Abstract
Peroxisomes play important roles in lipid metabolism. Surplus or damaged peroxisomes can be selectively targeted for autophagic degradation, a process termed pexophagy. Maintaining a proper level of pexophagy is critical for cellular homeostasis. Here, we found that endoplasmic reticulum (ER)–mitochondria contact sites are necessary for efficient pexophagy. During pexophagy, the peroxisomes destined for degradation are adjacent to the ER–mitochondria encounter structure (ERMES) that mediates the formation of ER–mitochondria contacts; disruption of the ERMES results in a severe defect in pexophagy. We show that a mutant form of Mdm34, a component of the ERMES, which impairs ERMES formation and diminishes its association with the peroxisomal membrane protein Pex11, also causes defects in pexophagy. The dynamin-related GTPase Vps1, which is specific for peroxisomal fission, is recruited to the peroxisomes at ER–mitochondria contacts by the selective autophagy scaffold Atg11 and the pexophagy receptor Atg36, facilitating peroxisome degradation.
Introduction
Peroxisomes are single-membrane organelles found in eukaryotes, and they are involved in several important aspects of cellular metabolism, including β-oxidation of fatty acids, a process providing a critical source of metabolic energy (Lodhi & Semenkovich, 2014; Sibirny, 2016; Smith & Aitchison, 2013). In mammals, long-chain fatty acids are converted to medium-chain fatty acids in peroxisomes before they are transported to mitochondria and broken down into carbon dioxide and water. In yeasts and plants, this process is carried out exclusively in peroxisomes (Sibirny, 2016). In addition, a by-product of the process, reactive oxygen species, may be detrimental to cells under certain circumstances. Therefore, to maintain cellular homeostasis, the quantity and quality of peroxisomes must be finely regulated, and autophagy plays an indispensable role in this process.
Autophagy is a cellular process that sequesters portions of the cytoplasm, which are subsequently delivered to the vacuole (in yeasts and plants) or lysosomes (in mammals) for degradation (Choi, Ryter, & Levine, 2013; Deretic & Levine, 2009; Klionsky & Codogno, 2013; Klionsky et al., 2003; Xie & Klionsky, 2007). Autophagy can work in either a nonselective or selective mode, depending on the extracellular stress (Jin, Liu, & Klionsky, 2013; Lynch-Day & Klionsky, 2010; Mijaljica et al., 2012). Nonselective autophagy is proposed to mediate the bulk degradation of cytoplasm and subsequent recycling of materials essential for cell survival under conditions of nutrient deprivation. Selective autophagy targets specific cargoes in different contexts. For example, if yeasts are grown on methanol (e.g., Pichia pastoris and Hansenula polymorpha) or oleic acid (Saccharomyces cerevisiae), peroxisomes proliferate; a subsequent shift to ethanol or glucose will result in the selective degradation of the excess peroxisomes.
Defects in peroxisome function can have serious medical consequences, as seen with Zellweger syndrome, Refsum disease, and rhizomelic chondrodysplasia punctata, which can result in severe developmental problems or even be lethal (Wanders et al., 1986). The accumulation of peroxisomal substrates or the absence of products that are normally generated through peroxisomal metabolic reactions can result in serious physiological challenges. Thus, it is critical to maintain functional peroxisomes. Conversely, the risks associated with defective organelles that generate reactive oxygen species and the energetic requirements for maintaining organelles that function properly mean that it is generally beneficial to eliminate them when they become superfluous. The selective autophagy process that recognizes and degrades surplus or damaged peroxisomes is termed pexophagy (Farre, Manjithaya, Mathewson, & Subramani, 2008; Motley, Nuttall, & Hettema, 2012; Till, Lakhani, Burnett, & Subramani, 2012). Therefore, a more complete knowledge of the molecular mechanisms of pexophagy and how pexophagy is regulated is critical to gain a more complete understanding of peroxisome-related diseases and potential therapeutic avenues.
Studies have suggested a general “cargo-ligand-receptor-scaffold” model to illustrate how selective autophagy works (Jin et al., 2013; Mijaljica et al., 2012). Ligands on cargoes recruit specific receptor proteins, which interact with a scaffold or Atg8 to link the cargo with the autophagy machinery, facilitating the selective engulfment of cargoes by phagophores. In yeast, Atg11 is identified as the scaffold for many selective autophagy processes, including pexophagy. In S. cerevisiae, Pex3 serves as the ligand on peroxisomes, and this protein interacts with the receptor Atg36 to mediate pexophagy (Motley et al., 2012). Atg11 and Atg36 were also recently shown to recruit the peroxisome fission complex containing the two dynamin-related GTPases, Dnm1 and Vps1, to the peroxisomes targeted for degradation to facilitate their sequestration by phagophores (Mao, Liu, Feng, & Klionsky, 2014).
Endoplasmic reticulum (ER)–mitochondria contacts are characterized as being critical for cellular physiology, especially mitochondrial biology, such as phospholipid and calcium exchange between mitochondria and the ER (Kornmann & Walter, 2010; Prudent & McBride, 2017; Rowland & Voeltz, 2012). The formation of the contacts is mediated by the ER–mitochondria encounter structure (ERMES), mainly consisting of Mmm1, Mdm10, Mdm12, and Mdm34 (Kornmann et al., 2009). Mmm1 is an integral ER membrane protein, whereas Mdm10 and Mdm34 are mitochondrial outer membrane proteins; Mdm12 is cytosolic. These components form the ERMES, connecting the ER and mitochondria. In mammalian cells, ER–mitochondria contacts have been suggested to be the sites where autophagosomes form, providing important membrane resources for autophagosome biogenesis (Hamasaki et al., 2013). In yeast cells, these contacts are crucial for mitophagy, the process that selectively targets mitochondria for autophagic degradation (Bockler & Westermann, 2014). In contrast, disruption of ERMES does not affect starvation-induced nonselective autophagy.
Here, we found that the ERMES components are required for efficient pexophagy. Atg11 and Atg36 recruit pexophagy-specific fission machinery to peroxisomes adjacent to ER–mitochondria contact sites during pexophagy. Moreover, an R349, 350A mutation of Mdm34 leads to diminished ERMES formation and impaired interaction between Mdm34 and Pex11, resulting in significant defects in pexophagy. Therefore, we propose that ER–mitochondria contacts sites are platforms and hubs for the recruitment of the pexophagy machinery to promote autophagic clearance of peroxisomes.
Results
ER–Mitochondria Contacts Contribute to Pexophagy
As mentioned above, in yeast cells, ER–mitochondria contacts are required for mitophagy (Bockler & Westermann, 2014). Recent studies observed that some peroxisomes are adjacent to ER–mitochondria contacts (Cohen et al., 2014; Mattiazzi Usaj et al., 2015). Accordingly, we asked whether ER–mitochondria contacts play a role in pexophagy. To answer the question, we first examined the localization of peroxisomes relative to ER–mitochondria contacts before and after pexophagy is induced. PEX14, which encodes a peroxisomal membrane protein, was chromosomally tagged with mCherry to visualize peroxisomes. Mdm34-green fluorescent protein (GFP), a mitochondrial component of the ERMES, was used to mark ER–mitochondria contacts; both Mdm34 and Mdm10 are mitochondrial, and in the present study, we used Mdm34 as the representative mitochondrial ERMES component. To induce pexophagy in S. cerevisiae, the cells were first grown in nutrient-rich medium containing glucose (YPD) to mid-log phase, then in medium with glycerol and a minimal level of glucose (SGd), before they were shifted to medium with oleic acid as the sole carbon source (YTO) to promote proliferation of peroxisomes; this specific growth regimen resulted in a large number of peroxisomes, which are otherwise of very low abundance in this organism (Hutchins, Veenhuis, & Klionsky, 1999). The cells were then shifted to conditions of nitrogen starvation in the presence of glucose (SD-N). Under this condition, the elevated population of peroxisomes is no longer necessary, and pexophagy is activated to degrade the surplus organelles.
When cells were grown in YPD, we observed a small number of peroxisomes with approximately 20% of the cells showing colocalization between Pex14-mCherry and Mdm34-GFP (Figure 1(a) and (b)). In YTO medium, there was a substantial increase in the total number of peroxisomes detected with Pex14-mCherry. In contrast, the number of Mdm34-GFP puncta per cell decreased, and a smaller percentage of cells showed colocalization of this marker with Pex14-mCherry compared with the cells growing in YPD (Figure 1(a) and (b)). On pexophagy induction following a 1-h shift to nitrogen starvation conditions, more than 50% of the cells showed peroxisomes adjacent to ERMES (Figure 1(a) and (b)). Similarly, when we looked at the localization of Mmm1-GFP, an ER membrane protein in the ERMES complex, relative to Pex14-mCherry, we observed significant colocalization between these two proteins after pexophagy induction ( Supplementary Figure S1A and S1B ). These observations suggest that ER–mitochondria contact sites very likely play a role in pexophagy.
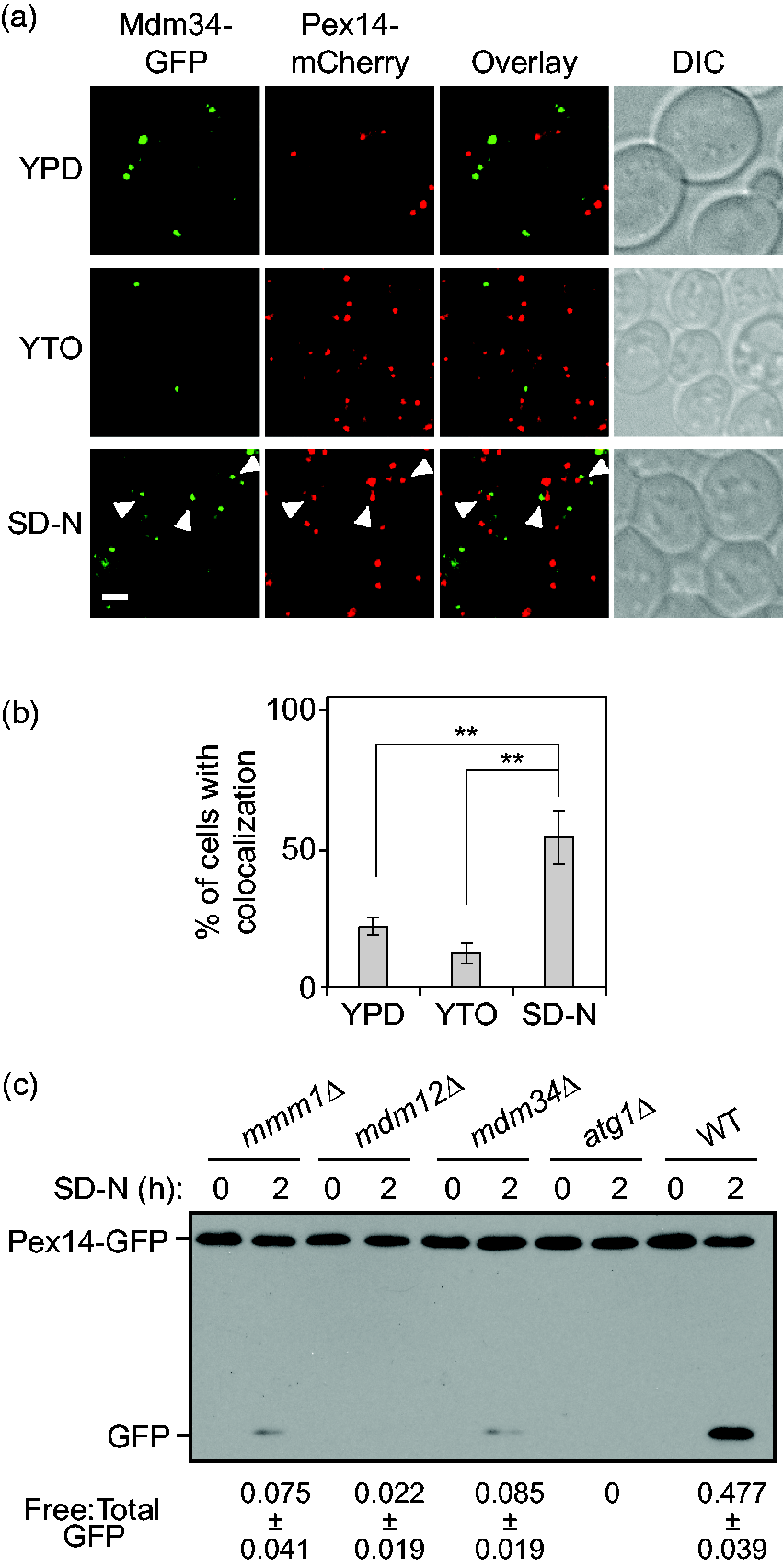
ER–mitochondria contacts contribute to pexophagy. (a) Pex14-mCherry Mdm34-GFP (XLY401) cells were cultured as described in the “Experimental Procedures” section to induce pexophagy. The cells collected from the cultures in YPD, or YTO and the cells starved in SD-N for 1 h were imaged by fluorescence microscopy. Single Z sections of representative images are shown. DIC = differential interference contrast. Scale bar: 2.5 µm. The arrowheads mark colocalizing GFP and mCherry puncta. (b) Quantification of the percentage of cells showing colocalization of mCherry and GFP puncta in panel (a). Error bars represent the standard deviation (SD) of three independent experiments. Two-tailed student’s t test was performed. **p < .01. (c) Pex14-GFP (TKMY67, WT), Pex14-GFP mmm1Δ (XLY403), Pex14-GFP mdm12Δ (XLY404), Pex14-GFP mdm34Δ (XLY405), and Pex14-GFP atg1Δ (TKMY72) cells were cultured as described in the “Experimental Procedures” section to induce pexophagy. The samples collected from cultures in YTO (SD-N 0 h) and 2 h after nitrogen starvation (SD-N 2 h) were trichloroacetic acid precipitated, lysed, subjected to SDS-PAGE, and analyzed by western blot. The ratio of free GFP to total GFP (free GFP+Pex14-GFP) was calculated for the samples. Average values ± SD of n = 3 independent experiments are shown as indicated. GFP = green fluorescent protein; SD-N = synthetic nitrogen starvation medium with dextrose; YPD = yeast extract, peptone, dextrose; YTO = yeast nitrogen base with oleic acid; WT = wild type.
Next, we wanted to disrupt the ER–mitochondria contacts and then determine whether pexophagy was affected. The contacts between ER and mitochondria are mediated by the four ERMES protein components: Mmm1, Mdm10, Mdm12, and Mdm34 (Kornmann et al., 2009). Mmm1 is an ER membrane protein, Mdm12 is a cytosolic component, and Mdm10 and Mdm34 are localized to the mitochondrial outer membrane. Deletion of individual genes encoding ERMES components leads to failure of establishment of ER–mitochondria contacts. To quantitatively monitor pexophagy activity in these mutant cells, we took advantage of the Pex14-GFP processing assay (Reggiori, Monastyrska, Shintani, & Klionsky, 2005). When pexophagy is induced, peroxisomes containing Pex14-GFP are transported into the vacuole for degradation. Pex14 is proteolytically degraded, whereas the GFP moiety is relatively resistant to vacuolar degradation and accumulates in the lumen. Analysis of the amount of free GFP by western blot can therefore be used as an indicator of pexophagy activity. In agreement with a proposed role for the ERMES in pexophagy, we observed a severe reduction in the generation of free GFP from the Pex14-GFP processing assay in the mmm1Δ, mdm12Δ, and mdm34Δ cells, compared with that seen in the wild type (Figure 1(c)). Previous study has indicated that deletion of MMM1, MDM12, or MDM34 does not affect nonselective autophagy (Bockler & Westermann, 2014). Taken together, these results suggest that ER–mitochondria contacts are indispensable for efficient pexophagy.
Transport protein particle complex III is an autophagy-specific guanine nucleotide-exchange factor for Ypt1, a GTPase required for ER-Golgi and Golgi trafficking as well as autophagy (Lynch-Day et al., 2010). Trs85 is the component in transport protein particle complex III that directs Ypt1 to the phagophore assembly site, where autophagosomes form in yeast. Based on the Pex14-GFP processing assay, the trs85Δ mutant displayed severe defects in pexophagy ( Supplementary Figure S1C ). This observation strengthens the point that the ER plays an important role in pexophagy.
Pexophagy Is Defective in the pex11Δ Mutant
A recent study reported that Pex11, an integral peroxisomal membrane protein, associates with Mdm34, based on both a bimolecular fluorescence complementation (BiFC) assay and the membrane yeast two-hybrid assay (Mattiazzi Usaj et al., 2015). This interaction contributes to the contacts between peroxisomes and ER–mitochondria. Decreased colocalization between peroxisomes and ERMES is observed in pex11Δ cells compared with that in wild-type cells when cultured in glucose medium (Mattiazzi Usaj et al., 2015). Therefore, we asked whether localization of peroxisomes in proximity to ERMES is also affected under pexophagy-inducing conditions in the pex11Δ cells. Indeed, we observed a significantly lower percentage of cells showing colocalization of Pex14-mCherry and Mdm34-GFP in the pex11Δ cells after the shift from oleic acid-containing medium to nitrogen starvation conditions (Figure 2(a) and (b)). Pex11 also regulates peroxisome fission (Thoms & Erdmann, 2005); we observed some large clusters of peroxisomes in the pex11Δ cells, but not in the wild-type cells, suggesting defects in peroxisome segregation (Figure 2(a)).

Pexophagy is defective in the pex11Δ mutant. (a) Pex14-mCherry Mdm34-GFP (XLY401, WT) and Pex14-mCherry Mdm34-GFP pex11Δ (XLY402) cells were cultured as described in the “Experimental Procedures” section to induce pexophagy. After nitrogen starvation for 1 h, the cells were collected and imaged by fluorescence microscopy. Single Z sections of representative images are shown. Scale bar: 2.5 µm. The arrowheads mark colocalizing GFP and mCherry puncta. The asterisks mark clusters of peroxisomes. (b) quantification of the percentage of cells showing colocalization of mCherry and GFP puncta in panel (a). Error bars represent the standard deviation (SD) of three independent experiments. Two-tailed student’s t test was performed. **p < .01. (c) Pex14-GFP (TKMY67, WT), Pex14-GFP pex11Δ (XLY406), and Pex14-GFP atg1Δ (TKMY72) cells were cultured as described in the “Experimental Procedures” section to induce pexophagy. The samples collected from cultures in YTO (SD-N 0 h) and 2 h after nitrogen starvation (SD-N 2 h) were trichloroacetic acid precipitated, lysed, subjected to SDS-PAGE, and analyzed by western blot. The ratio of free GFP to total GFP (free GFP+Pex14-GFP) was calculated for the samples. Average values ± SD of n = 3 independent experiments are shown as indicated. GFP = green fluorescent protein; SD-N = synthetic nitrogen starvation medium with dextrose; YTO = yeast nitrogen base with oleic acid; WT = wild type; DIC = differential interference contrast.
Based on the Pex14-GFP processing assay, the pex11Δ mutant displayed significant pexophagy defects (Figure 2(c)). However, because peroxisome fission is required for efficient pexophagy (Mao et al., 2014), we are not able to differentiate between a reduction in pexophagy that was due to defective peroxisome fission or to decreased localization of peroxisomes to ER–mitochondria contacts. To further explore whether localization of peroxisomes to ER–mitochondria contacts is required for pexophagy, we decided to more precisely characterize the residues involved in binding between Pex11 and Mdm34 in order to identify mutants that specifically diminish their interaction without affecting peroxisome fission.
An R349A R350A Double Mutant of Mdm34 Leads to Defects in Pexophagy
Based on the previous study by Mattiazzi Usaj et al. (2015), we tested the interaction between Pex11 and Mdm34 using the BiFC assay. In this assay, the Venus yellow fluorescent protein (vYFP) is split into two fragments, VN (N terminus of vYFP) and VC (C terminus of vYFP), which are tagged to two proteins of interest, respectively (Sung & Huh, 2007). VN and VC are brought into proximity with each other and emit a fluorescent signal if the proteins of interest interact with each other. Consistent with the previous study (Mattiazzi Usaj et al., 2015), we were able to detect a vYFP signal in the VN-PEX11 MDM34-VC cells, but not in the VN-PEX11 MDM12-VC cells, suggesting that Pex11 may specifically associate with Mdm34 (Figure 3(a)).

An R349A R350A double mutant of Mdm34 leads to defects in pexophagy. (a to e) The indicated cells were cultured in YPD to mid-log phase before they were collected and imaged by fluorescence microscopy. Single Z sections of representative images are shown. Scale bars: 2.5 µm. (a) VN-Pex11 Mdm12-VC (XLY407) and VN-Pex11 Mdm34-VC (XLY408). (b) VN-Pex11 Mdm34-VC (XLY408), VN-Pex11[Δ1-50] Mdm34-VC (XLY409), and VN-Pex11[Δ1-100] Mdm34-VC (XLY410). (c) VN-Pex11 Mdm34-VC (XLY408), VN-Pex11 Mdm34[ΔC50]-VC (XLY411), VN-Pex11 Mdm34[ΔC100]-VC (XLY412), and VN-Pex11 Mdm34[ΔC150]-VC (XLY413). (d) VN-Pex11 Mdm34-VC (XLY408, WT), VN-Pex11 Mdm34K347A-VC (XLY417), and VN-Pex11 Mdm34R349,350A-VC (XLY418). The percentage of cells showing a vYFP signal was quantified. Average values ± standard deviation (SD) of n = 3 independent experiments were shown as indicated. Mitochondria are stained with MitoTracker Red. (e) Mdm34-GFP (XLY420, WT), Mdm34K347A-GFP (XLY421), and Mdm34 R349,350A-GFP (XLY422). The percentage of cells showing GFP puncta was quantified. Average values ± SD of n = 3 independent experiments are shown as indicated. (f) Pex14-GFP Mdm34-VN (XLY428, WT), Pex14-GFP Mdm34K347A-VN (XLY429), and Pex14-GFP Mdm34R349,350A-VN (XLY430) cells were cultured as described in the “Experimental Procedures” section to induce pexophagy. The samples collected from cultures in YTO (SD-N 0 h) and 1 h after nitrogen starvation (SD-N 1 h) were trichloroacetic acid precipitated, lysed, subjected to SDS-PAGE, and analyzed by western blot. The ratio of free GFP to total GFP (free GFP+Pex14-GFP) was calculated. Average values ± SD of n = 3 independent experiments are shown as indicated. DIC = differential interference contrast; vYFP = Venus yellow fluorescent protein; GFP = green fluorescent protein; SD-N = synthetic nitrogen starvation medium with dextrose; WT = wild type; VN = N terminus of vYFP; VC = C terminus of vYFP.
There are two predicted transmembrane domains at the C terminus of H. polymorpha Pex11 ( Supplementary Figure S2A ; Williams et al., 2015). Based on the homology between H. polymorpha Pex11 and S. cerevisiae Pex11, we predict that two transmembrane domains are also present at the C terminus of the latter ( Supplementary Figure S2B ). To avoid affecting the localization of Pex11 on peroxisomes, we kept the C terminus intact and mapped the N terminus for regions mediating the interaction with Mdm34. Truncation of the first 100 N-terminal amino acids of Pex11 led to loss of the vYFP signal. This phenotype was not observed when amino acid residues 1 to 50 of Pex11 were deleted, indicating that residues 51 to 100 in Pex11 are required for the Pex11–Mdm34 interaction (Figure 3(b)). Similarly, analysis of the interaction between Pex11 and several truncation mutants of Mdm34 indicated that residues 343 to 359 in Mdm34 were necessary for the interaction (Figure 3(c) and data not shown).
Within the 343 to 359 amino acids region of Mdm34, there are nine positively charged (lysine or arginine) residues. We determined whether mutating these residues to alanine would affect the Pex11–Mdm34 interaction. Compared with the cells with wild-type Mdm34, among the mutants we tested, we found that the R349A R350A double mutation led to a lower percentage of cells showing a vYFP signal by the BiFC analysis, suggesting a potentially diminished Pex11 association with Mdm34R349,350A (Figure 3(d) and Supplementary Figure S2C ). In addition, the R349A R350A mutation of Mdm34 had no apparent major effects on mitochondrial morphology (Figure 3(d)). The other Mdm34 mutants we tested either did not affect the Pex11–Mdm34 interaction or severely disrupted the mitochondrial morphology (Figure 3(d) and Supplementary Figure S2C ). Therefore, we decided to focus on further analysis of the Mdm34R349,350A mutant. Although the Mdm34R347A mutant did not display significant defects in Pex11–Mdm34 interaction (Figure 3(d)), we also included this mutant for further analysis as a negative control.
The absence of any of the individual ERMES proteins affects assembly of the entire complex (Kornmann et al., 2009). Thus, GFP-tagged Mdm34 stains the mitochondria in ERMES mutant strains, instead of being in punctate form as in the wild type (Kornmann et al., 2009). When we compared the localization of WT Mdm34-GFP with the Mdm34R347A-GFP and the Mdm34R349,350A-GFP mutants, we observed a smaller percentage of cells with GFP puncta in the Mdm34R349,350A mutant but not in the Mdm34R347A mutant (Figure 3(e)). This result suggests that the formation of ERMES is partially defective in cells expressing the R349A R350A double-mutant form of Mdm34. Based on the Pex14-GFP processing assay, pexophagy activity was also significantly decreased in the cells expressing the Mdm34R349,350A mutant (Figure 3(f)). In contrast, in the Mdm34R347A mutant where ERMES complex and Pex11–Mdm34 interaction were not significantly affected, pexophagy activity was comparable to that in the wild-type cells. These data collectively suggest that the establishment of a functional ERMES complex and localization of peroxisomes adjacent to the ERMES are required for efficient pexophagy.
A Peroxisome Fission Complex Is Recruited Onto Peroxisomes Adjacent to ER–Mitochondria Contact Sites on Pexophagy Induction
We previously reported that, during pexophagy, the scaffold protein Atg11 and the pexophagy receptor Atg36 physically interact with the two dynamin-related GTPases, Dnm1 and Vps1 (Mao et al., 2014). These interactions facilitate pexophagy-specific fission to promote efficient engulfment of the organelles by phagophores. Utilizing the BiFC assay, we observed that Atg11 and Atg36 recruit Dnm1 and Vps1 to peroxisomes in close proximity to mitochondria (Mao et al., 2014); however, we did not know if there was a functional correlation. With identification of the role of ER–mitochondria contacts in pexophagy, we next decided to ask whether the fission machinery is recruited to peroxisomes adjacent to the contact sites when pexophagy is induced.
We used the recruitment of Vps1 by Atg11 and Atg36 as an example to test this hypothesis. Consistent with our previous study, a significant number of cells showed a vYFP signal in the VN-Atg11 or Atg36-VN cells that were transformed with a plasmid expressing Vps1-VC (Figure 4). These vYFP puncta indicate where Atg11-Vps1 or Atg36-Vps1 interaction occurs in the cells. After pexophagy induction, we observed a substantial increase in the colocalization between the vYFP signal and Mdm34-mCherry (Figure 4), suggesting the peroxisome fission machinery is recruited to ER–mitochondria contacts to facilitate pexophagy. Moreover, compared with the wild-type protein, a substantial decrease in the colocalization between Atg11-Vps1 interacting puncta and the Mdm34R349,350A-mCherry mutant was observed after pexophagy induction ( Supplementary Figure S3 ). Considering our observation that ERMES formation was impaired in the Mdm34R349,350A mutant (Figure 3(e)), these results imply that efficient ERMES complex formation is required for recruitment of the peroxisome fission machinery to the site of pexophagy.

A peroxisome fission complex is recruited onto peroxisomes adjacent to ER–mitochondria contact sites on pexophagy induction. VN-Atg11 Mdm34-mCherry (XLY431; a) or Atg36-VN Mdm34-mCherry (XLY432) cells (b) was transformed with pVps1-VC (416) and cultured as described in the “Experimental Procedures” section to induce pexophagy. The cells were starved for nitrogen for 1 h and imaged by fluorescence microscopy. Single Z-sections of representative images are shown. The arrowheads mark colocalizing BiFC and mCherry puncta. Scale bar: 2.5 µm. (c) Quantification of the percentage of cells showing colocalization of BiFC and mCherry signals in panels (a) and (b). To calculate the percentage, the number of cells showing the colocalization is divided by the total number of cells counted. Error bars represent the standard deviation (SD) of three independent experiments. Two-tailed student’s t test was performed. **p < .01. ***p < .001. DIC = differential interference contrast; SD-N = ■; YTO = ■; VN = N terminus of Venus yellow fluorescent protein; VC = C terminus of Venus yellow fluorescent protein.

A model for the role of ER–mitochondria contact sites in pexophagy. The ERMES is composed of Mmm1 (ER), Mdm12 (cytosolic), Mdm10, and Mdm34 (mitochondria). Pex11 interaction with Mdm34 facilitates peroxisomal localization to the ER–mitochondria contact sites. Pex3 binds Atg36, the pexophagy receptor, which binds the scaffold protein Atg11. The latter two proteins recruit the components Dnm1 and Vps1 to facilitate peroxisome fission. Atg11 promotes binding of Atg36 to Atg8–PE, which is present on the forming phagophore membrane. The interaction between Atg36 and Atg8–phosphatidylethanolamine, which has been shown previously (Motley et al., 2012), allows selective peroxisome sequestration. ER = endoplasmic reticulum.
Discussion
Recent studies have revealed many membrane contact sites between organelles, and they appear to play important roles in cellular physiology (Helle et al., 2013; Phillips & Voeltz, 2016). Along these lines, both ER–mitochondrial contacts and ER–plasma membrane contacts play critical roles in autophagosome biogenesis (Hamasaki et al., 2013; Nascimbeni et al., 2017). It was reported that in mammalian cells STX17-dependent localization of ATG14 onto ER–mitochondria contacts is required for autophagosome formation at the contact sites (Hamasaki et al., 2013). In yeast cells, ER–mitochondrial contacts are specifically required for mitophagosome formation and mitophagy but not starvation-induced nonselective autophagy (Bockler & Westermann, 2014). Here, we found that a substantial number of peroxisomes translocate to ER–mitochondrial contacts on pexophagy induction, and disruption of the contacts severely diminishes pexophagy activity. These findings suggest that ER–mitochondrial contacts are indispensable platforms for the efficient biogenesis of phagophores that target distinct cargoes under different contexts. Aside from the ERMES complex, the ER membrane protein complex proteins also contribute to the contacts between ER and mitochondria to facilitate lipid transfer between the organelles (Lahiri et al., 2014). It is likely that the ER membrane protein complex proteins also play roles in mitophagy and pexophagy.
It was also demonstrated that a pool of autophagosomes forms at ER–plasma membrane contact sites in mammalian cells (Nascimbeni et al., 2017). Extended synaptotagmin proteins, which are involved in tethering between the ER and plasma membrane, recruit VMP1 and the class III phosphatidylinositol 3-kinase complex to promote local phosphatidylinositol-3-phosphate synthesis and autophagosome biogenesis (Nascimbeni et al., 2017). In yeast cells, the tethering machinery between cortical ER and plasma membrane has been characterized (Manford, Stefan, Yuan, Macgurn, & Emr, 2012; Quon et al., 2018). It will be of interest to test whether ER–plasma membrane contacts also contribute to nonselective or selective autophagy pathways in yeast cells.
Pex11, a peroxisome integral membrane protein, is suggested to associate with Mdm34, a component of the ERMES mediating ER–mitochondria contacts, contributing to the localization of peroxisomes adjacent to these sites (Mattiazzi Usaj et al., 2015). Deletion of the PEX11 gene, however, does not result in a block of peroxisomes localizing to ER–mitochondria contacts, suggesting that some other unidentified protein(s) may also be involved in establishing the contacts between peroxisomes and the ERMES. Interestingly, Pex11 was also recently suggested to be involved in mediating the mitochondria–peroxisome contacts (Shai et al., 2018). Thus, the pexophagy defects in the pex11Δ mutant may also partly be due to decreased contacts between these organelles.
Further characterization of the interaction between Pex11 and Mdm34 indicates that residues 51 to 100 in Pex11 and residues 343 to 359 in Mdm34 are required for the Pex11–Mdm34 interaction. The Mdm34R349,350A mutant showed impaired ERMES formation and weaker Pex11–Mdm34 interaction. However, we cannot tell whether this mutation directly diminishes the Pex11–Mdm34 interaction or if the decreased ERMES formation in the mutant indirectly affected the interaction between the two proteins. Nonetheless, the observation of significant defects in pexophagy activity in the Mdm34R349,350A mutant suggests that establishment of ERMES and the localization of peroxisomes in close proximity to ER–mitochondria contacts are required for efficient pexophagy. Decreased ERMES formation may cause partial defects in lipid transfer between the ER and mitochondria, which may affect mitochondrial protein function. Thus, we cannot fully exclude the possibility that defective pexophagy activity in the Mdm34R349,350A mutant may be due to general mitochondrial defects.
Finally, deletion of MMM1 disrupts the ERMES but not peroxisome–mitochondria contacts (Mattiazzi Usaj et al., 2015). Therefore, the pexophagy defects observed in the mmm1Δ mutant suggest that the ER itself plays an important role in this process. Furthermore, the mdm12Δ and mdm34Δ mutants display stronger defects in pexophagy relative to mmm1Δ. Accordingly, we propose that ER–mitochondria contact sites are more critical than peroxisome–mitochondria contacts with regard to pexophagy.
Our previous study demonstrated that Atg11 and Atg36 recruit the peroxisomal fission complex to the peroxisomes targeted for degradation to facilitate the sequestration of the organelles (Mao et al., 2014). Here, we further showed that the fission machinery is recruited to the peroxisomes close to ER–mitochondria contacts. Overall, our results suggest a model where ER–mitochondria contacts are the major sites for recruitment of pexophagy machinery and for the formation of pexophagy-specific autophagosomes (Figure 5).
Experimental Procedures
Yeast Strains, Media, and Growth Conditions
Yeast strains used in this study are listed in Supplementary Table S1 . For nutrient-rich conditions, yeast cells were either grown in YPD medium (1% yeast extract, 2% peptone, and 2% glucose) or synthetic minimal medium with dextrose (SMD; 0.67% yeast nitrogen base, 2% glucose, and auxotrophic amino acids and vitamins as needed). To induce peroxisomal proliferation, cells were first grown in YPD or SMD-Ura to approximately 0.5 OD600 before they were shifted to glycerol medium (SGd; 0.67% yeast nitrogen base, 0.1% glucose, and 3% glycerol) for 16 h. Then, yeast extract and peptone were added into the cultures in SGd medium, and the cells were incubated for 4 h. The cells were then shifted to oleic acid medium (YTO; 0.67% yeast nitrogen base, 0.1% oleic acid, 0.1% Tween 40, and auxotrophic amino acids as needed) for 20 h. Pexophagy was induced by shifting the cells to nitrogen starvation medium containing glucose (SD-N; 0.17% yeast nitrogen base without ammonium sulfate or amino acids, and 2% glucose).
Plasmids
The plasmid pVps1-VC (416) was described previously (Mao et al., 2014).
Fluorescence Microscopy
For fluorescence microscopy, yeast cells were grown as described above to induce pexophagy. Samples were collected and then examined by microscopy (Delta Vision, Applied Precision) using a 100× objective, and pictures were captured with a CCD camera (CoolSnap HQ; Photometrics). Mitochondria were stained with MitoTracker Red CMXRos (Molecular Probes/Invitrogen, M7512).
Western Blot
The western blot was performed as described previously (Mao et al., 2014). Antibody to YFP (Clontech, 632381) was used to detect GFP-tagged proteins.
Supplemental Material
Supplemental material for Endoplasmic Reticulum–Mitochondria Contacts Are Required for Pexophagy in Saccharomyces cerevisiae
Supplemental Material for Endoplasmic Reticulum–Mitochondria Contacts Are Required for Pexophagy in Saccharomyces cerevisiae by Xu Liu, Xin Wen and Daniel J. Klionsky in Contact
Footnotes
Supplemental Material
Supplemental material for this article is available online.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by National Institute of General Medical Sciences grant GM053396.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
