Abstract
Objective
This study aims to comprehensively investigate the expression patterns of FK506-binding protein 3 (FKBP3) within various cancer subtypes and assess its clinical relevance alongside possible biological functions in lung adenocarcinoma (LUAD).
Methods
FKBP3 expression data, together with clinical data from hepatocellular carcinoma (LIHC) specimens, were sourced from The Cancer Genome Atlas (TCGA) database. A comparative evaluation of FKBP3 expression levels across tumor tissues and their corresponding normal controls was conducted using the Wilcoxon rank-sum test. Chi-square testing was used to investigate the relationship between FKBP3 expression levels and clinicopathological features. Evaluation of FKBP3’s prognostic relevance was executed through Cox regression and Kaplan-Meier survival analyses. Furthermore, the relationship linking FKBP3 with immune cell infiltration was investigated.
Results
Pan-cancer analysis revealed that FKBP3 exhibited markedly higher expression across multiple tumor types, showing statistically significant differences observed in bladder cancer (BLCA), breast cancer (BRCA), cervical cancer, colorectal cancer (COAD), esophageal cancer (ESCA), glioblastoma, head, and neck squamous cell carcinoma, kidney clear cell carcinoma (KIRC), LIHC, LUAD, lung squamous cell carcinoma (LUSC), kidney chromophobe (KICH), kidney papillary cell carcinoma, prostate cancer (PRAD), gastric cancer, and thymoma. Subsequent analysis concentrating on LUAD showed that FKBP3 expression levels exhibited marked elevation in both messenger RNA (mRNA) and protein levels in tumor tissues relative to normal tissues. Receiver operating characteristic (ROC) curve analysis revealed robust diagnostic potential of FKBP3 (area under the curve [AUC] = 0.894). Clinicopathological analysis showed that elevated FKBP3 expression levels significantly correlated with pathological N stage and residual tumor status (P < .05). Cox regression analysis established FKBP3 as an independent unfavorable prognostic factor regarding overall survival (OS) in LUAD subjects. A nomogram model was developed utilizing FKBP3 expression for prognostic prediction, which exhibited excellent calibration performance. Furthermore, Kaplan-Meier and time-dependent ROC analyses additionally validated the prognostic significance of FKBP3 in LUAD. Bioinformatics analysis indicated that FKBP3-related genes were markedly enriched within cancer-associated pathways, cellular senescence, and PD-L1/PD-1 signaling pathways. Immune infiltration analysis demonstrated that FKBP3 expression was inversely associated with most immune cells, particularly plasmacytoid dendritic cells (pDCs), mast cells, natural killer (NK) cells, and immature dendritic cells (iDCs), while exhibiting a positive correlation with Th2 cells. These findings indicate that FKBP3 may facilitate immune evasion through regulating the tumor immune microenvironment.
Conclusion
In summary, FKBP3 holds potential diagnostic, and prognostic value in LUAD and may play a role in regulating tumor immune processes, making it a promising therapeutic target.
Introduction
Lung cancer persists as a principal driver of cancer development and fatality across the globe, creating significant public health challenges. 1 lung adenocarcinoma (LUAD) emerges as the dominant variant within the disease spectrum, comprising nearly 50% of non-small cell lung cancer instances. 2 Evidence from the International Agency for Research on Cancer, a division operating under the World Health Organization, demonstrates that new LUAD case detection worldwide shows sustained annual escalation, with fatality rates maintaining considerably high values. These patterns underscore the urgent necessity for effective strategies in LUAD prevention, diagnosis, and treatment, making this a critical priority in oncology. 3
For LUAD therapy, new agents, such as immune checkpoint drugs (such as pembrolizumab and nivolumab) and epidermal growth factor receptor (EGFR), anaplastic lymphoma kinase (ALK), and other oncogenic driver-directed tyrosine kinase agents, have markedly enhanced survival for selected patient groups, holding optimistic prospects for therapy for LUAD. 4 Nonetheless, response to therapy differs greatly between individuals. The latter is mainly attributed to broad-range phenotypic and molecular heterogeneity in LUAD, leading to varied responses to therapy in tumor cells. 5 Additionally, issues that involve relapse of the tumor, spread to distant locations, metastasis, and drug resistance development remain crucial. Numerous individuals develop disease progression while experiencing response initiation periods, and the precise prediction of the clinical outcome is low. 6 Thus, determination of credible biomarkers that are capable of predicting patient outcome with exactitude and directing personalized therapy, and explanation of alternative targets for therapy, becomes vital for enhancing survival discovery in individuals with LUAD. 7
The FKBP (FK506-binding protein) protein family constitutes a group of peptidyl-prolyl cis-trans isomerases that are widely distributed across eukaryotic cells and are characterized by their distinct characteristic to bind the FK506 immunosuppressant. 8 The family members catalyze prolyl peptide bond cis-trans isomerization within polypeptide chains and thus participate in protein folding, transport, assembly, and degradation. The FKBP family members are significant and are involved in a variety of biological activities, including signal transduction, cell cycle regulation, and immune modulation. 9 There are over 20 FKBP family members identified currently, each one exhibiting a distinct pattern for expression and biological activity across various tissues and cell types. 10
Previous studies have shown that FKBP family components exert essential functions in tumor onset, development, invasiveness, and metastatic spread. 11 As an example, FKBP12, among the initially discovered components, controls cellular growth and metabolic processes through interaction with the mechanistic target of rapamycin complex 1 (mTORC1) complex, with aberrant expression being strongly associated with malignant advancement in various cancers. 12 Evidence shows that FKBP5 influences proliferative activity and metastatic behavior of tumor cells in breast cancer (BRCA) via regulation of the glucocorticoid receptor signaling pathway. 13 Being an important component within the FKBP family, FKBP3 has lately attracted growing interest in cancer research. 14 Recent findings suggest elevated FKBP3 levels within hepatocellular carcinoma (LIHC), where it enhances tumor cell growth and invasiveness through phosphatidylinositol 3-kinase/protein kinase B (PI3K/AKT) signaling pathway stimulation. 15 Within colorectal cancer (COAD) contexts, FKBP3 engages with β-catenin, amplifying Wnt/β-catenin signaling cascade functionality, consequently facilitating tumor emergence and advancement. 16 Such findings demonstrate FKBP3’s significant modulatory role during tumorigenesis mechanisms. 17
However, compared to other tumor types, research on FKBP3 in LUAD remains in its early stages. Currently, it is unclear whether FKBP3 expression is dysregulated in LUAD tissues, whether it is upregulated or downregulated, and how its expression levels are correlated with clinicopathological features, including tumor stage, lymph node metastasis status, patient age, and gender. Moreover, although numerous molecular markers have been identified for prognostic evaluation in LUAD, the prognostic value of FKBP3 has not been fully explored. Whether FKBP3 can serve as an independent prognostic indicator to assist clinicians in assessing patient outcomes remains to be determined.
A comprehensive investigation of FKBP3 expression patterns in LUAD, a systematic evaluation of its prognostic significance, and elucidation of its potential role in immune evasion are of great scientific importance for understanding the molecular mechanisms underlying LUAD development and progression. 18 On one hand, clarifying the expression pattern of FKBP3 and its link to clinicopathological characteristics may help identify novel molecular biomarkers for LUAD, providing a theoretical foundation for more accurate prognostic assessment and the development of personalized treatment strategies. On the other hand, uncovering the molecular mechanisms by which FKBP3 contributes to immune evasion could offer new targets for the development of FKBP3-based immunotherapeutic strategies, potentially overcoming current therapeutic limitations and enhancing therapeutic effects and quality of life in LUAD patients. Therefore, research on FKBP3 in LUAD holds significant clinical relevance and promising research potential.
Materials and Methods
Bioinformatics Analysis Based on the TCGA Database
Clinical information alongside FKBP3 expression data (in transcripts per million (TPM) format) from LUAD patients was obtained through the primary repository of The Cancer Genome Atlas (TCGA) (
Protein Expression and Immunohistochemical Analysis of FKBP3
Through the University of Alabama at Birmingham Cancer Data Analysis Portal (UALCAN) portal at the University of Alabama at Birmingham, Clinical Proteomic Tumor Analysis Consortium data enabled a thorough examination of protein expression variations of FKBP3 in LUAD compared to normal tissues.
21
The Human Protein Atlas database (
Biological Function of FKBP3 in Lung Adenocarcinoma
A protein-protein interaction (PPI) network with FKBP3 as its center was established through the Search Tool for the Retrieval of Interacting Genes/Proteins (STRING) online database for comprehending biological mechanisms and signaling cascades involving FKBP3 in LUAD, while network visualization was performed using Cytoscape software. 22 Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis was then conducted, incorporating FKBP3 together with its associated genes. 23
Statistical Analysis
R software (version 4.3.3) 24 was utilized for statistical computations and visual representations in this research. Tumor versus non-tumor FKBP3 levels were assessed via Wilcoxon rank-sum and signed-rank tests. Chi-square analysis explored FKBP3-clinicopathological relationships. The Kaplan-Meier methodology addressed survival outcomes. Cox regression models determined prognostic factors for LUAD survival, enabling nomogram construction. The ggplot2 package mapped FKBP3-immune cell associations. Two-tailed tests were applied throughout, with P < 0.05 denoting significance.
Results
Pan-cancer Expression Analysis of FKBP3
The study revealed marked FKBP3 upregulation in multiple tumor types, including bladder cancer (BLCA), cholangiocarcinoma (CHOL), BRCA, cervical cancer, COAD, esophageal cancer (ESCA), glioblastoma, head and neck squamous cell carcinoma, kidney clear cell carcinoma (KIRC), LIHC, LUAD, lung squamous cell carcinoma (LUSC), kidney chromophobe (KICH), kidney papillary cell carcinoma, prostate cancer (PRAD), gastric cancer, and thymoma. For other cancer types, FKBP3 expression exhibited no notable variations (Figure 1A).
Pan-cancer Expression Analysis of FKBP3.
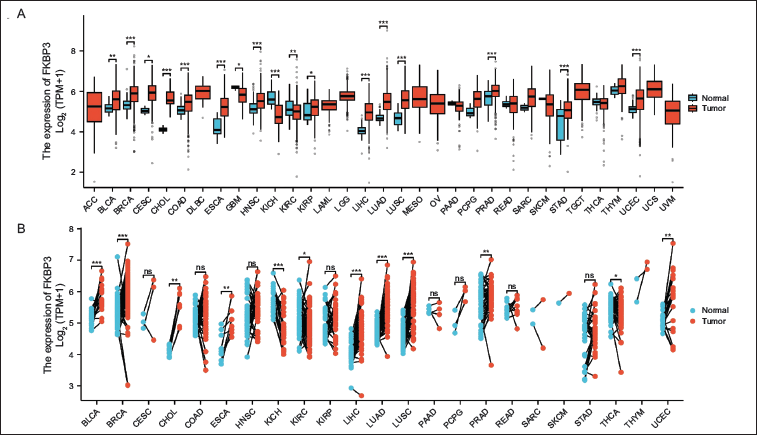
Moreover, comparative expression evaluation involving neoplastic and paired normal samples confirmed substantial FKBP3 elevation within BLCA, CHOL, BRCA, ESCA, KIRC, LIHC, LUAD, LUSC, KICH, PRAD, and thyroid cancer (Figure 1B).
FKBP3 Expression in LUAD
The TCGA database provided 516 tumor samples of LUAD together with 59 samples of normal tissue. The Wilcoxon rank-sum test was employed to assess FKBP3 mRNA levels between malignant and normal specimens. Cancerous specimens displayed markedly higher FKBP3 concentrations compared to normal counterparts (Figure 2A). Furthermore, the Wilcoxon signed-rank test applied to 59 paired sets of neoplastic and adjacent non-neoplastic specimens yielded consistent observations; neoplastic specimens demonstrated markedly increased FKBP3 mRNA concentrations (Figure 2B). Additionally, a receiver operating characteristic (ROC) curve was generated to evaluate FKBP3’s diagnostic performance in differentiating neoplastic and normal samples. The area under the ROC curve, or area under the curve (AUC), reached 0.894 (95% CI, 0.864-0.924), demonstrating high diagnostic capacity of FKBP3 for distinguishing neoplastic and normal tissues (Figure 2C). These results indicate that FKBP3 could function as a valuable diagnostic biomarker for LUAD. For additional confirmation of these findings, FKBP3 protein expression levels underwent examination through the UALCAN database. Findings demonstrated significantly elevated FKBP3 protein expression within LUAD tissues relative to normal tissues (Figure 2D). Immunohistochemical examination showed widespread FKBP3 expression throughout LUAD tissues, while normal tissues displayed merely sporadic expression patterns (Figure 2E-F).
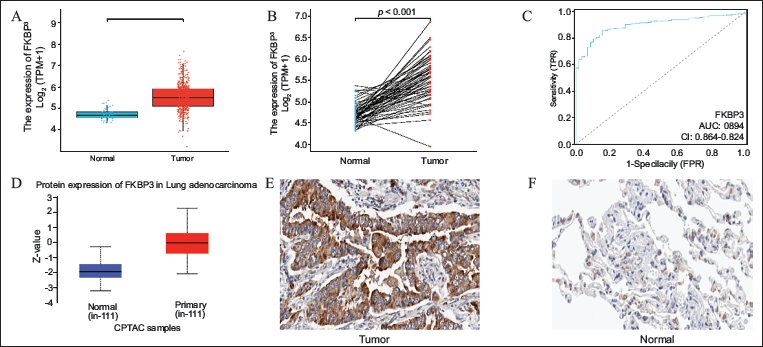
A: Comparison of FKBP3 mRNA expression between 516 tumor tissues and 59 normal tissues. B: Comparison of FKBP3 mRNA expression between 59 paired tumor and adjacent normal tissue samples. C: ROC curve showing the diagnostic performance of FKBP3 in distinguishing tumors from normal tissues in LUAD. D: UALCAN analysis showing significantly higher FKBP3 protein expression in LUAD tissues compared to normal tissues. E-F: Immunohistochemical analysis of FKBP3 expression in LUAD tissues. (Note: The original text contained incorrect references to MPV17 and FKBP5; these have been corrected to FKBP3 for consistency and accuracy.)
Correlation Between FKBP3 Expression and Clinicopathological Features
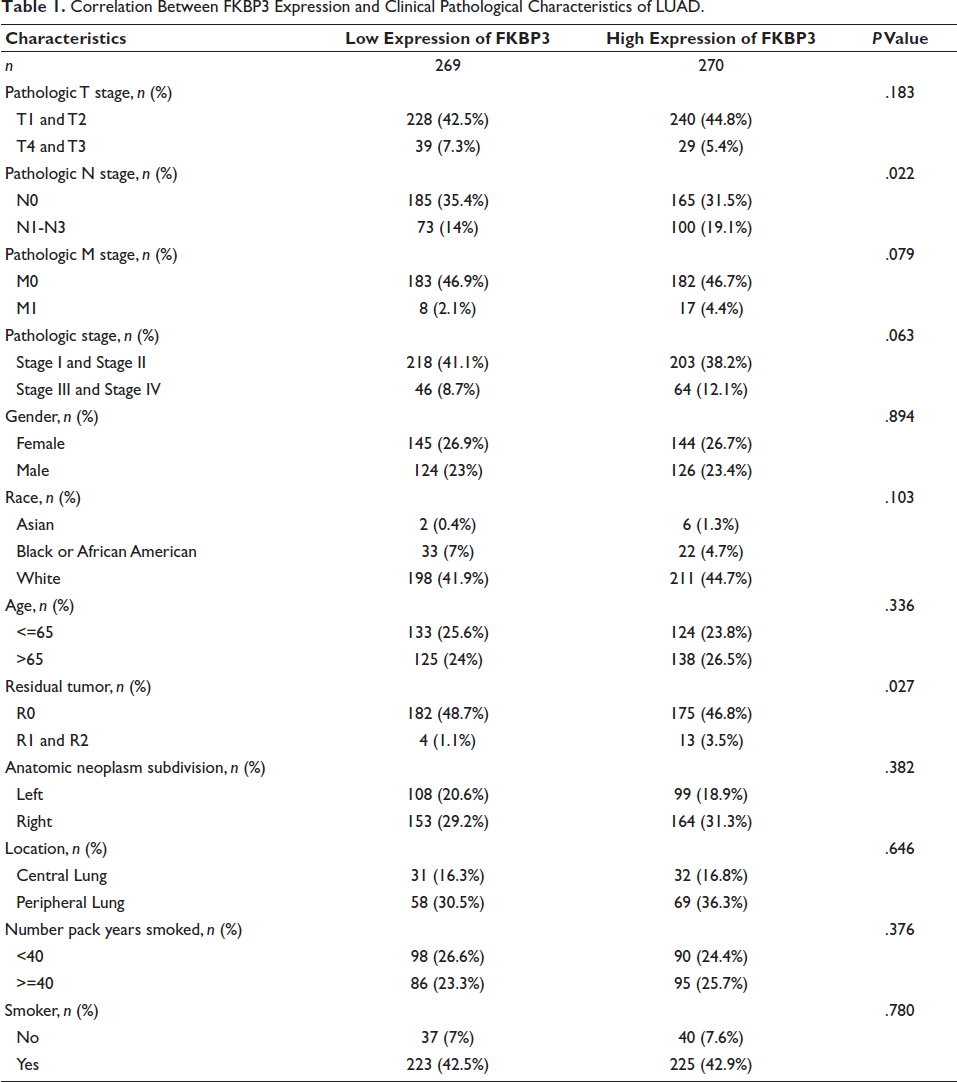
TCGA database supplied LUAD patient information encompassing pathological TNM stages, gender, race, age, residual tumor, anatomic subdivision, location, smoking history, and FKBP3 levels. Median FKBP3 values divided patients into high and low expression cohorts. Chi-square tests evaluated FKBP3-clinical parameter associations. Results demonstrated notable correlations between FKBP3 levels and pathological N stage (P = .022) along with residual tumor presence (P < .027) (Table 1).
Correlation Between FKBP3 Expression and Clinical Pathological Characteristics of LUAD.
FKBP3 is an Independent Risk Factor for Overall Survival in Lung Adenocarcinoma Patients
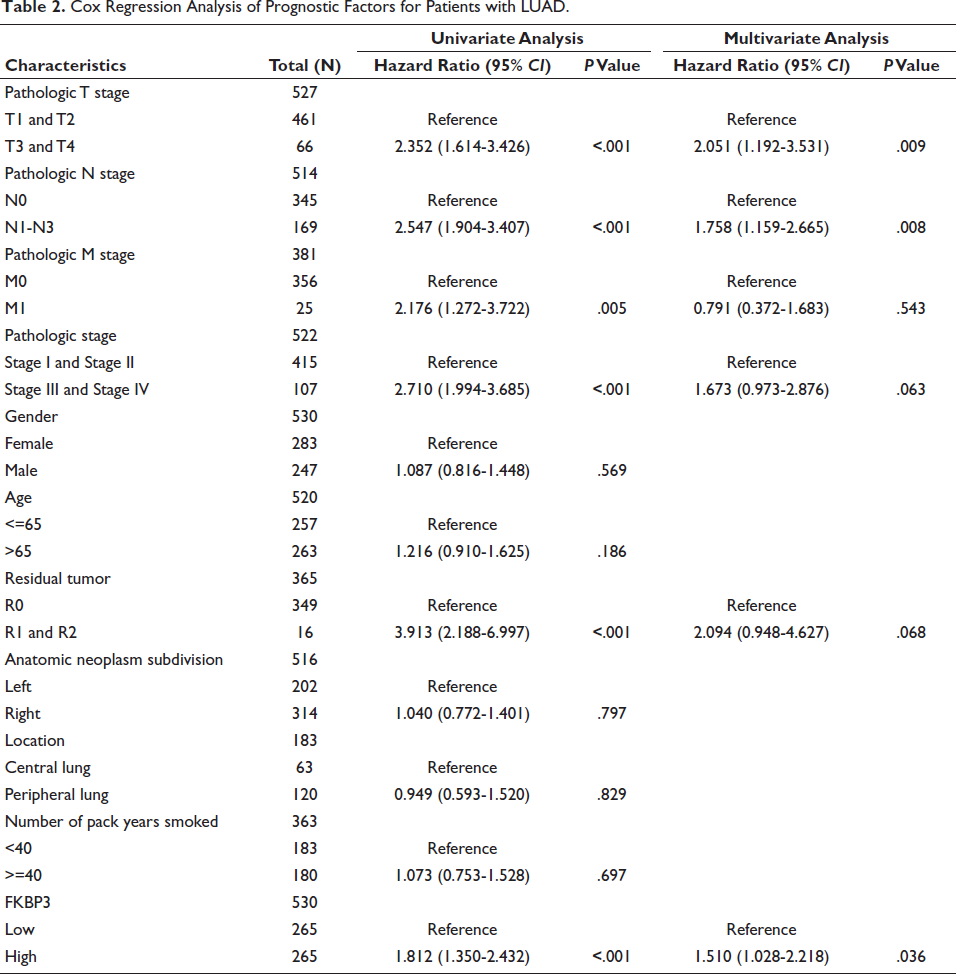
Univariate Cox regression analysis was initially employed to identify potential prognostic factors affecting overall survival (OS) in LUAD patients. Seven statistically significant factors emerged from this analysis: pathologic T stage (HR = 2.352 [95% CI: 1.614-3.426], P < .001), pathologic N stage (HR = 2.547 [95% CI: 1.904-3.407], P < .001), pathologic M stage (HR = 2.176 [95% CI: 1.272-3.722], P = .005), overall pathologic stage (HR = 2.710 [95% CI: 1.994-3.685], P < .001), residual tumor (HR = 3.913 [95% CI: 2.188-6.997], P < .001), and FKBP3 expression level (HR = 1.812 [95% CI: 1.350-2.432], P < .001), all showing statistical significance. Multivariate Cox regression analysis subsequently demonstrated that pathologic T stage (HR = 2.051 [95% CI: 1.192-3.531], P = .009), pathologic N stage (HR = 1.758 [95% CI: 1.159-2.665], P = .008), and FKBP3 expression level (HR = 1.510 [95% CI: 1.028-2.218], P = .036) served as independent prognostic factors regarding OS among LUAD patients (Table 2).
Cox Regression Analysis of Prognostic Factors for Patients with LUAD.
Construction and Validation of a Nomogram for Predicting Overall Survival in Lung Adenocarcinoma Patients
Multivariate Cox regression analysis led to the recognition of three independent prognostic factors, which facilitated the construction of a nomogram to predict OS rates at 1, 3, and 5-year intervals among LUAD patients (Figure 3A). Such a predictive instrument offers theoretical guidance for clinicians when evaluating patient prognosis.
Establishment and Verification of a Nomogram-based Prognostic System for Outcome Predictors in LUAD Subjects. A: Nomogram. B-D: Calibration Plots for the Nomogram’s Prediction of OS in LUAD Subjects Over 1, 3, and 5 Years.
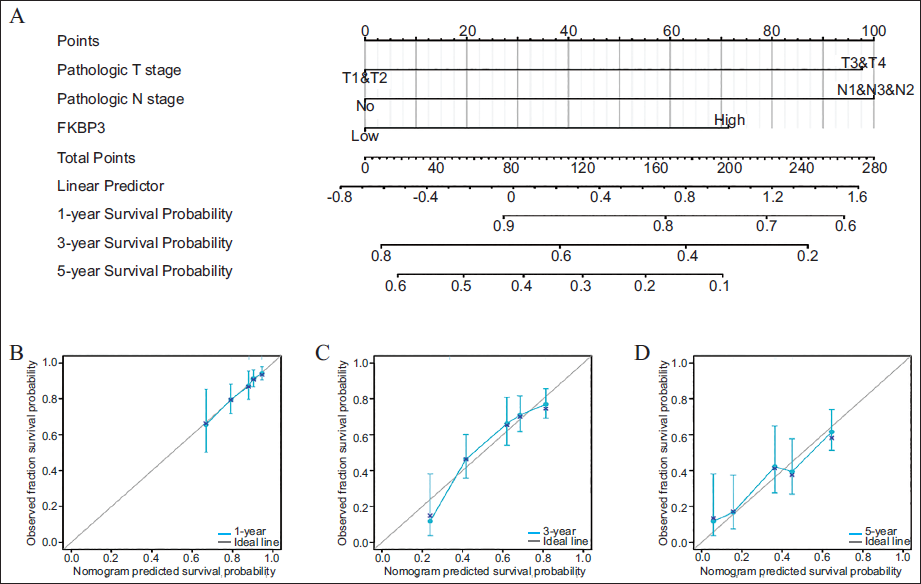
For evaluating the nomogram’s predictive accuracy, calibration curves were subsequently generated (Figures 3B-D). The x-axis displays predicted survival probabilities derived from the nomogram, whereas the y-axis shows actual observed survival probabilities (range, 0-1; equivalent to 0%-100%). The blue curve represents concordance between predicted and observed results, with the diagonal line indicating perfect prediction. The greater proximity of the blue curve to the diagonal line reflects the superior predictive accuracy of the model. In this analysis, the calibration curves showed good agreement with the ideal line, indicating that the nomogram demonstrated moderate accuracy in predicting patient prognosis.
FKBP3 as a Potential Prognostic Indicator for LUAD
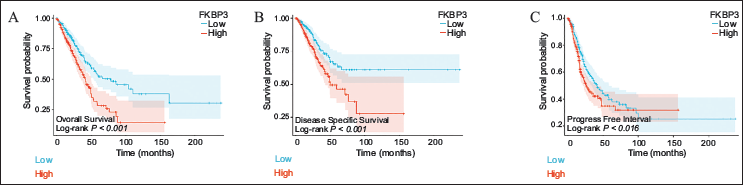
We subsequently assessed how FKBP3 expression influences patient survival outcomes along with its prognostic capacity through Kaplan-Meier survival curves and time-dependent ROC analysis. Findings demonstrated that patients harboring high FKBP3 expression displayed markedly reduced OS, disease-specific survival (DSS), and progression-free interval (PFI) relative to those with low FKBP3 expression (log-rank P < .001 for all comparisons) (Figures 4A and B), indicating elevated FKBP3 expression could correlate with unfavorable prognosis. Such results confirm the promise of FKBP3 as a prognostic biomarker for survival assessment among LUAD patients.
Biological Role of FKBP3 in Lung Adenocarcinoma
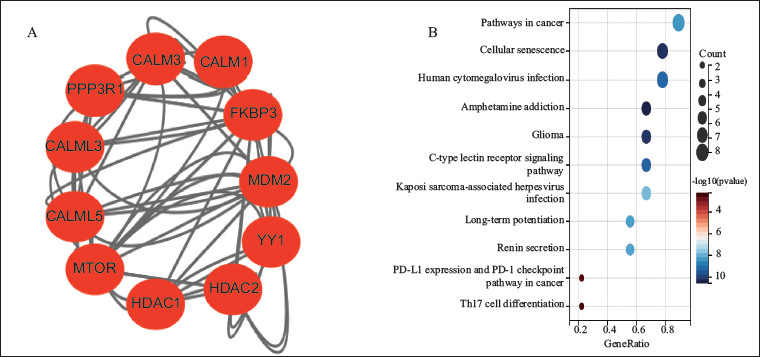
Construction of a PPI network involving FKBP3 and its interacting genes was accomplished through the STRING database, with visualization achieved via Cytoscape software (Figure 5A). Primary interaction partners comprised protein phosphatase 3 regulatory subunit B, alpha (PPP3R1), calmodulin 1 (CALM1), calmodulin 3 (CALM3), calmodulin 5 (CALM5), calmodulin like 3 (CALML3), murine double minute 2 (MDM2), YY1 transcription factor (YY1), mechanistic target of rapamycin (mTOR), and histone deacetylase 2 (HDAC2). Functional enrichment analyses for FKBP3-interacting genes revealed notable enrichment within pathways associated with immune evasion, including “Pathways in cancer,” “Cellular Senescence,” and “PD-L1 Expression and PD-1 Checkpoint Pathway in Cancer.” Additionally, these genes also showed enrichment in pathways such as “Glioma,” “C-type Lectin Receptor Signaling Pathway,” and “Human Cytomegalovirus Infection” (Figure 5B).
Analysis of FKBP3 and Immune Cell Infiltration
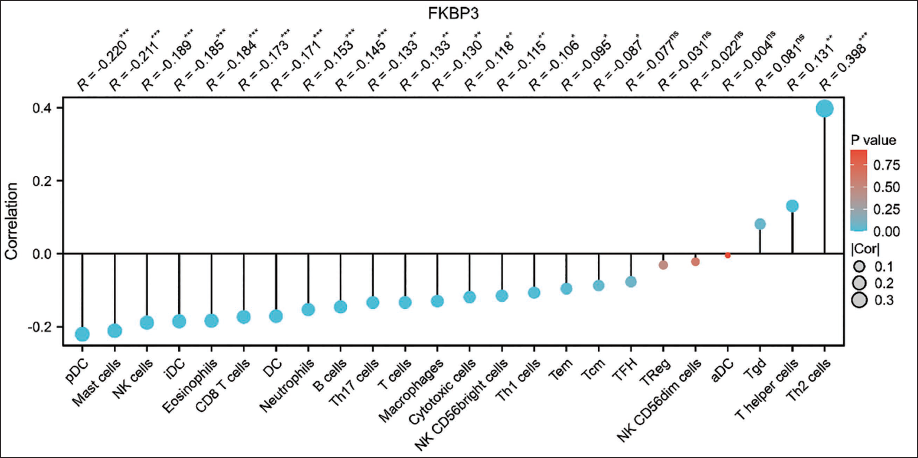
This study analyzed associations between FKBP3 expression and infiltration levels of various immune cells within the tumor microenvironment. Findings showed significant negative associations of FKBP3 expression with multiple immune cells, particularly plasmacytoid dendritic cells (pDCs) (R = −0.220, P < .001), mast cells (R = −0.211, P < .001), natural killer (NK) cells (R = −0.189, P < .001), immature dendritic cells (iDCs) (R = −0.186, P < .001), and eosinophils (R= −0.174, P < .001). These findings suggest FKBP3 may function in suppressing immune cell recruitment within the tumor immune microenvironment. In contrast, FKBP3 expression showed a positive association with Th2 cells (R = 0.398, P < .001), suggesting possible participation in immune escape processes (Figure 6). Together, these results emphasize FKBP3’s possible functional role in modulating tumor immunity.
Correlation Between FKBP3 Expression and Immune Cell Infiltration Levels in LUAD.
Discussion
This study systematically assessed the expression profiles of FKBP3 throughout different human malignancies, specifically examining its expression variations, clinical pathological correlations, prognostic implications, and underlying molecular pathways in LUAD. Findings revealed that FKBP3 is markedly elevated across various malignant entities, with notably elevated levels in LUAD, and demonstrates significant correlation with clinical outcomes. 25 These observations indicate that FKBP3 potentially exerts a crucial influence on the initiation and advancement of LUAD.
Initially, pan-malignancy evaluation demonstrated that FKBP3 displays elevated expression patterns across most solid malignancies, encompassing BLCA, BRCA, LIHC, and COAD. 26 These findings suggest that FKBP3 potentially possesses broad oncogenic capabilities during tumorigenesis and malignant advancement. These results align with earlier research demonstrating that additional FKBP family components, including FKBP5 and FKBP10, similarly display tumor-promoting characteristics across different malignancies. Being part of this protein family, elevated FKBP3 levels could facilitate malignant advancement via comparable molecular pathways. 27
Comprehensive examination of LUAD revealed that FKBP3 levels are substantially increased within malignant specimens compared to healthy tissues. These observations underwent confirmation through both mRNA data derived from the TCGA database and protein-level examination utilizing the UALCAN platform alongside immunohistochemical staining. ROC curve evaluation additionally established that FKBP3 possesses robust diagnostic capability, with an AUC of 0.894, suggesting potential clinical application for LUAD early identification. Remarkably, despite existing molecular markers currently utilized for LUAD diagnosis and prognostic assessment, universal indicators with broad applicability possessing elevated sensitivity and specificity remain limited. FKBP3 identification, therefore, offers an encouraging novel approach toward biomarker development within this domain.
Statistical analysis of clinicopathological features further indicated that FKBP3 expression is strongly correlated with lymph node metastasis (pathological N stage) and residual tumor status. Patients in the high-expression group displayed an increased incidence of lymph node involvement and residual tumors, suggesting that FKBP3 may contribute to local invasion and distant metastasis. This finding aligns with the observed interactions between FKBP3 and key signaling molecules such as mTOR, HDAC2, and MDM2 in the PPI network, further supporting its potential role in regulating cell proliferation and migration.
Regarding survival analysis, Kaplan-Meier curves demonstrated that elevated FKBP3 expression shows a notable correlation with reduced OS, DSS, and PFI among LUAD subjects. Cox regression analysis additionally validated that FKBP3 expression serves as an independent unfavorable prognostic indicator. Incorporating the development and validation of the nomogram model, FKBP3 manifests not merely as an individual molecular marker but also as a crucial component within a multifactorial framework. This emphasizes its capacity to offer a solid foundation for personalized risk stratification and treatment planning in LUAD patients.
In terms of functional mechanisms, the PPI network and KEGG pathway enrichment analysis revealed that FKBP3 and its interacting genes are mainly engaged in key biological processes such as the mTOR signaling pathway, HDAC-dependent epigenetic regulation, and general cancer-related pathways. Furthermore, FKBP3-related genes demonstrated enrichment within immune response pathways, including the PD-L1/PD-1 checkpoint pathway, indicating that FKBP3 might facilitate tumor progression through modulating immune evasion mechanisms. Remarkably, both HDACs and mTOR signaling represent well-established therapeutic targets for cancer treatment. Should FKBP3 actually participate in their upstream regulation or demonstrate co-expression with these pathways, it could create opportunities for innovative combination therapy strategies incorporating FKBP3-targeted interventions. 28
Immune infiltration analysis uncovered the potential function of FKBP3 in the tumor immune microenvironment. 29 Our results indicated that FKBP3 expression exhibited a significant negative correlation with various anti-tumor immune cells, encompassing NK cells, pDCs, mast cells, and iDCs, while demonstrating a positive correlation with Th2 cells. This immune cell infiltration pattern indicates that FKBP3 might assume a pivotal function in immune evasion within LUAD, possibly through inhibiting anti-tumor immune responses and facilitating immune tolerance. These results provide a novel predictive target for immunotherapy sensitivity, particularly within the current scenario where immune checkpoint inhibitors fail to benefit all patients.
Although this study presents relatively comprehensive and systematic results, certain limitations warrant recognition. Initially, the data originated predominantly from TCGA and other public databases, without validation across large-scale, multi-center clinical cohorts. Second, direct in vitro and in vivo experimental evidence remains absent to further clarify the mechanistic function of FKBP3 in LUAD. Moreover, while immune infiltration analysis indicates a regulatory function for FKBP3, the specific upstream regulators and downstream effectors participating in its immune-related pathways remain undefined.
Future studies should focus on the following aspects:
Validate the expression profile and clinical significance of FKBP3 in a larger cohort of human LUAD tissues using immunohistochemistry, in situ hybridization, and related techniques;
Employ gene knockdown or inhibition strategies, such as CRISPR/Cas9, shRNA, or small-molecule inhibitors, in cellular and animal models to assess the functional impact of FKBP3 on tumor proliferation, metastasis, and immune evasion;
Integrate cutting-edge technologies, including single-cell RNA sequencing and spatial transcriptomics, to comprehensively investigate the specific function of FKBP3 in tumor heterogeneity and immune modulation within the tumor microenvironment.
In conclusion, this study demonstrated that FKBP3 displays elevated expression within LUAD and correlates strongly with unfavorable clinicopathological characteristics and adverse prognosis. Additionally, the results clarify the potential mechanistic functions of FKBP3 within immune regulation and tumor-associated signaling pathways. These findings indicate that FKBP3 could serve not merely as a beneficial diagnostic and prognostic biomarker for LUAD, but additionally as a promising molecular target for future therapeutic and immunotherapeutic strategies. Consequently, FKBP3 demonstrates substantial scientific and translational clinical importance.
Footnotes
Authors’ Contribution
Min Liu: Responsible for conceptualization, drafting the original manuscript, and revising and editing the manuscript.
Guang Tian: Conducted investigation work, participated in drafting the original manuscript, and contributed to manuscript revision and editing.
Sangita Biswas: Performed investigation work, joined in drafting the original manuscript, and assisted with manuscript revision and editing.
Lifeng Zhao: Carried out investigation work, took part in drafting the original manuscript, and participated in manuscript revision and editing.
Min Liu and Guang Tian contributed equally to this work; Sangita Biswas and LifengZhao contributed equally to this work.
All authors have reviewed and approved the final published version of the manuscript.
Clinical Trial Number
Not applicable.
Consent for Publication
All authors have read and approved the manuscript and consent to its publication.
Data Availability Statement
Data used to support the findings of this study are available from public databases. If further raw data are needed for a more in-depth assessment of our study, please feel free to contact the corresponding author.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethics Approval
This study was exempt from ethical review, as all data were retrieved from public databases.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (No. 82460417).
