Abstract
Ontology embeddings map classes, roles, and individuals in ontologies into
Introduction
Several methods have been developed to embed Description Logic theories or ontologies in vector spaces (Chen et al., 2020a, 2021; Jackermeier et al., 2024; Kulmanov et al., 2019; Mondal et al., 2021; Özcep et al., 2023; Peng et al., 2022; Xiong et al., 2022). These embedding methods preserve some aspects of the semantics in the vector space, and may enable the computation of semantic similarity, inferring axioms that are entailed, and predicting axioms that are not entailed but may be added to the theory. For the lightweight Description Logic
Advances on different geometric embedding methods have usually focused on the expressiveness of the embedding methods; originally, hyperballs (Kulmanov et al., 2019) where used to represent the interpretation of concept symbols, yet hyperballs are not closed under intersection. Therefore, axis-aligned boxes were introduced (Jackermeier et al., 2024; Peng et al., 2022; Xiong et al., 2022). Furthermore,
We evaluate geometric embedding methods and incorporate deductive inference into the training process. We use the ELEmbeddings (Kulmanov et al., 2019), ELBE (Peng et al., 2022), and
Our main contributions are as follows:
We propose loss functions that incorporate negative samples in all normal forms and account for deductive closure during training. We introduce a fast approximate algorithm for computing the deductive closure of an We formulate evaluation methods for knowledge base completion that account for the deductive closure during evaluation.
This is an extended version of our previous work (Mashkova et al., 2024). Here, we include a more comprehensive treatment of computing the deductive closure and using the deductive closure with
Description Logic
Let
Normalized Forms of
Generalized Concept Inclusions (GCIs) and Role Inclusions (RIs).

Normalized Forms of

Here,
Normalization Rules for TBox for
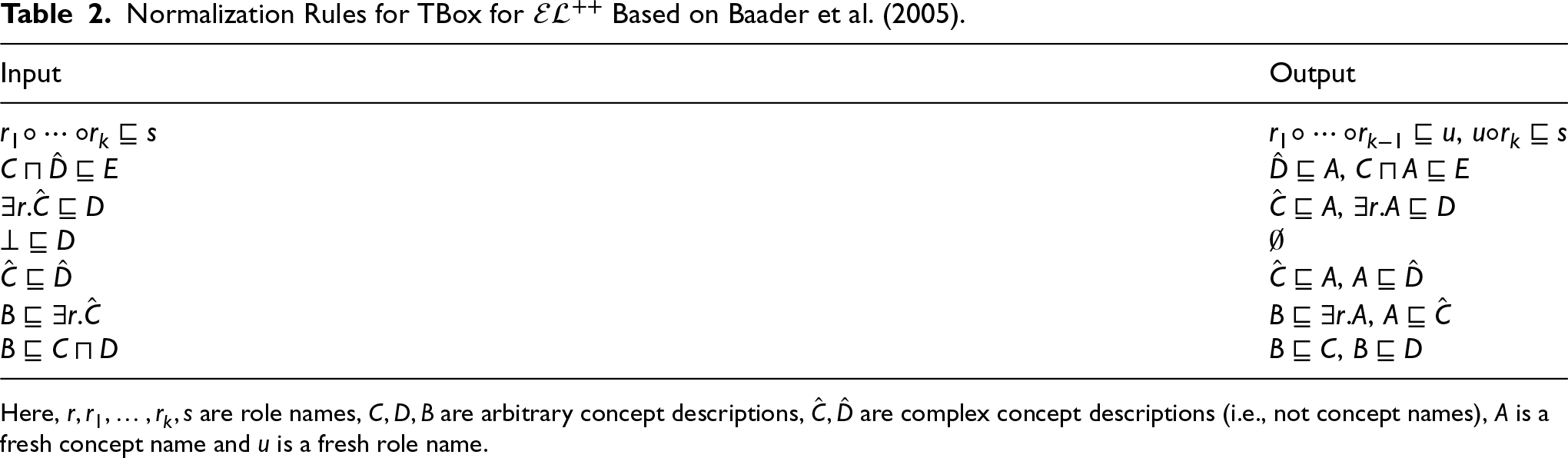

Here,
To define the semantics of an 
An interpretation
The task of knowledge base completion is the addition (or prediction) of axioms which hold yet are not represented in the knowledge base. We call the task “ontology completion” when exclusively TBox axioms are predicted. The task of knowledge base completion may encompass both deductive (Jiang et al., 2012; Sato et al., 2018) and inductive (Bouraoui et al., 2017; d’Amato et al., 2012) inference processes and give rise to two subtly different tasks: adding only “novel” axioms to a knowledge base that are not in the deductive closure of the knowledge base, and adding axioms that are in the deductive closure as well as some “novel” axioms that are not deductively inferred; both tasks are related but differ in how they are evaluated.
Inductive inference, analogously to knowledge graph completion (Chen et al., 2020b), predicts axioms based on patterns and regularities within the knowledge base. Knowledge base completion, or ontology completion, can be further distinguished based on the information that is used to predict “novel” axioms. We distinguish between two approaches to knowledge base completion: (1) knowledge base completion which relies solely on (formalized) information within the knowledge base to predict new axioms, and (2) knowledge base completion which incorporates side information, such as text, to enhance the prediction of new axioms. Here, we mainly consider the first case.
Deductive Closure
The deductive closure of a theory
Related Work
Graph-Based Ontology Embeddings
Graph-based ontology embeddings rely on a construction (projection) of graphs from ontology axioms mapping ontology classes, individuals and roles to nodes and labeled edges (Zhapa-Camacho & Hoehndorf, 2023). Embeddings for nodes and edge labels are optimized following two strategies: by generating random walks and using a sequence learning method such as Word2Vec (Mikolov et al., 2013) or by using Knowledge Graph Embedding (KGE) methods (Wang et al., 2017). These type of methods have been shown effective on knowledge base and ontology completion (Chen et al., 2021) and have been applied to domain-specific tasks such as PPI prediction (Chen et al., 2021) or gene–disease association prediction (Althagafi et al., 2024; Chen et al., 2020a). Graph-based methods rely on adjacency information of the ontology structure but cannot easily handle logical operators and do not approximate ontology models. Therefore, graph-based methods are not “faithful,” that is, do not approximate models, do not allow determining whether statements are “true” in these models, and therefore cannot be used to perform semantic entailment.
Geometric-Based Ontology Embeddings
Multiple methods have been developed for the geometric construction of models for the
Knowledge Base Completion Task
Several recent advancements in the knowledge base completion rely on side information as included in large language models (LLMs). Ji et al. (2023) explores how pretrained language models can be utilized for incorporating one ontology into another, with the main focus on inconsistency handling and ontology coherence. HalTon (Cao et al., 2023) addresses the task of event ontology completion via simultaneous event clustering, hierarchy expansion and type naming utilizing BERT (Devlin et al., 2019) for instance encoding. Li et al. (2024) formulates knowledge base completion task as a Natural Language Inference (NLI) problem and examines how this approach may be combined with concept embeddings for identifying missing knowledge in ontologies. As for other approaches, Mežnar et al. (2022) proposes a method that converts an ontology into a graph to recommend missing edges using structure-only link analysis methods, Shiraishi and Kaneiwa (2024) constructs matrix-based ontology embeddings which capture the global and local information for subsumption prediction. All these methods use side information from LLMs and would not be applicable, for example, in the case where a knowledge base is private or consists of only identifiers; we do not consider methods based on pre-trained LLMs here as baselines.
Negative Sampling and Objective Functions
Currently available geometric ontology embedding models which construct a model of an ontology by optimizing some objective function usually sample negative examples during training phase (Jackermeier et al., 2024; Kulmanov et al., 2019; Mohapatra et al., 2021; Mondal et al., 2021; Peng et al., 2022; Yang et al., 2025). This operation prevents overgeneralization of learned embeddings and trivial satisfiability in case a model collapses (Kulmanov et al., 2019; Yang et al., 2025) by incorporating additional constraints within a model. Ontology embedding methods select negatives by replacing one of the concepts with a randomly chosen one (either from the set of all concept names, or a subset thereof). ELEmbeddings, ELBE and
ELEmbeddings Negative Losses
For negative loss construction in ELEmbeddings, we employ notations from the ELEmbeddings method where
For ELEmbeddings, as reflected in Eq. (1), we use the original GCI1-BOT loss for disjoint classes; although non-containment of ball corresponding to 
The same logic applies for the negative loss in Eq. (2) where we minimize overlap between the translated ball corresponding to class 
Negative loss (3) is constructed similarly to the 
The second and the third part force the center corresponding to 
In the original method losses for axioms of type GCI0-BOT and GCI3-BOT force radii of unsatisfiable classes to become 

ELBE is a model that relies on boxes instead of balls. Here, similarly,
Following the same method of negative loss construction for ELEmbeddings, we use GCI1-BOT loss as a negative loss for 
Since axis-aligned hyperrectangles are closed under intersection, we also use GCI1-BOT for the intersection of boxes representing 
This property also allows us to interpret each negative sample for 
Other negative losses have the form similar to the ones constructed for ELEmbeddings:



Equations (13) and (14) are constructed in a similar fashion as for ELBE based on the GCI1-BOT loss which penalizes the element-wise distance 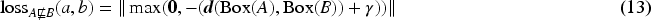

Negative losses 15–17 encourage boxes to be non-empty:



The GCI3 negative loss reflects the structure of the original GCI3 loss:
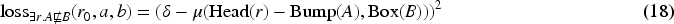
In the case of knowledge base completion where the deductive closure contains potentially many non-trivial entailed axioms, the random sampling approach for negatives may lead to suboptimal learning since some of the axioms treated as negatives may be entailed (and should therefore be true in any model, in particular the one constructed by the geometric embedding method). As an example, let us consider the simple ontology consisting of two axioms:
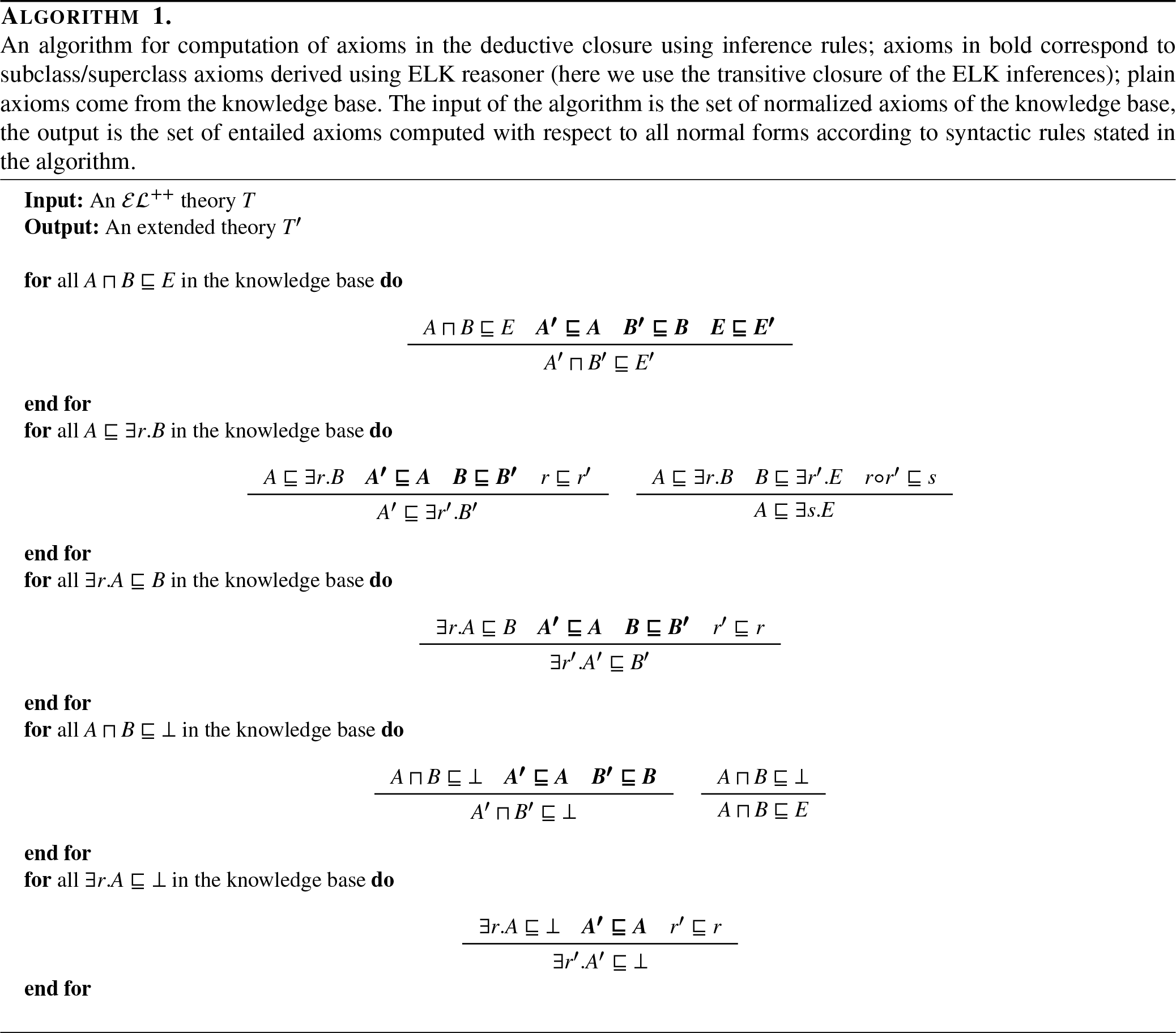

We suggest to filter selected negatives during training based on the deductive closure of the knowledge base: for each randomly generated axiom to be used as negative, we check whether it is present in the deductive closure and, if it is, we delete it.
In the task of knowledge base completion with many non-trivial entailed axioms, the deductive closure can also be used to modify the evaluation metrics, or define novel evaluation metrics that distinguish between entailed and non-entailed axioms. So far, ontology embedding methods that have been applied to the task of knowledge base completion have used evaluation measures that are taken from the task of knowledge graph completion; in particular, they only evaluate knowledge base completion using axioms that are “novel” and not entailed. However, any entailed axiom will be true in all models of the knowledge base, and therefore also in the geometric model that is constructed by the embedding method.
We suggest to filter entailed axioms from training or test sets when the aim is to predict “novel” (i.e., non-entailed) knowledge. The geometric embedding methods generate models making all entailed axioms true in all models. It is expected that methods explicitly constructing models preferentially make entailed axioms true and rank them higher than non-entailed axioms. If the evaluation is based solely on non-entailed axioms, it will consider all similar inferred axioms false, and to avoid this, we may filter such axioms from the ranking list. The more axioms are filtered, the more entailed axioms are predicted by a model.
Experiments
Datasets
Gene Ontology & STRING Data
Following previous works (Jackermeier et al., 2024; Kulmanov et al., 2019; Peng et al., 2022) we use common benchmarks for knowledge-base completion, in particular a task that predicts PPIs based on the functions of proteins. We also use the same data for the task of protein function prediction. For these tasks we use two datasets, each of them consists of the Gene Ontology (GO) (Consortium, 2015) with all its axioms, PPIs and protein function axioms extracted from the STRING database (Mering, 2003); each dataset focuses on only yeast proteins. GO is formalized using OWL 2 EL (Golbreich & Horrocks, 2007).
For the PPI yeast network we use the built-in dataset PPIYeastDataset available in the mOWL (Zhapa-Camacho et al., 2022) Python library (release 0.2.1) where axioms of interest are split randomly into train, validation and test datasets in ratio 90:5:5 keeping pairs of symmetric PPI axioms within the same dataset, and other axioms are placed into the training part; validation and test sets are made up of TBox axioms of type
Food Ontology & GALEN Ontology
Food Ontology (Dooley et al., 2018) contains structured information about foods formalized in
Evaluation Scores and Metrics
For GO & STRING data, we predict GCI2 axioms of type
The predictive performance is measured by the Hits@n metrics for 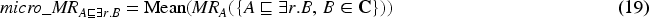



Additionally, we remove axioms represented in the train set or deductive closures (see Section 5) to obtain corresponding filtered metrics (FHits@n, FMR, FAUC). In related work focusing on knowledge graph completion or knowledge base completion tasks (Bordes et al., 2013; Kulmanov et al., 2019; Peng et al., 2022; Wang et al., 2014), filtered metrics are computed by removing axioms presented within the train set from the list of all ranked axioms. This filtration is applied to eliminate statements learnt by a model during training phase which are therefore likely to have lower rank and to evaluate the predictive performance of a model in a more fair setting.
All models are optimized with respect to the sum of individual GCI losses (here we define the loss in most general case using all positive and all negative losses):

All model architectures are built using the mOWL (Zhapa-Camacho et al., 2022) library on top of mOWL’s base models. All models were trained using the same fixed random seed.
All models are trained for 2,000 epochs for STRING & GO datasets and 800 epochs for the Food Ontology and GALEN datasets with batch size of 32,768. Training and optimization is performed using Pytorch with Adam optimizer (Kingma & Ba, 2015) and ReduceLROnPlateau scheduler with patience parameter
We evaluate whether adding negative losses for all normal forms will allow for the construction of a better model and improve the performance in the task of knowledge base completion. We test the effect of the expanded negative sampling and negative losses first on a small ontology that can be embedded and visualized in 2D space, and then on a larger application. We formulate and add negative losses for all normal forms given by equations (1)–(17).
First, we investigate a simple example corresponding to the task of protein function prediction using the ELEmbeddings model. Let us consider an ontology consisting of two axioms stating that there are two disjoint functions

ELEmbeddings example. Dashed circles represent translated classes by role vector corresponding to
Since we are interested in predicting not only axioms of type
Subsumption Prediction Experiments on Food Ontology.

‘‘l’’ corresponds to all negative losses, ‘‘l+n’’ means a model was trained using all negative losses and negatives filtering. For each model we report non-filtered metrics (NF) and filtered metrics with respect to the deductive closure of the train and the test set combined together (F). For macro_MR and micro_MR we additionally report the difference between filtered and non-filtered metrics (NF-F) to check how much of entailed knowledge is predicted on average. Values in
Subsumption Prediction Experiments on GALEN Ontology.

‘‘l’’ corresponds to all negative losses, ‘‘l+n’’ means a model was trained using all negative losses and negatives filtering. For each model we report non-filtered metrics (NF) and filtered metrics with respect to the deductive closure of the train and the test set combined together (F). For macro_MR and micro_MR we additionally report the difference between filtered and non-filtered metrics (NF-F) to check how much of entailed knowledge is predicted on average. Values in
Additionally, we evaluate the performance on a standard benchmark set for PPI prediction (see Table 5). For this task, the test axioms are of the type GCI2. We observe that ELEmbeddings and ELBE with negative losses for all normal forms integrated demonstrate superior performance compared to their initial configurations in terms of Hits@n metrics; it also allows
Protein–Protein Interaction Prediction Experiments on Yeast Proteins.

‘‘l’’ corresponds to all negative losses, ‘‘l+n’’ means a model was trained using all negative losses and negatives filtering. Non-filtered metrics are reported. Values in
To summarize the above mentioned observations, we note that in some cases additional negative losses may decrease the ability of models to predict new axioms and encourage models to predict entailed knowledge first (as, e.g., in protein function prediction case) thus leading to construction of a more accurate model of a theory. Since there is a tradeoff between prediction of novel and entailed knowledge, additional negative losses may demonstrate worse performance on novel knowledge prediction.
Using the example introduced above and the ELEmbeddings embedding model, we demonstrate that negatives filtration may be beneficial for constructing a model of a theory. Apart from axioms mentioned earlier, that is,

ELEmbeddings example. Dashed circles represent translated classes by role vector corresponding to
Tables 3–5 show results in the tasks of PPI and subsumption prediction. We find that excluding axioms in the deductive closure for negative selection slightly improves or yields similar results. One possible reason is that a randomly chosen axiom is very unlikely to be entailed since very few axioms are entailed compared to all possible axioms to choose from.
Because the chance of selecting an entailed axiom as a negative depends on the knowledge base on which the embedding method is applied, we perform additional experiments on Food Ontology with ELEmbeddings model where we bias the selection of negatives; we chose between 100% negatives to 0% negatives from the entailed axioms. We find that reducing the number of entailed axioms from the negatives has an effect to improve performance and the effect increases the more axioms would be chosen from the entailed ones (see Figure 3).

Metrics reported for biased fraction of random negatives combined with entailed axioms from the precomputed deductive closure. (a) H@1, H@10, H@100 and ROC AUC; (b) macro_MR.
We compute filtered metrics for the protein function and subsumption prediction tasks. Both of them account for entailed axioms prediction since if, for example,
For function prediction and subsumption prediction, we employ filtration of metrics based on the deductive closure of the train set and of the test set. Tables 3, 4 and 6 contain results for subsumption prediction on Food Ontology, subsumption prediction on GALEN ontology and function prediction on GO, respectively.
Protein Function Prediction Experiments on Yeast proteins.

‘‘l’’ corresponds to all negative losses, ‘‘l+n’’ means a model was trained using all negative losses and negatives filtering. For each model we report non-filtered metrics (NF) and filtered metrics with respect to the deductive closure of the train and the test set combined together (F). For macro_MR and micro_MR we additionally report the difference between filtered and non-filtered metrics (NF-F) to check how much of entailed knowledge is predicted on average. Values in
Our findings suggest that the baseline ELEmbeddings predicts primarily entailed axioms of GCI2 type, yet for GCI0 on Food Ontology the model predicts “novel” knowledge first whereas the model modifications with additional negative losses and negatives filtration derive entailed knowledge in the first place. For the GALEN ontology, however, the situation is similar to the protein function prediction case, that is, novel knowledge is predicted in the first place for modifications with additional negative losses and negatives filtration. This may indicate model construction where many classes overlap or “collapse” for all negative losses and negatives filtering case since the GALEN ontology does not contain disjointness axioms and consequently no classes will be separated by the model. The same holds for ELBE and
We evaluated properties of ELEmbeddings, ELBE and
We have introduced a method to compute the deductive closure of
Use of the deductive closure is useful not only in evaluation but also when selecting negatives. In formal knowledge bases, there are at least two ways in which negatives for axioms can be chosen: they are either non-entailed axioms, or they are axioms whose negation is entailed. However, in no case should entailed axioms be considered as negatives; we demonstrate that filtering entailed axioms from selected negatives during training improves the performance of the embedding method consistently in knowledge base completion (and, obviously, more so when entailed axioms are considered as positives during evaluation).
While we only report our experiments with ELEmbeddings, ELBE, and
Footnotes
Acknowledgments
This work has been supported by funding from King Abdullah University of Science and Technology (KAUST) Office of Sponsored Research (OSR) under Award No. URF/1/4355-01-01, URF/1/4675-01-01, URF/1/4697-01-01, URF/1/5041-01-01, and REI/1/5334-01-01. This work was supported by the SDAIA–KAUST Center of Excellence in Data Science and Artificial Intelligence (SDAIA–KAUST AI), by funding from King Abdullah University of Science and Technology (KAUST) – KAUST Center of Excellence for Smart Health (KCSH) under award number 5932, and by funding from King Abdullah University of Science and Technology (KAUST) – KAUST Center of Excellence for Generative AI under award number 5940. We acknowledge support from the KAUST Supercomputing Laboratory.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work has been supported by funding from King Abdullah University of Science and Technology (KAUST) Office of Sponsored Research (OSR) under Award No. URF/1/4675-01-01, URF/1/4697-01-01, URF/1/5041-01-01, REI/1/5235-01-01, and REI/1/5334-01-01. This work was supported by funding from King Abdullah University of Science and Technology (KAUST) -- KAUST Center of Excellence for Smart Health (KCSH), under award number 5932, and by funding from King Abdullah University of Science and Technology (KAUST) -- Center of Excellence for Generative AI, under award number 5940.
Notes
Appendix A GO & STRING Data Statistics,Train Part

| Dataset | GCI0 | GCI1 | GCI2 | GCI3 | GCI0_BOT | GCI1_BOT | GCI3_BOT | Classes | Roles | Test axioms |
|---|---|---|---|---|---|---|---|---|---|---|
| Yeast iw | 81,068 | 11,825 | 269,567 | 11,823 | 0 | 31 | 0 | 61,846 | 16 | 12,040 |
| Yeast hf | 81,068 | 11,825 | 290,433 | 11,823 | 0 | 31 | 0 | 61,850 | 16 | 1,530 |
Appendix B Food Ontology Statistics,Train Part

| GCI0 | GCI1 | GCI2 | GCI3 | GCI0_BOT | GCI1_BOT | GCI3_BOT | Classes | Roles | Test axioms |
|---|---|---|---|---|---|---|---|---|---|
| 21,795 | 1,267 | 10,719 | 897 | 0 | 495 | 0 | 24,969 | 43 | 5,752 |
Appendix C GALEN Ontology Statistics,Train Part

| GCI0 | GCI1 | GCI2 | GCI3 | GCI0_BOT | GCI1_BOT | GCI3_BOT | Classes | Roles | Test axioms |
|---|---|---|---|---|---|---|---|---|---|
| 27,339 | 15,613 | 29,618 | 15,615 | 0 | 0 | 0 | 49,223 | 888 | 667 |
Appendix D Hyperparameters
Dataset
Model
dim
lr
Yeast iw
ELEm
100
0.0001
-0.10
ELEm+l
50
0.0001
0.00
ELBE
200
0.0001
0.00
ELBE+l
200
0.0100
0.00
0.001
200
0.0010
0.01
1
0.05
200
0.0010
0.01
0.010
2
0.05
Yeast hf
ELEm
200
0.0001
0.01
ELEm+l
50
0.0001
-0.10
ELBE
200
0.0001
0.10
ELBE+l
200
0.0001
0.10
0.010
200
0.0100
0.10
4
0.20
200
0.0100
0.10
0.010
4
0.05
FoodOn
ELEm
400
0.0010
-0.10
ELEm+l
400
0.0010
-0.10
ELBE
200
0.0100
0.10
ELBE+l
200
0.0100
-0.01
0.001
100
0.0100
0.10
1
0.20
200
0.0010
0.10
0.010
4
0.10
GALEN
ELEm
400
0.0010
-0.10
ELEm+l
400
0.0010
-0.01
ELBE
100
0.0010
0.10
ELBE+l
200
0.0010
0.01
0.010
200
0.0010
0.00
4
0.05
200
0.0100
0.00
0.100
1
0.05
Appendix E Deductive Closure Computation Example
Let us add two more axioms to the simple ontology example from Section 6.4 about proteins
For GCI2 axioms
For GCI3 axioms
In this small protein function prediction example there are two disjointness axioms:
Appendix F Deductive Closure Computation Soundness
Let us show that each inference rule provides truth statements:
Let
Let
Let
Let
Let
Let
Let
Let
Let
Let
Let
Let
Let
Follows immediately from the fact that
Let
Follows immediately from the fact that
