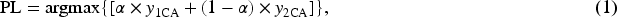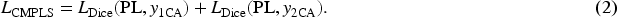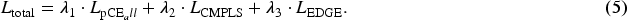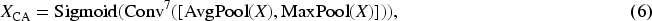Abstract
Scribble-supervised semantic segmentation has emerged as a promising alternative to fully supervised methods in medical imaging, owing to its low annotation cost and the inherent inclusion of contour information. However, the sparse supervision provided by scribble annotations, coupled with the underutilization of contour cues, often results in incomplete boundary representations in pseudo labels generated by the model. This limitation hinders the model’s ability to accurately capture complex anatomical structures, posing a significant challenge to achieving precise segmentation. To address this issue, we propose a novel three-branch network architecture. Built upon a multi-task learning framework, the model introduces a contour-assisted label supervision (contour auxiliary labels supervision) mechanism and a contour attention module to enhance the auxiliary decoder’s capability in extracting contour features. In addition, we design a contour mixed pseudo labels supervision strategy, which incorporates contour-enhanced representations into the pseudo-label generation process, thereby providing more informative and higher-quality supervision for scribble-based learning. We evaluate our method on the ACDC, MSCMR, and SegPC-2021 datasets. Experimental results demonstrate that our approach consistently outperforms state-of-the-art methods in terms of accuracy, robustness, and generalization. The scribble annotations and experimental code for the SegPC-2021 dataset are available at Github.
Introduction
Semantic segmentation has long been a critical task in medical image analysis. In clinical practice, segmentation accuracy is essential for both diagnosis and prognostic assessment. Recently, convolutional neural networks (CNNs) and transformer-based models have achieved remarkable success in medical image segmentation (Ronneberger et al., 2015; Valanarasu et al., 2021). Most of these models rely on pixel-level annotations from large-scale datasets that are densely and precisely labeled. However, manual annotation of medical images is both time-consuming and costly.
To address this challenge, researchers have increasingly explored new techniques that do not require large-scale, fully annotated datasets. Among them, weakly supervised learning has gained considerable attention as a promising alternative (Li et al., 2023; Luo et al., 2022; Obukhov et al., 2019; Wang et al., 2023a). Weakly supervised semantic segmentation typically leverages loosely annotated labels—such as points within regions of interest, scribbles, and bounding boxes—to train models (Chen & Sun, 2023; Cheng et al., 2021; Dai et al., 2015; Lin et al., 2016; Ying et al., 2023). These approaches significantly reduce annotation costs and alleviate the burden on clinical experts.
Scribbles inherently contain rich contour information and, compared to other forms of weak annotations such as points and bounding boxes, offer clearer guidance for medical image segmentation. As illustrated by the “scribble annotation” example in the figure, scribble supervision is a form of weak supervision in which annotators only need to draw a few strokes over the target regions. Scribble supervision has been widely applied in various medical imaging tasks, including cardiac magnetic resonance imaging (MRI) segmentation (Bernard et al., 2018; Zhuang, 2016, 2019) and uterine cancer MRI segmentation (Ying et al., 2023).
Although scribble annotations can substantially reduce reliance on expensive and time-consuming expert-labeled segmentation masks while providing rich contour information, existing studies have often underutilized this critical cue for segmentation. As a result, the models are hindered in their ability to learn the detailed visual features necessary for precise delineation.
To address this challenge, we propose a novel three-branch segmentation model. The model is designed to first independently learn contour representations through a contour-assisted label supervision (contour auxiliary labels supervision (CALS)) mechanism and a contour attention (CA) module, and then integrate these features into the pseudo-label generation process via a contour mixed pseudo labels supervision (CMPLS) strategy to produce more complete and structurally accurate pseudo labels.
Specifically, inspired by the principles of multi-task learning, we adopt a three-branch architecture consisting of a shared encoder and three decoders. Two of the decoders serve as primary branches, which learn from scribble annotations using the partial cross-entropy (pCE) loss. The third decoder, in the CALS branch, learns contour representations from auxiliary contour labels. These contour labels are generated during preprocessing using the Canny edge detection algorithm and are trained via an edge-based loss function. We observed that the output of the CALS decoder alone lacks sufficiently prominent contour representations, making it suboptimal when used in isolation. To remedy this, we introduce the CA module, which takes the CALS decoder’s output as input and applies a convolutional block attention mechanism to generate a refined contour feature map that emphasizes key boundary regions. This contour feature map is then fused with the outputs of the two primary decoders through element-wise multiplication and a dynamic mixing operation, resulting in pseudo labels that preserve more complete shape information.
Ultimately, the proposed method integrates both contour-assisted labels and enhanced pseudo labels for end-to-end training of the segmentation network. By effectively leveraging the contour information inherent in scribble supervision, our model achieves more accurate segmentation masks. The CALS mechanism contributes additional structural cues, the CA module guides the model’s attention toward critical contour regions, and the CMPLS strategy embeds these contour-aware features into the pseudo-label generation process. This joint strategy is designed to substantially improve the accuracy and reliability of segmentation in medical image analysis.
The main contributions of this paper are as follows:
We propose an innovative three-branch model combined with a CMPLS strategy to enhance the performance of scribble-based medical image segmentation. By effectively integrating the refined contour features learned from the auxiliary branch into the dynamic fusion of the primary branches, this model significantly improves the shape integrity of the generated pseudo labels. We introduce a CALS mechanism and a CA module to substantially enhance the segmentation performance of the model. The CALS mechanism employs a multi-task learning strategy to extract additional contour features from contour auxiliary labels. Concurrently, the CA module utilizes convolutional block attention to effectively sharpen the model’s focus and precision in capturing critical contour regions. To evaluate the performance of our model, we conducted extensive experiments on the SegPC-2021 dataset and two benchmark datasets: ACDC and MSCMR. The results indicate that our method not only outperforms existing scribble supervision techniques in terms of performance but also significantly narrows the gap with mask supervision methods, even surpassing their performance in certain cases.
Related Work
Pseudo Labels for Semantic Segmentation
Pseudo-labeling was originally developed in the context of semi-supervised learning. It typically involves using a base network pretrained on labeled data to generate predictions for unlabeled data, which are then treated as pseudo labels. These pseudo labels enable the incorporation of unlabeled samples into model training and have been widely adopted in tasks such as image classification and segmentation (Chen et al., 2021, 2020; Filipiak et al., 2021; Sohn et al., 2020; Teh et al., 2021; Wang et al., 2021b, 2023c; Yang et al., 2021; Zhu et al., 2024).
Inspired by pseudo-label techniques in semi-supervised learning, researchers have extended this concept to weakly supervised segmentation. Weakly supervised semantic segmentation leverages various forms of weak annotations (e.g., image-level tags, points, scribbles, and bounding boxes) to generate mask pseudo labels for images, simulating fully supervised segmentation during network training (Can et al., 2018; Lin et al., 2024; McEver & Manjunath, 2020; Wang et al., 2018). Current research in this area can be broadly categorized into four groups based on the type of supervision signals utilized: image-level tags supervision (Lin et al., 2024; Wang et al., 2018), point supervision (McEver & Manjunath, 2020; Ren et al., 2024), scribble supervision (Can et al., 2018; Lee & Jeong, 2020), and bounding box supervision (Dai et al., 2015).
It is worth noting that many weakly supervised methods rely on unsupervised segmentation techniques—such as class activation maps and superpixels (Dai et al., 2015; Lin et al., 2024; Ren et al., 2024; Wang et al., 2018)—or traditional algorithms such as random walk (Can et al., 2018; McEver & Manjunath, 2020) to generate pseudo labels. However, these approaches often suffer from poor generalizability in practical applications and struggle to adapt to the complexities of medical image segmentation. In contrast, our method generates pseudo labels with more complete shape representations for scribble supervision by dynamically fusing information from multiple decoders and incorporating the contour features learned by an auxiliary decoder.
Scribble-Supervised Semantic Segmentation
Building upon the main technical distinctions among various approaches and insights from previous studies (Li et al., 2023), scribble-supervised semantic segmentation methods can be broadly categorized into three main groups: (1) mix augmentation methods, (2) different scribble supervision loss functions, and (3) pseudo label generation methods.
(1)
(2)
(3)
Despite the achievements of previous methods in medical image segmentation, they are limited when generating pseudo labels due to the lack of complete shape features. To overcome this challenge, we integrated contour auxiliary labels extracted in the preprocessing stage into the network through multi-task learning, thereby enhancing the model’s ability to learn contour features. In addition, we introduced a CA module that further strengthens the contour features extracted from the images. By adopting a CMPLS strategy, we incorporated these contour features into the pseudo-labels generation process to produce high-quality pseudo-labels with richer shape information. This strategy not only improves the model’s ability to recognize the shape of the target structure but also significantly enhances the overall performance of the segmentation task.
Methodology
Overall Architecture
As illustrated in Figure 1, the proposed three-branch network architecture consists of a shared encoder
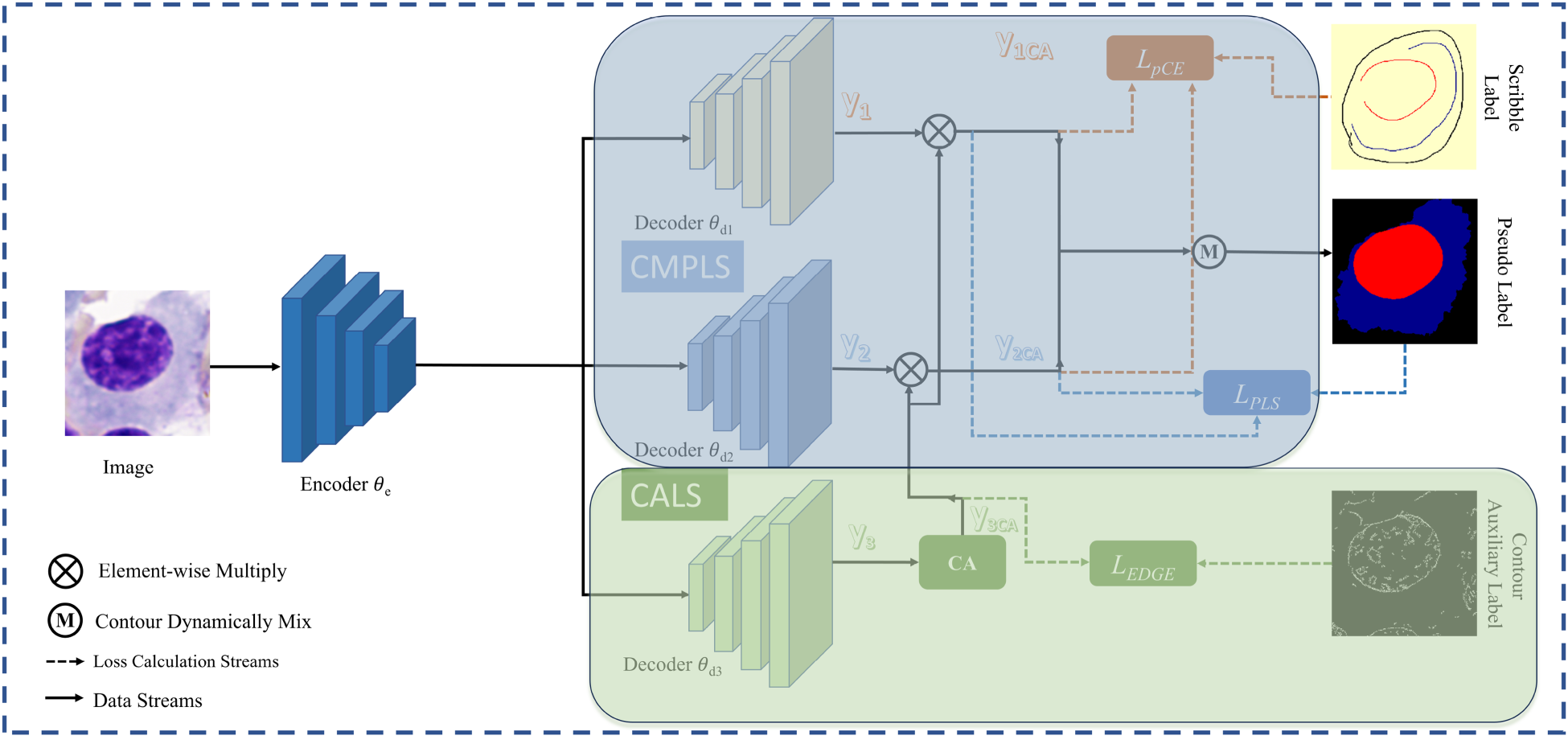
The Overall Structure is Shown in the Figure. The Framework Consists of an Encoder and Three Decoders, Decoder
If a single branch is used for segmentation while another serves as an auxiliary branch to learn contour information, the network may suffer from an inherent issue described in Huo et al. (2021), where each branch tends to preserve its own predictions, thereby hindering effective model updates. To address this, we adopt a dual-decoder structure inspired by Luo et al. (2022), in which two decoders with different perturbations independently learn segmentation predictions and are dynamically fused. Unlike (Luo et al., 2022), however, we introduce an additional CALS (contour-assisted label supervision) branch that learns supplementary contour features from perturbed inputs and contributes them to the pseudo-label supervision process. This design effectively reintroduces structurally meaningful perturbations, further decouples the learning pathways of different branches, and alleviates their tendency to reinforce initial predictions. During training, the integration of the CALS branch and the CA module progressively enhances the model’s ability to extract contour features, guiding its predictions to focus on critical structural regions.
Specifically, the decoders
In order to generate pseudo labels with more complete shape features, we adopt the CMPLS strategy. By incorporating additional shape information obtained from the CALS branch into the pseudo-label generation process of Luo et al. (2022), this strategy effectively enhances the model’s feature extraction and representation learning capabilities. As illustrated in Figure 1,
During training, decoders
The loss function used for training the CMPLS is defined as follows:
In equation (2),
Simultaneously, we utilize the loss function defined in equation (3) to learn target segmentation from the scribble annotations.
To provide additional contour features for pseudo labels and enhance the model’s ability to mine contour characteristics, we propose CALS. This mechanism consists of two key components: the supervision mechanism and the CA module, designed to optimize the model’s effectiveness in learning contour features.
Supervision Mechanism
To enhance the network’s ability to perceive the structure of segmentation targets, we introduced an auxiliary branch designed to learn contour features (as shown in the CALS section of Figure 1). Inspired by multi-task learning, this branch guides the model to progressively align the predicted contour features with the pseudo contour labels by calculating the loss between the processed model predictions and the contour pseudo labels.
Specifically, we employ the Canny algorithm (Canny, 1986) to extract the edges of the segmentation target structure from the image. Since an object’s contour can be viewed as a connection of its edges (Arbeláez et al., 2011), we treat the extracted edges as contours and store them, thereby avoiding additional annotation work. Practically, we only need to load the pre-saved contour maps alongside other image information. During training, the output of decoder
Throughout the end-to-end training process, our method gradually aligns the predicted boundaries with the true boundaries, thereby improving the model’s ability to perceive contours.
We utilize
Furthermore, the loss function for the entire network training is expressed in equation (5):
In our experiments, the values of
Applying different losses jointly to the network makes our network focus on both scribble labels, contour labels, and network-generated pseudo labels. In addition, the network also generates a pseudo label, which focuses more on contours, as a result, which is useful for the task of segmenting other elliptical or circular targets.
After introducing the CALS strategy, we observed a decline in convergence speed when the auxiliary branch relied solely on
The integration of the CA module with CALS not only enhances the auxiliary branch’s ability to perceive contour features but also accelerates its convergence while maintaining consistency with the contour auxiliary labels during training. Furthermore, the output
The module takes the output
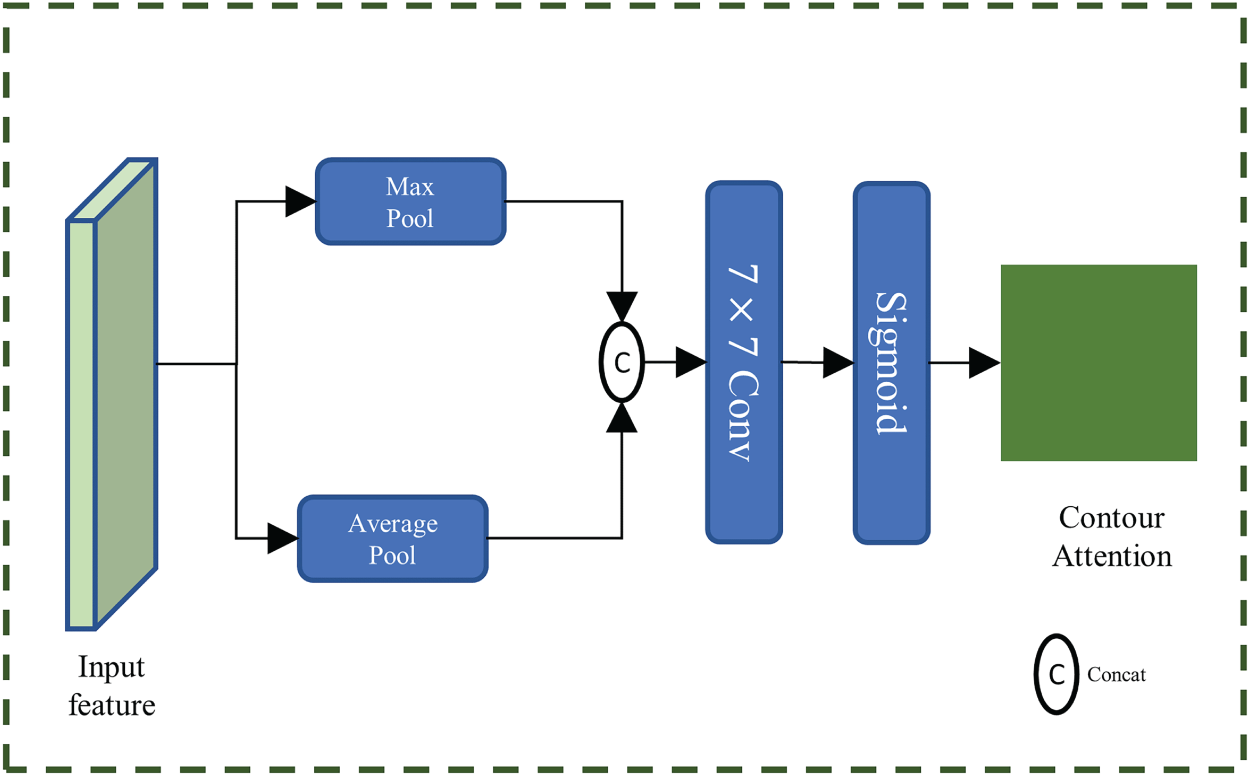
Schematic of the Contour Attention Module.
These weights are then element-wise multiplied with the outputs
To validate the effectiveness of our proposed method, we conducted a series of comprehensive experiments on the ACDC, MSCMR, and SegPC-2021 datasets. The ACDC and MSCMR datasets are the most commonly used benchmarks in the field of scribble-based semantic segmentation (Li et al., 2023, 2024; Liu et al., 2023; Luo et al., 2022; Zhang & Zhuang, 2022, 2023). The SegPC-2021 dataset is renowned for its complexity, focusing on multi-object segmentation in bone marrow cell analysis (Gupta et al., 2023; Qiu et al., 2022). Notably, precise segmentation of various tissues within the SegPC-2021 dataset is crucial for the diagnosis of multiple myeloma (MM; Gupta et al., 2023).
Datasets
The SegPC-2021 dataset is the most comprehensive publicly available dataset for plasma cell segmentation in MM (Gupta et al., 2023), comprising 775 bone marrow smear images captured by two cameras attached to a microscope. These bone marrow smears are well-known for their complex tissue structures (Jiang et al., 2022). The dataset includes precise mask annotations for both nuclei and cytoplasm, created by 10 experts, making it a highly complex multi-object segmentation dataset, particularly suitable for research in scribble-based segmentation. We first extracted individual cells from the dataset into separate images, ensuring that each image contains only one distinct and complete cell. Then, we used different colors in the scribble annotations to distinctly mark the nucleus, cytoplasm, and background of each cell. Following the guidelines in Valvano et al. (2021), we performed scribble annotations on all cells in the dataset, resulting in 2631 cell images with their scribble annotations, as shown in Figure 3(a). Each image contains two segmentation targets. Compared to cardiac segmentation datasets, these cell images have targets occupying a larger portion of the image, making them more sensitive to performance metrics, and each image contains two segmentation targets.
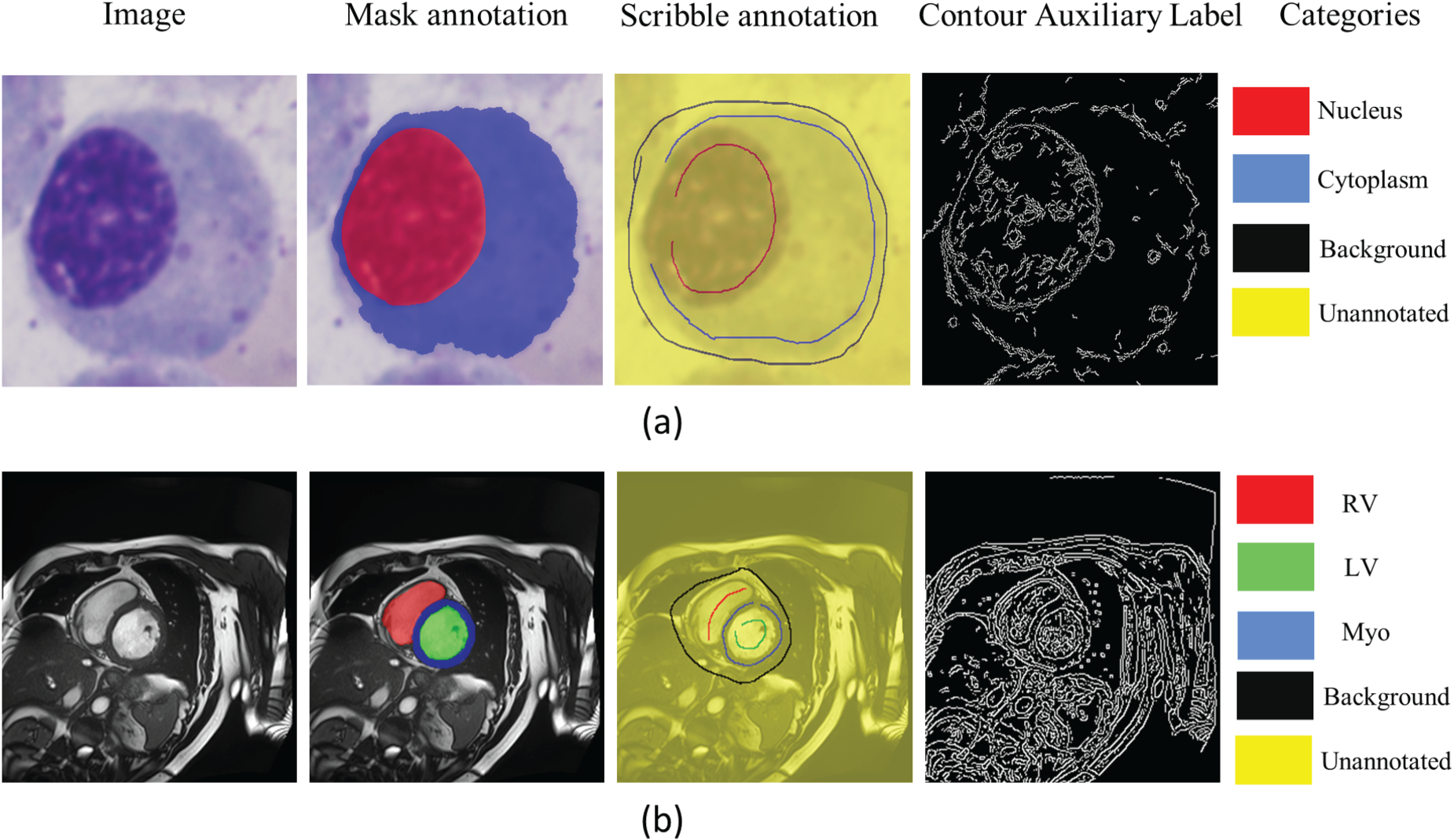
(a) An Example of the SegPC-2021 Dataset, and (b) an Example of the Publicly Available ACDC Dataset. Since MSCMR is Similar to ACDC, it is not Shown Here. In (a) and (b), the “Image” is the Raw Image, the “Mask Annotation” is the Mask Annotations Superimposed on the Raw Image, the “Scribble Annotation” is the Scribble Annotations Superimposed on the Raw Image, and the “Contour Auxiliary Label” is the Contour Auxiliary Labels Generated During the Preprocessing Stage. In the “Mask Annotation” and the ”Scribble Annotation” of (a), red, blue, black, and yellow Represent Nuclei, Cytoplasm, Background and Unannotated Pixels, Respectively. In the “Mask Annotation” and the “Scribble Annotation” of (b), Red, Green, Blue, Black, and Yellow Represent RV, LV, Myo, Background and Unannotated Pixels, Respectively. Note. RV = Right Ventricle; LV = Left Ventricle; Myo = Myocardium.
The ACDC dataset (Bernard et al., 2018) comprises cardiac MRI data from 100 patients, with expert annotations for the left ventricle (LV), right ventricle (RV), and myocardium (Myo). Each patient’s data includes images from both end-diastole and end-systole phases of the cardiac cycle. Based on previous studies (Luo et al., 2022; Valvano et al., 2021), we used a two-dimensional (2D) slice segmentation dataset instead of a three-dimensional volumetric segmentation dataset due to thickness considerations. The 2D slice dataset contains a total of 1,902 images. The majority of these images have three segmentation targets, while a few contain only two. The size of the segmentation targets varies across different images. Valvano et al. (2021) provided scribble annotations for each image, with examples of the mask annotations and scribble annotations shown in Figure 3(b).
The MSCMR dataset (Zhuang, 2016, 2019) consists of late gadolinium-enhanced MRI scans from 45 cardiomyopathy patients, with expert annotations for the LV, RV, and Myo provided for each scan. Enhanced MRI poses greater challenges for segmentation tasks compared to nonenhanced MRI (Zhang & Zhuang, 2022). Based on previous studies (Li et al., 2023; Liu et al., 2023), we utilized a 2D slice segmentation dataset, which includes a total of 686 images. Similar to the ACDC dataset, most images contain three segmentation targets, while a few have two, and the size of the segmentation targets varies across different images. Additionally, Zhang and Zhuang (2022) provides scribble annotations for 407 images from 25 scanned slices.
We adopted U-Net (Ronneberger et al., 2015) as the base segmentation network architecture and expanded it into a triple-branch network by adding two auxiliary decoders. A dropout layer (
We use the dice score coefficient (DSC) and 95% Hausdorff Distance (HD95) as performance metrics. In our study, we employed five-fold cross-validation on the SegPC-2021 dataset and the ACDC dataset, calculating the mean evaluation metrics for each fold to ensure the robustness and generalizability of the results. Our ACDC dataset partitioning follows the same approach as Luo et al. (2022). Given that the MSCMR dataset has scribble annotations for only 25 scans, it is impractical to perform five-fold cross-validation on all 45 scans, and using just 25 scans would result in a dataset that is too small. Therefore, we adhered to the dataset split method from Zhang and Zhuang (2022), using 25 scans for the training set, five scans for the validation set, and 15 scans for the test set.
To minimize the joint loss functions for model optimization, we employed an SGD optimizer with a weight decay of
Comparison With Other State-of-the-art Methods on the ACDC Dataset and MSCMR Dataset
To demonstrate the performance of our method, we compared CMPLS with various state-of-the-art (SOTA) methods. Our comparison methods include: Different scribble supervision loss function: pCE only (pCE) (Lin et al., 2016), uncertainty-aware self-ensembling and transformation-consistent model (USTM) (Liu et al., 2021), Mumford–Shah loss (MLoss) (Kim & Ye, 2019), GCRF lossGCRF (Obukhov et al., 2019), entropy minimization (EM) (Grandvalet & Bengio, 2004), and ZScribbleSeg (Zhang & Zhuang, 2023). Pseudo label generation methods: Using pseudo labels generated by random walker (RW) (Grady, 2006), Scribble2Label (S2L; Lee & Jeong, 2020), spatial–spectral mutual teaching and ensemble learning (S2ME; Wang et al., 2023a), DMPLS (Luo et al., 2022), Scribble walking and class-wise contrastive regularization (SC-Net; Zhou et al., 2023). Scribble with Vision-Class Embedding (ScribbleVC; Li et al., 2023), ScribbleMatch and reliable-guided pixel alignment (SRPA; Liu et al., 2023), and ScribFormer (Li et al., 2024). Mix augmentation methods: CycleMix (Zhang & Zhuang, 2022), TriMix (Zheng et al., 2022), and PCLMix (Lei et al., 2024).
Finally, we also studied the fully supervised method U-Net (Ronneberger et al., 2015).
On the MSCMR dataset, which primarily focuses on multi-sequence cardiac magnetic resonance (MR) images, our method consistently outperforms existing scribble-supervised segmentation approaches across all evaluation metrics. Notably, CMPLS achieves an average dice score improvement of 1.1% over the prior SOTA SRPA (88.5% vs. 87.4%). This performance gain is particularly evident in the segmentation of the Myo and RV, where precise boundary delineation is critical due to their complex and variable anatomical shapes. The superior performance in these categories demonstrates the effectiveness of our contour-aware pseudo-label strategy in capturing subtle structural variations and enhancing shape representation, which are typically underrepresented in weakly supervised settings.
In contrast, on the ACDC dataset, although CMPLS shows a slightly lower average dice score compared to TriMix (0.5% lower) and PCLMix (0.4% lower), it significantly outperforms both methods in terms of boundary accuracy, as indicated by the HD95 metric. Specifically, CMPLS reduces HD95 by 3.0 mm (from 5.9 mm to 2.9 mm) and 1.2 mm (from 4.1 mm to 2.9 mm) compared to TriMix and PCLMix, respectively. These results underscore the advantage of our method in promoting boundary precision, which is crucial for accurate structure delineation, especially in LV and Myo segmentation tasks—both of which involve complex edge geometries and thin-wall structures.
The discrepancy between dice and HD95 performance suggests that while other methods may achieve comparable volumetric overlap, they often struggle to accurately capture fine-grained boundary details, leading to larger localization errors. In contrast, CMPLS effectively leverages the contour features extracted by the CALS branch and integrates them through a dynamic contour-aware pseudo-labeling mechanism. This allows the network to learn more structure-sensitive features, thus maintaining high accuracy not only in the segmentation area but also in shape fidelity.
These findings highlight the robustness and generalization capability of our method across datasets with different characteristics—MSCMR with heterogeneous MR modalities, and ACDC with high-resolution cine MR images—as well as across structures with varying morphology and complexity. The results validate that contour-guided supervision, when effectively integrated, plays a pivotal role in improving segmentation quality under weak supervision scenarios.
Regarding fully supervised methods, the second part of Table 1 shows that the gap between weakly supervised and fully supervised methods is narrowing on the ACDC dataset. In Table 2, the second part indicates that our method significantly outperforms the basic fully supervised method on the MSCMR dataset. This showcases the great potential of CMPLS in medical image segmentation.
In the Experiments Conducted on the ACDC Dataset, We Compared Our Proposed Method With Other SOTA Methods in the Field of Weakly Supervised Learning.
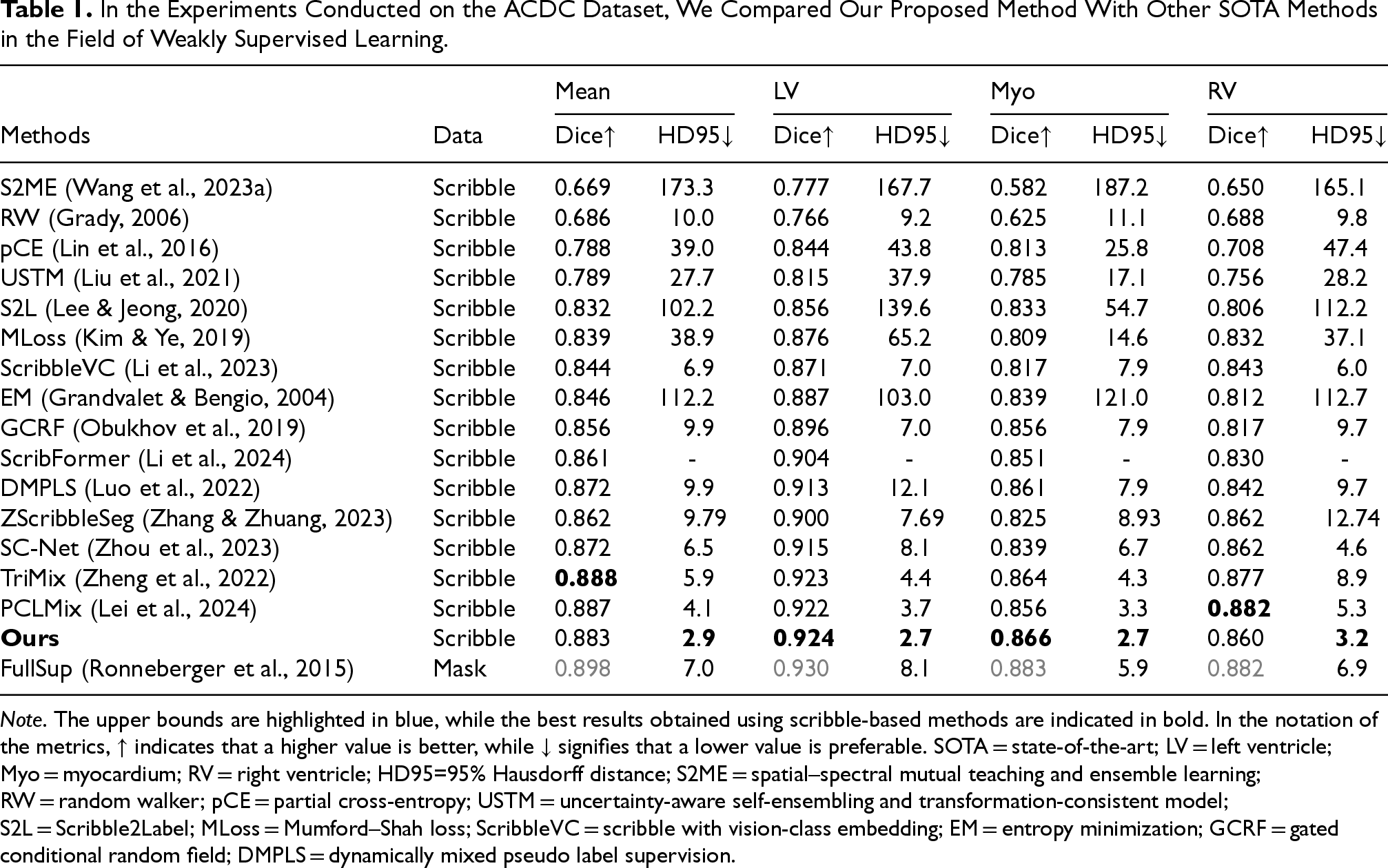
Note. The upper bounds are highlighted in blue, while the best results obtained using scribble-based methods are indicated in bold. In the notation of the metrics,
In the Experiments Conducted on the MSCMR Dataset, We Compared Our Proposed Method With Other SOTA Methods in the Field of Weakly Supervised Learning.
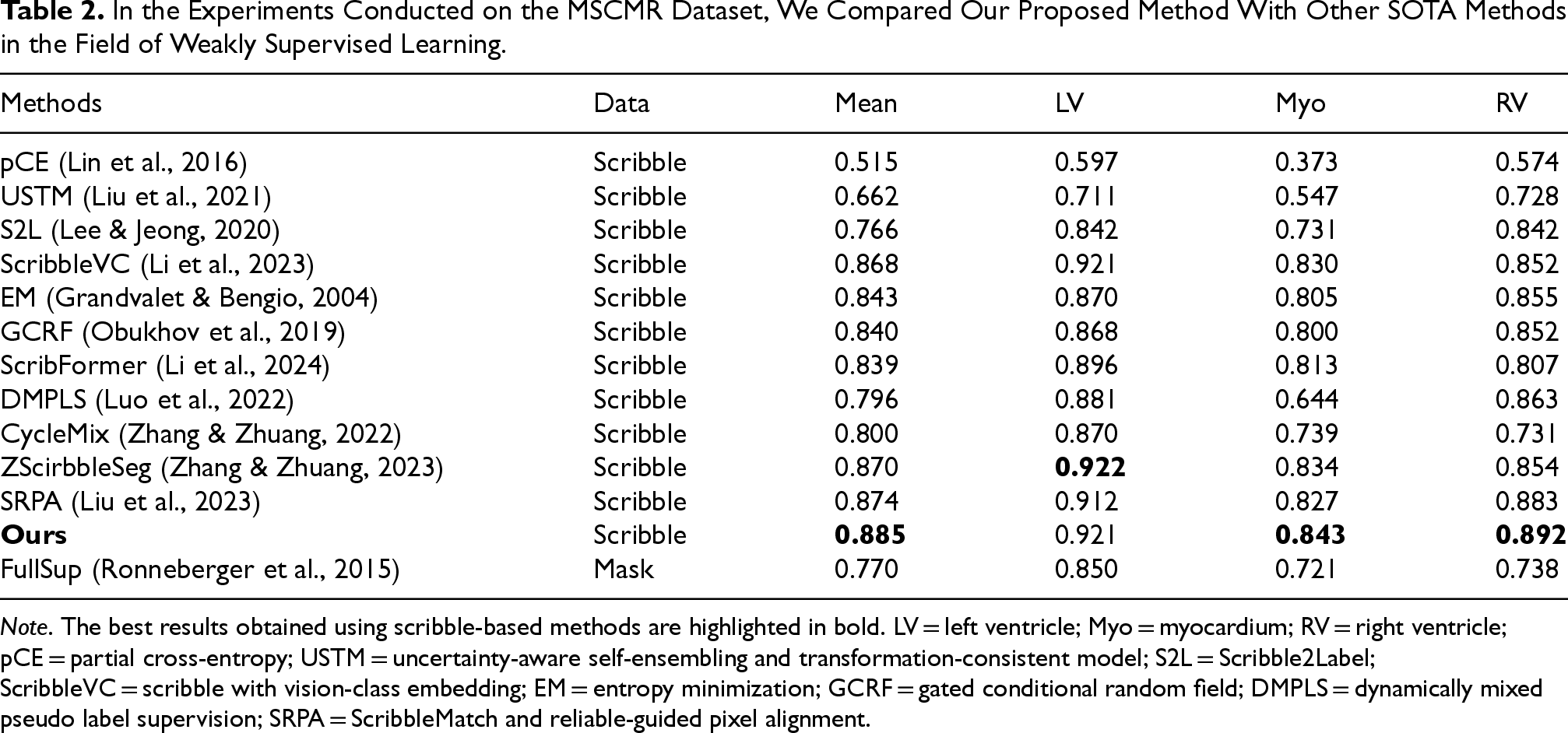
Note. The best results obtained using scribble-based methods are highlighted in bold. LV = left ventricle; Myo = myocardium; RV = right ventricle; pCE = partial cross-entropy; USTM = uncertainty-aware self-ensembling and transformation-consistent model; S2L = Scribble2Label; ScribbleVC = scribble with vision-class embedding; EM = entropy minimization; GCRF = gated conditional random field; DMPLS = dynamically mixed pseudo label supervision; SRPA = ScribbleMatch and reliable-guided pixel alignment.
Furthermore, comparing the S2ME method in Tables 1 and 3, we find that S2ME performs significantly better on the SegPC-2021 dataset (ranking fifth) than on the ACDC dataset (ranking last). We attribute this to the fact that the SegPC-2021 dataset consists of color images, while the ACDC dataset contains grayscale images. The S2ME method is based on spatial–spectral pseudo-labels generation, whereas our method demonstrates strong robustness and generalization capabilities.
Experimental Results Comparing Our Method With Other Methods on the SegPC-2021 Dataset, the Upper Limits are Highlighted in Blue, and the Best Results Obtained Using the Scribble Method are Highlighted in Bold.
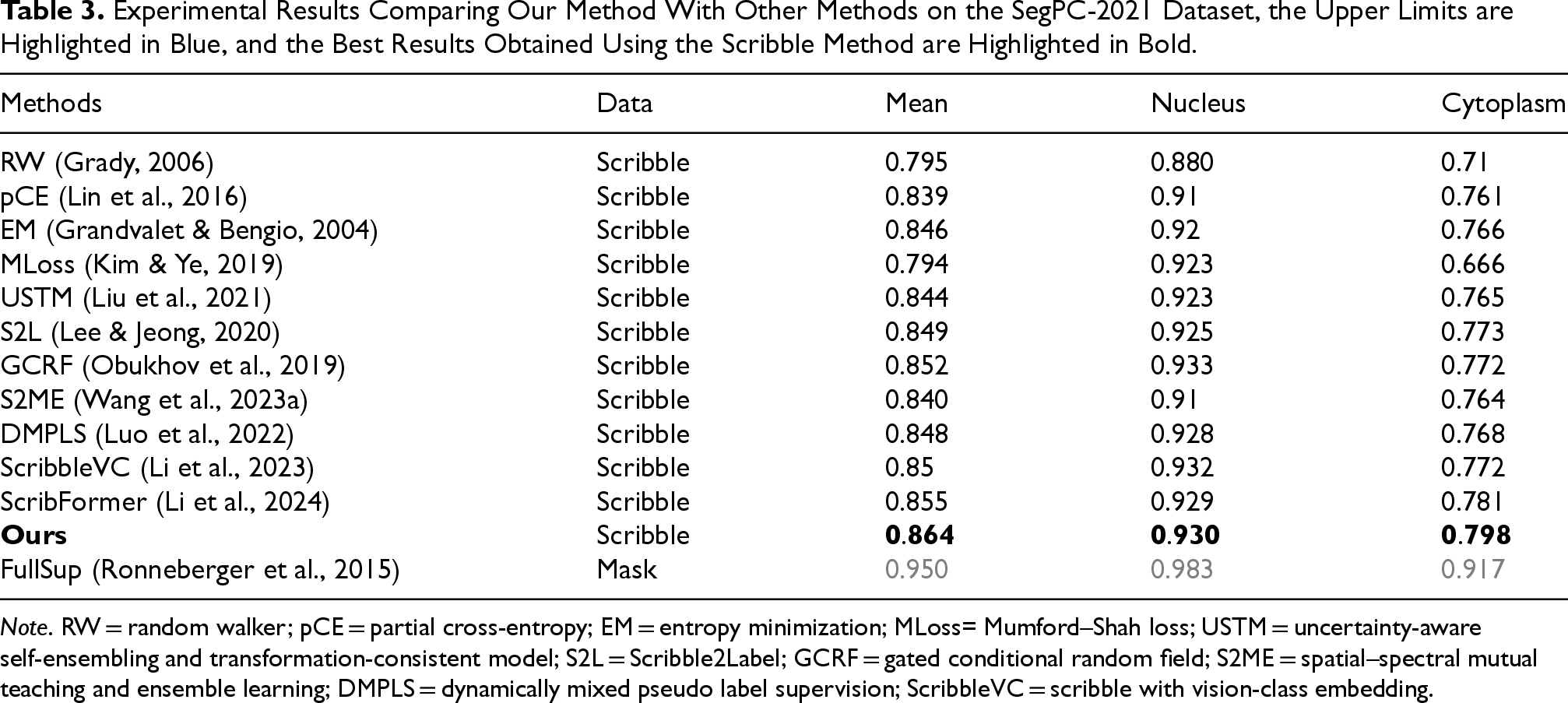
Note. RW = random walker; pCE = partial cross-entropy; EM = entropy minimization; MLoss= Mumford–Shah loss; USTM = uncertainty-aware self-ensembling and transformation-consistent model; S2L = Scribble2Label; GCRF = gated conditional random field; S2ME = spatial–spectral mutual teaching and ensemble learning; DMPLS = dynamically mixed pseudo label supervision; ScribbleVC = scribble with vision-class embedding.
Figure 4 shows a visual comparison of our method with other weakly supervised methods on the ACDC scribble dataset. As shown in Figure 4, our method exhibits significant advantages over other weakly supervised methods on the ACDC scribble dataset. First, in cases where the target region is surrounded by a very similar region (as shown in the first row of Figure 4), our method is able to accurately segment Myo and LV with even greater precision. in cases where the target region occupies a very small proportion of the graph (as shown in the second row of Figure 4), our method is able to segment the target region very accurately without interference from other regions. Finally, in cases where the imaging around the target region is blurred (as shown in the third and fourth rows of Figure 4), our method is still able to locate and segment the target region more accurately. Our method performs well in medical image segmentation, with the highest boundary clarity and accuracy in the segmentation results. Although the comparison methods perform better in some scenes, overall, they are slightly inferior to our method in terms of boundary accuracy and detail retention.
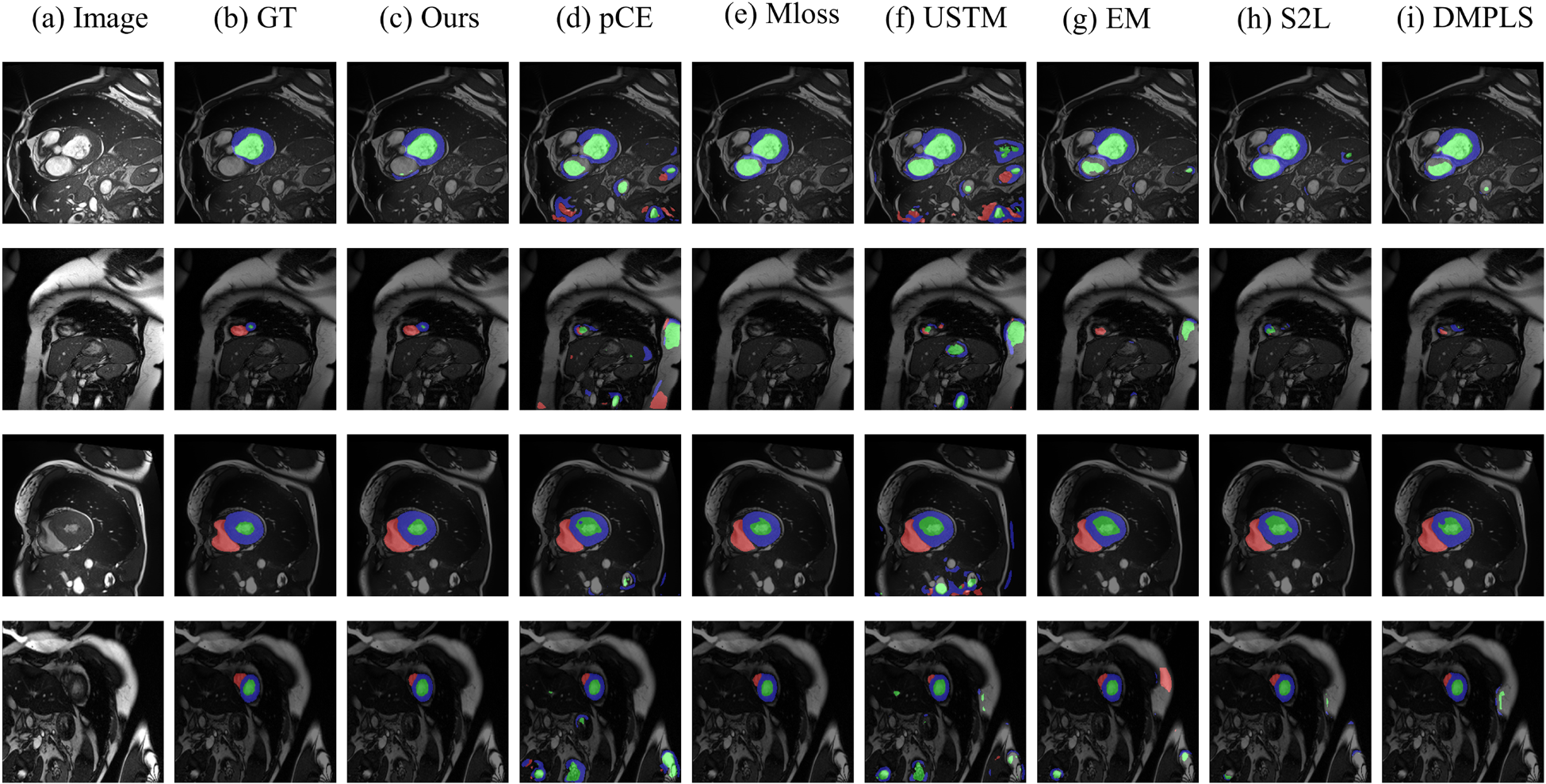
Intuitive Comparison of our Proposed Method With Other Weakly Supervised Methods on the ACDC Dataset. (a) Raw Image, (b) Ground Truth, (c) Our Method, (d) pCE, (e) MLoss, (f) USTM, (g) EM, (g) S2L, and (i) DMPLS. Apparently, Our Method can Better Identify the Cell Boundaries. The Prediction Result (c) is More Consistent With the Ground Truth Image. Note. pCE = Partial Cross-Entropy; MLoss = Mumford–Shah Loss; USTM = Uncertainty-Aware Self-Ensembling and Transformation-Consistent Model; EM = Entropy Minimization; S2L = Scribble2Label; DMPLS = Dynamically Mixed Pseudo-Label Supervision.
We compared CMPLS with other methods on the SegPC-2021 dataset, as shown in Table 3. In terms of scribble supervision, we found that CMPLS significantly outperforms other weak supervision methods in mean, nucleus, and cytoplasm metrics. Compared to the SOTA methods on other datasets, DMPLS and ScribbleVC, CMPLS surpasses them by 1.6% (84.8% vs. 86.4%) and 1.4% (85.0% vs. 86.4%), respectively. As for ScribFormer, which is the best performer on the SegPC-2021 dataset, our method surpasses it by 0.9% (85.5% vs. 86.4%).
Figure 5 shows a visual comparison of our method with other weakly supervised methods on the SegPC-2021 dataset. As shown, our method has significant advantages over other weakly supervised methods. First, in the case where the cytoplasm is fused with the background (second and third row of Figure 5), our method can distinguish the cytoplasm from the background more efficiently and segment the cytoplasm more accurately. Second, our method also performs well in dealing with color shifts due to overlapping (Figure 5 fourth row). Finally, in cases where the nucleus and cytoplasm are very close to each other (the first and fifth rows of Figure 5), our method achieves more accurate cell segmentation. Meanwhile, we can find that our method works extremely well in inscribing the boundary between the nucleus and the cytoplasm, while other methods tend to have many errors at the boundary.
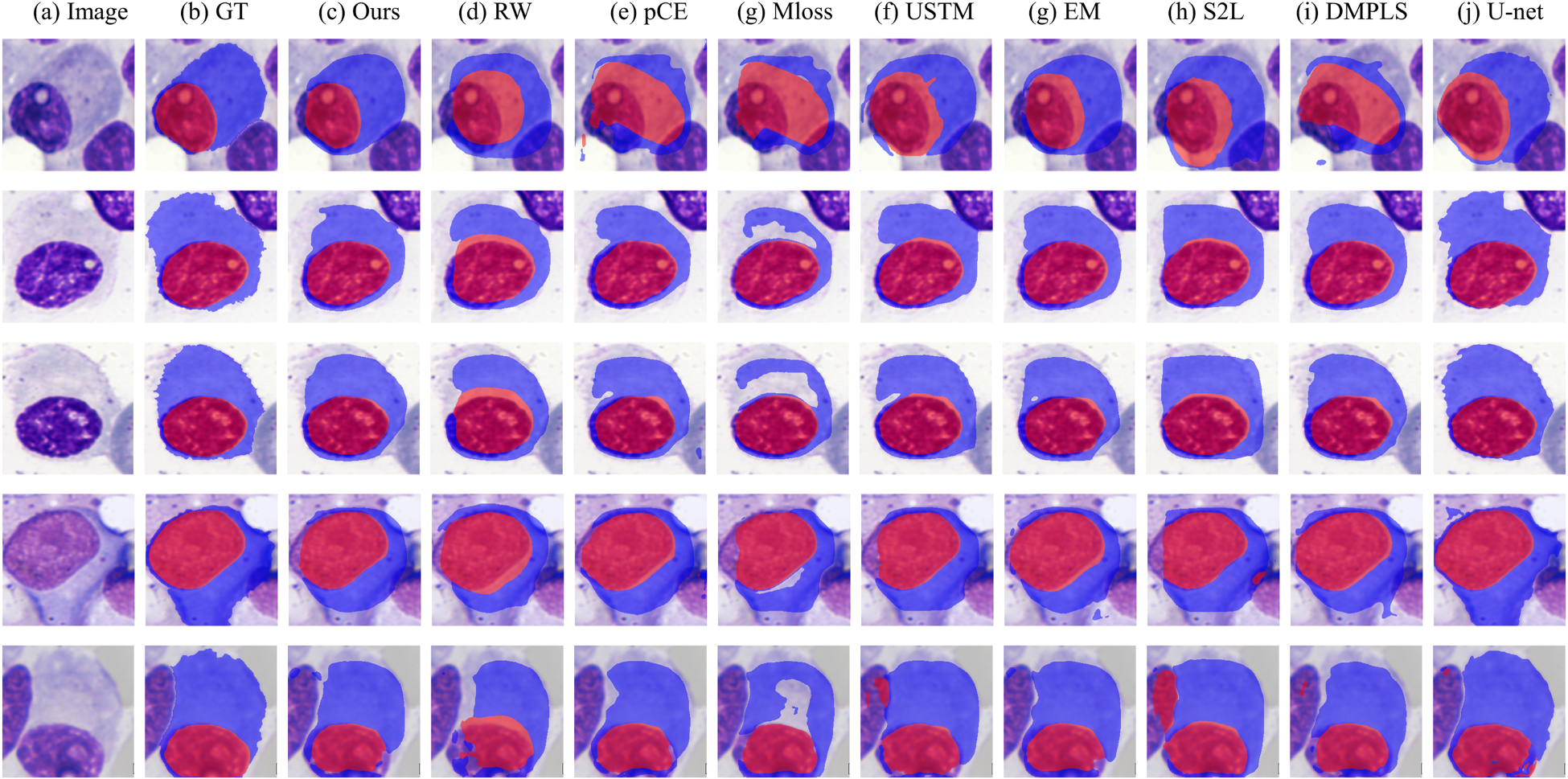
Intuitive Comparison of Our Proposed Method With Other Weakly Supervised Methods on the SegPC-2021 Dataset. (a) Raw Image, (b) Ground Truth, (c) Our Method, (d) RW, (e) pCE, (f) USTM, (g) EM, (g) S2L, (i) DMPLS, and (j) U-Net. Where (j) U-Net is the FullSup in Table 3. Apparently, Our Method can Better Identify the Cell Boundaries. The Prediction Result (c) is More Consistent with the Ground truth Image. Note. RW = Random Walker; pCE = Partial Cross-Entropy; USTM = Uncertainty-Aware Self-Ensembling and Transformation-Consistent Model; EM = Entropy Minimization; S2L = Scribble2Label; DMPLS = Dynamically mixe Pseudo Label Supervision.
However, when compared to the basic fully supervised model U-Net, we observed that weak supervision methods perform considerably worse on the SegPC-2021 dataset, especially in cytoplasm segmentation. We attribute this to the fact that, compared to MRI datasets, cells occupy larger areas in the images, involving more pixels in the region of interest, and the cell dataset has complex and highly irregular boundaries. Additionally, the staining of the cytoplasm is lighter than that of the nucleus. These factors contribute to the suboptimal performance of all scribble supervision methods on this dataset. Nevertheless, considering the enormous annotation cost associated with fully supervised methods (as illustrated in Figure 3(a), where full annotation requires substantial effort, especially for the cytoplasm boundary), our method achieves commendable performance with minimal annotation cost. This demonstrates the great potential of weak supervision methods in medical image segmentation.
To evaluate the effectiveness of contour pseudo-labels supervision (CALS) and CA, we performed ablation experiments on the SegPC-2021 dataset and the ACDC dataset. As shown in Table 4, the first baseline (“basic”) corresponds to the removal of CALS and CA from our method. The second baseline (“basic + CALS w/o ABL”) is equivalent to adding CALS to the first baseline, but removing the active boundary loss from the loss function. The third baseline (“basic + CALS”) is equivalent to adding the active boundary loss to the second baseline. Our method is equivalent to adding CA to the third baseline. The results in Table 4 show that the performance of the model improves with the successive addition of each module, effectively demonstrating the efficacy of CALS and CA. Compared to the base model, the addition of CALS and CA increased the mean DSC metrics by 1.6% on the SegPC-2021 dataset and 1.2% on the ACDC dataset.
Ablation Experiments.
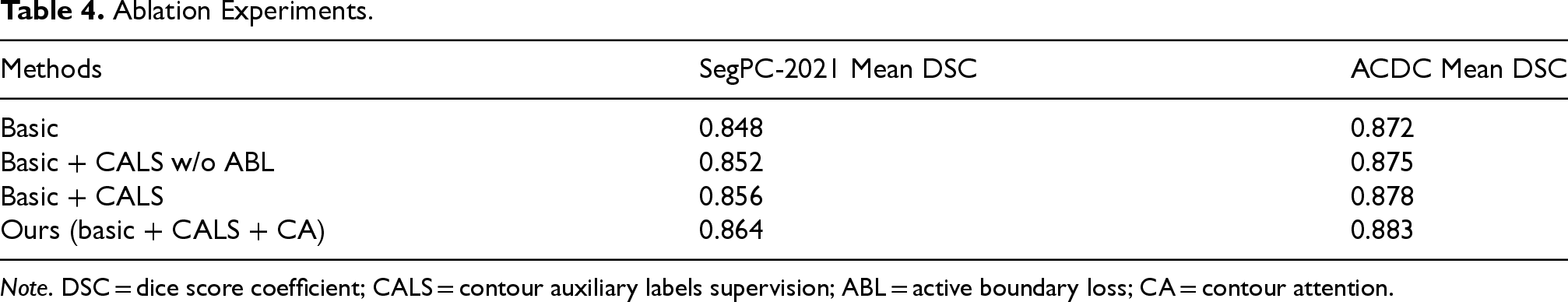
Ablation Experiments.

Note. DSC = dice score coefficient; CALS = contour auxiliary labels supervision; ABL = active boundary loss; CA = contour attention.
In order to evaluate the impact of the weight
Sensitivity Analysis of
In our proposed network, “contour pseudo label” plays a crucial role;
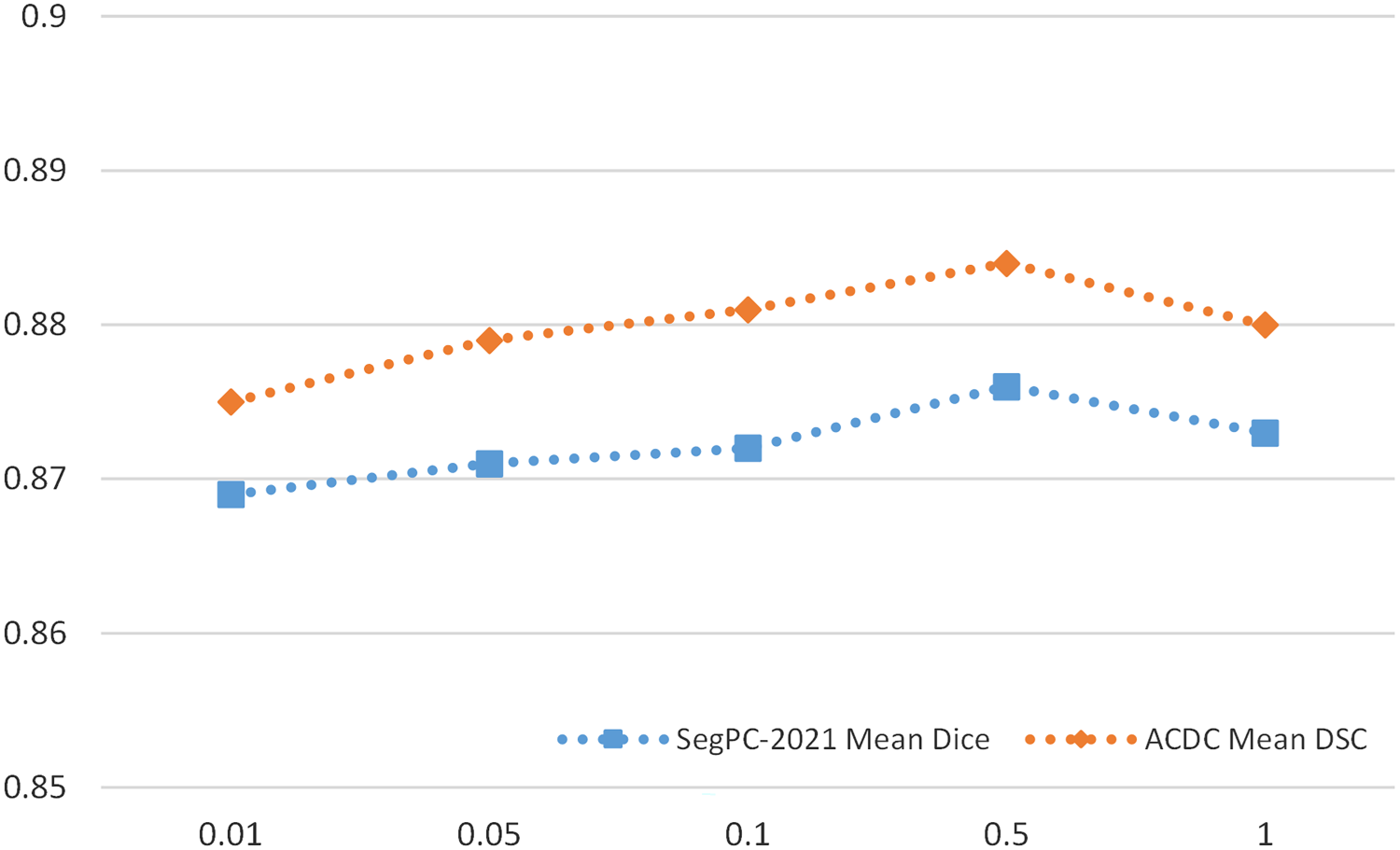
Sensitivity Analysis of Hyper-Parameter
The quality of the contour pseudo-label is crucial in our network. The Canny algorithm (Canny, 1986) has upper and lower thresholds, and we perform an inflation operation on the edges obtained by the Canny algorithm, implemented using the CV2.dilate function. Thus, we have three adjustable parameters to control the generated contour pseudo-label.
In order to obtain better results, we tried different parameter settings, using a triad combination of the low threshold, high threshold, and coefficient of expansion (denoted as L-U-Coe) for the SegPC-2021 dataset and the ACDC dataset. We investigated the changes in segmentation results when L-U-Coe took different values, and the results are shown in Figure 7. The results show that the best performance is achieved when the L-U-Coe value is set to 30-50-1. In addition, although anomalous changes were observed in both datasets at values of 40-60-1 and 50-60-2, respectively, our method was generally insensitive to changes in L-U-Coe values.
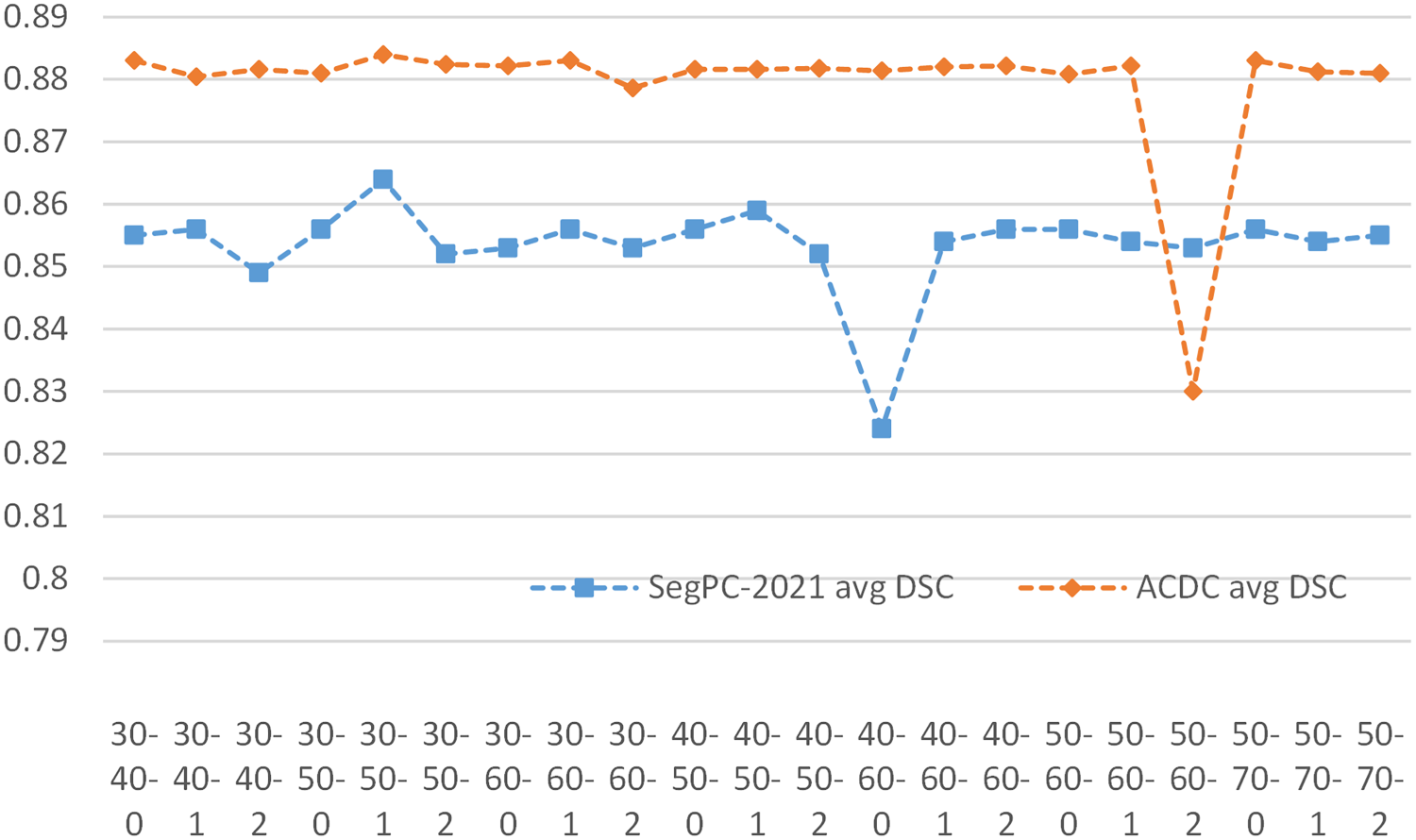
Sensitivity Analysis of Contour Pseudo Label.
In order to verify the higher quality of pseudo-labels generated by our method, we performed pseudo-label comparison experiments on the SegPC-2021 datasets as Figure 8 and Table 5.
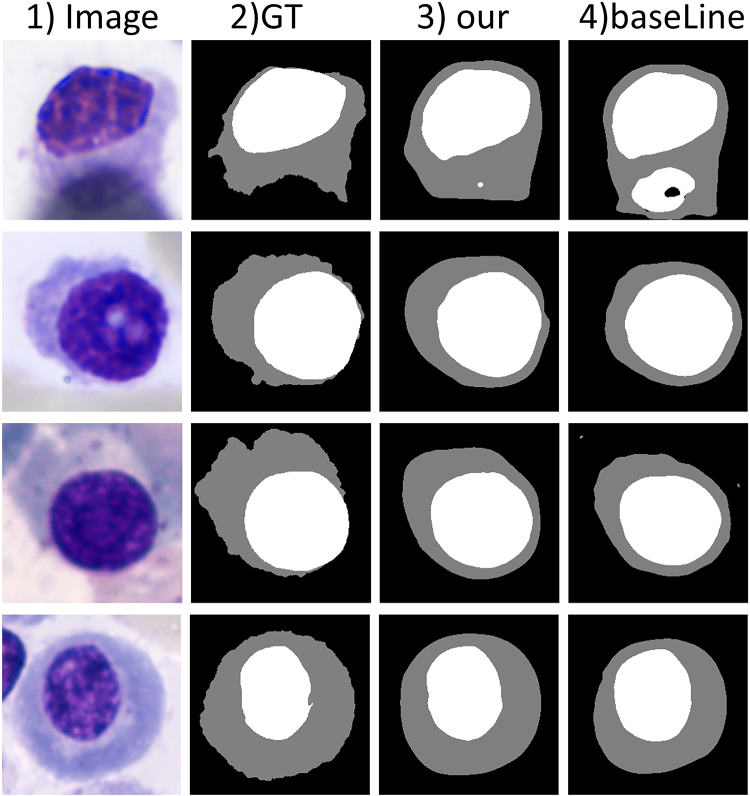
Intuitive Comparison of Pseudo Label, Where “Image” is the Raw Image, “GT” is the Real Label, “our” is our Model Generated Pseudo Labels, and “BaseLine” Removes CA and CALS Generated Pseudo Label From our Model. Note. CA = Contour Attention; CALS = Contour Auxiliary labels supervision.
Pseudo Labels Comparisons, The First Column Presents the DSC Results Calculated Between the Pseudo Label Generated by our Model Without CA and CALS and the Ground Truth Labels.
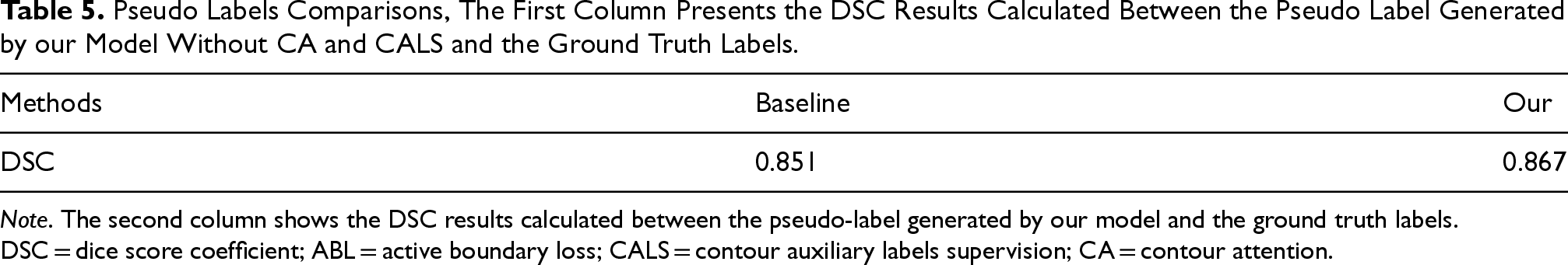

Note. The second column shows the DSC results calculated between the pseudo-label generated by our model and the ground truth labels. DSC = dice score coefficient; ABL = active boundary loss; CALS = contour auxiliary labels supervision; CA = contour attention.
Analyzing Figure 8 and Table 5, we can conclude that our model is closer to the ground truth labels compared to the baseline. This finding is crucial for guiding accurate segmentation of the nucleus and cytoplasm.
In this paper, we design a contour-aware scribble-supervised method for medical image segmentation. Our main idea is to enhance the model’s ability to accurately recognize cell contours and to compensate for the inherent limitations of scribble annotations in providing explicit contour information. We propose a contour pseudo-labels supervision module that uses contour pseudo-labels to guide the model in learning contour representations. We also propose the CA mechanism, which focuses the model’s learning on key contour regions, thereby ensuring more accurate segmentation results, especially at cell boundaries. Finally, we constructed an MM plasma cell scribble dataset based on the SegPC-2021 dataset and evaluated CMPLS on this dataset as well as two benchmark datasets. The results show that our method outperforms other weakly supervised methods based on scribble annotations.
In the future, we will explore comprehensive improvements to enhance the performance of scribble supervision methods on the SegPC-2021 dataset.
Footnotes
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Chongqing Natural Science Foundation Innovation and Development Joint Fund (CSTB2023NSCQ-LZX0109), Chongqing Technology Innovation & Application Development Key Project (cstb2022tiad-kpx0148), and Fundamental Research Funds for the Central Universities (No. 2022CDJYGRH-001).
Declaration of Competing Interest
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.