Abstract
VPS13 is a lipid transfer protein family conserved among Eukaryotes and playing roles in fundamental processes involving vesicular transport and membrane expansion including autophagy and organelle biogenesis. VPS13 folds into a long hydrophobic tunnel, allowing lipid transport, decorated by distinct domains involved in protein localization and regulation. Whereas VPS13 organization and function have been extensively studied in yeast and mammals, information in organisms originating from primary endosymbiosis is scarce. In the higher plant Arabidopsis thaliana, four paralogs, AtVPS13S, X, M1, and M2, were identified, AtVPS13S playing a role in the regulation of root growth, cell patterning, and reproduction. In this work, we performed phylogenetic, as well as domain and structural modeling of VPS13 proteins in Archaeplastida in order to understand their general organization and evolutionary history. We confirmed the presence of human VPS13B orthologues in some phyla and described two new VPS13 families presenting a particular domain arrangement: VPS13R in Rhodophytes and VPS13Y in Chlorophytes and Streptophytes. By focusing on Viridiplantae, we were able to draw the evolutionary history of these proteins made by multiple gene gains and duplications as well as domain rearrangements. We showed that some Chlorophytes have only three (AtVPS13M, S, Y) whereas some Charophytes have up to six VPS13 paralogs (AtVPS13M1, M2, S, Y, X, B). We also highlighted specific structural features of VPS13M and X paralogs. This study reveals the complex evolution of VPS13 family and opens important perspectives for their functional characterization in photosynthetic organisms.
Introduction
The VPS13 protein family was first discovered in 1977 in a mutant screen for loss of Carboxypeptidase Y (CPY) activity in yeast (Jones, 1977). Since, VPS13 proteins have been identified in many organisms and suspected to be present in all Eukaryotic cells. The yeast Saccharomyces cerevisiae has only one paralog (ScVPS13) whereas human has four (HsVPS13A/B/C/D). In the last decades, VPS13 functional analyses have demonstrated their role in a wide array of cellular processes including vesicular transport, autophagy, mitochondria homeostasis or lipid droplet (LD) biogenesis (reviewed in Dziurdzik and Conibear, 2021; Leonzino et al., 2021) and their mutations are associated to severe neurodegenerative diseases in human (reviewed in Ugur et al., 2020; Leonzino et al., 2021). This plethora of functions is in accordance with the wide variety of subcellular localizations observed at single organelles (ER, mitochondria, endosomes, LD, Golgi…), as well as at multiple contact sites between organelles (ER-mitochondria, ER-LD, mitochondria-endosomes…) (Dziurdzik and Conibear, 2021; Neuman et al., 2022). Recently, further structural, biochemical, and in silico approaches led to the characterization of VPS13 as lipid transfer proteins (LTPs) involved in bulk lipid transport and membrane expansion (Levine, 2022; Melia and Reinisch, 2022; Neuman et al., 2022). Indeed, VPS13 structural analyses by cryo-EM and 3D predictions showed that they form a long hydrophobic tunnel able to accommodate tens of glycerolipid molecules with a low specificity and that the VPS13 N-terminal (N-term) domain is able to transfer lipids between liposomes in vitro (Kumar et al., 2018; Li et al., 2020a; Melia and Reinisch, 2022; Neuman et al., 2022). VPS13 are massive proteins (more than 3000 amino acids (aa)) and their structure is organized around a repetition of Repeating β-Groove (RBG) domains forming a hydrophobic tunnel, likely mediating lipid transport, with addition of extra-domains inserted between RBG repeats and involved in the regulation of protein localization and function (Adlakha et al., 2022; Levine, 2022; Neuman et al., 2022; Dall’Armellina et al., 2023). The RBG domain core is composed of five antiparallel β-strands forming a concave β-sheet with a hydrophobic surface followed by a variable loop that can be disorganized or folded into a helix (Levine, 2022). The number of RBG domain repeats was proposed to determine VPS13 hydrophobic tunnel length, 12 repeats giving an average length of 20 nm (Levine, 2022; Neuman et al., 2022). The canonical domains found in VPS13 proteins are (1) a N-term Chorein, involved in protein localization and able to transport lipids in vitro, (2) a VPS13 adaptor binding (VAB) domain, involved in protein localization to specific organelles, (3) an Autophagy-related protein 2 C-terminus homology (ATG2_C) domain, likely involved in protein targeting via membrane binding, and (4) a C-term Pleckstrin Homology (PH) domain also involved in protein localization via protein–protein or protein–lipid interactions (Dziurdzik and Conibear, 2021; Levine, 2022; Neuman et al., 2022). The aforementioned domains are found in almost all the VPS13 proteins characterized to date. Other domains, such as WWE, Ubiquitin-associated (UBA), β-trefoil or β-sandwich domains, putatively involved in ubiquitination pathways or carbohydrate binding, are also found in HsVPS13s (Levine, 2022). Recently, clustering analyses of more than 1,000 VPS13 sequences from different organisms suggested that the last eukaryotic common ancestor (LECA) might harbor two VPS13 paralogs: ancestors of VPS13A/C/D and VPS13B (Levine, 2022). After gene duplication, VPS13D and then VPS13A and C might have emerged from the VPS13A/C/D ancestor. Interestingly, this analysis suggested that VPS13B was mainly found in Metazoa, but also in 25 protists and algae, suggesting it had been conserved in specific non-Metazoan phyla (Levine, 2022).
Whereas ScVPS13 and HsVPS13s are now well characterized, little is known about VPS13 proteins in organisms stemming from primary endosymbiotic events, that is, plants and algae. However, lipid trafficking in primary endosymbiosis derived organisms is highly complex due (1) to the presence of a chloroplast surrounded by two membranes (Petroutsos et al., 2014) and (2) to the adaptation to multiple environmental stresses triggering massive lipid remodeling events (Moellering and Benning, 2011; Michaud and Jouhet, 2019). Therefore, the characterization of LTPs in such organisms is an important step toward the understanding of their cell biology and their stress response adaptation. Arabidopsis thaliana possesses four paralogs, namely AtVPS13S, M1, M2, and X (Levine, 2022). Interestingly, AtVPS13M1 and M2 harbor several additional domains never observed previously in other VPS13 proteins: a C2 domain, an additional PH in N-term and three β-tripod domains that might be involved in protein localization mediated by protein–protein or protein–lipid interactions (Levine, 2022). AtVPS13S is the only characterized A. thaliana paralog at the time of writing. Initially named SHRUBBY (SHRY), AtVPS13S interacts with the transcription factor SHORT ROOT and plays a role in a pathway regulating root growth and radial patterning, even if the exact mechanism is not understood (Koizumi and Gallagher, 2013). Knock out atvps13S are also sterile, showing an important role in plant reproduction (Koizumi and Gallagher, 2013). In a recent pre-print, AtVPS13M2 was suggested to be involved in pollen germination and pollen tube elongation by regulating vesicular transport to the pollen tube tip (Tangpranomkorn et al., 2022, BioRxiv, DOI:10.1101/2022.11.01.514778). A VPS13-like gene has also been shown to play a role in apomixis, a plant reproductive process, in the angiosperm Taraxacum (Van Dijk et al., 2020). Apart from the above-mentioned studies, nothing is known about VPS13 paralogs (e.g., gene number, domain organization, functions) in organisms originating from primary endosymbiosis.
All photosynthetic organisms and some of their non-photosynthetic descendants stemmed from primary and secondary endosymbiotic events ((Keeling, 2010) and literature therein). Somewhere between 1600 (Yoon et al., 2004) and 900 million years ago (Ma) (Shih and Matzke, 2013), a cyanobacterium was engulfed into a Eukaryotic heterotroph and, instead of being digested, took permanent residence into the latter. This gave rise to the so-called primary chloroplasts of the three Archaeplastida lineages: the Viridiplantae, the Glaucophytes, and the Rhodophytes (red algae) (Figure 1) (Adl et al., 2012). From a Viridiplantae ancestor emerged three clades: the Prasinodermatophytes, the Chlorophytes (green algae), and the Streptophytes (plants), which is itself subdivided into Embryophytes (land plants) and Charophytes, algae being considered as land plants ancestors (Li et al., 2020b; Domozych and Bagdan, 2022; Bowles et al., 2023).

Schematic representation of Archaeplastida evolutionary history and VPS13 copy number and length of the sequences analyzed in this study. (A) A primary endosymbiosis of a cyanobacterium by a heterotrophic Eukaryote gave rise to three Archaeplastida lineages: the Viridiplantae, the Glaucophytes, and the Rhodophytes (red algae). From a Viridiplantae ancestor three clades emerged: the Prasinodermatophytes, the Chlorophytes (green algae), and the Streptophytes (plants), which is itself subdivided into Charophytes, algae being considered as ancestor of land plants, and Embryophytes (land plants). Within Charophytes, the Mesostigmaphyceae and Chlorokybophyceae groups are basal and Zynegmatophyceae is considered as a sister group of land plants. Copy numbers (B) and length (C) of VPS13 sequences in the different clades analyzed in this study. Data were generated from Dataset 1. The bar represents the median.
Here, we present a series of phylogenetic, bioinformatics, and structural modeling analyses aimed at deciphering the evolutionary history of VPS13 proteins in Archaeplastida in terms of copy number, domain organization and structure. We identified and described two new VPS13 types in Chlorophytes and Charophytes (VPS13Y) as well as in red algae (VPS13R). We also confirmed the presence of HsVPS13B orthologues in diverse groups of Viridiplantae and highlighted divergences in structure prediction. We finally focused on the Viridiplantae clade for which we were able to reconstruct VPS13 paralogs evolutionary history, domain organization and structure.
Results
Phylogeny of VPS13 Proteins in Archaeplastida
Sequences were retrieved by BLASTP searches (Altschul, 1997) in different databases using A. thaliana or HsVPS13B sequences as queries (see Material and Methods). Only sequences with at least one VPS13 canonical domain annotation were retrieved to populate a first dataset of 537 sequences (Table S1). However, numerous sequences were fragmented (size below 2000 amino acids), likely because of poorly assembled genomes and to errors performed by automatic genome annotation that failed to properly define the correct borders of such long proteins that contain many introns/exons (63 exons for AtVPS13M1, for example). An extreme case is illustrated in Figure S1 where a VPS13 gene in Coffea arabica on scaffold 440 is divided in at least seven successive gene models (evm.model.Scaffold_440.43 to evm.model.Scaffold_440.49) with predicted Vacuolar sorting associated protein 13 domains (IPR026847/PTHR16166) and translated to proteins ranging from 96 to 2207 aa. Some particularly long sequences were retrieved corresponding to different protein model predictions of the same gene. This is the case as example in Chlamydomonas reinhardtii in which two sequences, A0A2K3DAJ8 (8089 aa) and A8IIB7 (7192 aa), corresponding to the same VPS13 gene with different exon/intron border prediction, were retrieved (Table S1). In order to perform analyses with a significant number of likely full-length sequences from Archaeplastida, ≥2400 aa long sequences were selected to create a dataset containing 383 sequences (Dataset 1). Within this dataset, major variations in gene copy number and sequence length are observed among clades (Figure 1B and C). Nevertheless, these results have to be taken with caution as the dataset was not manually curated and hence some paralogs might be missing, might correspond to different models of the same gene, or might be incomplete. The highest number of VPS13 sequences is observed in the Charophyte Spirogloea muscicola (13 sequences) and the Embryophyte Tinopyrum intermedium (12 sequences) (Figure 1B). Limited variations of sequence length are observed for Embryophytes and the Glaucophytes in contrast to other clades (Figure 1C) and sequences more than 7000 aa long are recorded in some Chlorophytes (Figure 1C) as previously reported (Levine, 2022). Noteworthy, the 13 Glaucophyte sequences are very short (below 3000 aa), although according to domain analysis (see below), most of them are full length (i.e., presence of a N-term Chorein and a C-term PH domains).
A maximum likelihood (ML) phylogenetic analysis was then performed adding HsVPS13A through D and ScVPS13 to the dataset in order to infer their relationship with those from Archaeplastida. Because of the length of the alignment, 5,000 boostrap pseudoreplicates were performed and nodes supported by a bootstrap value above 4,000 (80%) were considered as significant. The unrooted tree presented in Figure 2 shows the complex evolutionary history of VPS13 proteins in each phylum. VPS13 proteins show high primary sequence diversity and no protein family present in all of the three Archaeplastida groups can be unambiguously defined. Embryophyte VPS13 proteins clearly split into four clades likely corresponding to the AtVPS13M1, M2, S, and X paralogs described in A. thaliana (Levine, 2022). The well-supported VPS13X and VPS13S clades (Figure 2, nodes no. 1 and 2, respectively) also contain Charophyte sequences, including Mesostigma viride and Chlorokybus atmophyticus, suggesting that VPS13X and S paralogs were present in Streptophytes ancestor. A structured clade that we named “VPS13M” (Figure 2, node no. 3) contains the Embryophytes VPS13M1 and M2 as well as Charophytes, Chlorophytes, and Prasinodermatophytes sequences. As expected, Chlorophytes and Prasinodermatophytes are basal to the Streptophytes group and have only one VPS13M. Some species have more VPS13M, such as Ostreococcus tauri or Chlorella sorokiniana, but sequence alignment showed that they likely correspond to different protein model predictions for the same gene. The Streptophytes analyzed seem to have a VPS13M1 and M2 paralogs except the Charophytes C. atmophyticus and M. viride that have one VPS13M branching before the VPS13M1/M2 split (Figure 2, node no. 4). This implies that the Viridiplantae ancestor had one VPS13M which was duplicated after the emergence of Chlorokibophyceae and Mesostigmaphyceae.

Maximum likelihood phylogenetic analysis of VPS13 proteins from Archaeplastida. Organism names are colored according to their taxonomic clade. The analysis was performed with 5000 boostrap pseudoreplicates and nodes supported by more than 4000 boostraps (80%) are highlighted by a pink dot. The supported nodes discussed in the text are highlighted by a pink circle and a number. The main VPS13 groups that will be discussed in this work (VPS13S, X, M (M1 and M2), R, Y, B and Glaucophyte) are highlighted. Only the groups VPS13M, X, R, and VPS13S in Streptophytes are well supported by this analysis. The other groups are more supported by the Bayesian analyses performed thereafter.
Another supported clade contains only Rhodophytes’ sequences and was named “VPS13R” (Figure 2, node no. 5). Unsupported (bootstrap values below 80%) clades include (1) Glaucophytes, (2) a variably supported clade clustering several Prasinodermatophytes, Chlorophytes, and Charophytes sequences, later named “VPS13Y” (see below), (3) a clade, basal to VPS13S, composed of Prasinodermatophytes and Chlorophytes, and (4) two orphan clades (Figure 2). Interestingly, whereas ScVPS13 and HsVSP13A/C clustered together and VPS13D branched alone, HsVPS13B clustered with sequences from Charophytes, Chlorophytes and from the Bryophyte Sphagnum fallax, nonetheless with a weak support, corroborating the finding that VPS13B might be present in a small subset of organisms besides Metazoa (Levine, 2022). Therefore, VPS13B was likely present in the LECA.
In order to more confidently trace the evolution of the VPS13 paralogs in Archaeplastida, a Bayesian and a ML analyses were performed on a reduced dataset of 104 sequences composed of two Glaucophytes, three Rhodophytes, two Prasinodermatophytes, one organism in each of the five Charophyte groups pictured in Figure 1, four Chlorophytes and eight Embryophytes (including lower and higher plants, one gymnosperm and angiosperms, monocotyledons, and dicotyledons) as well as the human and yeast VPS13 sequences (Dataset 2). All the sequences in Dataset 2 were manually curated to eliminate redundant sequences. As VPS13 primary sequences can be highly divergent but share a conserved overall 3D structure and domain arrangement, we mathematically formalized the domain occurrence for each sequence and included such information in the Bayesian analysis (see Material and Methods). Remote homology analyses using HHPred (Zimmermann et al., 2018) were performed for all the sequences included in the dataset in order to identify putative domains. Most of the canonical VPS13 domains such as Chorein, VAB, or ATG2_C were found with additional domains for some sequences that will be further discussed below (Table S2). VPS13B clade was used to root the trees (Figure S2 and S3). These analyses confirmed that no group including sequences from the three Archaeplastida phyla (i.e., Viridiplantae, Glaucophyte, Rhodophyte) was found, showing that VPS13 paralogs have highly diverged. Eight clades, namely M, S, X, B, R, Glaucophyte, and opisthokont were highlighted by both the Bayesian (Figure S2) and the ML (Figure S3) analysis. However, the analyses do not allow to establish any relationship among the different VPS13 groups. The opisthokonts’ sequences HsVPS13A/C/D and ScVPS13 produced a separate clade whereas VPS13B clade included a wide variety of organisms (Chlorophytes, Charophytes, and one Embryophyte). Despite a low support (PP = 0.72 (Figure S2) and 16.5% bootstraps (Figure S3)), those sequences clustered together in all the analyses performed and the Chara braunii sequence GBG81180 was also identified as a VPS13B orthologue in (Levine, 2022) further suggesting the presence of this paralog in Viridiplantae. A high selection pressure may occur on VPS13B sequences during evolution in contrast to other VPS13 paralogs. However, VPS13B was possibly lost independently several times during evolution (see below) because it is currently only present in a small number of phyla other than Metazoa. These analyses confirmed the existence of a clade grouping several Rhodophyte sequences, identifying a Rhodophyte-specific VPS13 protein. The cluster was named “R” (for Rhodophyte) (Figure S2 and S3), suggesting a high conservation of VPS13R orthologues during Rhodophyte evolution. VPS13R is not the only VPS13 found in red algae, indeed, other Rhodophyte sequences produced orphan clades (Figures S2 and S3). Such proteins may have specifically evolved in the different Rhodophyte lineages. VPS13R sequences show a common domain organization with the presence of a β-sandwich, and sometimes a UBA domain, downstream of the C-term PH domain (Figure S2). The β-sandwich and UBA domains have been acquired independently during evolution because they are also found in HsVPS13B and D, respectively, but upstream of the VAB. Glaucophyte sequences mostly clustered together, nonetheless with a low support (PP = 0.77 (Figure S2) and 45.3% (Figure S3)). VPS13 paralogs in this clade maintained a very simple organization and only the canonical Chorein, VAB, ATG_C, and PH domains are found (Figure S2). VPS13S, M, Y, and X clades are present only in Viridiplantae and will be discussed below.
Bayesian Analysis of Viridiplantae VPS13 Paralogs
From the results presented above, Viridiplantae VPS13 family showed an interesting evolutionary history, hence a reduced dataset (Dataset 3, Table S3, 79 sequences) including Embryophytes, Streptophytes, Chlorophytes, and Prasinodermatophytes was produced and analyzed as described above. Human and yeast sequences were excluded to avoid destabilization of the analysis. The tree presented in Figure 3 shows that VPS13 protein sequences clustered into five robustly supported (PP = 1.00) clades, namely M, in turn divided into three sub-clades (M, M1, and M2), S, X, B, and Y. VPS13Y has never been reported before. We named it “Y” because no clear and unambiguous homology with other known VPS13 protein groups was revealed by this analysis. Within each VPS13 clade, the organization of the branches is in agreement with the expected taxonomy: Prasinodermatophytes and Chlorophytes are always basal to Streptophytes within which Charophytes are basal to Embryophytes, with non-vascular plants being well separated from vascular plants, showing a linear evolution of VPS13 proteins in each phylum. The only exception is VPS13B which groups incongruously with the expected taxonomy, suggesting an unresolved evolutionary pattern.
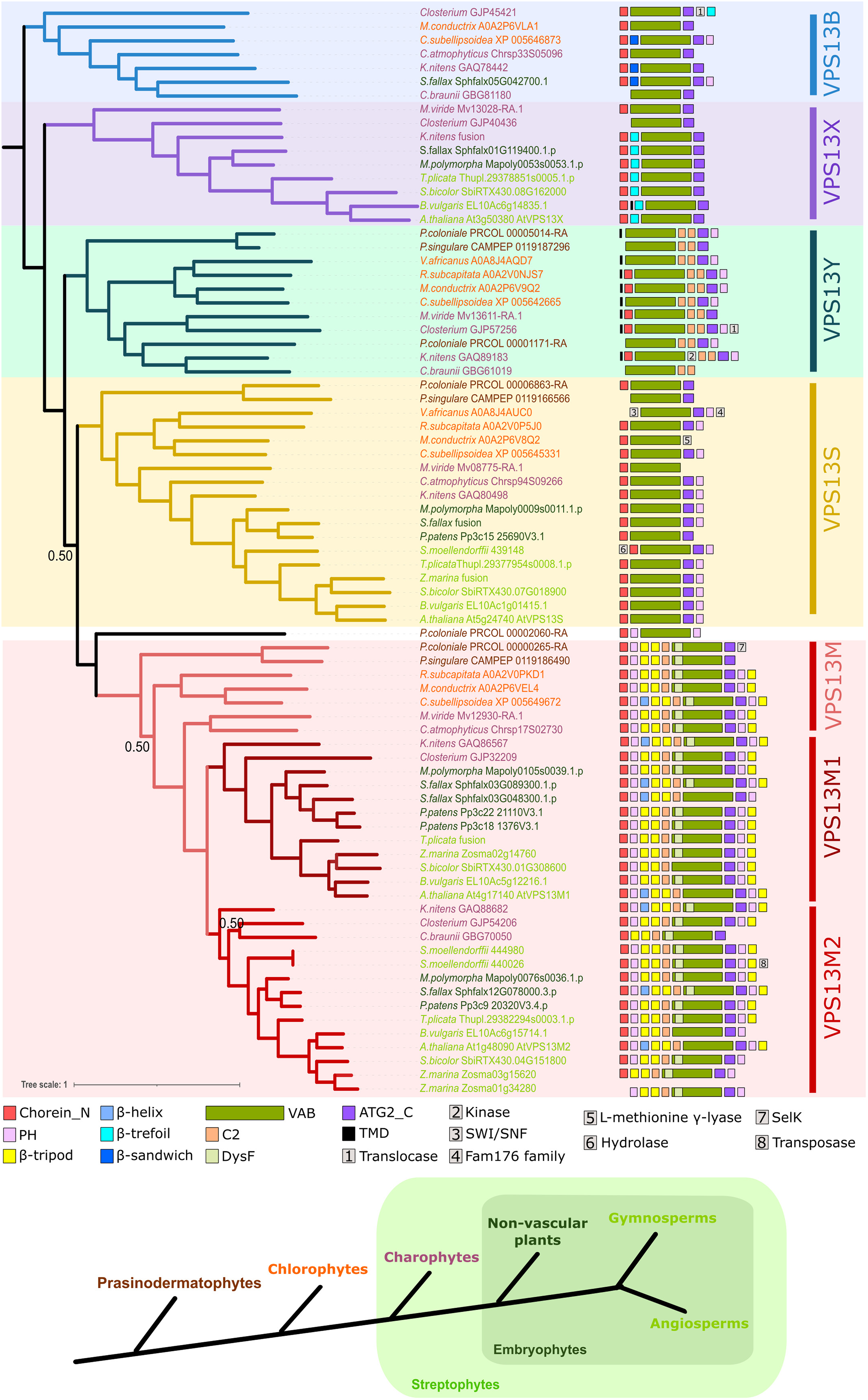
Bayesian phylogenetic analysis of VPS13 proteins in Viridiplantae. The domain organization, predicted using HHPred, was included as a character in the analysis and is represented on the right. Recurrent domains are represented by a colored square whereas others are represented by a number. Only the Bayesian posterior probability values below 0.95 are presented on the tree. Supported groups of VPS13 paralogs are highlighted in colors. Taxonomic clades to which each sequence belongs is indicated by a color code and evolutive relationships between them is summarized on a scheme below the tree. ATG2_C = Autophagy-related protein 2 C-terminus homology; PH = Plekstrin Homology; TMD = transmembrane domain, VAB = VPS13 adaptor binding.
The VPS13M and S were found in all species analyzed showing that they were conserved through the evolution of Viridiplantae. VPS13S was reported as single copy in all the organisms analyzed. These analyses confirmed the presence of only one VPS13M gene in Chlorophytes and basal Charophytes (M. viride and C. atmophyticus) whereas other Streptophytes have two paralogs (M1 and M2), showing that VPS13M was likely duplicated before the emergence of Klebsormidiophyceae. Overall, only one VPS13M1 and M2 paralogs were found with some exceptions. A duplication of VPS13M1 paralog early in the evolution of Bryophytes has occurred because the two mosses Sphagnum fallax and Physcomitrium patens possess two VPS13M1. VPS13M2 duplicated likely independently in different phyla during evolution because two paralogs were found in the monocotyledon Zostera marina as well as in the basal vascular plant Selaginella moellendorffii. VPS13X clade contains sequences belonging only to Streptophytes, including M. viride. Despite our advanced BLASTP searches, no VPS13X homologous sequences could be found in any Chlorophyte genome, suggesting that VPS13X arose early in the evolution of Streptophytes. Interestingly, the VPS13Y clade contains Prasinodermatophytes, Charophytes and Chlorophytes proteins only, revealing that VPS13Y was likely present in the Viridiplantae most recent common ancestor (MRCA) and lost before the emergence of Embryophytes.
The results of our analysis focused on Viridiplantae confirmed that VPS13B orthologues are likely present only in a small subset of organisms spread on different Viridiplantae clades putting forward that a VPS13B paralog was initially present and independently lost several times during evolution (Levine, 2022). Four Charophytes species out of the five analyzed here possess a VPS13B. The putative Closterium VPS13B orthologue sequence (GJP45421) does not always cluster with other VPS13B (Figure 2 and Figure S3) but was also clustering with HsVPS13B in the analysis performed by Levine (2022), supporting its affiliation to the VPS13B group. Among Chlorophytes, VPS13B was detected in both Trebouxiophyceae species analyzed (Coccomyxa subellipsoidea and Micractinum conductrix), but not in the Chlorophyceae (Volvox africanus and Raphidocelis subcapitata). In addition, a VPS13B orthologue was detected in the moss Sphagnum fallax (Embryophyte). In order to better understand when VPS13B has been lost in the different Viridiplantae phyla, Position-Specific Iterative Basic Local Alignment Search Tool (PSI-BLAST) (Altschul, 1997) searches were performed in Viridiplantae using either HsVPS13B or the six VPS13Bs found in this study as queries on NCBI (Sayers et al., 2022). Only a few Trebouxiophyceae (Coccomyxa, Micractinum, Trebouxia, Auxenochlorella, Chlorella) sequences were retrieved (Supplemental File S1), confirming that VPS13B gene might have been conserved in this lineage during evolution. One sequence was also found in Tetraselmis striata (Figure 2) and in a basal Chlorophyte group, namely the Pedinophyceae, further suggesting that VPS13B was likely present in the MRCA of Chlorophytes and not gained by horizontal gene transfer in Trebouxiophyceae. The only other Embryophyte species possessing a VPS13B orthologue is Sphagnum magellanicum, indicating that VPS13B was maintained only in this plant genus.
Overall, our results suggest that the Viridiplantae MRCA had four VPS13 paralogs: VPS13M, S, B, and the newly identified Y. In Chlorophytes, VPS13B was lost in different clades but retained in several Trebouxiophyceae and likely in a small number of other groups, such as Pedinophyceae. VPS13X appeared early in the evolution of Streptophytes, its putative origin will be discussed below. After the emergence of Mesostigmaphyceae and Chlorokybophyceae, VPS13M was duplicated to give rise to VPS13M1 and M2. Thus, Streptophytes’ MRCA likely had five VPS13 paralogs: VPS13M, S, Y, X, and B, this hypothesis being supported by the presence of the five paralogs in the Charophytes M. viride and Klebsormidium nitens, the latter also having the VPS13M1 and M2 paralogs arising from VPS13M duplication. Then, VPS13Y was likely lost early at the emergence of Embryophytes. The presence of a VPS13B protein in two Sphagnum species suggests that a VPS13B paralog was present in the MRCA of Embryophytes. Then, VPS13B gene was lost in most of the Embryophyte phyla but maintained in some Bryophytes like Sphagnum. Thus, our results support the hypothesis that the Embryophytes’ MRCA had five VPS13 paralogs, namely VPS13M1, M2, S, X, and B. Whereas our analysis confirmed the presence of HsVPS13B orthologues in non-metazoans (Levine, 2022), we were not able to establish significant relationships among the other ancestral Viridiplantae VPS13 groups (i.e., M, S, X, and Y). Of note, one Prasinoderma coloniale sequence (PRCOL_00002060-RA) does not clearly clustered with one VPS13 group, being sometimes basal to VPS13M group (Figure 3 and Figure S4) or branching with other groups (Figure 2, and Figure S2 and S3). This sequence has a peculiar domain organization with a PH domain upstream of the VAB (Figure 3). Whether this paralog is an ancestral one lost during evolution in other phyla or belongs to a specific Prasinodermatophyte VPS13 group remains to be determined.
Expression of the Different VPS13 Paralogs in Viridiplantae
Analysis of A. thaliana transcriptomic data available in the Bio-Analytic Resources for plant biology (BAR) browser (Winter et al., 2007) showed that the four AtVPS13 paralogs are expressed (data not shown). We next focused on species having the less widespread paralogs VPS13B, Y, and X. For the Chlorophyte C. subellipsoidea, putative mRNA sequences corresponding to the four VPS13 paralogs M, S, Y, and B were found in NCBI and a BLASTN search was performed against an Expressed Sequence Tags (ESTs) cluster available on the Joint Genome Institutes (JGI) (Nordberg et al., 2014). EST sequences were found with a high confidence for the four C. subellipsoidea VPS13 paralogs showing that they are all expressed and likely functional (Supplemental File S2). Most of the ESTs aligned with the 3’ end of the transcripts, but hits were also found all along the sequences (Figure S5). This analysis also revealed errors in the predicted VPS13 gene models as several ESTs support a different intron/exon organization. In addition, an AGO1 domain was initially predicted at the C-term of CsVPS13B protein but EST analyses clearly show that the transcript ends after the PH domain (Figure S5). Thus, ESTs were used to curate CsVPS13 paralog sequences and obtain more accurate mRNA and protein sequences for further analyses (see below) (Supplemental File S2).
We also investigated the VPS13 paralogs expression in two Charophyte species used in this study (C. braunii and K. nitens) as well as in S. fallax. Assembled transcriptomes were found on NCBI for Chara braunii (Bioproject PRJNA492241), Klebsormidium flaccidum (Bioproject PRJN242253) and two Sphagmun species: S. recurvum (Biosample UHLI) and S. palustre (Biosample RCBT). TBLASTN searches were performed using respective VPS13 protein sequences as queries. Interestingly, transcripts were identified with a high confidence for all the paralogs (e-value = 0, Table S4). In addition, transcripts covering almost the entire sequences were found for the six VPS13 paralogs in K. flaccidum. KfVPS13M2 and Y transcripts were likely full length whereas the others are split into different transcripts likely due to errors in the assembly. Transcripts corresponding to the five S. fallax VPS13 paralogs were detected in both S. recurvum and S. palustre, confirming the presence and the expression of VPS13B in the Sphagnum clade. In addition, the two SfVPS13M1 isoforms are also present and expressed in both Sphagnum species, showing that they remained functional after VPS13M1 gene duplication. In C. braunii, only three paralogs (VPS13B, M2, and X) were previously identified by BLASTP searches and transcripts were found for all with a high confidence (Table S4). We also found other putative hits of shorter length (around 500 aa) and lower confidence that might correspond to other VPS13 paralogs initially undetected by BLASTP searches. Thus, TBLASTN searches on C. braunii transcriptome were performed using all VPS13 paralogs from K. nitens and Closterium as queries. This analysis allowed the detection of putative CbVPS13X paralogs identified in both TBALSTN search and covering around 2000 aa of KnVPS13X. A putative VPS13S was detected only with KnVPS13S but with a low coverage (559 aa). However, no putative M1 sequences were retrieved, maybe due to lower expression level, poor transcript assembly, or the absence of this paralog in C. braunii.
Overall, this analysis showed that all the VPS13 paralogs discovered with our phylogenetic analyses were expressed in the Chlorophyte C. subellipsoidea, the Charophytes K. nitens, and C. braunii, and in the Bryophytes Sphagnum spp, showing that they might be all functional. In addition, transcriptomic analysis was used to refine VPS13 sequences in C. subellipsoidea and to find putative orthologues in C. braunii not detected by protein searches, illustrating the interest of analyzing transcriptomic data in the analysis of huge and complex proteins such as VPS13.
Domain Organization of VPS13 Proteins in Viridiplantae
To go further into the putative evolution of VPS13 proteins, we analyzed their domain organization and structure from modeling. VPS13 protein structure is organized around a repetition of RBG domains forming a hydrophobic tunnel, likely mediating lipid transport, with addition of extra-domains inserted between RBG domains and involved in the regulation of protein localization and function (Adlakha et al., 2022; Levine, 2022). In order to understand if the general organization of Viridiplantae VPS13 proteins, in particular of VPS13X, VPS13B, and the newly identified VPS13Y paralogs, is conserved, a subset of key organisms (A. thaliana, S. fallax, C. subellipsoidea, and K. nitens) in which we confirmed the expression of the paralogs were chosen to carry out RBG domain predictions. RBG domains were predicted using HHPred (Zimmermann et al., 2018) and, as queries, RBG domain sequences previously identified in A. thaliana as well as in HsVPS13B for VPS13B orthologues (see Material and Methods) (Levine, 2022).
VPS13S
A. thaliana VPS13S is composed of 12 RBG repeats accompanied by the canonical VPS13 domains only (Levine, 2022), like most of the VPS13S proteins identified in our analysis, suggesting that the VPS13S ancestor had the following domain organization: Chorein-VAB-ATG2_C-PH (Figure 4). Additional domains were found in N- or C-term but they could stem from gene model prediction errors as exemplified above for CsVPS13B AGO1 domain in C-term. As an example, a Hydrolase domain was found in N-term of S. moellendorffii VPS13S or a methionine γ-lyase domain in the C-term of Micractinum conductrix (Figure 3). An extensive analysis of transcriptomic data, if available, might shed some light on the actual occurrence of such unexpected domains. RBG domains analysis showed that VPS13S of K. nitens (4022 aa) and C. subellipsoidea (3698 aa) also contain 12 RBG domains, even if they are longer than AtVPS13S (3466 aa) therefore an increase in protein size is likely not correlated to an increase of RBG repeats or tunnel length (Figure 4). Overall, the domain organization of VPS13S proteins seems to be conversed across the Viridiplantae phylum without addition of any new domain. It is noteworthy that a similar basic organization is found in the single VPS13 paralog in yeast (Levine, 2022).
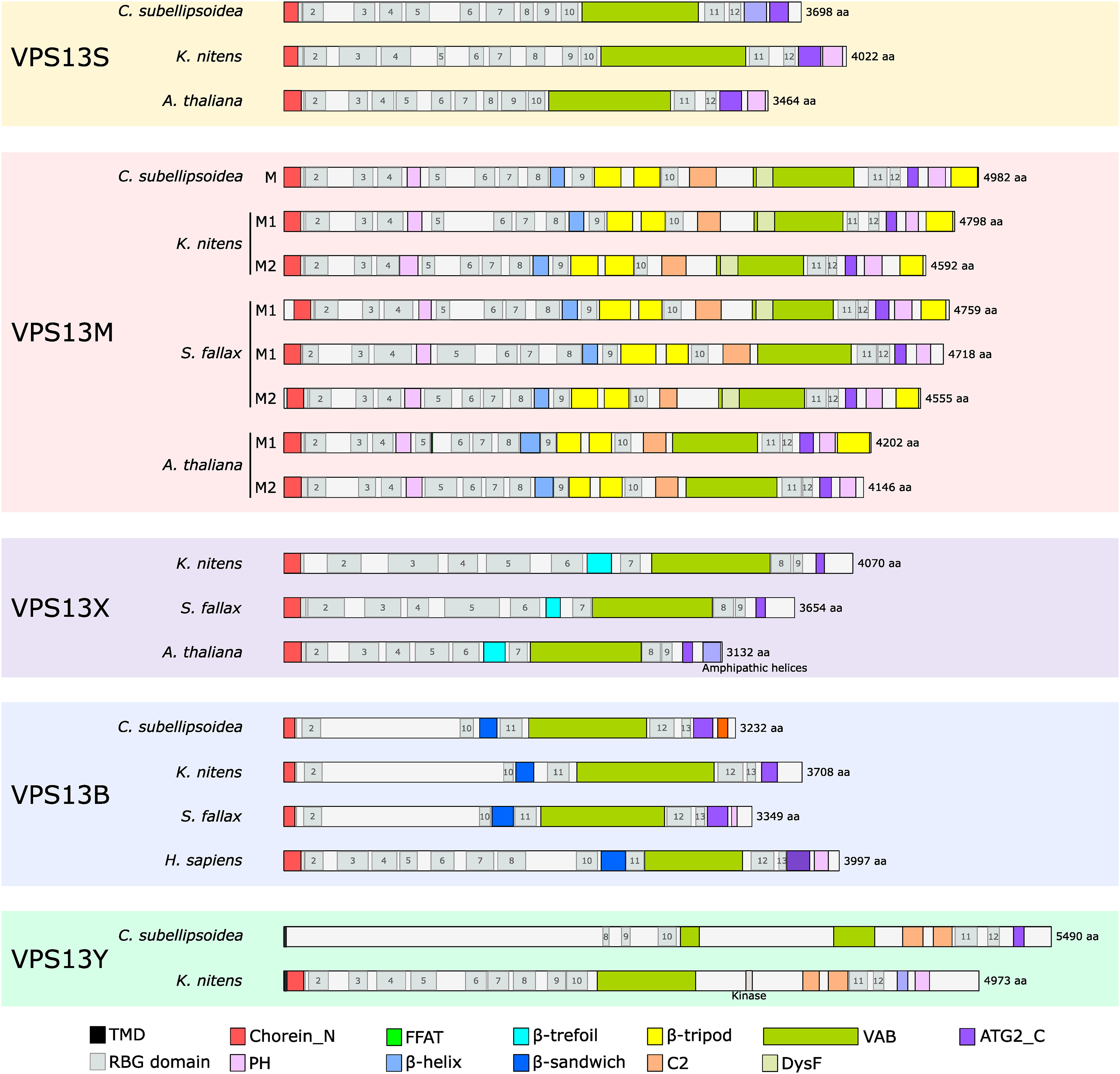
RBG domain analyses in Viridiplantae VPS13 paralogs. On each sequences, domains and RBG repeats detected using HHPred were indicated, apart for AtVPS13 and HsVPS13B sequences for which the information was retrieved from Levine (2022). Protein size is indicated on the right and expressed as amino acids (aa). ATG2_C = Autophagy-related protein 2 C-terminus homology; FFAT = two phenylalanine in an acidic track; PH = Plekstrin Homology; TMD = transmembrane domain; VAB = VPS13 adaptor binding.
VPS13M
A. thaliana VPS13M1 and M2 proteins are composed of 12 RBG domains and are characterized by the occurrence of up to seven additional domains (Figures 3 and 4) (Levine, 2022). Indeed, a (1) PH domain is inserted between RBG4 and 5 in addition to the canonical one located downstream of the C-term ATG2_C domain, (2) two β-tripod domains in tandem are inserted between RBG9 and 10 and a third one is present at the C-term end of most of the VPS13M paralogs, (3) a C2 domain is inserted right upstream of the VAB domain, (4) a β-helix domain, and (5) a Dysferlin/Pex24 (DysF, IPR010482) domain is predicted within the VAB of several VPS13M orthologues. The PH domain located in the N-term part of the protein was identified in almost all the VPS13M sequences analyzed, suggesting it was present in the Viridiplantae VPS13M ancestor (Figure 3). The only organisms lacking this PH domain in the VPS13M2 N-term are the Embryophyte Zostera marina and the Charophyte C. braunii. This might be linked to a higher sequence divergence that did not allow HHpred to predict this domain or to a loss in some lineages. The tandemly repeated β-tripod domains are present in all the VPS13M, suggesting their presence in the Viridiplantae MRCA. Interestingly, only Angiosperms’ VPS13M2 lack the C-term β-tripod domain (Figure 3). Noteworthy, this domain is present in all VPS13M1, in Chlorophytes’ VPS13M as well as in Charophytes’ VPS13Ms excluding C. braunii VPS13M2, lower plants and the Gymnosperm Thuja plicata. A loss of the C-term β-tripod domain in Angiosperms is conceivable, possibly to allow the emergence of a new function (see Discussion). Interestingly, the two Prasinodermatophytes VPS13Ms do not possess a predicted β-tripod domain in C-term, neither a PH domain, suggesting a particular evolution of the C-term in this phylum or an error in gene model annotation. The additional C2 domain upstream of the VAB is predicted in all phyla, including Prasinodermatophytes, suggesting it was present in the VPS13M ancestral protein. In the analysis performed by Levine (2022), a β-helix domain was identified between RBG8 and 9. This domain was also detected when HHpred predictions were performed on a reduced portion of the proteins (between 150 and 180 aa) and was confirmed by AlphaFold2 (Jumper et al., 2021; Mirdita et al., 2022) structural prediction for a small subset of VPS13M, M1, and M2 (see below and Figure 5). Thus, the β-helix domain was likely present in all VPS13M and in the common ancestor of VPS13M. Noteworthy, the DysF domain is predicted upstream of or overlapping the VAB domain for almost all the VPS13M orthologues, excluding A. thaliana (Figures 3 and 4). Structural analyses (see below) showed that this domain is indeed inserted in the second module of the VAB domain, highlighting an additional feature of VPS13M sequences. Because of its presence in all the Viridiplantae phylum, it was likely present in the VPS13M ancestor and lost in some lineages such as in A. thaliana. A FFAT (two phenylalanines in an acidic tract) motif is also present in AtVPS13M1, but not AtVPS13M2 (Slee and Levine, 2019), downstream of RBG5 but we were not able to perform automatic analysis to easily detect FFAT motifs in other VPS13M orthologues in order to assess its conservation. All VPS13M proteins analyzed are composed of 12 RBG domain repeats, showing the high conservation of the VPS13M clade (Figure 4).

Structural modeling and characteristics of VPS13S, M1, and X in Arabidopsis thaliana. The 3D structures were obtained mainly using AlphaFold2 (Jumper et al., 2021) and RoseTTAFold (Baek et al., 2021) for AtVPS13M1. Overlapping fragments were aligned on RBG repeats and assembled using ChimeraX. (A) Predicted structures of the full length AtVPS13S, M1, and X proteins. β-strands, α-helices, and loops are colored in yellow, red, and green, respectively. The different VPS13 domains are highlighted by different colors. VPS13 proteins folded into a long tunnel around which accessory domains are organized. (B) Representation of the AtVPS13S, M1, and X RBG repeats only showing the formation of the canonical VPS13 tunnel shape. For VPS13X, the tunnel is bended at the level of RBG5 colored in red. (C) Hydrophobic (in orange) and hydrophilic (in blue) surface representation of the RBG repeats structure of (B), highlighting the hydrophobic surface of the tunnel (arrows).
Overall, our results indicate that VPS13M ancestor had 12 RBG domains and the following organization: Chorein-PH-β_helix-β_tripod-β_tripod-C2-DysF-VAB-ATG2_C-PH-β_tripod. VPS13M was likely duplicated shortly after the emergence of the Charophyceae of the Mesostigmaphyceae and Chlorokybophyceae groups, giving rise to VPS13M1 and M2. Finally, the C-term β-tripod domain was then lost in Angiosperms’ VPS13M2.
VPS13X
VPS13X has a particular structure in A. thaliana as it is composed of only nine RBG repeats and does not possess a canonical PH domain in C-term but amphipathic helices instead (Levine, 2022). Also, a β-trefoil domain was found between RBG6 and 7. Most of the VPS13X proteins analyzed have a similar organization, apart in Closterium where no Chorein and β-trefoil domains were detected, maybe because we did not retrieve a full-length protein (Figure 3). K. nitens and S. fallax VPS13X are also composed of nine RBG domains and are significantly longer than AtVPS13X (3132 aa), being 4792 and 3654 aa long, respectively (Figure 4). The origin of VPS13X is mysterious (see Discussion) but we can assess that it appeared early in the Streptophyte lineage.
VPS13B
In human, VPS13B is composed of 13 RBG repeats and contains the four VPS13 canonical domains and a β-sandwich domain inserted between the RBG10 and 11 (Levine, 2022; Dall’Armellina et al., 2023). In Viridiplantae, the Chorein, VAB, and ATG2_C are also found. The PH domain was found only in two sequences (C. subellipsoidea and S. fallax) and the β-sandwich was found by a closer HHPred search and structural prediction performed in C. subellipsoidea, S. fallax, and K. nitens. The β-sandwich may be considered as an ancestral domain maintained in Viridiplantae VPS13B orthologues (Figure 4). RBG domain prediction shows that RBG1, 2, 10, 11, and 13 were easily detected using HsVPS13B RBG sequences as queries. Surprisingly, we failed to detect any RBG domains in the N-term portion of the proteins between RBG2 and 10 in contrast to HsVPS13B (Figure 4). We hypothesized that this might be linked to a higher divergence of RBG repeats in this region in Viridiplantae and performed a HHPred search for RBG domains using RBGs from AtVPS13 paralogs without any success. Neither specific domains nor motifs were found. Structural predictions also failed to reveal clear RBG domain folding (see below). As AlphaFold2 predictions can be challenging for these proteins (Neuman et al., 2022), we performed secondary structure predictions for C. subellipsoidea, K. nitens, and S. fallax (from N-term to the VAB domain) using Quick2D, which gathers predictions from several softwares (Zimmermann et al., 2018). This analysis showed the presence of several predicted disordered regions as well as some potential β-strands upstream of the RBG10 for CsVPS13B and in KnVPS13B (Supplemental File 3), corroborating the hypothesis that the region between RBG2 and 10 might not fold into the well-described RBG domains. For C. subellipsoidea, we were able to detect ESTs and to curate the VPS13B sequence to obtain the probable expressed protein (see above) confirming that the unstructured region exists and is not linked to automatic annotation errors. In contrast to CsVPS13B and KnVPS13B, the secondary structure analysis predicted several successive β-strands followed by a helix in SfVPS13B sequence, suggesting the occurrence of four additional domains folded into β-sheets followed by helices not predicted by HHpred or AlphaFold2 (Supplemental File 3).
Overall, the VPS13B C-term part downstream of the RBG10 as well as the Chorein and RBG2 domains seem to be conserved through evolution whereas the region between RBG2 and 10 are highly divergent in terms of RBG repeats and structure in Viridiplantae compared to HsVPS13B. From this analysis, it is difficult to infer the putative ancestral state (with 13 RBGs as in human or six as in photosynthetic organisms, see Discussion) of VPS13B orthologues.
VPS13Y
The newly identified VPS13Y was found in Prasinodermatophytes, Chlorophytes, and Charophytes only. VPS13Y characteristic feature is the presence of two C2 domains in tandem downstream of the VAB domain (Figures 3 and 4). VPS13 canonical ATG2_C and PH domains in C-term are also found in most of the VPS13Y proteins. No Chorein domain was detected by HHPred in N-term for some species, likely linked to errors in gene model prediction or sequence divergence. Interestingly, using TMHMM2.0 software (Krogh et al., 2001), a transmembrane (TM) helix was predicted at the N-term for most of the VPS13Y sequences. No TM domains were predicted in other VPS13 paralogs apart in Beta vulgaris VPS13X, implying that membrane anchoring might be a VPS13Y group specific feature. Thus, the ancestral VPS13Y protein might present the following domain organization: TM-Chorein-VAB-C2-C2-ATG2_C-PH. An additional Kinase domain was found in an extra-region located between the VAB and the first C2 KnVPS13Y. In K. nitens, we were able to predict the presence of 12 RBG domains (Figure 4), some of them being more divergent and predicted with the help of structural modeling. Unexpectedly, we failed to predict RBG domains in the N-term part of CsVPS13Y, even using KnVPS13Y RBG domains as a query (Figure 4). Thus, we performed secondary structure as well as structural prediction as described for VPS13B paralogs. Both methods predicted the presence of helices as well as disordered regions without β-strands, even in the N-term, in accordance with the absence of predicted Chorein domain by HHPred (see below and Supplemental File 3). The presence of the predicted TM domain, like in other VPS13Y orthologues, and transcriptomic data validate that the sequence is full length, raising question about the function of CsVPS13Y as an LTP. Similarly, the Prasimoderma coloniale VPS13Y (PRCOL_00005014-RA) sequence has two predicted TM domains in N-term, confirming that the sequence is full length, but no Chorein domain is predicted. Thus, VPS13Y C-term seems to be well conserved whereas the N-term is more divergent, as observed for VPS13B.
VPS13 Protein Structures in Viridiplantae
In order to further investigate the structural organization of the different Viridiplantae VPS13 paralogs, structural predictions were performed with ColabFold, an online AlphaFold2 tool (Jumper et al., 2021; Mirdita et al., 2022) as well as with RoseTTAFold (Baek et al., 2021). Structural prediction of overlapping VPS13 fragments was performed and aligned along the RBG domains, which were robustly predicted, and assembled using ChimeraX (Pettersen et al., 2021) (see Material and Methods). We succeeded in obtaining full-length structures for AtVPS13S, X, and M1. AtVPS13M2 structural prediction was not performed because of its close relationship to AtVPS13M1. AtVPS13M1 structural prediction was a challenging task because of the multiple additional domains that often led to steric clashes when different fragments were assembled. Thus, a combination of six fragments predicted with AlphaFold2 and also RoseTTAFold (Baek et al., 2021) were used in order to build a putative structure without steric clashes.
Structural prediction performed using the sequence from A. thaliana shows that VPS13S, M, and X harbor the well-described tunnel structure composed of RBG repeats with the canonical VAB, ATG_C and PH domains located at the C-term extremity of the tunnel (Figure 5A, Supplemental Files S4-6). In AtVPS13M1, whereas the PH1 domain and the FFAT motif are located within the first third of the tunnel, the other domains are all close to the C-term part. The different domains protrude from the tunnel without disturbing its structure and are linked by flexible loops. Thus, the orientation of the domains with respect to each other should be taken with caution, as it may not reflect the true organization of AtVPS13M1. In addition, their position might be modified by binding to partners. Accessory domains are suspected to play a role in VPS13 protein binding to membranes and positioning between two membranes. Thus, the particular location of the PH1 and C2 domains, known to bind lipids, in the central part of the tunnel raised questions about their mode of action. In VPS13X, the additional amphipathic helices and the β-trefoil domain are located close to the C-term part of the tunnel, likely for partners binding at the membrane surface.
Tunnel length of AtVPS13S and M1 is around 20.7 and 22.2 nm respectively, in accordance with what has been described for other VPS13 proteins harboring 12 RBG units (Figure 5B) (Neuman et al., 2022). As expected because of the presence of nine RBG repeats, VPS13X is shorter, measuring around 16.1 nm. Surprisingly, whereas AtVPS13S and M1 form a rather straight tunnel, AtVPS13X tunnel is bended, forming an almost 90° angle between the N- and the C-term tunnel extremities. Such bending was obtained whatever the overlapping fragments used for the predictions and assembly. The tunnel curvature seems to occur at the level of the RBG5 unit, likely because it is composed of seven instead of five β-strands (Levine, 2022). In addition, AtVPS13X tunnel is wider with an average width of 4.4 nm in contrast to AtVPS13S and M1 that are around 4 nm. Thus, the VPS13X protein harbors an unusual structure with a shorter, wider, and bended tunnel. Hydrophobicity analysis reveals the presence of an hydrophobic surface, likely involved in lipid transport, all along the tunnel concave face for the three VPS13s (Figure 5C), highlighting the conservation of the biochemical properties of Viridiplantae VPS13M, X, and S paralogs compared to opisthokonts’ VPS13s (Neuman et al., 2022; Dall’Armellina et al., 2023).
We attempted to predict the structure of VPS13Y from K. nitens and C. subellipsoidea. For KnVPS13Y, the N-term segment 1–1200 folded into a canonical hydrophobic tunnel, as described above (Figure S6), in accordance with the prediction of RBG repeats in KnVPS13Y (Figure 4). However, we were not able to assemble the full-length structure due to the presence of long unstructured regions in the tunnel and between the VAB and the C2 domains. In contrast, CsVPS13Y did not fold into a β-stranded structure but rather into a low confidence structure mainly composed of helices (Figure S6). Thus, it seems that VPS13Y has differentially evolved in the Viridiplantae phylum, keeping a canonical structure likely involved in lipid transport in K. nitens whereas diverging in C. subellipsoidea.
The analyzed VPS13B sequences did not contain well-predicted RBGs and as expected, did not fold into the well-known VPS13 hydrophobic tunnel in C. subellipsoidea, K. nitens, and S. fallax (Figure S7). The Chorein-RBG2 domains fold into the canonical β-sheet structure with an N-term helix, but the rest of the sequence is predicted to fold mostly into α-helices instead of β-sheets (Figure S7).
One characteristic feature of VPS13 proteins is the presence of the VAB domain involved in VPS13 targeting to membrane organelles (Bean et al., 2018; Dziurdzik et al., 2020). In yeast and humans, the VAB domain is composed of six repeated modules harboring a similar β-sandwich structure (Adlakha et al., 2022; Dall’Armellina et al., 2023). In yeast, the modules are composed of nine β-strands (Adlakha et al., 2022). We performed structural prediction of the VAB domains of each representative VPS13 paralogs (Figure 6). For VPS13Ms, we also included the C2 domain upstream of the VAB. Most of the sequences folded into a six module-structure composed of 5 to 10 β-strands, most of them being composed of 7 or 8 strands. The presence of α-helices is predicted in the module 4 of some paralogs such as AtVPS13S and X (Figure 6). The modules size ranges from 98 to 311 aa with an average length of 147 aa. Surprisingly, whereas six modules are clearly predicted for AtVPS13S, X, KnVPS13Y and SfVPS13B, only five are found in AtVPS13M1 and M2. Some β-strands, that might be reminiscent to the module 1 (Figure 6), are predicted upstream of the module 2. In order to better understand the structure of VPS13M VAB domains, structural predictions were also performed for the monocotyledon Zostera marina VPS13M1, the Charophyte K. nitens VPS13M1 and M2, the Chlorophyte C. subellipsoidea and the Prasinodermatophyte P. singulare VPS13M. It should be noticed that all these sequences have a predicted DysF domain, unlike the A. thaliana sequences (Figure 3). All the VPS13M sequences folded into a five-modules VAB domain with three to five β-strands upstream (Figure 6 and Figure S8). The C2 forms a domain clearly independent from the VAB. Interestingly, the predicted DysF forms a distinct domain inserted in the VAB module 2 (Figure 6). Its presence in all the Viridiplantae phylum is indicative of an ancestral domain lost in some species, such as A. thaliana in both VPS13M1 and M2 proteins. In yeast and mammals, a conserved Asn residue, important for VPS13 binding to partners, is present close to the N-term of each module (Bean et al., 2018; Dziurdzik et al., 2020; Dall’Armellina et al., 2023). Therefore, we aligned the sequences of all the VAB modules of the 12 VPS13 analyzed in Figures 6 and Figure S8, including the β-strands upstream of VPS13M module 2 (72 sequences in total). The results show that an Asn residue is indeed found within the seven first aa in almost all the modules (Supplemental Figure S9) with the exception of, as expected, most of the β-strands located upstream of VPS13M module 2 and, more surprisingly, in the module four of the three VPS13M1 analyzed. As the conserved Asn is found in VPS13M2 as well as in CsVPS13M and PsVPS13M module 4, its absence could be a feature of VPS13M1 acquired after VPS13M duplication. VPS13Y VAB domains from C. subellipsoidea and K. nitens harbor canonical six-module structures (Figure 6 and Figure S8). Noteworthy is the presence of long, mostly unfolded, sequences in modules 3 and 4 of CsVPS13 (Figure S8) which explain the prediction of a fragmented VAB by HHPred (Figure 4). S. fallax and C. subellipsoidea VPS13B VAB also fold into a six-module unit in contrast to K. nitens VPS13B for which no clear structure was predicted (Figure 6 and Figure S8). Thus, while AlphaFold2 and RoseTTAFold failed to predict a canonical tunnel structure for the N-term part (Figures S6 and S7), the presence of a canonical VAB domain in CsVPS13Y, CsVPS13B, and SfVPS13B suggests that they are indeed VPS13 proteins.

Structural modeling and characteristics of the VAB domain of Viridiplantae VPS13 paralogs. The sequences used are: Arabidopsis thaliana VPS13S (AtVPS13S), VPS13X (AtVPS13X), VPS13M1 (AtVPS13M1), and VPS13M2 (AtVPS13M2), Coccomyxa subellipsoidea VPS13M (CsVPS13M), Klebsormidium nitens VPS13Y (KnVPS13Y), VPS13M1 (KnVPS13M1), and VPS13M2 (KnVPS13M2), Prasinoderma singulare VPS13M (PsVPS13M), and Sphagnum fallax VPS13B (SfVPS13B). The 3D structures were obtained mainly using AlphaFold2 (Jumper et al., 2021) but also RoseTTAFold (Baek et al., 2021) for PsVPS13M. The C2 domain of VPS13M paralogs were included in the prediction. The different modules of the VAB domains are represented by colors and numbered from 1 to 6, the C2 and DysF domains found in VPS13Ms are in raspberry and limon, respectively.
Discussion
Evolutionary History of VPS13 Proteins Viridiplantae
In this work, we performed phylogenetic analyses of VPS13 proteins in Archaeplastida. Our results revealed the complex evolutionary history of this protein family with several losses, acquisitions, and duplications. As VPS13 proteins are mainly characterized by a specific domain organization and structural similarities rather than conservation of the amino acid sequences, in addition to ML analyses, we performed Bayesian analyses in which domains were inferred as characters in a reduced dataset of Rhodophytes, Glaucophytes, and Viridiplantae and then, only in Viridiplantae where VPS13 paralogs seemed more conserved. Our results showed that one VPS13 group, VPS13B, mainly represented in Metazoa (Velayos-Baeza et al., 2004; Levine, 2022), is found also in different phyla of Viridiplantae and was likely present in the LECA corroborating Levine's hypothesis (Levine, 2022). In Viridiplantae, four groups, VPS13M, S, X, and Y were clearly identified and interestingly, none of the Rhodophytes’ and Glaucophytes’ sequences clustered with these groups suggesting a high divergence of VPS13 sequences after the primary endosymbiosis. Rhodophytes’ sequences produced one robust clade that we named VPS13R grouping proteins with a similar domain organization. The other red algal sequences, though, are spread in different clades of the tree. Glaucophytes’ sequences clustered together in the different analyses performed but with a weak support and no clear groups or domain organization was detected, likely because of the reduced dataset available for this clade.
The evolutionary history of VPS13 proteins in Viridiplantae is more linear and summarized in Figure 7. The Viridiplantae MRCA had likely four VPS13 paralogs: VPS13S, M, Y, and B with the domain organization presented in Figure 7. VPS13B paralogs disappeared in Prasinodermatophytes and most of the Chlorophyte lineages apart from the Trebouxiophyceae and some others. Of note, in P. coloniale, two VPS13Y paralogs seem to be present as well as another sequence, without any clear link with the described Viridiplantae VPS13s, suggesting that Prasinodermatophytes might have acquired other groups of VPS13s. Early in the emergence of Streptophytes, VPS13X appeared. VPS13M gene was then duplicated to give rise to VPS13M1 and M2 groups shortly before the emergence of the Klebsormidiophyceae. This hypothesis is supported by the presence of five paralogs (VPS13B, S, M, X, and Y) in M. viride and six (VPS13B, S, M1, M2, X, and Y) in K. nitens. VPS13Y paralog was lost before the emergence of land plants around 500–600 Ma (Becker, 2013), and VPS13B was then lost in almost all lineages except the Bryophyte Sphagnum which maintained a VPS13B gene. Thereafter, VPS13M1 was duplicated in Bryophytes and the Angiosperms’ VPS13M2 proteins lost their C-term β-tripod domain (Figure 7). In addition, VPS13M2 was also duplicated in some angiosperm species (S. moellendorffii and Z. marina). These results illustrate the complex evolution of VPS13 proteins in terms of paralog number (gain and loss) and domain organization.

Schematic representation of VPS13 evolutionary history in Viridiplantae. VPS13 paralogs with their domain organization are represented in the different phyla analyzed in this study as well as the probable paralogs in most recent common ancestors (MRCA) of Viridiplantae and Streptophytes. Red and yellow dots indicate events of gene or domain losses and acquisitions, respectively. The MRCA of Viridiplantae had likely four VPS13 paralogs: VPS13S, M, Y, and B with the presented putative domain organization. After the separation between Chlorophytes and Streptophytes, VPS13B paralogs disappeared in most of the Chlorophyte lineages but the Trebouxiophyceae. Early in the emergence of Streptophytes, VPS13X appeared. VPS13M gene was duplicated to give rise to VPS13M1 and M2 groups after the emergence of Mesostigmaphyceae and Chlorokybophyceae. VPS13Y paralog was lost before the emergence of land plants and VPS13B was then lost in almost all lineages but the Bryophyte Sphagnum which maintained a VPS13B gene. Thereafter, the Angiosperms’ VPS13M2 proteins lost their C-term β-tripod domain. VPS13M2 duplication, not represented in this scheme, also occurred in some Embryophyte species (Zostera marina and Selaginella moellendorffii). ATG2_C = Autophagy-related protein 2 C-terminus homology; PH = Plekstrin Homology; TMD = transmembrane domain; VAB = VPS13 adaptor binding.
Characteristics of VPS13 Proteins in Viridiplantae
The retrieval of putative full-length VPS13 sequences was a challenging task because some genomes were poorly annotated and because automatic annotation software often failed to predict correct intron/exon borders for such long genes and some of them appeared split up. Extra unusual domains in N- or C-term are also frequently observed, likely linked to error in gene border annotations. Thus, for such genes, analysis of transcriptomic data, as we did for C. subellipsoidea, is the best method to obtain accurate VPS13 gene models. By selecting sequences longer than 2400 aa, we retrieved 383 sequences covering the three Archaeplastida lineages and showed that the gene number is highly variable ranging from 1 to 13. As mentioned above, the lower copy number is probably underestimated because we failed to detect split up genes. In Viridiplantae, our most curated dataset, the VPS13 gene copy number varies from three in Chlorophyceae to six in the Charophyte K. nitens and the Bryophyte S. fallax (Figure 7). Huge variations in sequence length were also observed.
By analyzing RBG repeats, we showed that sequence length is not always correlated to an increase in RBG repeats. Tunnel length and the addition of new domains are not directly correlated to the protein sequence length neither, as exemplified for VPS13X proteins ranging from 3122 to 4070 aa (Figure 4) though harboring the same RBG number and domain organization. Our analysis showed that general VPS13 protein domain and structure organization is widely conserved among Viridiplantae with the occurrence of a N-term Chorein, a VAB and an ATG2_C domain, most of them also having a C-term PH domain (Figure 3). Structural prediction revealed that the VAB domain structure composed of six repeated modules with an invariant Asn residue in N-term (Adlakha et al., 2022; Dall’Armellina et al., 2023) is also conserved in Viridiplantae VPS13s with some variations in VPS13M paralogs that will be discussed below. Additional domains, such as β-tripod, β-helix, β-sandwich, C2, or UBA domains, are found in different VPS13 groups and are likely real and functional. Others are only present in sparse sequences and might be linked to annotation errors. RBG repeats are also predicted for most of the paralogs we have analyzed, maximum number being 12 for Viridiplantae. In accordance with the presence of RBG domains, structural predictions showed that Viridiplantae VPS13s form long tunnels with a hydrophobic surface as described in opisthokonts (Neuman et al., 2022; Dall’Armellina et al., 2023). Additional localizations and/or functions might have arisen from the addition of new domains, not found in opisthokonts, such as β-tripod or C2 domains.
VPS13S
VPS13S has the simplest organization and it is tempting to speculate that VPS13S proteins harbor an ancestral structure and domain organization. It is particularly worth mentioning that VPS13S domain organization is highly similar to the one found in yeast ScVPS13. Both have 12 RBG repeats and harbor the same canonical functional domains. ScVPS13 is the only VPS13 protein found in S. cerevisiae and its presence is important for several basic cellular processes like prospore membrane expansion (Park and Neiman, 2012; Park et al., 2013) or autophagy (Dabrowski et al., 2023). It is therefore tempting to hypothesize that at least one simple form of VPS13 protein without additional non-canonical domains or alternative RBG domain organization is required in an organism to fulfill basic cellular functions. This hypothesis is corroborated by the fact that AtVPS13S plays important roles in Arabidopsis reproduction as its absence causes male as well as female sterility (Koizumi and Gallagher, 2013). AtVPS13S also interacts with the transcription factor SHORT ROOT and plays a role in a pathway regulating root growth and radial patterning (Koizumi and Gallagher, 2013). However, the exact mechanism by which AtVPS13S acts on this pathway is still elusive. AtVPS13S seems to act in gibberellic acid (GA) signaling, downstream of its synthesis (Koizumi and Gallagher, 2013). As GA is a mostly hydrophobic phytohormone, it is conceivable to suppose that AtVPS13S could be involved in its transport. AtVPS13S probably has other functions in cells, as this paralog is well conserved in Viridiplantae and GA signaling has emerged in vascular plants (Nishiyama et al., 2018).
VPS13M
VPS13M has the most complex organization with seven domains added during evolution, likely to regulate its localizations and/or functions. In VPS13M ancestor, three β-tripod domains, an additional PH, a C2 domain, a β-helix, as well as a DysF domain were present (Figure 4). A similar β-tripod domain was described in bacterial insecticidal proteins and shown to be involved in specific cell surface receptor recognition prior to the formation of membrane pores mediated by other protein domains (Zaitseva et al., 2019; Levine, 2022). Interestingly, the β-tripod domain seems to be widespread in Viridiplantae as 1085 out of the 1743 identified sequences containing a β-tripod domain, were from Viridiplantae (Zaitseva et al., 2019). Thus, the addition of β-tripod domains in VPS13M proteins might be involved in the regulation of protein localization and/or interaction with protein partners. A similar role could be attributed to the additional PH domain in N-term as well as to the C2 domain downstream the VAB domain, which could be involved in protein–protein or protein–lipid interactions. A β-helix was also found in VPS13M and it was hypothesized that it could be linked to the ubiquitination pathway because of its occurrence in F-box proteins (Levine, 2022). A certain degree of divergence was also revealed in VPS13M VAB domains by structural predictions (Figure 6). Indeed, only five full modules instead of six were identified and a DysF domain is inserted in the second module. The function of DysF domains is not clearly established but it is found in several proteins in yeast and humans involved in membrane remodeling (Bulankina and Thoms, 2020; Ferreira and Carvalho, 2021). In yeast, the DysF domain of Pex30 is essential for its function in LD biogenesis and at different contact sites (Ferreira and Carvalho, 2021). Therefore, the DysF domain could give to VPS13M new location determinants or new functions. The DysF domain was curiously lost in some sequences including in A. thaliana VPS13M1 and M2. In addition, in the VPS13M1 paralogs, no Asn residue was detected at the N-term of the VAB module 4. Asn residues from RBG5 and 6 are important for ScVPS13 and HsVPS13D partner binding (Dziurdzik et al., 2020), whereas HsVPS13B lacks this invariant residue in modules 2 and 3 (Dall’Armellina et al., 2023). Thus, the impact of the lack of Asn residue in the VPS13M VAB module 4 remains to be investigated. Structural prediction of AtVPS13M1 shows that it harbors the canonical hydrophobic tunnel structure with all the extra-domains protruding outside without affecting its folding (Figure 6). Questions remain about the organization of the extra-domains in relation to each other. In addition, these domains are likely involved in partners and membrane binding to regulate VPS13M localization and function. Although VPS13s in opisthokonts and yeast have simpler domain organizations, the mechanisms involved in the regulation of VPS13 localization at different sites is already complex (Bean et al., 2018; Dziurdzik and Conibear, 2021). Thus, it will be of high interest to investigate how VPS13M paralogs can integrate all the cellular signals to fulfill their function.
The evolutionary origin of VPS13M was not inferred from our analyses, but the complex domain organization depicted above is already found in Prasinodermatophytes. Up to now, β-tripod-containing VPS13 proteins were identified only in Viridiplantae, neither Rhodophytes or Glaucophytes possess β-tripod domains in VPS13 proteins. This suggests that the β-tripod domain emerged after the separation of Rhodophytes and Viridiplantae 1.5–2 Ma (Bowles et al., 2023). In addition, our analysis did not reveal any relationships among VPS13M and another VPS13 groups. As it possesses 12 RBG repeats like VPS13S, it would be parsimonious to hypothesize that it could come from a duplication of VPS13S ancestor followed by the rapid acquisition of the extra-domains.
From an interrogation of the InterPro database (Paysan-Lafosse et al., 2023), we found that IPR009291 domain (Vacuolar protein sorting-associated protein 62, aka β-tripod domain) is present in ≈6000 sequences belonging mainly to Viridiplantae (≈3000 sequences) and fungi (≈2000 sequences). In fungi, none of the sequences containing a β-tripod domain was associated to a known VPS13 domain. A few other lineages harboring β-tripod domain containing proteins, such as the Chlorarachniophytes Lotharella or the Rhodophyte Porphyridium purpureum, did not show association with VPS13 domains either. This makes the association of the β-tripod with the VPS13 canonical domains a Viridiplantae-specific feature. In order to check whether the β-tripod domain was retained through the secondary green endosymbiotic event, we retrieved sequences from Euglena gracilis and Bigelowiella natans (Supplemental File S7). However, none of them contained a predicted β-tripod domain, suggesting that VPS13M paralogs were not retained after the secondary endosymbiosis. Thus, the origin of the Viridiplantae VPS13M paralog with its high complexity remains unknown.
Our analyses revealed the highly dynamic and still ongoing evolution of the VPS13M group. VPS13M was duplicated early in the Streptophyte lineage into VPS13M1 and M2. Other duplication events were observed like VPS13M1 in Bryophytes and VPS13M2 in Z. marina and S. moellendorffii (Figure 3). In most of the phyla, both proteins maintained a conserved domain organization with 12 RBG and the domains described above. Interestingly, some VPS13M have also lost their DysF domain inserted in the VAB, like A. thaliana VPS13M1 and M2. In Angiosperms, the C-term β-tripod domain was lost, likely to allow the emergence of a new function. In a recent pre-print, functional analysis of AtVPS13M1 and M2 suggested that AtVPS13M2, but not M1, could play a role in pollen germination and pollen tube growth (Tangpranomkoorn, 2022, BioRvix, DOI:10.1101/2022.11.01.514778). A rapid polarized extension of the pollen tube is required in order to deliver sperm cells to the ovule. Its extension requires a massive vesicular lipid and protein transport to the pollen tube tips to allow the delivery of components required to sustain its growth, including lipids (Adhikari et al., 2020). AtVPS13M2 accumulates at the pollen tube tip secretory vesicles and might play a role in the establishment of polarity and in cell wall deposition during pollen tube growth (Tangpranomkoorn, 2022, BioRvix, DOI:10.1101/2022.11.01.514778). Pollen tube growth is common in Angiosperms but also found in some Gymnosperm species that have usually shorter pollen tubes that grow more slowly (Fernando et al., 2005; Adhikari et al., 2020). The only Gymnosperm species included in our dataset is Thuja plicata in which pollen germination and pollen tube growth was reported (Land, 1902). TpVPS13M2 still has a C-term β-tripod domain, thus it is difficult to speculate about the link between the presence of this domain and an essential role in the pollen germination process. An extended investigation of VPS13M2 sequences and a functional analysis in Gymnosperms might help to understand when the C-term β-tripod domain was lost and the reason for the VPS13M2 specialization in Angiosperms.
VPS13B
From previous analyses of VPS13 RBG domain organization, it was suggested that the LECA likely had two paralogs: VPS13B and an ancestral VPS13 which gave rise to the other groups of VPS13 (Levine, 2022). Our analyses further confirmed the ancestry of VPS13B as it was found in some Viridiplantae. However, VPS13B is maintained in a small subset of phyla, raising questions about the function of this protein. In human, VPS13B mutations are associated to the Cohen Syndrome, a developmental disorder associated with variable clinical phenotypes such as microcephaly, pigmentary retinopathy or hypotonia (Douzgou and Petersen, 2011). At the cellular level, VPS13B localizes to the Golgi and is involved in endosome tethering (Seifert et al., 2011; Koike and Jahn, 2019). Roles in the maintenance of Golgi integrity (Seifert et al., 2011; Da Costa et al., 2020), autophagy and neurite outgrowth (Seifert et al., 2015) as well as in acrosome formation in mice (Da Costa et al., 2020), were demonstrated. Most of these functions are associated to extensive membrane trafficking. Whereas maintenance of Golgi and endosome tethering functions are basal processes likely required for the maintenance of all Eukaryotic cells, VPS13B function might be essential in some developmental processes specific to animals such as brain development or acrosome formation during spermatogenesis, explaining its maintenance in this clade. However, we could not find evident functions that might be specific to the small subset of Viridiplantae species in which VPS13B orthologues are present, suggesting that new functions might have evolved independently in each phylum. We were able to find transcripts supporting the expression of VPS13B in C. subellipsoidea and in two Sphagnum species, proving their expression and functionality, possibly as LTPs because of the conservation of the Chorein + RBG2 domain, able to bind lipids in vitro (Kumar et al., 2018).
Another intriguing point is the domain organization of VPS13B orthologues. Whereas the Chorein and RBG2 domains as well as the C-term part of VPS13B orthologues, including the VAB domain, appeared well conserved in terms of domain and structure, we failed to detect seven of the RBG domains predicted in HsVPS13 between RBG2 and RBG10 (Levine, 2022; Dall’Armellina et al., 2023). In contrast, this region is mostly composed of helices and unstructured loops, raising questions about their general structure and function conservation (Figure S7). A reduced number of RBG domains might imply a reduction of the tunnel length, maybe to accommodate the protein to contact sites of shorter length. However, if this is the case, why was the sequence maintained between the Chorein and the VAB domains during evolution? This might be the sign that VPS13B is slowly losing its bridge-like LTP function in Viridiplantae. Alternatively, the structure might have been modified to accommodate for the interaction with other protein partners or to gain new functions. A closer structural analysis coupled with functional studies by reverse genetics approaches might help to decipher the role of VPS13B paralogs in non-Metazoa and their evolutionary history.
From our analyses, two main hypotheses can be posed concerning the evolution of VPS13B: (1) the VPS13B ancestor had six RBG repeats which were then multiplied in Humans to obtain a VPS13B with 13 RBG repeats or (2) VPS13B ancestor encompassed 13 RBG repeats that have progressively diverged in Viridiplantae. The second hypothesis seems more probable because an ancestor with six RBG would imply the addition of a 1000-aa sequence, not clearly folded, in Viridiplantae VPS13B. The analyses of the N-term domain organization of a higher number of VPS13B orthologues, including basal Metazoan, might help to depict the organization of ancestral VPS13B and their evolution.
VPS13X
The origin of VPS13X is puzzling as its sequence and domain organization is highly divergent compared to other paralogs. In agreement with this, VPS13X was only recently identified by remote homology analyses (Levine, 2022) and not in previous studies using BLASTP searches (Velayos-Baeza et al., 2004; Michaud et al., 2017; Leterme and Michaud, 2022). VPS13X is composed of nine RBG domains, having the smallest number of repeats in currently characterized VPS13s, and is the only group of VPS13 in which a C-term PH domain is absent, amphipathic α-helices are found instead (Levine, 2022). Thus, VPS13X looks like a basal ancestral protein, with an extra β-sandwich domain between RBG6 and 7. However, we were not able to detect any VPS13X orthologues in species other than Streptophytes, even by PSI-BLASTP using either Charophytes or Embryophytes VPS13X as queries (data not shown), hinting that VPS13X appeared soon after the divergence of Streptophytes and Chlorophytes or that it was lost in the genome of all the other species sequenced so far. As it was not found in other organisms, the hypothesis of a horizontal gene transfer is also unlikely. A previous extended RBG analysis proposed that RBG3 and 6 domains are unrelated to specific other known VPS13 RBG domains, but might have evolved from penultimate domains, and that RBG5 is unique to VPS13X as it is composed of seven instead of five β-strands (Levine, 2022). Structural predictions show that VPS13X forms a shorter, larger, and bended hydrophobic tunnel in contrast to other VPS13 (Figure 4). In addition, our analysis did not reveal any robust relationship with other VPS13 from Archaeplastida. Thus, if VPS13X arose from a VPS13 paralog gene duplication, a rapid evolution of the gene occurred with a loss of PH and RBG domains. Proteins can mutate at different rates depending on several factors like selection pressure (such as sperm proteins (Torgerson et al., 2002)), folding stability and chaperoning efficiency (Agozzino and Dill, 2018). A new protein stemming from a gene duplication may have folding instabilities and be more prone to mutation. An extreme consequence would be the loss of the protein. In comparison, VPS13M1 and M2 genes appeared by duplication of VPS13M ancestor almost at the same time as VPS13X appeared after the divergence of Streptophytes and Chlorophytes. Nonetheless, a high similarity is still observed between VPS13M1 and M2, which is not the case of VPS13X. An alternative hypothesis might be that VPS13X arose from VPS13B. In fact, VPS13B proteins harbor a β-sandwich fold between RBG10 and 11 and VPS13X proteins have a β-trefoil fold inserted between RBG6 and 7 (structurally equivalent positions). Interestingly, the β-sandwich and the β-trefoil folds are both structurally related as it has been proposed that β-trefoils might have evolved from β-sandwiches (Longo et al., 2022). Furthermore, HsVPS13B and AtVPS13X both have central RBG domains that share low similarity to the penultimate RBG domains, in contrast with other previously analyzed VPS13 paralogs (Levine, 2022). It is also noteworthy that the AtVPS13X RBG5 is composed of seven β-strands instead of the usual five, which might be a relic of the RBG domain loss necessary for the transition from 13 to 9 RBG repeats in VPS13X. Additionally, the C-term PH domain is only weakly predicted in VPS13B proteins while it is absent in VPS13X proteins. It is therefore possible that VPS13X proteins derived from VPS13B proteins by losing several central RBG domains, by converting the β-sandwich fold into a β-trefoil fold and by losing the C-term PH domain. However if so, this event implies the duplication of VPS13B gene early in Streptophytes as both VPS13B and X are found in Charophytes and in the Bryophyte Sphagnum. Then, VPS13X was maintained whereas VPS13B was lost. At this stage, the number of VPS13B sequences obtained from Streptophytes is too low to perform an informative comparative analysis of VPS13X and B structures and sequences. Finally, VPS13X might also have emerged from the fusion of a C-term VPS13 paralog and the N-term of another RBG protein family, such as SHIP164 or Tweek, present in A. thaliana (Levine, 2022), but an extensive analysis of these families in photosynthetic organisms may be required to corroborate or deny this hypothesis.
The function of VPS13X is currently unknown but structural predictions suggest that it might also play a role in lipid and membrane trafficking processes. In accordance with the reduced number of RBG domains, AtVPS13X forms a shorter hydrophobic tunnel, likely around 16 nm, which is predicted to be larger and bended (Figure 4), inferring that it may play a role at contact sites other than those of the longer VPS13 paralogs (around 20 nm) and between membranes that are perpendicularly apposed. Its occurrence only in Streptophytes suggests a role in a process specific to this clade. Expansion of LTPs in plants is observed for several families, the strongest example being the apparition of a plant-specific LTP family: the non-specific (ns) LTPs. This family emerged in Embryophytes after the split with Charophytes and more than 100 genes are currently found in genomes of higher plants, highlighting the extremely rapid expansion of this family (Edstam et al., 2011). Noteworthy, the number of other LTP homologous families, such as the StARTs (steroidogenic acute regulatory protein-related lipid transfer) or the ORPs (Oxysterol-binding protein related proteins), is often higher in plants compared to yeast or animals (Leterme and Michaud, 2022). This is likely linked to the emergence of new cell functions and structures such as chloroplasts, plasmodesmata, or complex cell walls. Indeed, the synthesis of plant cell wall structures, such as cutin, wax, or sporopollenin, requires a massive secretion of hydrophobic components and many nsLTPs have been involved in such processes (Edstam et al., 2011; Edqvist et al., 2018). In addition, the terrestrialization of plants was accompanied by their ability to adapt to a wide variety of environmental variations and biotic stresses (Bowles et al., 2023). Plants have evolved several mechanisms involving massive membrane remodeling in order to adapt to nutrient stress or to establish symbiosis with microorganisms (Moellering and Benning, 2011; Michaud and Jouhet, 2019). Genome analyses of different Charophytes revealed that they already encode several key genes involved in land plant specific processes, including symbiosis with microorganisms (Nishiyama et al., 2018; Domozych and Bagdan, 2022; Bowles et al., 2023). Thus, VPS13X might have emerged in Charophytes to ensure a key function in Streptophyte evolution likely linked to lipid and membrane trafficking. No functional studies have been carried out yet in A. thaliana or other species and this will be a key step to understand the role and evolution of the VPS13X family.
VPS13Y
VPS13Y is a newly described group of VPS13 characterized by the presence of a putative TM domain in N-term end as well as two C2 domains downstream of the VAB. Whereas the N-term region of KnVPS13Y is predicted to fold into a β-sheet tunnel, as expected for a VPS13 protein, such folding was not predicted by structural nor by secondary structure analyses for CsVPS13Y (Figure S6). Instead, it folds into helices, as observed for some VPS13B paralogs, raising questions about VPS13Y function and conservation. Inferring a function to this group is a challenging task as no other described VPS13 has a similar domain organization. C2 domains are involved in protein–lipid binding and as VPS13Y proteins putatively have N-term TM domain, they might act as membrane tethers, like Extended-synaptotagmin LTPs (Saheki and De Camilli, 2017). Because of the absence of the Chorein domain, it is also possible that CsVPS13Y has lost its ability to act as a LTP.
VPS13Y has been detected in Prasinodermatophytes, Chlorophytes, and Charophytes, suggesting it was present in the Viridiplantae MRCA and lost in Embryophytes, the reason being unknown.
Overall, our results provided the first extensive description of VPS13 paralogs in Archaeplastida and revealed a conservation of their general organization. It also highlighted the complex evolutionary history of Viridiplantae VPS13 proteins, with several paralogs and losses and appearances of functional domains. Future efforts are now required in order to decipher the localization, structure, and function of the different VPS13 paralogs in Archaeplastida.
Material and Methods
Sequence Retrieval and Selection
For Viridiplantae, all the sequences from Embryophytes (land plants) were retrieved from the Phytozome database (Goodstein et al., 2012), which contains assembled genomes from more than 300 Viridiplantae species. For Glaucophytes, Rhodophytes, Chlorophytes, Prasinodermatophytes, and Charophytes, sequences were retrieved from five databases: Uniprot (The UniProt Consortium et al., 2023), Joint Genome Institue (JGI) (Nordberg et al., 2014), Phycocosm (Grigoriev et al., 2021), Eukprot V3 (Richter et al., 2022), and the National Center for Biotechnology Information (NCBI) (Sayers et al., 2022). To find as many VPS13 homologs as possible, BLAST searches (Altschul et al., 1990) were primarily performed using the well-described Arabidopsis thaliana sequences (M1: At4g17140, M2: At1g48090, S: At5g24740, and X: At3g50380) as queries. For Embryophytes, BLASTPs were performed with AtVPS13M1 and AtVPS13X aa sequences, whereas for other organisms, BLAST searches were performed from the four A. thaliana sequences because of the putative higher divergence of sequences compared to the plant ones. As orthologues of HsVPS13B were recently supposed to be present in some protists (Levine, 2022), HsVPS13B was BLASTP searched as well. Sequences with at least one VPS13 domain annotation, such as VPS13_N, VAB, Chorein or VPS13_C, were retrieved to generate data in Table S1. Then, a curation was performed by selecting sequences that were at least 2400 aa-long to constitute a dataset of 383 sequences (with addition of the four HsVPS13 and ScVPS13), used for ML phylogenetic analyses (Dataset 1). Some sequences were split up in two or three pieces and were merged to obtain likely full-length sequences: Closterium fusion = GJP45422 + GJP45421 + GJP45420; Klebsormidium nitens fusion = GAQ85408 + GAQ85409; Sphagnum fallax fusion = Sphfalx12G108100.1 + Sphfalx12G108200.1, Thuja plicata fusion = Thupl.29379161s0002.1.p + Thupl.29379161s0001.1.p. Then, a subset of organisms was chosen in each taxonomic clade to obtain a reduced dataset of 104 sequences to perform ML and Bayesian phylogenetic analysis (Dataset 2). We selected species for which we had the highest number of sequence and eliminate protein models potentially emerging from the same gene but corresponding to splicing variant by multiple alignments of sequences within each species. When two proteins likely originating from the same gene were discovered, the longest isoform was kept for the analysis. Presence of at least two canonical VPS13 domains (Chorein, VAB, ATG2_C, PH) was also used as a criterion to select sequences. HsVPS13A/B/C/D and ScVPS13 were included in this dataset. A final dataset, containing only Viridiplantae sequences of Dataset 2, was created in order to perform ML and Bayesian phylogenetic analysis in this phylum, excluding HsVPS13A/B/C/D and ScVPS13 sequences (Dataset 3, 79 sequences).
Domain and RBG Predictions
Domain prediction was performed using HHPred (Zimmermann et al., 2018) on all the sequences of Dataset 2 in order to infer domain information in the Bayesian analysis. The full-length sequences were used for analysis and domains predicted with at least 50% confidence were kept. Because of the huge protein size, HHPred usually failed to predict all the putative domains and 2nd and 3rd analysis were performed in regions in which no domains were predicted at a first place. Once again, domains predicted with at least 50% confidence were kept. Domains found in the sequences are summarized in Tables S2 and S3 in the form it was coded for the Bayesian analysis.
RBG domain prediction was performed by HHPred in the targeted sequences by using as queries RBG domain sequences of AtVPS13 paralogs as well as HsVPS13B described by Levine (2022). For CsVPS13Y, KnVPS13Y predicted RBG domains were also used. When a low similarity was found, some RBG were confirmed by performing structural predictions with ColabFold (Mirdita et al., 2022).
Maximum Likelihood Analyses
ML analyses were performed with datasets 1, 2, and 3. The full-length protein sequences were aligned with the software MAFFT (Katoh and Standley, 2013) performing 16 iterations of refinement. The obtained alignment files were processed with the trimAL (Capella-Gutiérrez et al., 2009) program to remove all sites with a percentage of gaps higher than 10% (gap score = 10). After trimming, the size of sequences in the final alignments is around 900–1000 aa and include mainly amino acids from conserved domains (Chorein-RBG2-RBG10-VAB-RBG11-RBG12-ATG_C-PH). The phylogenetic reconstruction was performed with the software FastME (Lefort et al., 2015) using the LG model and 5000 bootstrap iterations. Nodes with at least 4000 boostraps (80%) were considered as well supported.
Bayesian Inference
Two independent Metropolis-coupled Markov Chain Monte Carlo (MC3) analyses were performed using the final curated datasets in the MrBayes3.2 software (Ronquist et al., 2012). Two partitioned datasets were used for Bayesian inference, each including a binary column indicating the result of domain annotation reported above and a second partition for the amino acid sequence, for the Archaeplastida (Table S2) and for Viridiplantae (Table S3) containing 104 (Standard:1-42,PROTEIN:43-1166) and 79 (Standard:1-25,PROTEIN:26-1332) entries, respectively. The domain annotation was mathematically formalized into a 1/0 code (presence = 1, absence = 0) and each domain accounted as a position in the alignment. One cold chain and either seven (temperature parameter 0.01, Dataset 2) or three “heated” (temperature parameter 0.1, Dataset 3) chains drove the analyses. The analyses were allowed to switch among all the substitution models implemented in the software to identify the best model without any a priori (aamodelpr = mixed). 90,000 and 35,000 generations with sampling every 100 generations were set for Dataset 2 and Dataset 3, respectively. The standard deviation of the split frequency between the two parallel analyses at the end of each analysis were 0.068996 and 0.006487 for Dataset 2 and Dataset 3, respectively. The first 25% of the trees produced were discarded in order to let the analyses stabilize (burnin = 0.25). The convergence of the runs was estimated and the potential scale reduction factor (PSRF + ) were all around 1 for all the parameters indicating that the associated chains converged to one target distribution. Large PSRF (PSRF + >1) indicate convergence failure. Nodes with a posterior probability ≥0.95 were considered as well supported. Alignments in fasta and trees in newick format are available upon request.
Expression Analysis
For C. subellipsoidea, BLASTN searches on JGI were performed against the EST cluster available using the different VPS13 paralog hypothetical mRNA sequences as queries (CsVPS13M: XM_005649615.1, CsVPS13S: XM_005645274.1, CsVPS13Y: XM_005642608.1, CsVPS13B: XM_005646816.1). ESTs with at least 90% identity were retrieved and aligned with the corresponding hypothetical mRNA sequences using SnapGene® software version 5.0.8. Then, in order to obtain curated version of the mRNA and protein sequences, modification on hypothetical mRNAs were performed by incorporating variations (insertions, deletions, substitutions) observed within at least two ESTs.
C. braunii (Bioproject PRJNA492241), K. nitens (Bioproject PRJN242253), Sphagmun recurvum (Biosample UHLI), and Sphagnum palustre (Biosample RCBT) transcriptome assemblies were retrieved from NCBI. TBLASTN searches were performed using respective VPS13 protein sequences as queries and sequences that covered at least 10% of the query and with an e-value below 0.001 were retrieved (maximum 20 hits). Then, according to the percentage of identity to the VPS13 paralogs used as query, we annotated each hit in order to determine which paralogs are expressed in each species.
Structural Predictions
Structural predictions were performed using ColabFold, an easily accessible web interface to perform AlphaFold2 3D predictions (Jumper et al., 2021; Mirdita et al., 2022). RoseTTAFold (Baek et al., 2021) was also used for some complex structure, such as AtVPS13M1, or some VAB sequences that were not properly folded with AlphaFold2 predictions. For full-length structures determination, overlapping fragments of maximum 1400 aa were designed with overlapping RBG repeats at each extremity for structural prediction. Fragments structure were aligned on overlapping RBG and joined between the two most-well aligned residues using ChimeraX (Pettersen et al., 2021). Four fragments were assembled for AtVPS13S (1-1107; 1108-1801; 1802-2840; 2841-3464) and AtVPS13X (1-989; 990-1676; 1677-2622; 2623-3132). For AtVPS13M1, six fragments were predicted (1-955; 956-1837; 1838-2414; 2415-3469; 3470-3737; 3738-4202) and assembled in order to avoid steric clashes between all the additional domains. Assembled structure of AtVPS13S, M1, and X are provided in Supplemental Files S4, S5 and S6, respectively. VAB modules sequences were determined from the structures. Structures were visualized and annotated in PyMOL (The PyMOL Molecular Graphics System, Version 1.2r3pre, Schrödinger, LLC).
Supplemental Material
sj-pptx-1-ctc-10.1177_25152564231211976 - Supplemental material for Phylogenetic and Structural Analyses of VPS13 Proteins in Archaeplastida Reveal Their Complex Evolutionary History in Viridiplantae
Supplemental material, sj-pptx-1-ctc-10.1177_25152564231211976 for Phylogenetic and Structural Analyses of VPS13 Proteins in Archaeplastida Reveal Their Complex Evolutionary History in Viridiplantae by Sébastien Leterme, Olivier Bastien, Riccardo Aiese Cigliano, Alberto Amato and Morgane Michaud in Contact
Supplemental Material
sj-docx-2-ctc-10.1177_25152564231211976 - Supplemental material for Phylogenetic and Structural Analyses of VPS13 Proteins in Archaeplastida Reveal Their Complex Evolutionary History in Viridiplantae
Supplemental material, sj-docx-2-ctc-10.1177_25152564231211976 for Phylogenetic and Structural Analyses of VPS13 Proteins in Archaeplastida Reveal Their Complex Evolutionary History in Viridiplantae by Sébastien Leterme, Olivier Bastien, Riccardo Aiese Cigliano, Alberto Amato and Morgane Michaud in Contact
Supplemental Material
sj-docx-3-ctc-10.1177_25152564231211976 - Supplemental material for Phylogenetic and Structural Analyses of VPS13 Proteins in Archaeplastida Reveal Their Complex Evolutionary History in Viridiplantae
Supplemental material, sj-docx-3-ctc-10.1177_25152564231211976 for Phylogenetic and Structural Analyses of VPS13 Proteins in Archaeplastida Reveal Their Complex Evolutionary History in Viridiplantae by Sébastien Leterme, Olivier Bastien, Riccardo Aiese Cigliano, Alberto Amato and Morgane Michaud in Contact
Supplemental Material
sj-docx-4-ctc-10.1177_25152564231211976 - Supplemental material for Phylogenetic and Structural Analyses of VPS13 Proteins in Archaeplastida Reveal Their Complex Evolutionary History in Viridiplantae
Supplemental material, sj-docx-4-ctc-10.1177_25152564231211976 for Phylogenetic and Structural Analyses of VPS13 Proteins in Archaeplastida Reveal Their Complex Evolutionary History in Viridiplantae by Sébastien Leterme, Olivier Bastien, Riccardo Aiese Cigliano, Alberto Amato and Morgane Michaud in Contact
Supplemental Material
sj-pdb-5-ctc-10.1177_25152564231211976 - Supplemental material for Phylogenetic and Structural Analyses of VPS13 Proteins in Archaeplastida Reveal Their Complex Evolutionary History in Viridiplantae
Supplemental material, sj-pdb-5-ctc-10.1177_25152564231211976 for Phylogenetic and Structural Analyses of VPS13 Proteins in Archaeplastida Reveal Their Complex Evolutionary History in Viridiplantae by Sébastien Leterme, Olivier Bastien, Riccardo Aiese Cigliano, Alberto Amato and Morgane Michaud in Contact
Supplemental Material
sj-pdb-6-ctc-10.1177_25152564231211976 - Supplemental material for Phylogenetic and Structural Analyses of VPS13 Proteins in Archaeplastida Reveal Their Complex Evolutionary History in Viridiplantae
Supplemental material, sj-pdb-6-ctc-10.1177_25152564231211976 for Phylogenetic and Structural Analyses of VPS13 Proteins in Archaeplastida Reveal Their Complex Evolutionary History in Viridiplantae by Sébastien Leterme, Olivier Bastien, Riccardo Aiese Cigliano, Alberto Amato and Morgane Michaud in Contact
Supplemental Material
sj-pdb-7-ctc-10.1177_25152564231211976 - Supplemental material for Phylogenetic and Structural Analyses of VPS13 Proteins in Archaeplastida Reveal Their Complex Evolutionary History in Viridiplantae
Supplemental material, sj-pdb-7-ctc-10.1177_25152564231211976 for Phylogenetic and Structural Analyses of VPS13 Proteins in Archaeplastida Reveal Their Complex Evolutionary History in Viridiplantae by Sébastien Leterme, Olivier Bastien, Riccardo Aiese Cigliano, Alberto Amato and Morgane Michaud in Contact
Supplemental Material
sj-docx-8-ctc-10.1177_25152564231211976 - Supplemental material for Phylogenetic and Structural Analyses of VPS13 Proteins in Archaeplastida Reveal Their Complex Evolutionary History in Viridiplantae
Supplemental material, sj-docx-8-ctc-10.1177_25152564231211976 for Phylogenetic and Structural Analyses of VPS13 Proteins in Archaeplastida Reveal Their Complex Evolutionary History in Viridiplantae by Sébastien Leterme, Olivier Bastien, Riccardo Aiese Cigliano, Alberto Amato and Morgane Michaud in Contact
Supplemental Material
sj-xlsx-9-ctc-10.1177_25152564231211976 - Supplemental material for Phylogenetic and Structural Analyses of VPS13 Proteins in Archaeplastida Reveal Their Complex Evolutionary History in Viridiplantae
Supplemental material, sj-xlsx-9-ctc-10.1177_25152564231211976 for Phylogenetic and Structural Analyses of VPS13 Proteins in Archaeplastida Reveal Their Complex Evolutionary History in Viridiplantae by Sébastien Leterme, Olivier Bastien, Riccardo Aiese Cigliano, Alberto Amato and Morgane Michaud in Contact
Supplemental Material
sj-xlsx-10-ctc-10.1177_25152564231211976 - Supplemental material for Phylogenetic and Structural Analyses of VPS13 Proteins in Archaeplastida Reveal Their Complex Evolutionary History in Viridiplantae
Supplemental material, sj-xlsx-10-ctc-10.1177_25152564231211976 for Phylogenetic and Structural Analyses of VPS13 Proteins in Archaeplastida Reveal Their Complex Evolutionary History in Viridiplantae by Sébastien Leterme, Olivier Bastien, Riccardo Aiese Cigliano, Alberto Amato and Morgane Michaud in Contact
Supplemental Material
sj-xlsx-11-ctc-10.1177_25152564231211976 - Supplemental material for Phylogenetic and Structural Analyses of VPS13 Proteins in Archaeplastida Reveal Their Complex Evolutionary History in Viridiplantae
Supplemental material, sj-xlsx-11-ctc-10.1177_25152564231211976 for Phylogenetic and Structural Analyses of VPS13 Proteins in Archaeplastida Reveal Their Complex Evolutionary History in Viridiplantae by Sébastien Leterme, Olivier Bastien, Riccardo Aiese Cigliano, Alberto Amato and Morgane Michaud in Contact
Supplemental Material
sj-fas-12-ctc-10.1177_25152564231211976 - Supplemental material for Phylogenetic and Structural Analyses of VPS13 Proteins in Archaeplastida Reveal Their Complex Evolutionary History in Viridiplantae
Supplemental material, sj-fas-12-ctc-10.1177_25152564231211976 for Phylogenetic and Structural Analyses of VPS13 Proteins in Archaeplastida Reveal Their Complex Evolutionary History in Viridiplantae by Sébastien Leterme, Olivier Bastien, Riccardo Aiese Cigliano, Alberto Amato and Morgane Michaud in Contact
Supplemental Material
sj-fas-13-ctc-10.1177_25152564231211976 - Supplemental material for Phylogenetic and Structural Analyses of VPS13 Proteins in Archaeplastida Reveal Their Complex Evolutionary History in Viridiplantae
Supplemental material, sj-fas-13-ctc-10.1177_25152564231211976 for Phylogenetic and Structural Analyses of VPS13 Proteins in Archaeplastida Reveal Their Complex Evolutionary History in Viridiplantae by Sébastien Leterme, Olivier Bastien, Riccardo Aiese Cigliano, Alberto Amato and Morgane Michaud in Contact
Supplemental Material
sj-fas-14-ctc-10.1177_25152564231211976 - Supplemental material for Phylogenetic and Structural Analyses of VPS13 Proteins in Archaeplastida Reveal Their Complex Evolutionary History in Viridiplantae
Supplemental material, sj-fas-14-ctc-10.1177_25152564231211976 for Phylogenetic and Structural Analyses of VPS13 Proteins in Archaeplastida Reveal Their Complex Evolutionary History in Viridiplantae by Sébastien Leterme, Olivier Bastien, Riccardo Aiese Cigliano, Alberto Amato and Morgane Michaud in Contact
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work is supported by the French National Research Agency in the framework of the “investissement d’avenir” program (ANR-15-IDEX-02) and of the ANRJCJC MiCoSLiT (ANR-19-C13–0013) to MM. It also received funding from GRAL, a program from the Chemistry Biology Health (CBH) Graduate School of University Grenoble Alps (ANR-17-EURE-0003).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
