Abstract
Current high-throughput screening assay optimization is often a manual and time-consuming process, even when utilizing design-of-experiment approaches. A cross-platform, Cloud-based Bayesian optimization-based algorithm was developed as part of the National Center for Advancing Translational Sciences (NCATS) ASPIRE (A Specialized Platform for Innovative Research Exploration) Initiative to accelerate preclinical drug discovery. A cell-free assay for papain enzymatic activity was used as proof of concept for biological assay development and system operationalization. Compared with a brute-force approach that sequentially tested all 294 assay conditions to find the global optimum, the Bayesian optimization algorithm could find suitable conditions for optimal assay performance by testing 21 assay conditions on average, with up to 20 conditions being tested simultaneously, as confirmed by repeated simulation. The algorithm could achieve a sevenfold reduction in costs for lab supplies and high-throughput experimentation runtime, all while being controlled from a remote site through a secure connection. Based on this proof of concept, this technology is expected to be applied to more complex biological assays and automated chemistry reaction screening at NCATS, and should be transferable to other institutions.
Graphical Abstract
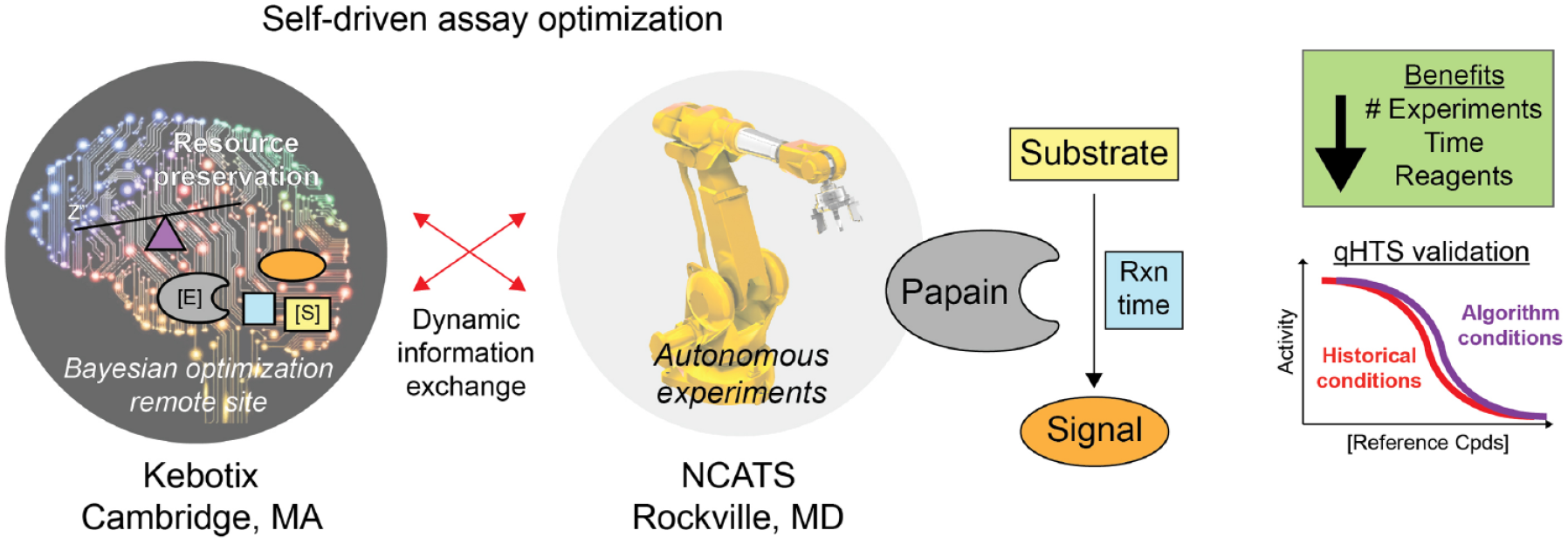
Introduction
There is great interest in applying artificial intelligence (AI) and machine learning (ML) to various phases of preclinical drug discovery. 1 This includes chemistry reaction screening, hit selection, and even molecular design.2,3 Newer Bayesian approaches—including those that leverage high-throughput experimentation and robotics—have recently been applied to chemical reaction screening with significant improvements in optimization efficiency.4,5 It is anticipated that AI/ML approaches can enhance the efficiency of solving many of the multifactorial problems encountered in drug discovery. This includes biological assay development, a process in which it can take experienced biologists months to even years to develop and validate a robust, pathophysiological relevant assay. 6
One reason for the length of assay optimization is that scientists must optimize multiple and often competing factors, with many of the variable interactions difficult to predict a priori. This includes the concentration of key reagents (substrate, enzymes, cofactors), temperature, buffer/media composition, timing, and other factors. Assay optimization can utilize several strategies, including brute force, design of experiment (DOE), or approaches informed from historical experience or fundamental principles. Brute-force approaches, which test as many variable permutations as possible, and DOE approaches, which apply statistics to systematically determine the relationship between a set of variables affecting an output, have been used in high-throughput screening (HTS) assay optimization.7–10 Both suffer from a lack of feedback and adaptivity. This means that they each specify the entire set of experiments to be conducted a priori, which means they waste resources on low-performing experiments rather than concentrating attention on high-performing experiments. Additionally, this means they each scale poorly with higher dimensions, as the number of experiments necessary increases exponentially. The cost of performing all these experiments can eventually become infeasible due to costs from reagents, time, and automated system modifications. While valuable, relying solely on historical and fundamental principles has the potential for bias, and such preconceptions about how best to optimize an assay may not be best applicable to novel biological systems.
The National Center for Advancing Translational Sciences (NCATS) ASPIRE (A Specialized Platform for Innovative Research Exploration) Initiative seeks to combine advances in automated chemistry, high-throughput biological annotation, and AI/ML to accelerate preclinical drug discovery.11–13 Another important design element of the ASPIRE Initiative is to facilitate close, real-time collaboration between NCATS and our decentralized network of intramural (National Institutes of Health [NIH]) and extramural partners. To this end, part of this collaboration component is expected to involve off-site access and control of NCATS laboratory systems. With respect to off-site laboratory control, the COVID-19 pandemic and subsequent public health measures like physical distancing have demonstrated the limitations of relying on scientists to be physically present in laboratories. To assess the feasibility and utility of integrating AI/ML methods in our preclinical drug and probe discovery center, we developed an autonomous Bayesian assay optimization system. Using a facile cell-free fluorometric assay for the protease papain, we demonstrate that such an autonomous, remotely controlled system can provide an efficient, robust set of assay conditions capable of identifying bioactive small molecules with similar performance to conventional approaches.
Materials and Methods
Papain Cell-Free Enzymatic Assay
Screening was performed at the NCATS screening facility.
14
Compounds were prepared and stored as 10 mM DMSO stock solutions. To test for compound inhibition of the protease papain, a cell-free papain enzymatic activity assay was adapted from published protocols.15,16 A detailed standardized protocol is provided for both the Bayesian optimization experiments and the historical experiments (
Table 1
).
17
A general description for both experimental protocols is as follows: 3 µL of 1.33X papain in assay buffer (100 mM sodium acetate, pH 5.5, 5 mM
Standardized Cell-Free Papain Assay Protocols for Bayesian Optimization and Historical Condition Experiments a .
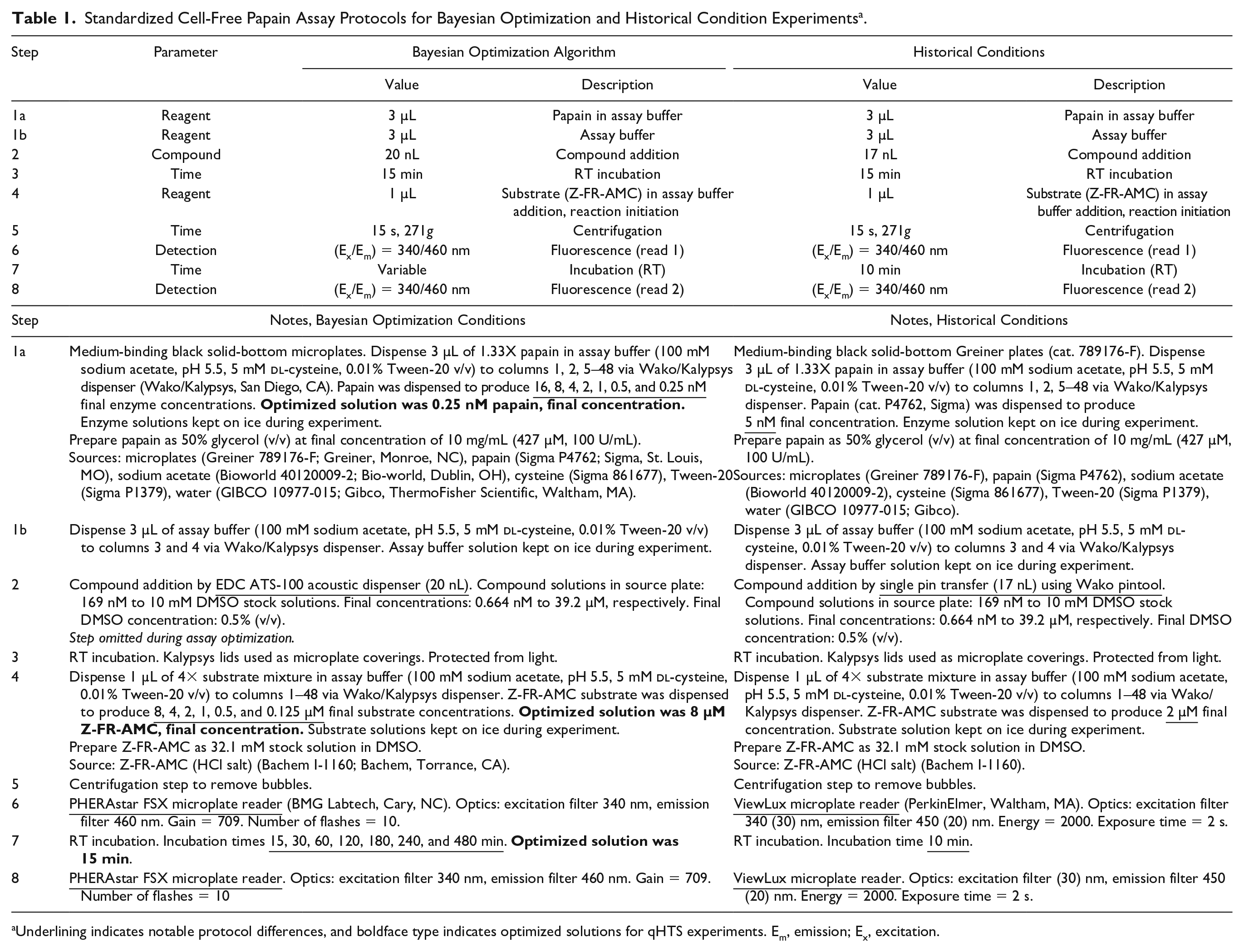
Underlining indicates notable protocol differences, and boldface type indicates optimized solutions for qHTS experiments. Em, emission; Ex, excitation.
Three multihead liquid dispensers were configured to facilitate autonomous and continuous experiments, including a microplate washing step for microplate reuse and minimization of plastic consumption ( Suppl. Table S1 ). Three critical assay design parameters, based on the NCATS Assay Guidance Manual, were selected for optimization: reaction time, substrate concentration, and enzyme concentration.18,19 For this study, one experimental condition was tested per microplate.
Computation
Kebotix’s Bayesian optimization software was adapted and deployed to provide suggested experimental conditions for the papain biochemical assay. Given the asynchronous quasi-batching of results, the following protocol was used: if there were any new results since the last suggestion made, and at least five results collected overall, a suggested assay protocol would be made using a sequential version of the Kebotix Bayesian optimization algorithm. Otherwise, such a suggestion would be generated randomly from the untested parameter choices. An assay optimization score was then calculated based on maximizing desirable features: sufficiently high Z′, reduction in reagent use, and shorter protocol time (eq 1).
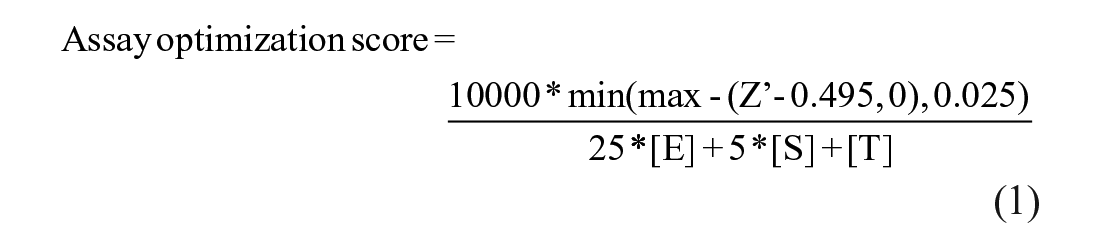
[E] represents the nanomolar concentration of enzyme, [S] represents the micromolar concentration of substrate, [T] represents the incubation time in minutes, and Z′ is a calculated metric of assay quality using the positive and neutral plate controls. 20 The denominator of the score represents a notion of the “cost” of the experiment, where smaller costs are preferred, and the coefficients reflect the relative material, absolute concentrations, and time costs of the different components (e.g., the enzyme was roughly 5000-fold more expensive than the substrate). A similar heuristic trade-off was made between these material costs and the incubation time based on the experimenters’ judgment. Individual coefficients for enzyme, time, and substrate were chosen (rather than ratios) to better account for total “cost” in situations where ratios might be identical but total reagents are different. The two thresholds of 0 and 0.025 applied to the quantity Z′ − 0.495 reflect the judgment that Z′ values below 0.495 are all equally undesirable, Z′ values above 0.52 (0.495 + 0.025) are all equally desirable, and those between 0.495 and 0.52 are partially desirable. Finally, the multiplicative factor of 10,000 simply translated the score into the low single digits for ease of human understanding (with no effect on the optimization algorithm).
Given the results from previous assays, the Kebotix Bayesian optimization builds a Gaussian process model to predict the scores of untested assay conditions, with uncertainty. It then identifies the conditions that maximize a linear combination of the predicted score and the uncertainty in that prediction. This amounts to an “optimistic estimate” (upper confidence bound) on the score, and it is meant to balance the goals of exploitation and exploration. In addition, given the functional form of the assay optimization score above, the algorithm calculates the maximum possible score for each set of conditions, which would be achieved if Z′ > 0.52, and restricts its search to conditions whose maximum possible score is higher than the current maximum, performing constrained optimization over this narrower space of cheaper experiments. This ensures that the upper confidence bounds are potentially attainable; in other words, no experiments are suggested by the optimizer that, even if they successfully resulted in a large value of Z′, could not possibly improve on the current optimum. Kebotix has plans to release a commercial system under the ChemOS trademark in the near future, likely with accompanying academic discounts. At present, public Bayesian optimization packages can provide some support to problems such as biological assay optimization.
Four variations in the experimental optimization runs were performed in which all experiments within a run were started every 10 or 20 min, and a logarithmic transform was applied before passing the options (concentrations and durations) to the optimization algorithm. The two timings were meant to illustrate performance given different trade-offs between the number of experiments and overall time spent: more frequent experiments result in fewer feedback cycles and more experiments necessary to optimize; less frequent experiments lead to more time spent on the optimization. A logarithmic transform was also evaluated for its effect on the optimization algorithm: the options for both concentrations and time were roughly exponentially scaled (i.e., 15, 30, 60, 120 min), and we envisioned that equally spacing these options would make for a smoother and therefore easier-to-optimize objective function.
Brute Force and Live Optimization Experimental Parameters
A total of 512 assay optimization scores were experimentally generated by randomly sampling each permutation (“brute force”) with the following experimental conditions: final enzyme concentration (0, 0.25, 0.5, 1, 2, 4, 8, and 16 nM), final substrate concentration (0, 0.125, 0.25, 0.50, 1, 2, 4, and 8 µM), and incubation time (5, 15, 30, 60, 120, 180, 240, and 480 min). Brute-force microplates with zero-value conditions (0 nM enzyme, 0 µM substrate, 5 min of incubation) were used as experimental controls but were excluded from subsequent analyses. Microplates utilizing 0.250 µM substrate concentrations were excluded due to technical issues. Removing these options leaves seven allowable final enzyme concentrations (0.25, 0.5, 1, 2, 4, 8, and 16 nM), six allowable final substrate concentrations (0.125, 0.5, 1, 2, 4, and 8μM), and seven allowable incubation times (15, 30, 60, 120, 180, 240, and 480 min), for a total of 7*6*7 = 294 total experimental conditions. This formed the space of possibilities for both simulations and the live Bayesian optimization experiments.
Data Simulations
Simulations were performed using the brute-force data collected to identify typical behavior of variations on the optimization algorithm. In these simulations, the timings associated with the experimental process were mimicked, with the results of each suggestion being drawn from the brute-force data upon completion. The following 10 variations were considered: new experiments started every 10, 20, 30, or 60 min or sequentially, and with and without the logarithmic transform applied to the options. Two scenarios were considered. In the first (complete simulations), the optimization would proceed until the optimum conditions were suggested (or completion), immediately stopping, and approaches would be assessed by how many experiments they required to find the optimum. In the second (budget simulations), the optimization would proceed for the designated budget of either 40 or 20 microplates and then stop, and variations would be assessed according to whether the optimum was discovered. In each of these scenarios, and for each of the 10 variations considered, 1000 simulations were run. Results were then compared with the expected performance of a randomly ordered brute-force run. Simulations were programmed in Python 3.8.
QHTS Data Analysis and Statistics
Reference papain inhibitors ( Suppl. Table S2 ) were tested in 11-point serial threefold titration (0.664 nM to 39.2 µM final concentrations). The Library of Pharmacologically Active Compounds (LOPAC1280) was tested in quantitative HTS (qHTS) format in 7-point serial fourfold titration (31 nM to 50 µM final concentrations). 21 Data from each assay were normalized to intraplate controls (neutral control, DMSO, microplate columns 1 and 2; positive control, enzyme-free, microplate columns 3 and 4). The aforementioned intraplate controls were used for the calculation of the Z′ factor for each microplate. The fluorescence intensity difference between read 2 (postreaction) and read 1 (shortly after substrate addition) was used to compute reaction progress. Concentration–response curves (CRCs) were fitted and classified as described previously, with active compounds producing curve classes of −1.1, −1.2, −2.1, or −2.2. 21 IC50 values were calculated and plotted using GraphPad Prism (version 9.1.0, La Jolla, CA) and the sigmoidal dose–response (variable slope) equation. Data were curated using the Palantir Technologies (Washington, DC) data integration Foundry platform (NIH Integrated Data Analysis Platform [NIDAP]), which is configured to ingest all HTS results generated at NCATS and harmonized these data with other sources, such as ChEMBL and OrthoMCL. All qHTS screening results are publicly available at PubChem (AIDs 1645873 and 1645872). The chemical structures were standardized using the LyChI (Layered Chemical Identifier) program (version 20141028, github.com/ncats/lychi). P values were one-sided and used a Bonferroni correction.
Results
Assay Overview
A cell-free assay for papain protease activity was chosen for proof of concept. This assay was ideal because its low cost and simplicity make it suitable for many experiments (brute force), and previous NCATS experience would allow for historical comparisons.15,16 In this fluorescence intensity assay, papain enzymatically cleaves the quenched AMC fluorophore from a dipeptide substrate to generate nonquenched free AMC fluorophore whose fluorescence intensity is proportional to papain activity in the absence of compound-mediated interferences ( Fig. 1 and Table 1 ). Historically, the assay utilized low nanomolar enzyme concentrations (5 nM), low micromolar concentrations of substrate (2 µM), and short incubation times (10 min). The assay was incorporated in the NCATS HTS facility, which allowed for robotic liquid and compound dispensing, microplate handling and fluorescence reading, and microplate washing and reuse.
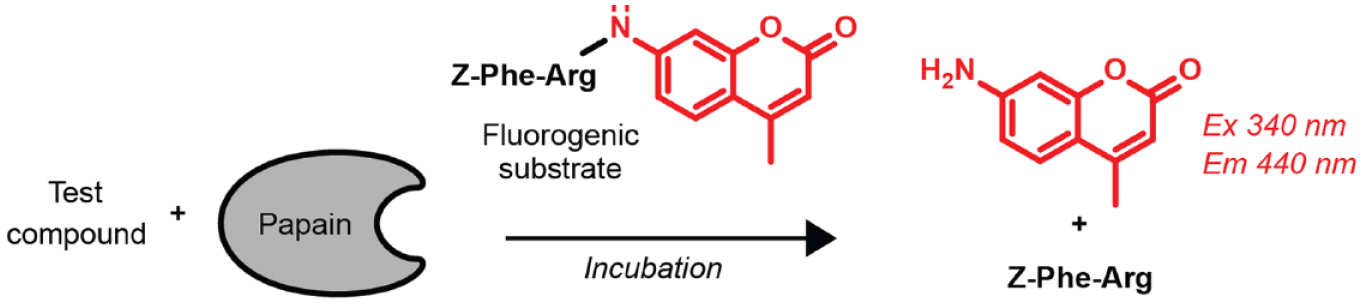
Papain assay schematic. Papain hydrolyzes the AMC fluorophore from the dipeptide substrate, which leads to an increase in blue fluorescence. Inhibitors of papain activity are expected to decrease the fluorescence intensity readout. See also Table 1 for standardized protocols of Bayesian algorithm-derived and historical protocols.
System Communication
The NCATS HTS system composed of an integrated, automated robotic platform is based on a dynamic and asynchronous scheduling methodology in which assay plates act as the input to the system, and each plate may have associated control and compound plates as required. To execute an assay on a microplate, the system scheduler relies on wait queues and mutexes (“mutual exclusion”; locks to enable resource availability-based synchronization). At the start of an experimental run (which can contain multiple microplates running in parallel), all microplates are placed in a first-in, first-out (FIFO) wait queue. A wait queue is a queue of microplates waiting for a lock, which in this case represents control of a peripheral device on the screening system. For each peripheral device, when its associated lock is unlocked, the microplates acquire the lock in the order of the queue. Microplates still in the queue waiting to use a locked peripheral device are in a blocked state until the lock is released. Each step, or a series of linked steps, in a method has an associated mutex.
This HTS system can monitor all events occurring on the platform for each microplate and every associated method step, can allow multiple processes to run in parallel, and can launch new processes directed from external applications. To incorporate AI/ML-driven experimentation from extramural collaborators (i.e., collaborating organization external to the NCATS HTS facility), a messaging technique was required such that an external informatics platform could initiate experiments to be performed on the NCATS HTS system, with the resultant data then sent back for analysis to the initiating platform. To enable this messaging, a RabbitMQ server was deployed at NCATS and was made public facing to allow external messaging. Several steps were implemented to ensure data security, including the deployment of the server within the NIH firewall demilitarized zone (DMZ), the utilization of Hypertext Transfer Protocol Secure (HTTPS) communication, Transport Layer Security (TLS) protocol version 1.2, and other technical controls.
Advanced Messaging Queuing Protocol (AMQP) was used to provide a platform-agnostic method for ensuring information is safely transported between applications, among organizations, within mobile infrastructures, and across the Cloud. Having an open standard was important to effectively communicate with a variety of different platforms, including those that might wish to remotely initiate experiments on the NCATS platform. The remote access feature was incorporated primarily to accommodate NCATS’s decentralized network of collaborators. This AMQP approach is based on Command Query Responsibility Segregation (CQRS), which is a pattern that segregates the operations that read data (queries) from the operations that update data (commands) by using separate interfaces. This means that the data models used for querying and updates are different (SFTP, REST API, etc.). The screening platform can be considered the “query” portion by reading the results of an experiment, while Kebotix is the “command” portion, issuing new experiments to be performed ( Fig. 2 ). In the NCATS system, this informatics platform produces a message that routes through the exchange to be placed within the experimental queue at the HTS site. This message initiates an experiment and consists of a unique identifier for experiment tracking and, in this specific report, details on the desired enzyme concentration, substrate concentration, and incubation time. Using LabView, 22 the HTS system consumes this message from an experimental queue and generates a method specific to it based on the experimental conditions requested to be programmatically launched as an assay on the platform. During the experiment, any plate read steps that generate data trigger the HTS system to generate a new message (containing the unique identifier and resultant data) that is added to the results queue. Upon completion of an experiment by a microplate, another message is produced by the HTS system and sent to the results queue to let the informatics platform know that an experiment with a given microplate is done. These messages in the results queue are then consumed by the informatics platform such that data generated by the experiments run on the HTS system can be analyzed and any respective models used to generate new experiments can be updated. The informatics platform could then programmatically initiate new experiments using new microplates by producing another message to be sent to the experiment queue. This process would continue until some stop criterion for the entire run is met.
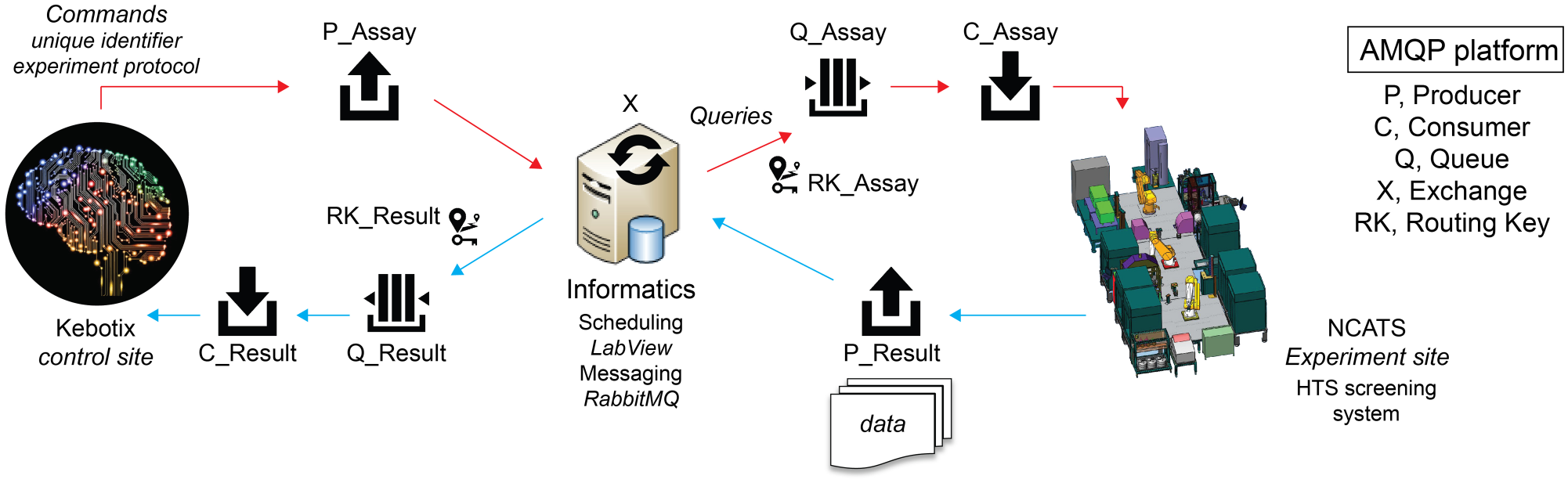
Design of autonomous assay optimization platform. The system utilizes AMQP to facilitate communication between the NCATS HTS system, a shared informatics resource, and the Kebotix extramural Bayesian optimization command platform. Producer (P), an application that sends messages; Queue (Q), a buffer that stores messages; Consumer (C), an application that receives messages; Exchange (X), a router that receives messages from P and pushes them to C; Routing Key (RK), a relationship that specifies that a Q is interested in messages from an X.
The goal of sending commands from remote locations is to initiate experiments on demand, which is possible since AMQP is event-driven, meaning commands issued by Kebotix initiate a new experiment, assuming there are resources available to perform the request. An asynchronous operation feature was incorporated in which synchronization is based on resource availability, not time. This means that multiple commands can be issued and multiple queries can be in process simultaneously, as the duration of a particular experiment or time to process the results and make new predictions will be variable. Overall, this is a flexible, scalable communications method to enable event-driven, asynchronous execution of assays from potentially multiple system controllers (in this study, Kebotix only initiated the experiments). The asynchronous feature allows controllers to run multiple microplates or completely independent experiments in parallel if resources allow.
Brute-Force Biological Assay Optimization
To help assess the effectiveness of the Bayesian assay optimization approach, the assay optimization parameters were tested sequentially using a brute-force approach. Based on the experimental workflow and automation capacity, successive microplates required a minimum time interval of 10 min ( Fig. 3A ). The system capacity was 20 microplates available for performing experiments at a given moment. This capacity was chosen to balance throughput and resource availability (i.e., too many microplates would overwhelm the robotic and plate washing system) and limit microplate consumption. Brute force was chosen as a comparator because it can be used for high-throughput optimization in automated HTS laboratories. Notably, this was a completely autonomous process that required no human intervention and allowed for multiple other independent projects to be run on the same testing platform throughout the duration of the experiment.
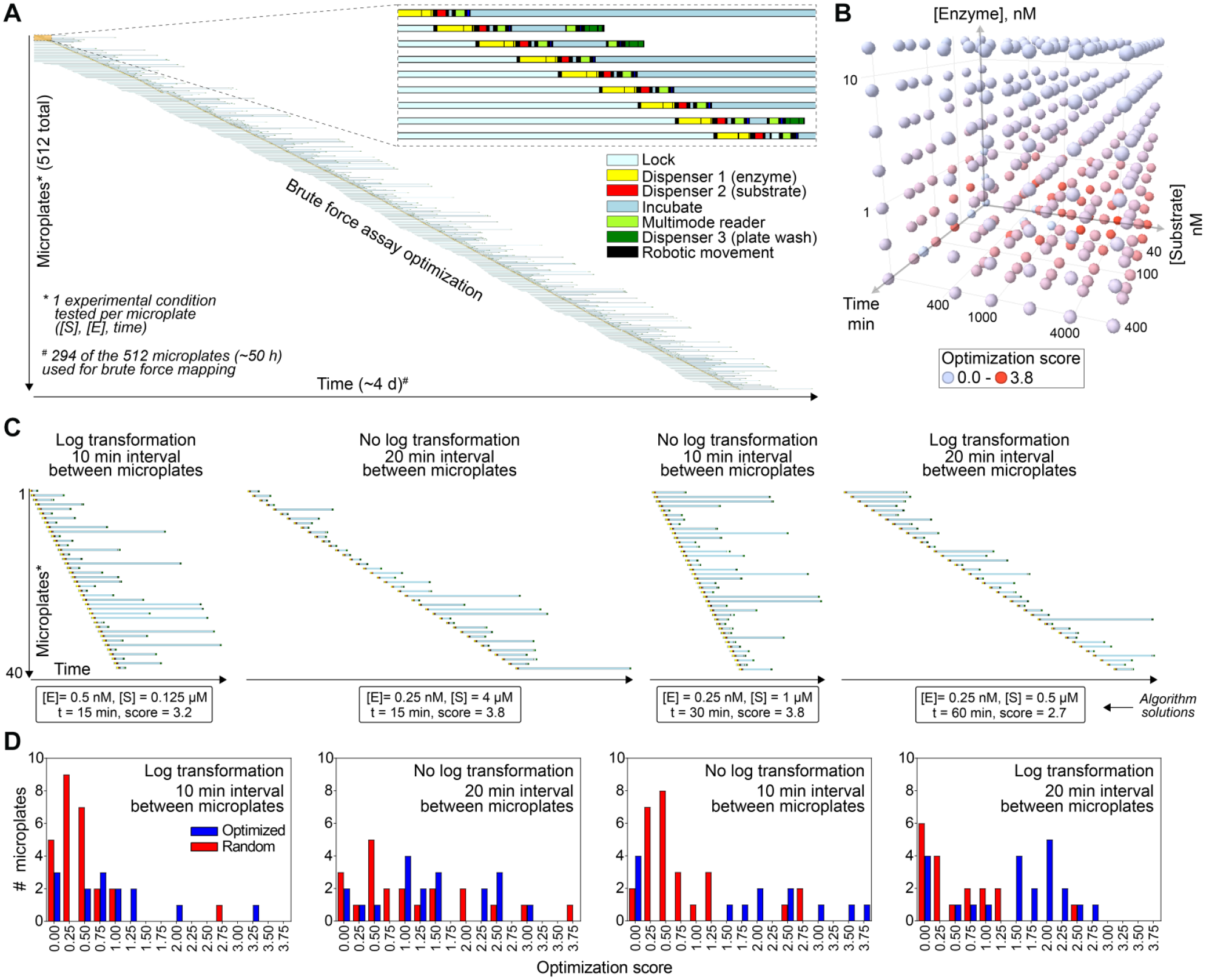
Optimization of papain biochemical assay by brute force and Bayesian optimization approaches. (
After excluding experimental controls (no enzyme, substrate, or minimal incubation time) and conditions that failed due to dispensing failures (see Methods), testing each permutation required 294 individual microplates (approximately 50 h of runtime) ( Fig. 3A ). Each microplate tested one unique experimental condition, and the entire process which included all experimental controls and technical failures used approximately 4 days of automated experimentation time. The assay optimization score (eq 1) was evaluated at all experiments. The best experimental conditions were variable substrate concentrations (low micromoles), low nanomolar enzyme concentrations (0.25 to 0.5 nM), and short incubation times (15 to 30 min) ( Fig. 3B ). Notably, these conditions are in good agreement with the historical testing conditions ( Table 1 ).15,16
Bayesian Assay Optimization
The Kebotix Bayesian optimization algorithm was used to determine the optimal assay conditions while blinded to the brute-force results. To facilitate future improvements in the algorithm for assay optimization, the algorithm was performed in four independent runs, each with a unique combination of parameter transformations (log or nonlog) and interval time between microplates (10 or 20 min). The time interval was chosen based on the experimental workflow and automation capacity. The actual computation requires only milliseconds and was not a rate-limiting step in this experimental setup. Each independent run was allowed a maximum of 40 unique microplates to test. Again, each microplate assessed one unique experimental condition. Not surprisingly, the runs with increased time intervals between microplates required more time compared with the runs with shorter intervals between microplates ( Fig. 3C ). Each run arrived at a different optimal experimental condition ( Fig. 3C ). For the enzyme concentration, each run suggested a final concentration of either 0.25 or 0.5 nM papain. There was more variability in suggested final substrate concentrations and incubation times, ranging from 0.125 to 4 µM peptide and from 15 to 60 min, respectively. Interestingly, the two suggestions with the highest optimization scores (3.8) had opposing balances of substrate and incubation times (4 µM substrate and 15 min incubation vs 1 µM substrate and 60 min incubation).
Live optimization runs illustrate the performance of the optimization algorithm in action. While the true optimum is unknown without all suggestions having been attempted, a simple comparison of the performance of the randomly generated suggestions and those generated by the Bayesian optimization can illustrate the performance of the optimizer ( Fig. 3D ). Across the four runs, the median optimized suggestion achieved an assay optimization score of 1.4, while random suggestions achieved optimization scores above 1.4 in only 12 of 92 (13%) attempts, fourfold less frequently. Even accounting for the arbitrariness of this distribution-based threshold, this difference is significant (p = 3 × 10−6). This demonstrates that the optimizer can enrich for high-quality experiments and provide biologists with more viable choices for selecting final conditions.
Simulations
Ground-truth data generated by the brute-force approach enabled further evaluation of the overall platform by simulation. Individual components of the adaptive learning process itself, like logarithmic data transformation and the timing of experimental queues, could also be evaluated by simulation. Two types of simulations were performed: complete simulations and budget simulations. Complete simulations assessed the number of microplates needed to identify the optimal assay condition (allowing up to the full 294 possible microplates used for the brute-force experiments), while budget simulations assessed how often a run could identify the optimal assay condition with 40 or less microplates.
In complete simulations, variations of the optimizer both with and without the logarithmic transform found the optimum in only 11 to 21 microplates on average, which is a 7- to 13-fold improvement on randomly ordered brute-force simulations, with expected variation according to the frequency of experiments ( Fig. 4A ). For reference, brute force finds the optimal conditions on average at the midway point of a complete simulation (after 147.5 microplates out of a possible 294), and finds the optimum in 40 of 294 (13.6%) budget simulations. This illustrates the trade-off between time and experiment costs: running more frequent experiments (shorter intervals between consecutive microplates) saves time, while running less frequent experiments requires fewer to find the optimum. The improvement using log transform was minimal (mean difference less than 1 microplate), indicating the transform was not important for the algorithm to find the optimum.
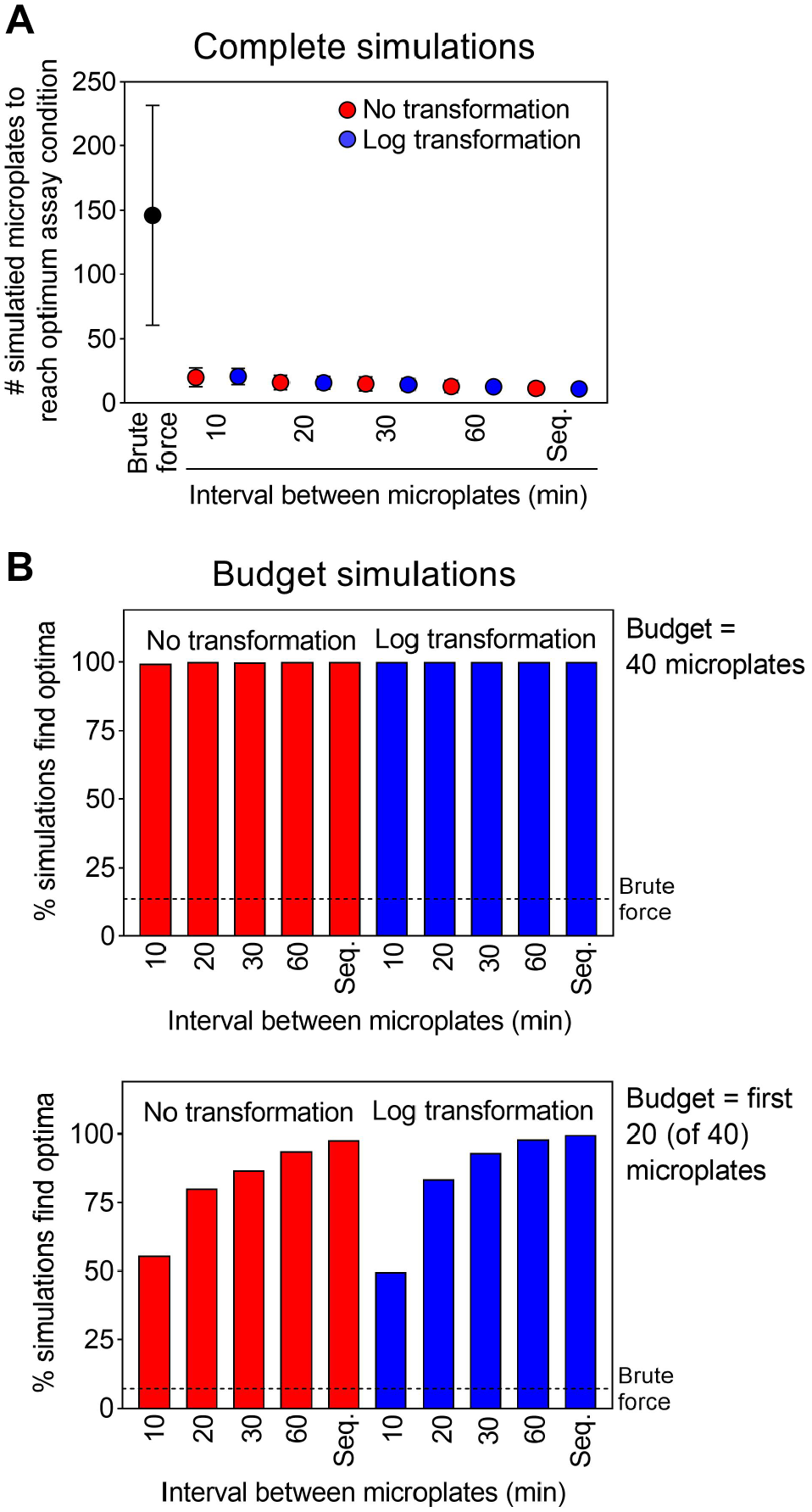
Evaluation of adaptive learning and optimization processes using brute-force-derived simulation data. Simulations compared adaptive learning to brute force, and the effects of time delays between subsequent microplates for nontransformed and log-transformed data. (
In budget-based simulations where each run was allotted up to 40 microplates, the optimization algorithm was able to identify the optimum at least 99.4% of the time in all variations, sevenfold more frequently than the brute-force approach with p values less than 10−846 ( Fig. 4B ). When simulating a more stringent budget (20 microplates), all variations were able to identify the optimum at least 49.7% of the time, and as frequently as 99.8% for sequential microplates applying the logarithmic transform, offering a similar 7- to 15-fold improvement on brute force ( Fig. 4B ). In both simulation formats, extending the time between microplates beyond 10 min decreased the number of microplates needed to find the optimum ( Fig. 4 ). These simulations demonstrate the efficiencies of adaptive learning for assay optimization and should help in future experiments with this platform.
Validation of Optimized Conditions with Reference Small Molecules and Pilot qHTS
Select compounds were tested for inhibition of papain enzymatic activity to further assess the robustness and applicability of the algorithm-derived solutions. At this stage, an additional compound transfer step was added to the assay protocol, and CRCs were automatically analyzed by an in-house informatics platform. A series of nearly two dozen reference papain inhibitors spanning nanomolar to micromolar potencies were selected based on in-house data from a papain counterscreen used during a qHTS campaign targeting the protease cruzain.15,16 These reference compounds were tested using the historical counterscreen conditions ( Table 1 ) and the optimal conditions determined from the algorithm. At this point, an independent run of the Bayesian optimization system was performed using 10 min time intervals between successive microplates, with logarithmic transform, and with the previous 20 microplate capacity.
While there was some systematic bias observed (Bland–Altman bias = 0.3 ± 0.2), there was gross concordance for the IC50 values determined using the historical conditions and those identified by the algorithm (r2 = 0.93) ( Fig. 5A ). Notably, there were some differences in CRC shape between the two assay conditions, which could be due to differences in reagent concentrations or other technical factors such as workflow timing or variations in liquid dispensing ( Fig. 5B ).
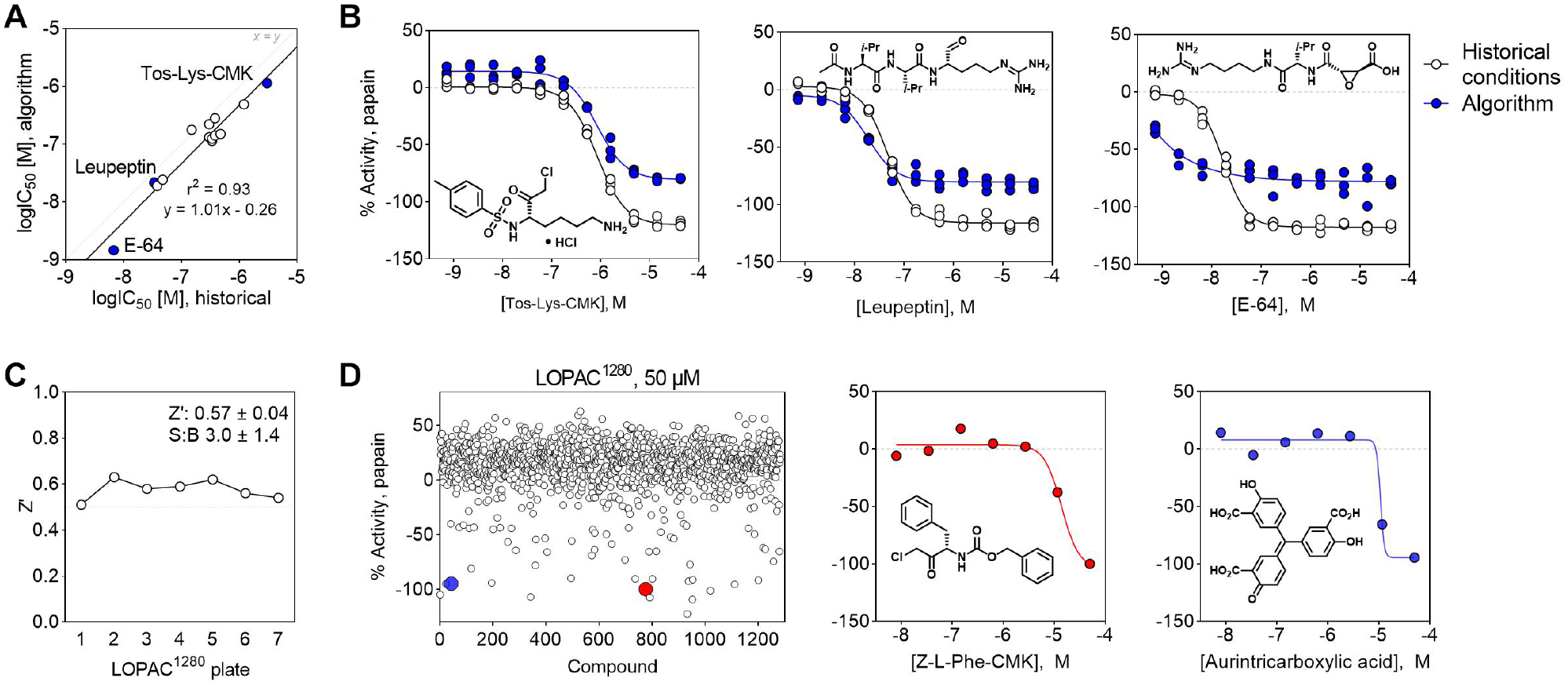
Validation of the algorithm-optimized experimental conditions for a cell-free papain inhibition assay by small molecules. (
The applicability of algorithm-optimized assay conditions was also assessed by a pilot qHTS using the LOPAC1280. The Kebotix conditions showed acceptable assay quality metrics using the LOPAC1280 in qHTS format (mean Z′ = 0.57 ± 0.04) (
Fig. 5C
). The pilot qHTS resulted in 54 compounds (4.2%) exhibiting papain inhibition based on curve class. This autonomously optimized method could identify putative small-molecule inhibitors of papain enzymatic activity using the LOPAC1280 in qHTS format, including a likely covalent inhibitor Z-
Discussion
We developed a cross-platform Bayesian optimization system and applied it to autonomous biological assay optimization. As proof of concept, the system was able to efficiently optimize a papain biochemical enzymatic assay. The optimized assay conditions were then validated by testing a collection of reference papain inhibitors and a pilot qHTS library. These assay conditions provided practical solutions, as testing with the AI/ML-derived assay conditions recapitulated the performance of inhibition data from historical protocols, and could identify a series of inhibitions among the LOPAC1280.
Several notable features can be achieved by utilization of the developed platform. First, the proposed approach can function continuously without human supervision once reagents are prepared and system checks are performed. Second, by applying the asynchronous, event-driven method, the screening facility was able to perform other tasks on unrelated projects at the same time, including other qHTS campaigns. Third, the unique messaging system implemented in the proposed platform allows us to run the physical experiments in one place (Rockville, MD) while being controlled by an off-site location (Cambridge, MA) through a secure electronic connection. This last feature may be especially beneficial in situations with limited on-site availability (e.g., pandemic-related restrictions).
There are several advantages of this Bayesian optimization methodology relative to traditional and brute-force approaches. These algorithmic approaches can be more efficient than traditional high-throughput assay optimization approaches such as brute force, resulting in cost savings. This is especially important for expensive reagents where the number of optimization experiments is cost-prohibitive and for precious reagents such as difficult-to-expand or rare cell lines. Even for cheap experiments, they also do not require the involvement of the researcher in the optimization run itself, which provides additional savings when paired, as in this example, with an autonomous experimentation platform. Finally, while not demonstrated in this work, they offer a better scaling than traditional methods at high dimensions, which also present additional challenges for humans to analyze.
The Bayesian optimization approach also had some notable limitations, though many of these limitations can likely be overcome in future iterations. We can distinguish two types of limitations: those that were necessary for achieving the full comparison to demonstrate performance of the Bayesian optimization approach relative to brute force, and those which would apply even without the need to run the brute-force experiments as a comparison.
For examples of the first type, it was necessary to only optimize across a small number of dimensions (n = 3), to allow the brute-force experiments to all be run in a reasonable time frame. That exhaustive search was only necessary in this case to provide an instance of the objective function on the complete optimization space to allow for simulations to demonstrate the improved performance of the algorithm. In real use, we would expect a larger number of parameters to be varied without the need to exhaustively attempt all possible combinations merely for simulation purposes. A second example of this type is the limitation of the options to particular choices. For instance, nothing about the experiment necessitates that 30 min and 60 min are possible durations but 45 min is not. We chose to limit the options to these choices to allow the brute force to exhaustively test all possibilities, but in actual use, duration could be a continuous variable with only lower and upper limits defined.
One limitation of the second type is that the varying concentrations of substrate and enzyme were achieved by premixing the batches of each at the selected concentrations. This aided in the development, but during the optimization run, some concentrations will tend to be chosen more frequently than others, resulting in wastage. This limitation could be overcome with dynamic dilution approaches, which would also allow concentrations to be varied continuously, as with duration as discussed above. A final limitation of the second type is that the space of all possibilities must be defined at the outset of the optimization. This is shared with other traditional approaches like DOE, but more manual optimization approaches may choose to vary a different set of parameters midway through the run based on the results thus far.
Future work will focus on applying this technology to more complex biological systems such as multistep biochemical assays and cellular assays. Additional optimization parameters can include variables such as reaction solution composition (buffer, pH, detergent, chelation agents, reducing agents, salts, cofactors, decoy proteins), key reagents (enzyme batches and sources, antibodies, cell lines), reaction conditions (temperature), and processing steps (washes, reader settings), among others. The assay optimization score can also be improved or tailored to specific applications. For example, while extremely useful for many assays, the Z′ metric threshold of 0.5 may not be suitable for certain assays. 23
A primary focus of this work compared the Bayesian algorithm to brute force. Brute force was chosen as the comparator because automated laboratories (such as the NCATS HTS facility) can and do exploit this nonadaptive approach for optimization. Potential future work could be direct comparison to other optimization methods such as DOE, simulated annealing, or alternative Bayesian algorithms.
We hypothesize that this approach could be rendered more efficient for assay optimization by including multiple experimental conditions per assay plate. If the optimization score can be satisfactorily estimated using only a portion of the wells on a given plate, multiple conditions could be combined on the same plate, although some factors (e.g., time) would need to be shared. Adapting this platform to reaction screening is another future endeavor, especially in the context of the NCATS ASPIRE Initiative, which includes a significant automated chemistry component. Other potential applications of this technology could involve hit selection for confirmatory testing and in iterative screening approaches. It is therefore anticipated that the continued development of this integrated, AI-driven autonomous system will enhance the efficiency of assay development and other complex tasks in early preclinical drug discovery at NCATS. Through dissemination of lessons learned and best practices, the scientific community at-large should also benefit.
Footnotes
Acknowledgements
The authors acknowledge the following financial support: intramural program of NCATS and NIH. The content of this publication does not necessarily reflect the views or policies of the Department of Health and Human Services, nor does mention of trade names, commercial products, or organizations imply endorsement by the U.S. Government.
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: D.S. and C.K. are founders of Kebotix and S.E. is an employee of Kebotix.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
