPT, subsequent single nucleotide polymorphism array diagnosis showed non-mosaic fetal T18. This incongruity suggests a potential discrepancy between fetal and placental karyotypes, and could be explained by a mosaic placenta with a non-mosaic fetus. The 402 false positive samples may have arisen owing to factors, such as a low fetal fraction and fetal or placental mosaicism. However, a further detailed analysis in subsequent investigations is necessary to validate these possibilities.
In this cohort, most true positive cases of RATs were mosaicism, which is consistent with previous studies. 17 The clinical utility of RAT analysis in NIPT is an area of research that still requires further investigation. Among the pathogenic CNVs detected in our study, 22q11 deletion syndrome was the most common. This finding is consistent with the highest prevalence of 22q11 deletion syndrome in MMSs. Notably, the number of observed MMS cases might have been underestimated because of NIPT’s low sequence depth and the late onset of some conditions beyond our follow-up period.
With China’s two-child and three-child policies, maternal and infant safety has gained increased attention, especially regarding birth defects. Many studies have shown that advanced maternal age is a risk factor for chromosomal aneuploidies.18,19 In our cohort, the group with advanced maternal age showed the highest positive rates for three major aneuploidies and SCAs (Table 2). Specifically, the positive rate for T21 was nearly three times that of those without clinical indications, while the positive rate for T18 exceeded three times. These findings are consistent with previous research in China,20,21 emphasizing an elevated risk in fetuses carried by older pregnant women. Notably, IVF-ET pregnancies had the lowest positive rates for all abnormalities, which indicated a lower risk associated with this method. Abnormal ultrasound findings had the highest rate of MMSs, which is consistent with a previous study. 22 This finding suggests that abnormal ultrasound findings may indicate CNVs. Based on our results and existing knowledge, an expanded NIPT approach may be necessary to detect CNVs in pregnant women with an abnormal ultrasound soft index. Additionally, our findings indicate that screening for chromosomal abnormalities is essential even in the general risk population without specific clinical indications.
Our study showed a clear association between the incidence of chromosomal abnormalities and maternal age. The positive rates of chromosomal abnormalities for T21, T18, and T13 increased steadily with maternal age, with a sharp rise occurring after 35 years of age (Figure 2), which is consistent with our advanced maternal age group findings mentioned above. Remarkably, the incidence of T21 showed a distinct elevation with increasing maternal age compared with other chromosomal aberrations. This phenomenon may be attributed to the survival advantage of T21 fetuses over other trisomies because aneuploidies involving other autosomal chromosomes often lead to an early fetal demise. The rare cases of RATs in our cohort may have contributed to this finding. The positive rate for SCAs also increased from 18 to 40 years old, with a small decrease after 40 years old, possibly due to the limited sample size. MMSs were more prevalent in younger populations, suggesting the need for focused attention even in younger pregnant women. These findings provide a stratified perspective on different chromosomal abnormalities in relation to maternal age, providing evidence for clinical practice regarding pregnancy and prenatal testing.
The fetal fraction, which represents the proportion of cell-free fetal DNA in maternal blood samples, plays a crucial role in NIPT. Our study showed that the fetal fraction was positively correlated with gestational age and negatively correlated with the BMI in pregnant women, consistent with numerous previous studies. 23 Interestingly, we observed relative stability in the fetal fraction during a gestational age <20 weeks. One possible explanation for this finding is that maternal weight tends to increase during the second trimester of pregnancy. Additionally, in our study, the fetal fraction was higher in twin pregnancies than in singleton pregnancies. Our findings could provide guidance for the clinical interpretation of NIPT results. The fetal fraction was lower in cases of false positives than cases of true positives. A low fetal fraction can induce false positives in NIPT. When the fetal fraction is low, the proportion of trisomic DNA (trisomic fraction) may artificially appear higher. This finding occurs because a lower amount of fetal DNA makes accurately distinguishing between normal and trisomic DNA levels more challenging. In NIPT, a higher trisomic fraction is typically indicative of a trisomy such as Down syndrome (trisomy 21). However, with a low fetal fraction, the observed higher trisomic fraction might be misleading, resulting in a false positive. Therefore, a low fetal fraction reduces the test’s ability to accurately measure the true proportion of trisomic DNA, leading to potential misclassification.
Our study showed the performance of NIPT in detecting fetal chromosomal aneuploidies and MMSs in a large cohort of pregnant women in Sichuan, China. This study of 118,969 women represents one of the largest single-site evaluations of NIPT screening. This large sample size enabled us to identify potentially abnormal samples within a substantial cohort of diverse pregnancies undergoing NIPT through cell-free DNA sequence analysis. Using sequencing-based approaches, NIPT achieved high sensitivity and specificity for trisomies 21, 18, and 13. Detection rates for SCAs, RATs, and MMSs were also remarkably robust. Our findings indicate that NIPT can serve as an effective first-tier screening tool in Sichuan Province to identify clinically actionable chromosomal abnormalities with a low false positive rate. Moreover, our findings indicate that assessing the relative positive rates of each chromosome could offer useful insight into the distribution of aneuploidy within tissues, thereby facilitating accurate resolution of clinical prevision. Broader implementation of primary aneuploidy screening may improve prenatal detection rates in this region. Further follow-up studies are required to determine the effect of NIPT on downstream diagnostic testing and pregnancy outcomes.
Despite our large sample size and comprehensive analysis, the performance of SCAs and MMSs cannot be guaranteed to be accurate. Unlike trisomies, which are more easily identifiable before birth, many symptoms associated with SCAs and MMS are challenging to detect in early infancy. Therefore, the performance of SCAs and MMSs observed in this study may deviate from the true situation. Nonetheless, this possibility does not hinder the provision of data guidance for current clinical applications of prenatal screening and NIPT.
Supplemental Material
sj-pdf-1-imr-10.1177_03000605241274584 - Supplemental material for Performance evaluation of noninvasive prenatal testing on 24 chromosomes in a cohort of 118,969 pregnant women in Sichuan, China
Supplemental material, sj-pdf-1-imr-10.1177_03000605241274584 for Performance evaluation of noninvasive prenatal testing on 24 chromosomes in a cohort of 118,969 pregnant women in Sichuan, China by Shengfang Qin, Yitong Zhao, Fengqiu Deng, Mengjia Yan, Na Xi, Chun Chen, Ping Wei, Zhiling Wu and Xueyan Wang in Journal of International Medical Research
Footnotes
Acknowledgements
We acknowledge Beijing USCI Medical Laboratory for laboratory processing, data analysis, and assistance with the manuscript.
Author contributions
XW: concept, design, data analysis, discussion, writing, and revision of the manuscript. SQ and YZ: statistical analysis, results, data analysis, and writing. FD and CC: data collection and statistical analysis. MY and NX: data collection and revision. PW and ZW: writing and revision of the manuscript.
Data availability statement
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Declaration of conflicting interest
The authors declare that there is no conflict of interest.
Funding
This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors.
Supplementary material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
