Abstract
Aneuploidy is caused by problems during cellular division and segregation errors during meiosis that lead to an abnormal number of chromosomes and initiate significant genetic abnormalities during pregnancy or the loss of a fetus due to miscarriage. Screening and diagnostic technologies have been developed to detect this genetic condition and provide parents with critical information about their unborn child. In this review, we highlight the complexities of aneuploidy as a disease as well as multiple technological advancements in testing that help to identify aneuploidy at various time points throughout pregnancy. We focus on aneuploidy diagnosis during preimplantation genetic testing that is performed during in vitro fertilization as well as prenatal screening and diagnosis during pregnancy. This review focuses on DNA-based analysis and laboratory techniques for aneuploidy detection through reviewing molecular- and engineering-based technical advancements. We also present key challenges in aneuploidy detection during pregnancy, including sample collection, mosaic embryos, economic factors, and the social implications of this testing. The goal of this review is to synthesize broad information about aneuploidy screening and diagnostic sample collection and analysis during pregnancy and discuss major challenges the field is still facing despite decades of advancements.
Introduction
Aneuploidy is linked to a variety of diseases and conditions during tumor growth in cancer,1,2 the development of a fetus during pregnancy, 3 and more. These conditions involve the presence of unbalanced genomes in cells due to copy number variations depending on lost or gained chromosomes, and can also lead to imbalances in protein stoichiometry.4,5 Notably, aneuploidy is a result of errors during cell division, including mitotic errors such as spindle-assembly checkpoint (SAC) pathway mutations, premature loss of chromatid cohesion, atypical kinetochore attachment, or extra centrosomes. 4 Segregation errors during meiosis can also cause aneuploidy cell formation; these errors include missegregation of homologous chromosomes during meiosis I or of sister chromatids during meiosis II. 4 When gametes develop these errors and are then fertilized, this can lead to embryos with one or three copies of a specific chromosome, instead of the normal two copies. Aneuploidy occurs in around 0.4%–0.6% of newborns and is responsible for more than 50% of spontaneous abortions or miscarriages.3,6,7 Many of these chromosomally abnormal conceptions are associated with increased maternal age, which has become a more relevant factor over time as it is becoming more common for women to be pregnant later in life. For example, the proportion of pregnancies in women 35–39 and 40–44 years of age has increased 9% and 15%, respectively, from 2007 to 2016 in the United States alone.3,7,8
While aneuploid embryos do lead to a disproportionate amount of miscarriages, there are a limited number of aneuploidies that are still compatible with human life. 5 The most common of those aneuploidies is trisomy 21, also known as Down syndrome, accounting for more than 50% of aneuploidy cases, which include live births, fetal deaths, and pregnancy terminations. 3 Next is trisomy 18 (Edward’s syndrome) at 15% and trisomy 13 (Patau’s syndrome) at 5% of cases, which generally result in only a few months of survival for the baby.3,5 Lastly, sex chromosome aneuploidies (45, X and 47, XXX, XXY, and XYY) account for about 12% of these aneuploidy cases. 3 Trisomy 21 occurs once in every 750 live births and involves an extra copy of the more than 300 genes located on chromosome 21. 9 This genetic alteration can lead to a variety of phenotypic changes, including learning difficulties, cardiac deficiencies, distinctive facial features, and leukemia, among others. 10 One possible explanation as to why fetal development is still possible for trisomy 13, 18, and 21 cases is that they have the fewest number of transcripts that they encode on their respective chromosomes, limiting the consequences that can occur from a lost or extra chromosome copy compared with others.
Ultimately, there is a need for proper detection of aneuploidy during embryo development to inform parents of the health of the developing baby. Aneuploidy detection can occur at various time points during pregnancy using prenatal testing or even before pregnancy during in vitro fertilization (IVF) if preimplantation genetic testing for aneuploidy (PGT-A) is performed. PGT-A is used for patients with recurrent pregnancy loss or implantation failures that may be caused by a woman’s decline in fertility as a result of increased maternal age or a number of other factors. 7 PGT-A is used in conjunction with IVF to select the embryos with the best potential chance of survival and normal development. 7 A trophectoderm (TE) biopsy is used for sample collection during PGT-A which removes cells for genomic analysis that eventually become the placenta and ensures that the developing embryo in the inner cell mass is not disturbed. 11 Genomic analysis is used to make an even more informed decision as to which embryos are the best to transplant; it also allows for more single-embryo transfers, which reduces the chance of twins or other sets of multiple babies. 7 This is beneficial because it can lead to fewer preterm births and low birthweight infants and provides a better chance that a healthy pregnancy will be achieved sooner. 7 PGT-A is highlighted as one aneuploidy diagnostic method in Table 1 .
Selected Techniques for Aneuploidy Diagnosis Testing during Pregnancy.
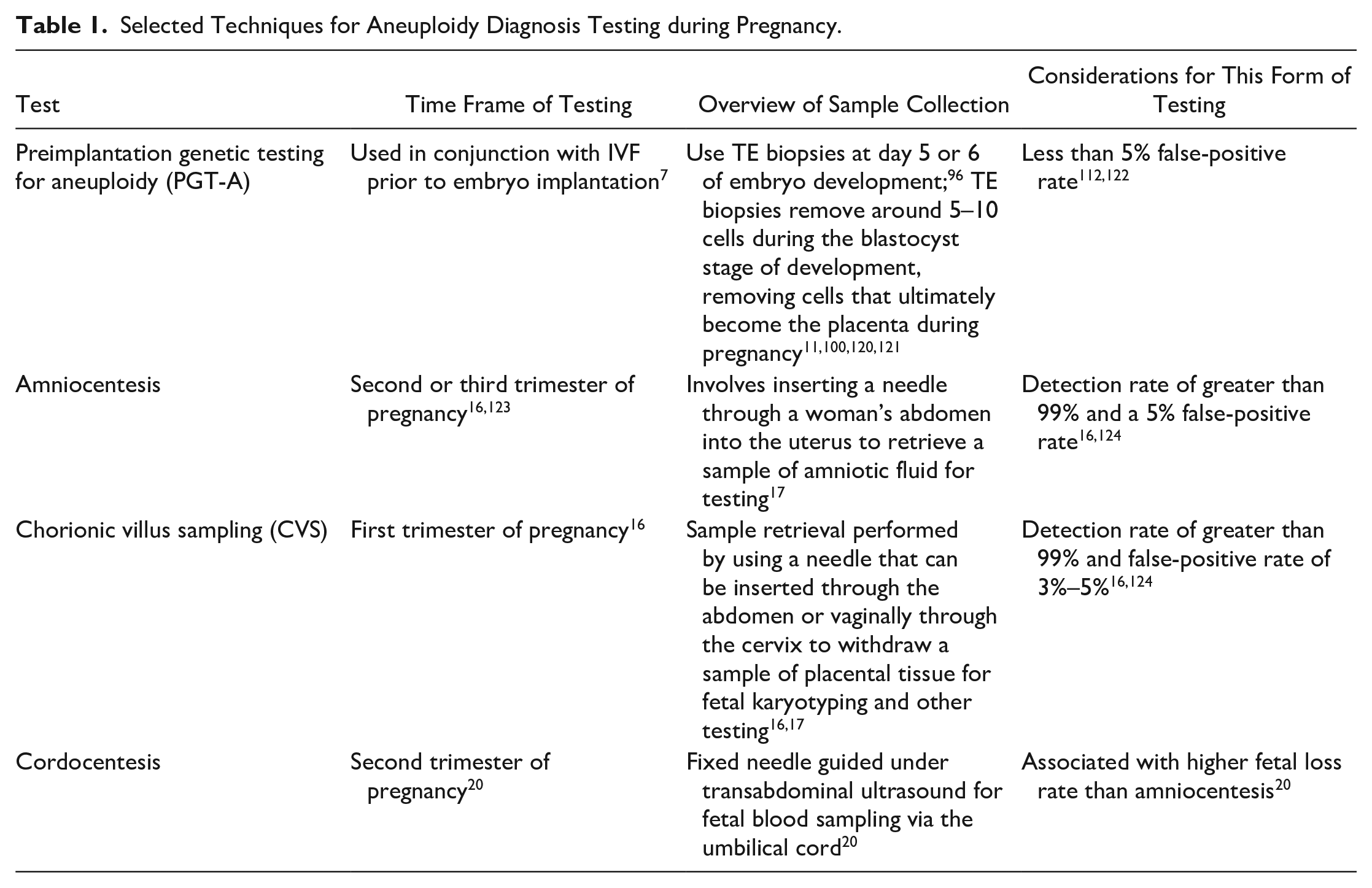

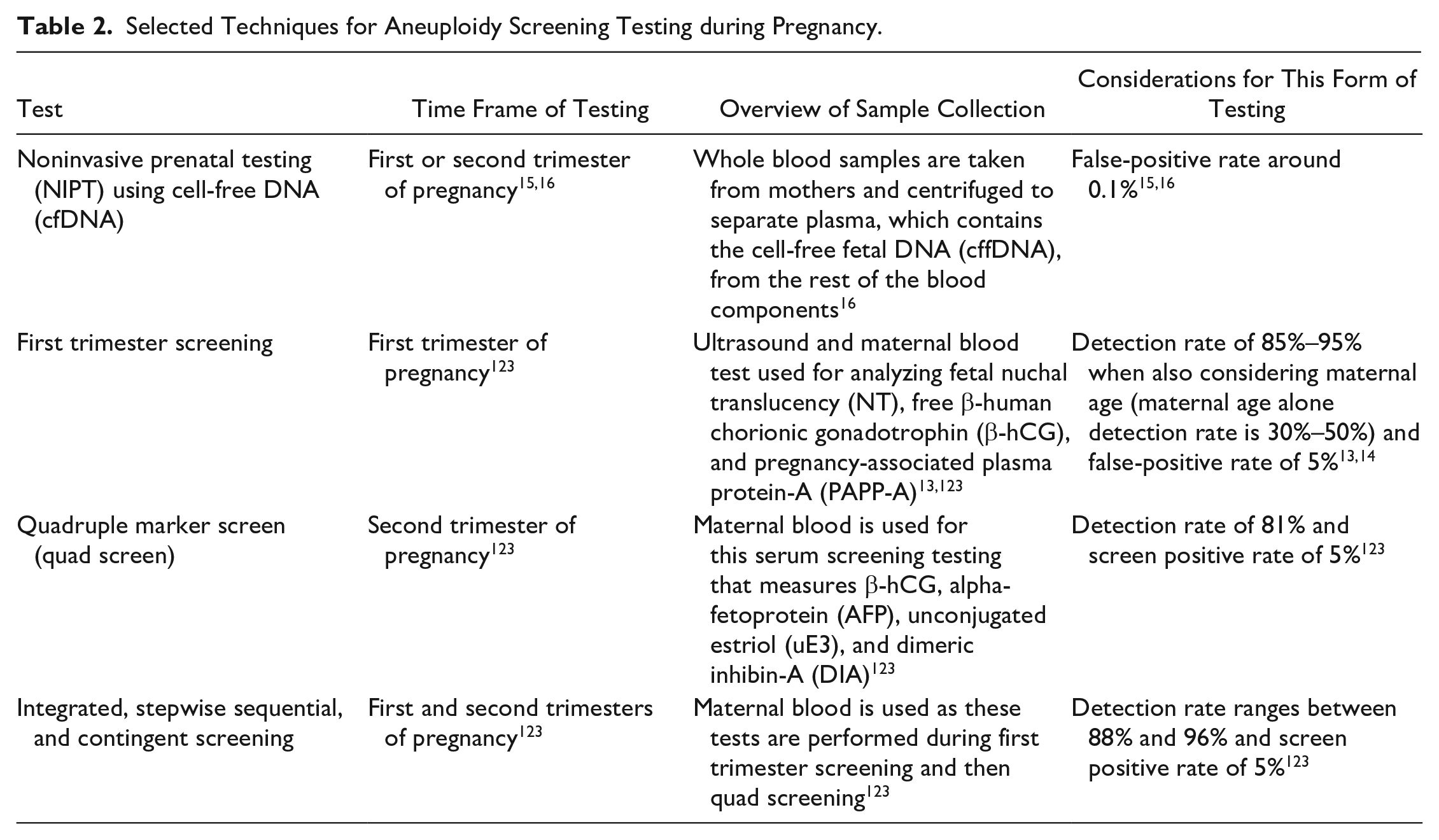
Another point in the pregnancy progression where aneuploidy detection occurs is in prenatal testing. This is done using a variety of different sample collection and analysis techniques. Since 2007, the American College of Obstetricians and Gynecologists (ACOG) has advised that all pregnant women be offered prenatal screening for fetal aneuploidy, but at least 20% of women elect for no aneuploidy screening, even without financial barriers.3,12 Some screening methods that have been or are currently available for aneuploidy detection are first trimester screening; quadruple marker screening; integrated, stepwise sequential, and contingent screening; and noninvasive prenatal testing (NIPT).13,14 Screening techniques cannot technically provide an aneuploidy diagnosis but are crucial in informing patients of the possibility of having an affected fetus and whether further diagnostic testing should be performed. 15 Table 2 features some key characteristics of each of these screening methods, while DNA-based laboratory methods are discussed in greater detail in this work. In addition to screening options during prenatal testing, there are diagnostic options that may be recommended when a pregnancy is at a higher risk for aneuploidies. 14 Chorionic villus sampling (CVS) and amniocentesis are both invasive diagnostic tools and were previously associated with a risk of miscarriage at a rate between 0% and 2%.16,17 However, more recent studies have challenged that figure and concluded that neither CVS nor amniocentesis is associated with an increased risk of miscarriage or stillbirth, especially when compared with a control group of women with similar risk profiles.18,19 Cordocentesis is one more, less common, prenatal diagnostic procedure that has been used when diagnostic information cannot be gathered by amniocentesis or CVS. 20 Table 1 shows more information about these diagnostic tests that can be used for aneuploidy detection during pregnancy.
Selected Techniques for Aneuploidy Screening Testing during Pregnancy.

Overall, these techniques show many sample retrieval strategies for PGT-A and prenatal testing for aneuploidy diagnostics and screening. This review aims to discuss multiple molecular- and engineering-based sample analysis approaches that have been used for this important detection technology with a major focus on DNA-based analysis. Additionally, this review discusses specific challenges that researchers, clinicians, and expecting parents face when using these different types of tests, including sample collection, mosaic embryos, economic factors, and social implications of aneuploidy detection testing.
Molecular-Based Approaches for Detecting and Studying Aneuploidy
Following sample collection there are a variety of techniques that can be used for the detection and analysis of fetal aneuploidy on a genetic level in both PGT-A and prenatal testing. First, we discuss molecular-based approaches that have been used and adapted for these forms of testing.
FISH, SNP Arrays, and aCGH
One cell-based molecular technique of historic significance in the aneuploidy detection field to begin with is fluorescent in situ hybridization (FISH). FISH works by hybridizing highly specific probes to different chromosomes that can be used to identify the number of copies of a given chromosome in a nucleus. 21 This technique has been performed using cells from chorionic villus and amniotic fluid samples during prenatal testing and was once seen as a major step in the field because it could be performed more rapidly than other existing techniques, such as cytogenetic analysis.21,22 In 2013, Rosner et al. performed a retrospective study on 470 patients who had CVS and FISH detection within 1 day and had a 90% sensitivity and 99.4% specificity in their results. 23 FISH has also been performed on PGT-A samples and was extremely beneficial for patients of advanced maternal age with repetitive implantation failure. 24 In a prospective, randomized controlled trial performed by Rubio et al. in 2012, they found a significant increase in live birth rates per patient when FISH analysis was performed on chromosomes 13, 15, 16, 17, 18, 21, 22, X, and Y compared with patients with no PGT-A analysis. 24 While FISH has been a crucial technique used in aneuploidy detection, many DNA-based technologies continue to outperform this technique due to the low resolution, limits on how many chromosomes can be analyzed, and the accuracy of the technique compared with others. 25
In addition to detection, this molecular technique can also be used to study diseases caused by aneuploidy, such as Down syndrome. Korbel et al. used FISH as a technique to aid in creating a high-resolution genetic map of Down syndrome phenotypes based on analyzing 30 subjects who carry rare segmental trisomies of different regions of chromosome 21. 26 This research group mapped segmental trisomies at exon-level resolution to find regions of the genome that cause eight different phenotypes associated with Down syndrome. 26 This study shows the advantages of using advanced genomics with a rare patient population to study copy number variation and complex diseases that were not previously feasible.
In addition to FISH, there are array-based, molecular-based technologies for aneuploidy detection. Array-based technological advancements in PGT-A chromosome testing have included array comparative genomic hybridization (aCGH) and single-nucleotide polymorphism (SNP) arrays. These techniques use whole-genome amplification (WGA) of genomic DNA, which has the potential to introduce small sample quality issues with potential defects in testing these samples. 7 Following WGA, aCGH works by loading the fluorescently labeled amplified DNA and reference DNA to a microarray where the DNA from the two sources then compete to hybridize to thousands of DNA probes associated with sequences on different chromosomes. 7 Following hybridization, the fluorescent signal is read, and a chromosomal copy number of the DNA is calculated based on the reference DNA. 7 In a study performed by Kan et al. of 220 amniotic fluid or chorionic villi samples, all but one triploidy case were concordant with conventional cytogenetic results and 7 of 220 samples also had copy number variants detected with aCGH that were not found in the cytogenetic results. 27 Large clinical cohort studies using microarray analysis have also been performed. Notable studies by Shaffer et al. and Wapner et al. included 2858 and 4406 women, respectively.28,29 In the report by Shaffer et al., it was found that microarray analysis was able to identify clinically significant genomic alterations in 6.5% of the total cases with one or more abnormal ultrasound findings, where most of these cases were less than the resolution of karyotyping. 29 Similarly, in the work by Wapner et al., additional, clinically significant cytogenetic information was also identified by chromosomal microarray analysis that was not present with karyotyping. 28 Large clinical studies such as these provide evidence for substratification of specific anomalies that can be successfully detected by these forms of analysis.
aCGH has also been a widely used tool to analyze samples during PGT-A, and many retrospective cohort studies have been performed comparing next-generation sequencing (NGS) and aCGH results during PGT-A since aCGH is a primary test performed while NGS continues to grow in its use in the field. One finding in a cohort study of 548 sample analyzed by NGS and 368 by aCGH was that there was a significantly higher implantation rate in the NGS group at 71.6% versus 64.6% in the aCGH group. 30 Additionally, the ongoing pregnancy/live birth rate for the NGS group was at 62%, again significantly higher than the aCGH group in this study at 54.4%. 30
SNP arrays are another array-based analysis method for aneuploidy detection that use fluorescently labeled probes that bind to thousands of SNPs throughout the genome, which are used to identify alleles that vary within a population—each person has two allele copies, one on each chromosome. 7 SNP-based testing has been utilized as a high-resolution genome scan during prenatal screening. 31 Parental DNA may also be used during SNP array analysis to identify inherited DNA and help to recognize allele dropout where alleles do not amplify and are therefore undetected during analysis.32,33 Faas et al. performed a study using a 250,000-SNP array and was able to identify microdeletions, inherited copy number variants, copy number variants with uncertain clinical relevance, and homozygous stretches through using this array-based method providing information beyond aneuploidy analysis. 31 Zimmermann et al. also performed targeted SNP-based analysis using cell-free fetal DNA (cffDNA), where five chromosomes in 145 samples that passed a DNA quality test were correctly called, for a total of 725 of 725 correct calls. 34 Additionally, Curnow et al. utilized targeted SNP-based analysis to identify triploid, unrecognized twin, and vanishing twin pregnancies. 35 The study included 30,795 clinical cases for NIPT for fetal whole-chromosome aneuploidies and successfully identified any previously unrecognized cases. 35 Another form of targeted chromosomal analysis is digital analysis of selected regions (DANSR) assays. Juneau et al. used the DANSR targeted approach with microarrays and NGS and found that the microarray method provided faster and more accurate results for the cffDNA measurements. 36 Stokowski also performed targeted cell-free DNA (cfDNA) analysis using DANSR with microarray quantitation and was able to identify 107 of 108 trisomy 21 cases, 29 of 30 trisomy 18 cases, and 12 of 12 trisomy 13 cases, with a specificity of 100% for all chromosomes. 37
SNP arrays are also used during PGT-A and have been found to be as accurate as metaphase karyotyping, which is the technical standard for chromosome analysis. 38 In a study performed analyzing 1800 IVF cycles with 24-chromosome SNP arrays, Simon et al. determined that SNP-based PGT-A could reduce the negative effects that maternal age has on IVF outcomes. 38 They also found that through using their testing method, there was also no statistically significant difference in pregnancy outcomes between single-embryo transfer and double-embryo transfer cycles. 38
PCR-Based Technology
PCR has been the gold standard molecular-based test for many diagnostic fields, from infectious diseases 39 to precision medicine. 40 It is an enzyme-driven process that can replicate a few copies of DNA molecules into large quantities because the amount of DNA in a starting sample is generally too low for accurate measurement. 39 Quantitative PCR (qPCR), quantitative fluorescence-PCR (QF-PCR), and real-time PCR are all terms for different PCR-based tests for detection and quantification. 40 Table 3 provides the pros and cons of PCR-based technologies that can be considered by researchers and clinicians. Researchers have used qPCR during PGT-A to provide a result within 4 h and allow for the transfer of fresh embryos. 7 This has been done using multiplexed qPCR to amplify sequences on different chromosomes and calculate the chromosomal copy number efficiently with an error rate of 1%. 7 qPCR has been verified extensively for chromosomes with common aneuploidies, including chromosomes 13, 18, 21, X, and Y. However, in one randomized controlled trial it was even shown that PGT-A using qPCR can be performed for all 24 chromosomes in a 4 h window with 41 of 42 samples consistent with a karyotype diagnosis or 42 of 42 after a minimum threshold for concurrence is applied. 41 In another study, by Scott et al., comparing IVF outcomes between groups who did or did not undergo PGT-A using qPCR, they found that the group with qPCR-based PGT-A had an implantation rate of 66.4% versus 47.9% in the control group, and delivery rates of 84.7% versus 67.5% in the control group, showing a statistically significant difference. 42
Pros and Cons Analysis of DNA-Based Molecular Techniques.
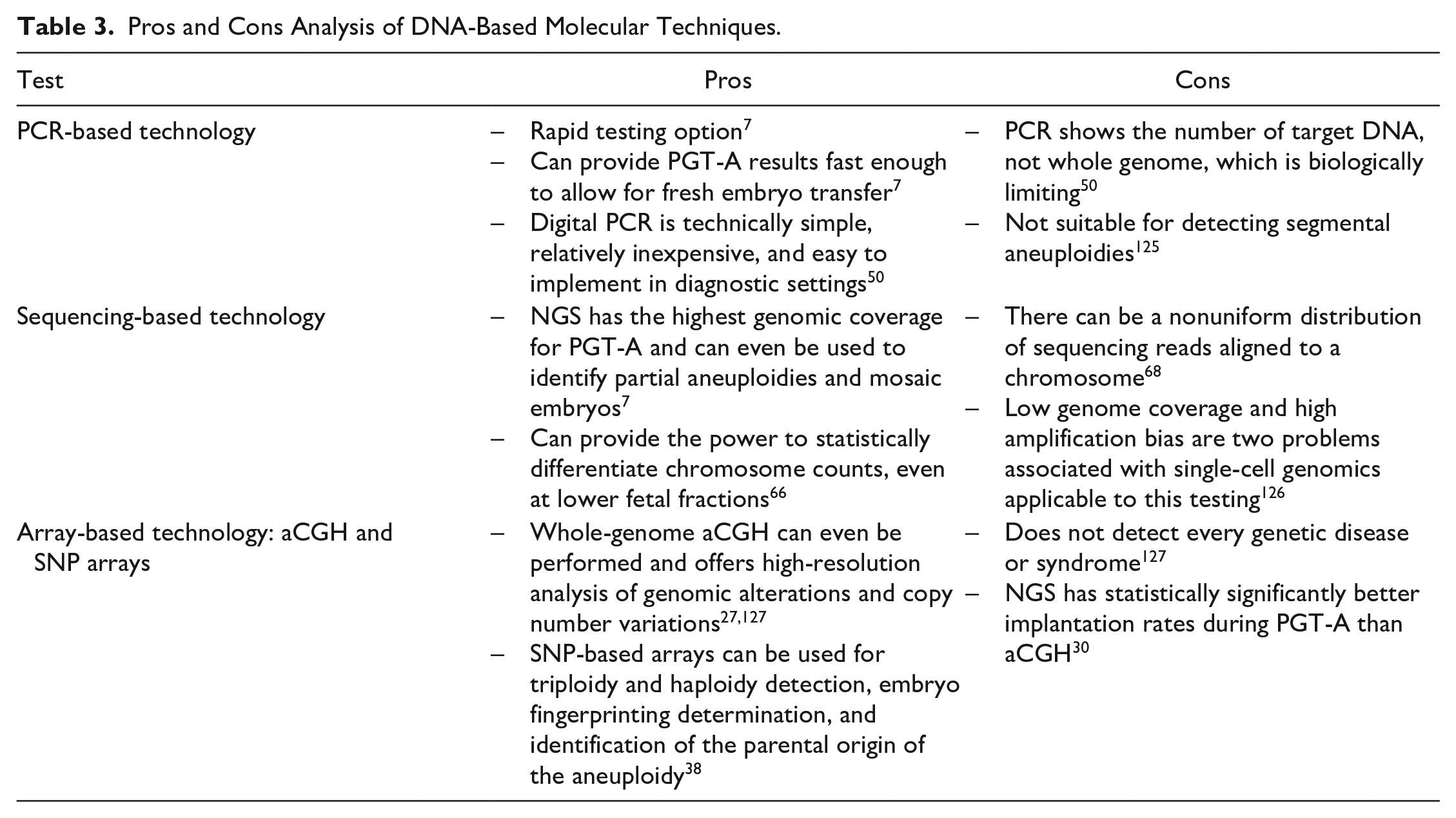
QF-PCR has also been adapted for prenatal screening of large cohorts (7720 and 9080 prenatal samples43,44) for trisomies 13, 18, and 21. Mann et al. reported that out of the 7720 prenatal samples they tested, 97% of them received a report on the business day after the sample was received with no misdiagnosis for nonmosaic trisomies. 43 This shows how PCR-based technologies can provide answers quickly and accurately. QF-PCR has also been used for rapid prenatal testing by targeting segmental duplications in the genome, known as SD-QF-PCR.45,46 This procedure co-amplifies sections of DNA duplicated within the genome on two different chromosomes using a single pair of fluorescent primers. The two chromosomes read as two different sizes when analyzed by capillary electrophoresis 45 or melt curve analysis 46 and result in a relative dosage to determine aneuploidy samples. Digital PCR (dPCR) is another form of PCR that can be used in prenatal testing for aneuploidies such as trisomy 21 using cffDNA. 47 dPCR utilizes multiplex PCR analysis on diluted nucleic acid samples whose positive amplification signal is read as single-template molecules. 47 This is then used to determine the template concentration in the original samples while not relying on dose–response relationships like reporter dyes used in qPCR. 47 A study by Lo et al. showed that through using dPCR on noninvasive trisomy 21 testing, they were able to accurately detect trisomy 21 in samples containing 25% fetal DNA in 2007, where current dPCR technology can analyze samples at less than 4% fetal DNA.47,48 In a study by Tan et al., they showed that dPCR can be used for multiplex droplet dPCR during NIPT using universal locked nucleic acid probes to test for fetal aneuploidy. 49 They were able to accurately diagnose 60 of 60 clinical maternal plasma samples using their accessible, cost-effective method. 49 El Khattabi et al. also validated their dPCR methodology using multiplex PCR to show the correct function of their probes for technical optimization, followed by validating clinical samples for trisomy 21 with fetal DNA fractions as low as 5%. 50
Sequencing-Based Technology
Following the completion of the Human Genome Project in 2003, sequencing has taken over many diagnostic fields, including those related to aneuploidy. NGS is one form of sequencing that has drawn attention because of the amount of detailed information it provides about the genomic sample. 51 Table 3 also includes pros and cons of sequencing-based technology as it is considered for genomic analysis during aneuploidy detection testing. The procedure for NGS on Illumina platforms uses WGA, DNA fragmentation, and library preparation, which involves adapter ligation and PCR amplification. 7 Examples of WGA techniques that are used to perform copy number variation detection on single or limited cell numbers, including aneuploidy detection, are PicoPLEX DNA-Seq, DOPlify, REPLI-g, and Ampi-1 WGA. 52 These methods and others vary in amplification technique (i.e., multiple displacement amplification vs PCR), genome coverage, representation bias, error rates, yield, and robustness. 52 Following DNA library preparation and sequencing, the data are analyzed by comparing the result to a reference genome where researchers are able to gather the chromosomal copy number information and aneuploidy diagnosis. 7 With the continued technological expansion of NGS, Fiorentino et al. were able to develop a method for 24-chromosome aneuploidy screening for use in PGT-A. 53 A retrospective study was performed to compare the NGS-based method to existing karyotyping of single cells or aCGH data. 53 They found an NGS specificity for aneuploidy call or individual chromosome copy number reading of 99.98% and a sensitivity of 100%, as well as a specificity and sensitivity of 100% for aneuploid embryo calls. 53
NGS has also been used in PGT-A with human embryonic blastocoel fluid (BF) or blastocyst culture conditioned medium (BCCM) as a starting sample instead of a TE biopsy. 54 BF is fluid found inside the embryo that can be aspirated out and is less invasive than a biopsy because no cells are removed, while BCCM is fluid found in the culture plate during IVF and can be easily collected for sample analysis. 55 The retrieval of BF and BCCM is less invasive and technically challenging than a TE biopsy, making it easier and more cost-effective to use for testing. 54 In one example, results of NGS with these starting samples from Kuznyetsov et al. show 47 of 47 of the samples amplified successfully, and the concordance rates of whole-chromosome copy number per sample between this method and using TE biopsies as a starting sample were 100% and 98.2% per single chromosome. 54 In another, more recent study by Kuznyetsov et al. using cell-free embryonic DNA found in spent embryo culture medium (SCM) and BF, the concordance rate per sample for ploidy status compared with the TE biopsy was 97.8%. 56 These alternative starting samples for PGT are known as noninvasive PGT (niPGT). In a recent review of the implications of this form of sample testing for PGT, it was found that BF or SCM can offer DNA suitable for genetic analysis, but there are still mixed conclusions regarding whether it can consistently provide accurate clinical diagnostic information. 57 Li et al. found that the results of their analysis with SCM differed from the TE biopsy samples, 58 and Jiao et al. reported 90% and 86% clinical concordance rates between BCCM and TE biopsy, respectively, compared with the blastocyst-stage embryo. 59 Several other studies have also used BF or SCM for niPGT-A using NGS and found different levels of success with these alternative starting samples.55,58,60–64 While there are many economic and practical reasons for testing from these samples, the reliability, concordance of genetic results with that of the embryo, and DNA quality all pose issues and support the need for further analysis of this starting sample. 57 Lastly, the sensitivity and specificity of NGS for PGT-A have also been helpful in detecting chromosomally mosaic embryos that have two or more cytogenetically distinct cell lines. 65 High-resolution NGS is sensitive enough to detect mosaicism when 20% or more of the cells in the biopsy are aneuploidy—meaning one of five cells in a five-cell biopsy. 7
NGS has also been used in prenatal screening to count or genotype different cffDNA fragments, even at lower fetal fractions. 66 The fetal fraction is the fraction of cfDNA that originates from the fetus/placenta as opposed to the maternal cfDNA that is always circulating. 66 More specifically, cffDNA comes from cytotrophoblastic cells in the placenta where apoptosis is a main mechanism controlling the release of fetal DNA. 67 Using NGS, cfDNA is aligned to each chromosome in order to determine these important counts. 68 Researchers have noticed cases of nonuniform distribution of sequencing reads aligned to a chromosome that they hypothesize is related to the GC content of the cfDNA, which affects PCR and bridge amplification on Illumina platforms. 68 Regardless of these problems, NGS-based technologies such as VeriSeq (Illumina, Inc., San Diego, CA) have been used to screen pregnant women for fetal aneuploidy of chromosomes 21, 18, 13, X, and Y at a minimum of 10 weeks’ gestation, with only 2 h of hands-on time for a technician. 69 The MiSeq and NextSeq platforms from Illumina have also been successfully used with cffDNA for aneuploidy detection, with a 10.9% failure rate in 3594 MiSeq cases and 8.7% failure rate in 1000 NextSeq cases. 70 Targeted NGS has also been utilized to analyze selected genomic regions of interest for NIPT. 71 In a report by Koumbaris et al., a blind study showed 100% diagnostic sensitivity while being more cost-effective as an NGS-based method for NIPT. 71 Overall, NGS is a powerful tool that provides specific chromosomal copy number analysis and is only predicted to grow in its use for aneuploidy detection.
Bioinformatic Methods to Study Aneuploidy
Aneuploidy diseases can also be studied using bioinformatic techniques, such as genome-wide association studies (GWAS), that depend on molecular-based technologies and are able to analyze genetic samples to learn more about diseases. In late 2019, Chernus et al. published the first GWAS to identify genes associated with maternal nondisjunction of chromosome 21 as a way to investigate predisposing factors to this aneuploidy causing error. 72 The study utilized genotyping of 749 live birth offspring with trisomy 21 and 1437 parents to identify mothers with nondisjunction errors in their oocyte. 72 The GWAS analysis found many new potentially associated loci with nondisjunction errors, and candidate gene analysis was also performed, which found candidate genes that were strongly associated with maternal nondisjunction of chromosome 21. 72 This study goes beyond detecting aneuploidy and aims to learn more about what may cause aneuploidy embryos so that the knowledge on this genetic irregularity can continue to expand and help inform patients. Additionally, other studies have also used GWAS and candidate gene analysis to investigate different phenotypes associated with aneuploidy-related diseases, such as Alzheimer’s disease,73,74 congenital heart defect, 75 and Down syndrome-associated atrioventricular septal defect, which is a heart abnormality that people with Down syndrome have a 2000-fold higher risk of having. 76 More recently, Starostik et al. even used published single-cell RNA sequencing data of 74 human embryos to better understand aneuploidy during early human development. 77 This bioinformatic analysis uncovered mosaic aneuploidies in 59 of 74 samples and, by reconstructing histories of chromosome segregation, found that 55 samples had mitotic aneuploidies and 23 had meiotic aneuploidies. 77 One more finding from this study was that aneuploidy cells upregulated immune response genes and downregulated proliferation, metabolism, and protein processing genes. 77 Studies such as these that dive deeper into the genetics of people or families with Down syndrome or other aneuploidy diseases can lead to a lot of progress in the understanding of these fields and data collected during NGS.
Engineering-Based Approaches for Detecting Aneuploidy
In addition to molecular-based improvements to aneuploidy testing, there have been many advancements to the engineered technologies that have been used in conjunction with the molecular tests. Here, we present some of the advancements in fluidic, mechanical, and electrical engineering work that have contributed immensely to the field of aneuploidy detection.
Microfluidic-Based Approaches
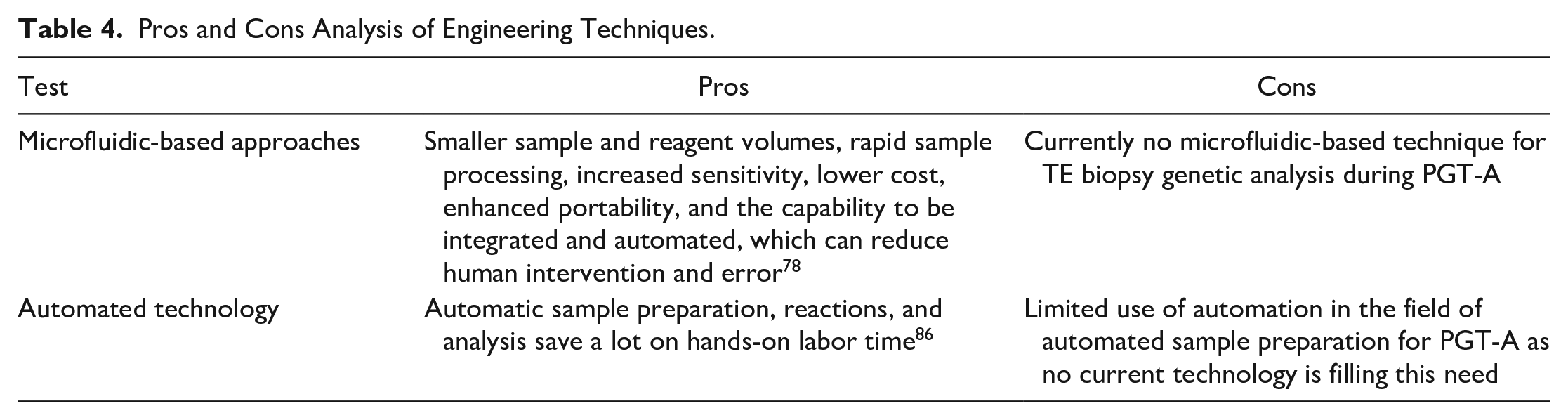
Microfluidic technologies involve controlling and manipulating fluids and particles at the micrometer scale and have numerous benefits compared with working with larger volumes; these are outlined in Table 4 . 78 Microfluidics have been explored for scientific and technological advancements in aneuploidy detection for human-assisted reproduction and mammalian gamete and preimplantation embryo biology. 79 However, there is currently no microfluidic-based technique for TE biopsy genetic analysis during PGT-A. 79 Nevertheless, there are currently multiple noninvasive, microfluidic-based techniques that analyze biochemical signals to aid in embryo selection during IVF, as opposed to using genetic analysis. 79 One study, by O’Donovan et al., used a microfluidic chip to monitor oxygen consumption of embryos to assess their metabolism and aid in embryo selection prior to implantation during IVF. 80 While this is not a genetic-based aneuploidy detection method, it is still used to assist in the embryo selection process to increase the chances of a successful pregnancy, just as aneuploidy testing does.
Pros and Cons Analysis of Engineering Techniques.

Meanwhile, microfluidic devices have been utilized in prenatal testing for fetal aneuploidy detection. For example, Fan et al. designed a microfluidic dPCR system to be used for rapid, allele-independent molecular detection of aneuploidy using amniocentesis and CVS starting samples. 81 Microfluidic dPCR was able to accurately detect 40 of 40 fetal trisomy cases across chromosomes 21, 18, and 13, with each detection taking less than 6 h. 81 Conversely, Winter et al. used a slanted inertial microfluidic device to isolate circulating extravillous trophoblasts from maternal peripheral blood during prenatal testing. 82 With between one and five of these fetal cells per milliliter of blood, their microfluidic device was able to enrich for this rare cell type with 79% recovery and perform successful aneuploidy detection. 82
Additionally, Ho et al. created a microfluidic-based, same-day prenatal diagnostic FISH assay chip. 83 Nanostructured titanium dioxide coated the chips to facilitate cell adhesion; thus, pretreatment and hybridization occurred in the microchannels of their device, with on-chip detection showing results concordant with off-chip karyotypes within 3 h of sampling using 20-fold fewer probes within the microfluidic FISH system. 83 One more microfluidic-based system for aneuploidy analysis utilizes silicon-based nanostructured microfluidics for the capture of circulating fetal nucleated red blood cells (fnRBCs) and extravillous cytotrophoblasts (EVTs) for noninvasive prenatal testing. 84 Huang et al. used specific antibodies in conjunction with the microfluidic system for efficient capture of the targeted cells in maternal blood, 14–22 fnRBCs/4 mL and 1–44 EVTs/4 mL. 84 Samples then went through genetic analysis for diagnosis using a variety of techniques, including FISH, aCGH, and NGS. Microfluidics have been used in the development of rapid molecular prenatal diagnostics since microchips have the ability to save time and costs by reducing the amount of reagents used, protocol times because reagents are more concentrated in microchannels, and thus turnover times for results. 83
Automated Technology
Automation has also provided important benefits to the progress of technological and engineering advancements in aneuploidy detection. Numerous research and commercial labs have been pursuing advancements related to automated sample analysis using techniques like NGS in prenatal screening and PGT-A during aneuploidy testing. 85 Other groups are still investigating alternative ways to automate sample preparation, detection, and analysis, though. One alternative automated prenatal screening option has been contributed by Van Opstal et al. and uses multiplex ligation-dependent probe amplification (MLPA) with one PCR for quick, easy, automated DNA isolation and multiplex analysis of amniotic fluid samples. 86 They found 100% sensitivity and specificity for common aneuploidies using this automated technique compared with FISH karyotypes. 86 Their method used automatic sample preparation, reactions, and analysis, saving a lot on hands-on labor time. 86 Automated microscopy has also been used in FISH analysis 87 and to analyze circulating fetal cells as a potential for chromosomal analysis. 88 Automated FISH analysis was implemented because the manual process is both expensive and labor-intensive, requiring a high skill level and subjective interpretation of signals. 87 Wauters et al. developed an automated FISH examination platform for amniotic fluid cells and was able to correctly diagnose 146 of 146 samples compared with karyotype results with three aneuploidy cases. 87 More recently, cfDNA has also been subjected to innovations in automated analysis using NGS during NIPT. 89 Crea et al. utilized the ion proton sequencing platform to screen for fetal trisomy 13, 18, and 21 with 100% sensitivity and specificity for 110 samples. 89 cfDNA has also been incorporated into an automated, digitally enabled assay for aneuploidy screening by Dahl et al., who used molecular probe technology to label target chromosomes and then enriched for single molecules for imaging by passing them through a nanofilter. 90 They classified 30 of 30 trisomy 21 pregnancies correctly using their core Vanadis NIPT assay 90 and are affiliated with Vanadis Diagnostics, a PerkinElmer company, showing the translation of this form of automated screening from research labs to larger biotechnology companies.
Challenges and Considerations for the Field of Aneuploidy Detection during Pregnancy
Fetal Fraction during NIPT
The field of aneuploidy detection has had numerous advancements over the life span of this form of testing. With these advancements, though, have come many challenges, which continue to push research in the field forward. One of the first challenges that may be faced in PGT-A and prenatal testing is retrieving a starting sample that is large and pure enough for accurate analysis. Although simple to retrieve, one of the more challenging starting samples to use for testing has been cffDNA during NIPT. This is because the average fetal fraction of cffDNA in maternal plasma can be less than 3% to up to 30%, with an average range of 10%–15% between 10 and 20 gestational weeks. 66 Many studies have shown that accuracy in testing results increases as fetal fraction increases and that samples with fetal fractions of less than 3%–4% are not suitable for aneuploidy screening tests. 66 In order to detect a trisomy sample, the method must be able to distinguish a 50% increase in the amount of cffDNA from a specific chromosome, which has placed many limitations on detection methods, since it is challenging to identify this difference in the fetal DNA when the fetal fraction is such a small percentage. There are many biological influences on fetal fraction from both fetoplacental and maternal factors. 91 Fetoplacental influences on fetal fraction include gestational age, crown-rump length, mosaicism, fetal aneuploidy, triploidy, and multiple pregnancy. 91 Maternal factors that affect fetal fraction consist of maternal weight, maternal autoimmune disease, low-molecular-weight heparin, assisted reproductive technology conception, maternal age, and more. 91 Many researchers have identified maternal weight as one of the most clinically significant influences on fetal fraction due to there being a fixed amount of fetal DNA in maternal plasma, but an increased maternal weight increases the maternal circulatory volume, thereby diluting the cffDNA in the system.66,91 Vora et al. even reports that total cfDNA in maternal plasma at the time of the baby’s birth is directly proportional to the mother’s body mass index (BMI). 92 This indicates that an increase in the total amount of cfDNA will also decrease the fraction of cffDNA, further complicating aneuploidy testing by decreasing the starting sample. 92 Another factor that can alter the analysis of cffDNA is maternal malignancy, which can affect the copy number variation profile in NIPT.93,94 Cell-free tumor DNA can be detected in plasma just as cffDNA is during NIPT. 94 Tumor DNA can include duplications and deletions across the genome, which can then alter the ratios between patient and reference DNA and provide test failures or unusual results. 94 A systematic review of literature reporting about false-positive or false-negative NIPT results showed that 15% of the explained false-positive cases were due to maternal malignancy out of the 22 studies with 206 discordant cases analyzed. 95 This figure does not directly reflect true clinical frequency but identifies maternal malignancy as a potential reason for false-positive NIPT. 95
Embryo Mosaicism
Another sample-related factor that can impact aneuploidy detection during prenatal screening or PGT-A is the degree of mosaicism a sample has. Mosaicism occurs when there are both normal and aneuploidy cell lines in one embryo. 96 In NIPT, mosaicism will affect the sensitivity and specificity of screening, and in PGT-A mosaic embryos show decreased implantation rates and increased miscarriage rates compared with euploid or healthy embryos.7,66,91 During PGT-A, the two cell types in one embryo can often cause aneuploidy misdiagnosis since a cell biopsy taken at an early stage may not accurately represent the ploidy status of the cells in the whole embryo. 96 According to previous studies on PGT-A, it was suggested that more than one cell should be biopsied for correct diagnosis, 97 which is important since the prevalence of mosaicism has been reported to be as high as 20% of the embryo. 98 In a study by Gleicher et al., mathematical modeling was even employed to predict that a biopsy of at least 27 TE cells is needed to reach minimal diagnostic predictability. 99 The two models presented in the study assume that there is an even distribution of mosaicism in the TE, although mosaicism is usually clonal, and challenge the clinical utilization of current PGT-A sampling, which generally uses a six-cell biopsy. 99
Reciprocal errors are another biological challenge associated with mosaicism that must be overcome, which involves one monosomy cell line for a chromosome and one trisomy cell line for that same chromosome in one embryo. In this case, when the sample is pooled together to be analyzed at once, these aneuploidies would go undetected by canceling each other out and lead to a false-negative aneuploidy diagnosis. 98 There are also analytical challenges associated with mosaicism, as statistical smoothing and copy number assignment algorithms are generally not updated to accurately call out mosaic chromosomes. 98 Additionally, there are limitations regarding how to translate information about mosaicism to the clinic so that patients can understand the effects of particular chromosomal mosaicism on implantation, miscarriage, and having a healthy child.98,100 Groups such as the Practice Committee of the American Society for Reproductive Medicine (ASRM), Genetic Counseling Professional Group (GCPG), Preimplantation Genetic Diagnosis International Society (PGDIS), and Society for Assisted Reproductive Technology (SART) have all worked to provide statements or reports regarding the clinical management of mosaic and aneuploidy embryo transfer to address these limitations.101–103 There have also been alternative statements presented in response, for example, by the International Do No Harm Group in IVF (IDNHG-IVF), regarding concerns of increasing added procedures to IVF. 104 In addition to statements by groups or professional societies, continued research has been pursued regarding the safety of these embryo transfers by groups, including Grati et al., who devised an evidence-based scoring system for prioritizing mosaic aneuploid embryos for transfer. 105 Overall, the attention to this topic further emphasizes the need for efficient, accurate diagnostic technology with targeted answers relevant to the clinic, especially as more clinics are choosing to transfer mosaic embryos if patients do not have any euploid embryos they can transfer and they cannot undergo another cycle of IVF. 7
In prenatal diagnosis, chromosomal mosaicism presents a similar challenge during screening and often appears as full aneuploidies. 106 Depending on the prenatal testing performed, it is possible to find true fetal mosaicism (TFM) or confined placental mosaicism (CPM). Amniocentesis can be used to diagnose TFM, while a test like NIPT could find TFM and CPM. 106 One problem with CPM is that in around 1% of cases, the CPM result would show an aneuploidy cell line based on reading cytotrophoblasts during cffDNA testing, while the fetus does not, causing a false-positive result. 106 Overall, mosaicism needs to be considered in both PGT-A and NIPT prenatal testing, especially to make sure that patients are receiving accurate results.
Economic Considerations
In addition to challenges related to the samples being tested, another major challenge associated with aneuploidy detection and patients’ access to testing is cost. Since the ACOG has advised that all pregnant women be offered prenatal screening for fetal aneuploidy, the majority of women do receive this testing, at least in the United States. 12 As there are a variety of prenatal tests that can be performed, test selection depends on factors such as a woman’s prior risk, stage of pregnancy, access to healthcare services, cost, and patient preference. 107 The economic concerns related to offering specific types of testing have been investigated by many different groups since different tests incur different costs based on reagents and equipment needed, as well as the amount of time the test takes. Gekas et al. investigated the cost-effectiveness and detection of Down syndrome using rapid testing versus karyotyping and found that rapid testing methods can detect common aneuploidies faster and are less expensive than karyotyping. 108 Through comparing the global costs of karyotyping and QF-PCR, the authors found that, on average, karyotyping costs $0.97 million (CAD) more per year than QF-PCR over a variety of screening strategies. 108 Additionally, they found that the cost-effective ratio—net cost divided by changes in health outcome—for QF-PCR was $24,084 per Down syndrome case detected, versus $27,898 for karyotyping. 108 From this information, the authors even suggested that it may be appropriate to question the additional costs of karyotyping, which may be especially important in the United States, where guidelines recommend that karyotyping be performed as a follow-up to a rapid aneuploidy diagnosis (RAD) when needed.
Additionally, with prenatal testing Song et al. also studied the clinical and economic consequences of fetal trisomy 21 screening with NIPT using cfDNA in high-risk pregnancies. 109 When comparing NIPT to first trimester screening and integrated screening (INT), which both look at serum markers, they found NIPT was more clinically effective at detecting trisomy 21 and less costly than the other two options even at a base price of $795. 109 Kaimal et al. also had a similar finding that for women 40 years or older, using cfDNA for primary screening is both optimal and cost-effective with a cost-effective ratio of $73,154 per quality-adjusted life-year. 110 Lastly, one more study done by Benn et al. corroborated the economic benefits to using NIPT for screening over other techniques. 107 They evaluated the economic value of replacing conventional fetal aneuploidy screening with NIPT and found that it would reduce healthcare costs if it could be performed for $744 or less in the general pregnancy population. 107
Investigating costs and cost–benefit analysis is just as important when it comes to IVF and PGT-A. IVF treatment numbers have increased fourfold over the last 20 years to 160,000 treatment cycles annually as of 2018. 111 At an out-of-pocket cost ranging between $10,000 and $15,000 per treatment cycle, many states in the United States are turning to policy changes to mandate insurance coverage, which would reduce prices to $2000 to $3000. 111 Unfortunately, with the large cost of IVF on its own, cost also remains a significant limiting factor to using PGT-A in every IVF patient. 7 Although these forms of testing can add an additional cost burden, Neal et al. found that it is actually more cost-effective over time for IVF patients with more than one embryo to perform PGT-A than it would be to use other non-genetic-based tools to select an embryo for implantation. 112 With 8998 patients from 74 IVF centers, the study found that using PGT-A could save $931 to $2411, depending on the number of embryos screened. 112 Additionally, performing PGT-A reduced the treatment time by up to 4 months, and these patients experienced fewer failed embryo transfers and clinical miscarriages. 112 Collins et al. also had a similar result from their decision analytic model for using PGT-A during IVF to increase live birth rates in women over 37. 113 They found that the cost of only IVF was $19,415 per cycle, with a live birth rate of 13.4% and an incremental cost-effective ratio of $145,063. 113 This compared to when preimplantation genetic screening was also performed during IVF, which increased live birth rates by 4.2% at an additional cost of $4509, leading to an incremental cost-effective ratio of $105,489 per additional live birth. 113 Thus, although there is a cost associated with these additional techniques, it is important to consider the cost benefits patients will see over time, especially in reducing treatment times and improving results.
Social Implications of Testing
Another important consideration for challenges in the field of aneuploidy detection is the social implications associated with the testing. While ensuring safe and accurate tests is important, it is just as imperative to consider the effect this testing could have on people both physically and mentally. In 2019, Labonté et al. performed a scoping review of the evidence on psychological and social consequences to performing NIPT. 15 The findings of this studied showed that short-term anxiety for women decreased with a negative NIPT test result and that regret associated with decisions made was generally low. 15 Conversely, Green et al. found that up to 30% of women with a positive aneuploidy result showed regret about performing a screening test. 114 One interesting note from Labonté et al. was that there remains to be a study on the long-term effects of NIPT or on women’s partners’ outcomes, as the short-term effects of the testing are currently the only metric being investigated. 15 This is a place where the field can continue to add value and important results for parents before they undergo these procedures.
Another social implication of this prenatal testing, specifically NIPT, is learning the sex of the baby, which could expose societal issues where one sex is preferred over the other. 115 Farrell et al. utilized focus groups of pregnant women and women who had delivered in the past year to learn more about their opinion on this capability of NIPT, among other factors. 116 They found that there were cases where women did not know they would learn the sex of the fetus from NIPT until they were asked if they wanted to know, whereas others wanted NIPT specifically to learn the fetal sex. 116 One participant in the study cautioned that expecting mothers should be made aware of the purpose of the testing and the possibility of learning about genetic abnormalities from the testing. 116
This idea brings up an important point that as aneuploidy detection technology continues to advance, it is suggested by more doctors, and more patients are getting testing, it is also important that the level of education and counseling for this testing also grow. This counseling should specify the risks, benefits, and limitations of each kind of screening or diagnostic test so that pregnant women can make the best decision for their health and their baby’s. As there may be a perceived pressure to undergo NIPT or other testing based on recommendations, this is even more of a reason why a realistic scope of the psychological and social consequences should be fully expressed before testing. 15 In addition to the educational value of this counseling, it also allows for reproductive autonomy. In a 2009 Research Highlight by Nature Review Genetics, the authors argue that prenatal testing is unique in that there is currently no cure to the genetic conditions being tested for and diagnosed; thus, they pose that the mother’s only options are to “accept the child’s impairment or to terminate the pregnancy.” 117 While testing for aneuploidy is an important tool, the reproductive autonomy it provides also poses a third option for pregnant mothers, which is to not perform the testing to begin with. With the use of informed consent, where women need to provide permission to perform this testing, this should always be an option, although it has been found that it often is not as these tests have become more routine.117,118 When given the option to undergo testing, a cohort study published in 2016 showed that with more than 28,000 pregnancies screened with cfDNA, less than 40% of those women who screened positive chose to have invasive prenatal diagnosis as a follow-up test. 3 In the Netherlands, when a nationwide implementation study on NIPT, called TRIDENT-2, was offered as a first-tier test for all pregnant women in 2017, only 42% of women selected to undergo the testing. 119 These statistics, and many more, show that a woman’s decision for testing during pregnancy cannot be assumed. It is the job of researchers, clinicians, and genetic counselors to provide a thorough analysis of benefits and risks for patients for different testing options so that they can best understand how this testing is meant to be helpful and informative regarding the health of their child if they choose to undergo testing. 118 Additionally, results of the testing must be managed since many gene variants with unclear or unknown significance (known as “variants of uncertain significance”) during DNA sequencing of an unborn fetus can provide information to patients that is difficult to interpret. 118 As there is currently an insufficient number of experts in prenatal genetics to counsel all pregnant women pre- and posttesting, this further emphasizes the need for more educational resources and informed consent. 118
In the case that a screening or diagnostic test does lead to a positive aneuploidy result, this testing can give women and families the opportunity to seek out support and references as soon as possible. The National Down Syndrome Society (NDSS) is one example of a support network for disabilities caused by aneuploidy. NDSS provides access to Down syndrome parent support groups with more than 375 affiliates, as well as advice for parents when learning of a positive Down syndrome diagnosis. Other resources include Dads Appreciating Down Syndrome (DADS), specific to fathers of Down syndrome children, and Know Me First, which has educational resources for parents when they receive a Down syndrome diagnosis. Knowledge of these resources can be just as important as knowledge of different forms of testing during pregnancy and can help mothers being tested too. Overall, this testing has the ability to inform and empower expecting mothers by advising them regarding their child’s health and tools and resources out there that they can look to for support, helping not only babies but also women.
Conclusion
This review paper analyzed a variety of different sample collection and analysis techniques for PGT-A and prenatal testing for aneuploidy, specifically focusing on laboratory-based technologies. While CVS and amniocentesis are extremely accurate diagnostic procedures, the invasive nature of these techniques and chance for miscarriage have pushed the field toward NIPT as a highly accurate screening tool. Although challenges have been exposed for NIPT, such as low fetal fractions and mosaic embryos, economic considerations show that this test is more valuable over other screening options. Additionally, with the growth in analysis tools such as dPCR and NGS, NIPT sample analysis will consistently improve as a reliable and safe testing method. This review also aimed to explain that the progress in engineering can go along with molecular analysis progress to decrease human interaction during testing and increase the accuracy of results. Lastly, social implications were discussed in tandem with the scientific analysis to explain how education and genetic counseling are necessary in this field of science and medicine.
Footnotes
Acknowledgements
We would like to acknowledge the support for this review paper from leading researchers in prenatal screening and PGT-A from Brown University and PerkinElmer.
Declaration of Conflicting Interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: AT is a paid scientific advisor/consultant and lecturer for PerkinElmer.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The authors acknowledge financial support from Brown University and PerkinElmer.
